Cell-free DNA as a potential biomarker of differentiation and toxicity in cardiac organoids
Figures
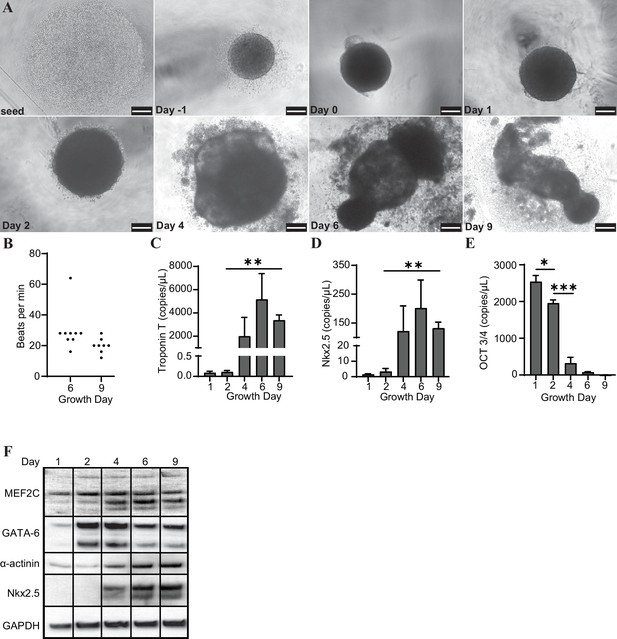
Cardiac organoids derived from H9 embryonic stem cells exhibit characteristic morphological changes and express markers of differentiation.
(A) Brightfield images of cardiac organoids during growth and differentiation. Scale bars represent 100 µm. (B) Quantification of beats per minute across three separate biological replicates for growth day 6 (n = 9 organoids total) and day 9 (n = 7 organoids total). ddPCR of Troponin T (C), Nkx2.5 (D), and Oct3/4 (E) in cardiac organoids at different timepoints during differentiation, normalized to GAPDH. (F) Western blot showing protein expression of MEF2C, GATA-6, α-actinin, and Nkx2.5 relative to GAPDH in cardiac organoids at different timepoints. Graphs show average concentrations + SD. Samples were compared using an unpaired t-test with Welch’s correction. *p<0.05; **p<0.01; ***p<0.001; ns, nonsignificant.
-
Figure 1—source data 1
Cardiac organoids derived from H9 embryonic stem cells exhibit characteristic morphological changes and express markers of differentiation.
- https://cdn.elifesciences.org/articles/83532/elife-83532-fig1-data1-v2.zip
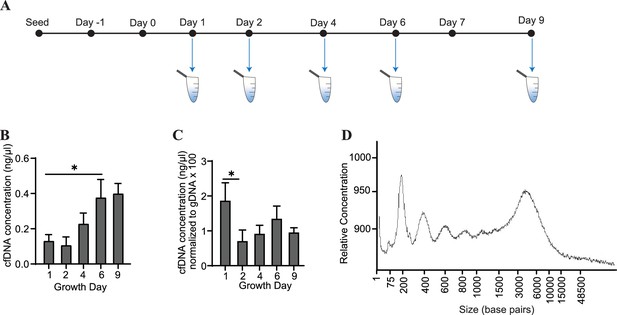
Cardiac organoids release cfDNA during their development.
(A) Schematic illustrating timepoints of cfDNA collection in cardiac organoids. (B) Shown are cfDNA concentrations taken from n = 3 biological replicates during cardiac organoid differentiation. Data points represent the average of 2–3 technical replicates per biological replicate. (C) Cardiac organoid-derived cfDNA concentrations from panel (B), normalized to gDNA concentrations (multiplied by 100 to facilitate axis readability). (D) Electropherogram showing fragment sizes of cfDNA derived from mature cardiac organoids on day 9. Graphs show average + SD. Samples were compared using an unpaired t-test with Welch’s correction. *p<0.05; **p<0.01; ***p<0.001; ns, nonsignificant.
-
Figure 2—source data 1
Electropherogram showing fragment lengths of cfDNA derived from cardiac organoids at different time points during development.
- https://cdn.elifesciences.org/articles/83532/elife-83532-fig2-data1-v2.zip
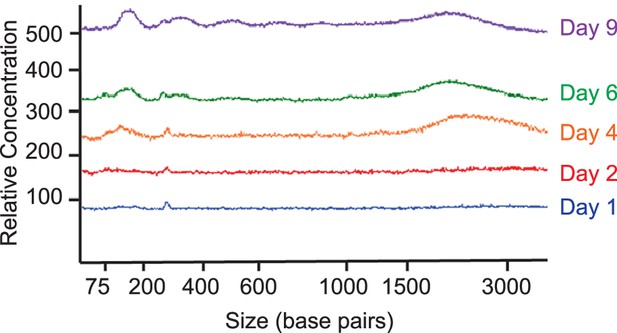
Overlay of representative electropherograms showing fragment sizes of cfDNA derived from cardiac organoids on different growth days.
-
Figure 2—figure supplement 1—source data 1
Electropherogram showing fragment lengths of cfDNA derived from cardiac organoids at different time points during development.
- https://cdn.elifesciences.org/articles/83532/elife-83532-fig2-figsupp1-data1-v2.zip
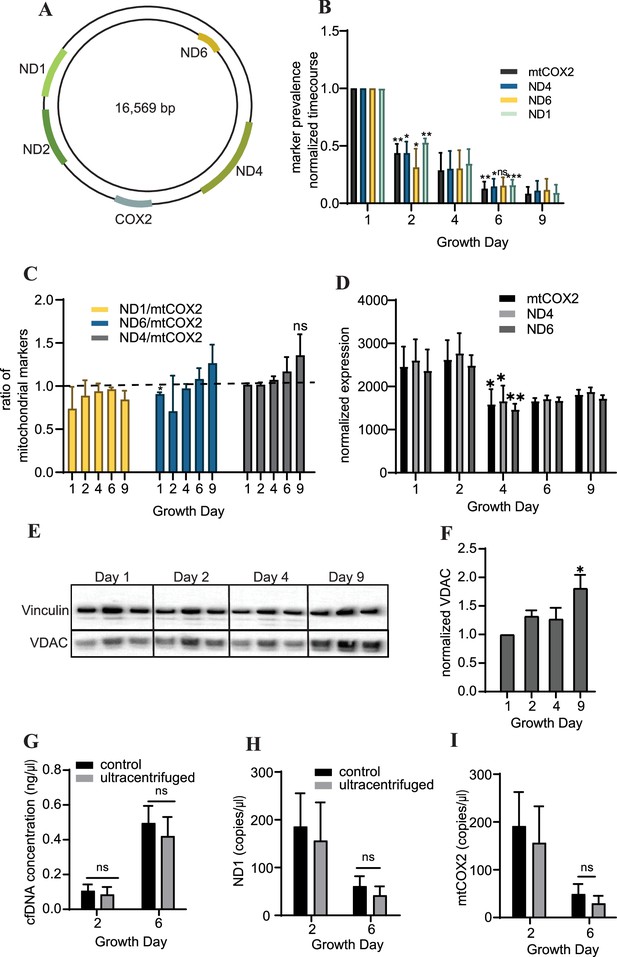
Cardiac organoids exhibit a time-dependent decrease in cell-free mitochondrial DNA abundance during growth.
(A) Schematic showing the location of markers examined on the mitochondrial genome, adapted from Figure 1 of Uhler and Falkenberg, 2015. (B) Relative abundance of mitochondrial markers ND1, ND4, ND6, and mtCOX2 in cardiac organoid-derived cfDNA, obtained using ddPCR. Shown are the average copies/μl of n = 3 biological replicates, each normalized to the values on day 1, to correct for batch variation. Samples on day 2 were compared to day 1 using a one-sample t-test comparing to a hypothetical value of 1.0. Day 6 samples were compared to day 2 values using an unpaired t-test with Welch’s correction. *p<0.05; **p<0.01; ***p<0.001; ns, nonsignificant. (C) Ratio of the values obtained in panel (B) for each marker to mtCOX2. Significant deviations from a ratio of 1.0 indicating unequal marker abundance were determined using a one-sample t-test comparing to hypothetical value of 1.0. Shown are the averages of n = 3 biological replicates, + SD. (D) Expression of mitochondrial DNA markers at the tissue level in cardiac organoids during differentiation obtained using ddPCR, normalized to β-actin. (E) Western blot showing protein expression of the mitochondrial protein VDAC/Porin and loading control vinculin in n = 3 biological replicates of cardiac organoids taken at different timepoints during differentiation. (Vinculin loading control is the same for Figure 4D.) (F) Quantification of VDAC protein expression from the western blot in panel (E), normalized to vinculin. Samples were further normalized to day 1 and compared using a one-sample t-test with hypothetical value of 1.0 to test for significant deviation from day 1. (G) Concentration of cfDNA extracted from media conditioned by cardiac organoids on growth days 2 or 6 after ultracentrifugation (100,000 × g for 1 hr) compared to control (no ultracentrifugation). Abundance of mitochondrial markers ND1 (H) and mtCOX2 (I) in cfDNA derived from ultracentrifuged media conditioned by cardiac organoids during development, obtained using ddPCR. Graphs show average concentrations + SD. Samples were compared using an unpaired t-test with Welch’s correction. *p<0.05; **p<0.01; ***p<0.001; ns, nonsignificant.
-
Figure 3—source data 1
Cardiac organoids exhibit a time-dependent decrease in cell-free mitochondrial DNA abundance during growth.
- https://cdn.elifesciences.org/articles/83532/elife-83532-fig3-data1-v2.zip
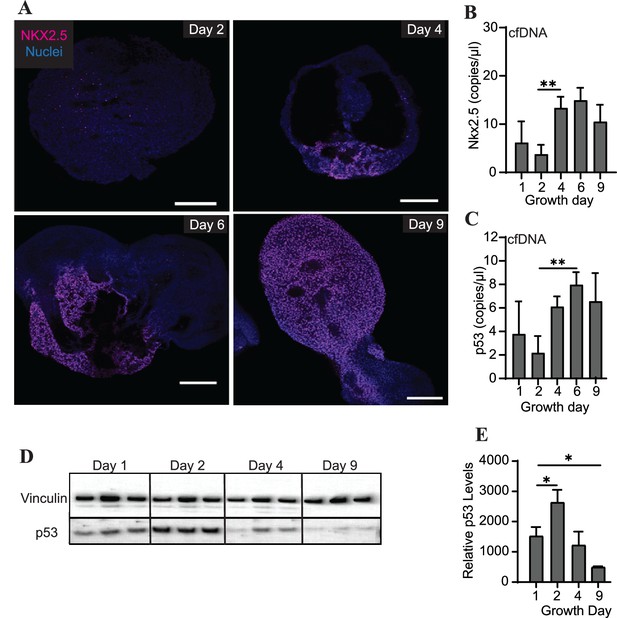
Abundance of specific gDNA sequences is reflective of tissue-level changes in marker expression.
(A) Immunofluorescence staining of Nkx2.5 in cardiac organoids during differentiation. Scale bars represent 200 μm. (B) Nkx2.5 copies/μl in 1 ng of cardiac organoid-derived cfDNA at different time points during differentiation obtained using ddPCR, taken from n = 3 biological replicates. (C) p53 copies/μl in 1 ng of cardiac organoid-derived cfDNA at different time points during differentiation obtained using ddPCR, taken from n = 3 biological replicates. (D) Western blot showing p53 protein levels in cardiac organoids during differentiation. (Vinculin loading control is the same for Figure 3E.) (E) Quantification of the western blot in panel (D), normalized to Vinculin (n = 3 biological replicates). Graphs show average concentrations + SD. Samples were compared using an unpaired t-test with Welch’s correction. *p<0.05; **p<0.01; ***p<0.001; ns, nonsignificant.
-
Figure 4—source data 1
Copy number of Sox17 or Oct3/4 in cfDNA derived from cardiac organoids on different growth days during development.
- https://cdn.elifesciences.org/articles/83532/elife-83532-fig4-data1-v2.zip
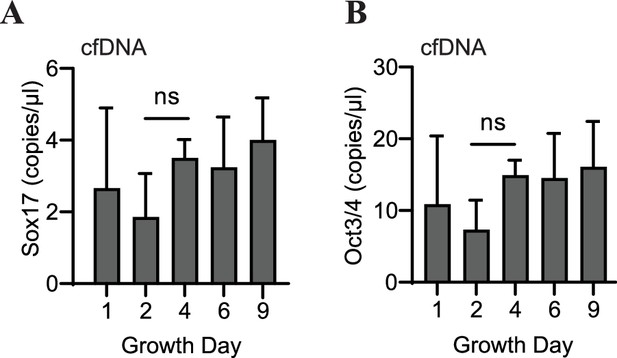
Abundance of specific gDNA sequences is reflective of tissue-level changes in marker expression.
Sox17 (A) or Oct3/4 (B) copies/μl in 1 ng of cardiac organoid-derived cfDNA at different time points during differentiation obtained using ddPCR, taken from n = 3 biological replicates. Graphs show average concentrations + SD. Samples were compared using an unpaired t-test with Welch’s correction. *p<0.05; **p<0.01; ***p<0.001; ns, nonsignificant.
-
Figure 4—figure supplement 1—source data 1
Copy number of Sox17 or Oct3/4 in cfDNA derived from cardiac organoids on different growth days during development.
- https://cdn.elifesciences.org/articles/83532/elife-83532-fig4-figsupp1-data1-v2.zip
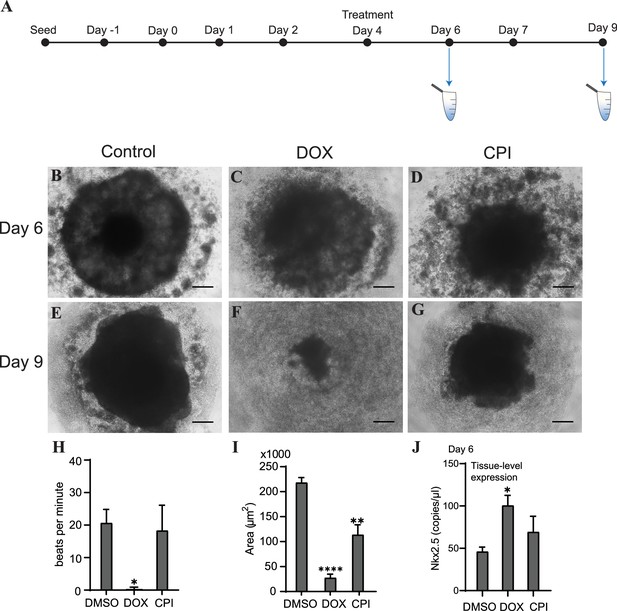
Doxorubicin (DOX) causes severe cardiac organoid malformation in comparison to CPI.
(A) Schematic showing drug treatment and collection times of cfDNA during cardiac organoid growth. Representative images of tissues on day 6: control (B), DOX-treated (C), and CPI-treated (D). Representative images of tissues on day 9: control (E), DOX-treated (F), and CPI-treated (G). Scale bars represent 100 µm. (H) Beats per minute measured on growth day 9 for tissues treated with either DOX or CPI compared to DMSO-treated control. Shown are the average counts for 3–5 tissues each across three independent replicates. (I) Average tissue size of mature cardiac organoids treated with DOX or CPI measured on growth day 9. Shown are the average counts across three independent replicates (3–5 organoids per replicate). (J) ddPCR showing tissue-level expression of Nkx2.5 normalized to TBP on growth day 6 in cardiac organoids treated with DOX or CPI. Graphs show average concentrations + SD. Samples were compared using an unpaired t-test with Welch’s correction. *p<0.05; **p<0.01; ***p<0.001; ns, nonsignificant. (Tissue morphology in response to treatment with DOX later in organoid growth is shown in Figure 5—figure supplement 1.)
-
Figure 5—source data 1
Doxorubicin causes severe cardiac organoid malformation in comparison to CPI-203.
- https://cdn.elifesciences.org/articles/83532/elife-83532-fig5-data1-v2.zip
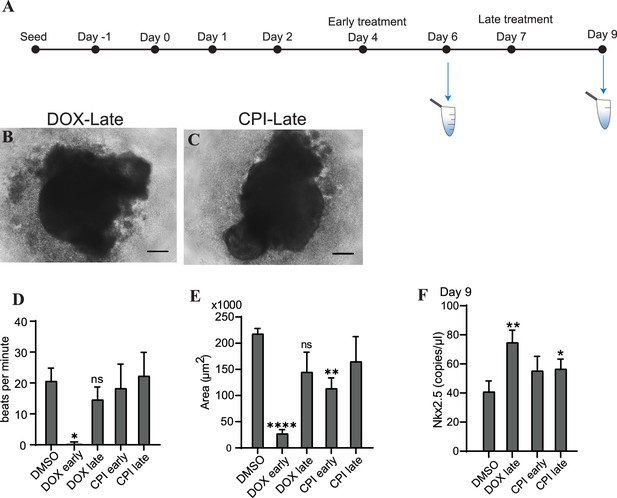
Toxicity of DOX and CPI-203 in cardiac organoids is impacted by time of exposure.
(A) Schematic showing treatment and cfDNA collection times during the growth of cardiac organoids. Representative images of mature cardiac organoids on day 9 treated with doxorubicin (DOX) (B) or CPI (C) on day 7 (late treatment). Scale bars represent 100 µm. (D) Beats per minute measured on growth day 9 for tissues treated with either DOX or CPI either early (day 4) or late (day 7) in development. Shown are the average counts across three independent replicates (3–5 organoids per replicate). (E) Average tissue size of mature cardiac organoids on growth day 9 treated with DOX or CPI either early (day 4) or late (day 7) during development. Shown are the average measurements across three independent replicates (3–5 organoids per replicate). (F) Abundance of Nkx2.5 normalized to TBP at the tissue-level in mature cardiac organoids on growth day 9 treated with DOX or CPI on either days 4 (early treatment) or day 7 (late treatment) shown via ddPCR. (The samples treated with DOX on day 4 are not shown because they were severely toxic by day 9 and there was not enough tissue for RNA collection.) Graphs show average concentrations + SD. Samples were compared using an unpaired t-test with Welch’s correction. *p<0.05; **p<0.01; ***p<0.001; ns, nonsignificant.
-
Figure 5—figure supplement 1—source data 1
Toxicity of DOX and CPI-203 in cardiac organoids is impacted by time of exposure.
- https://cdn.elifesciences.org/articles/83532/elife-83532-fig5-figsupp1-data1-v2.zip
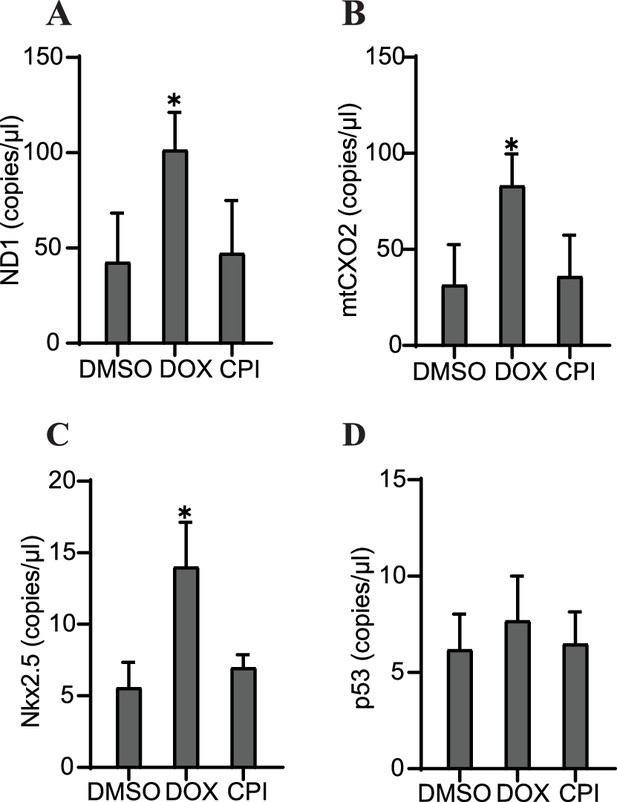
Specific cfDNA sequences may be predictive of toxicity in cardiac organoids.
Abundance of ND1 (A), mtCOX2 (B), Nkx2.5 (C), or p53 (D) in 0.5 ng cfDNA collected from cardiac organoids on growth day 6 treated with doxorubicin (DOX) or CPI, obtained using ddPCR. Graphs show average concentrations + SD. Samples were compared using an unpaired t-test with Welch’s correction. *p<0.05; **p<0.01; ***p<0.001; ns, nonsignificant. (Concentration of cfDNA and additional analysis of cfDNA sequences upon treatment with DOX later in growth are shown in Figure 6—figure supplement 1.)
-
Figure 6—source data 1
Specific sequences of cfDNA may be predictive of toxicity in cardiac organoids.
- https://cdn.elifesciences.org/articles/83532/elife-83532-fig6-data1-v2.zip
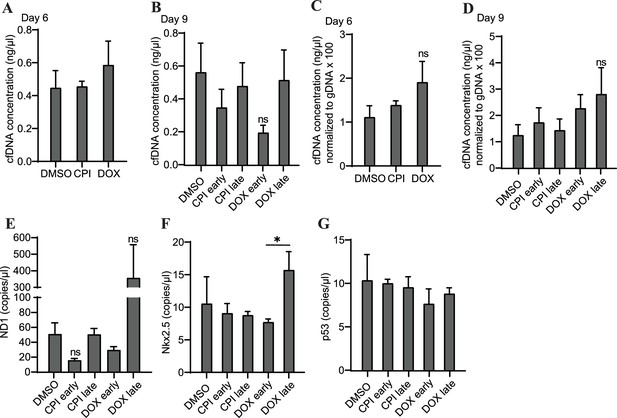
Concentration of cfDNA in response to early or late exposure to DOX or CPI-203.
(A) cfDNA concentrations from cardiac organoids on day 6 treated with either doxorubicin (DOX) or CPI. (B) cfDNA concentrations from cardiac organoids on day 9 treated with either DOX or CPI on growth days 4 (early treatment) or 7 (late treatment). (C) The cfDNA concentrations in panel (A) normalized to gDNA (multiplied by 100 to facilitate axis readability). (D) The cfDNA concentrations in panel (B) normalized to gDNA (multiplied by 100 to facilitate axis readability). Abundance of ND1 (E), Nkx2.5 (F), and p53 (G) in cfDNA collected from mature cardiac organoids on day 9, treated with DOX or CPI on either day 4 (early treatment) or day 7 (late treatment), shown via ddPCR. Graphs show average concentrations + SD. Samples were compared using an unpaired t-test with Welch’s correction. *p<0.05; **p<0.01; ***p<0.001; ns, nonsignificant.
-
Figure 6—figure supplement 1—source data 1
Concentration of cfDNA in response to early or late exposure to DOX or CPI-203.
- https://cdn.elifesciences.org/articles/83532/elife-83532-fig6-figsupp1-data1-v2.zip





