The deep-rooted origin of disulfide-rich spider venom toxins
Figures
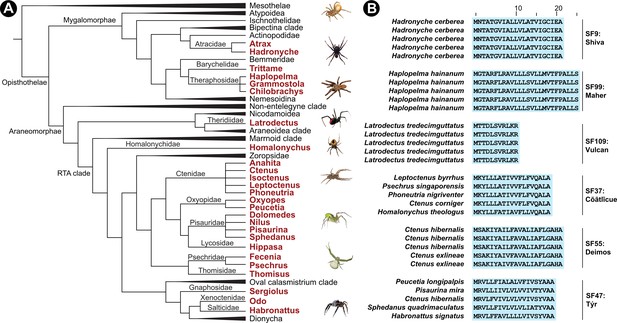
Schematic representation of Araneae phylogeny and their venom superfamilies.
Panel A here shows a cladogram of Araneae with lineages under investigation indicated in red. In panel B, representative signal peptide alignments of toxin superfamilies are shown with sequence conservation of >90% highlighted in blue.
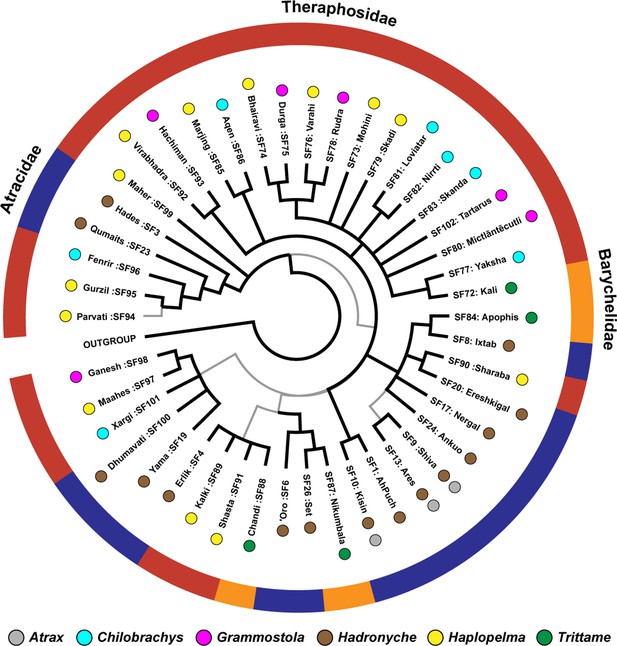
The Bayesian phylogeny of mygalomorph spider venom toxin superfamilies.
This figure represents the Bayesian phylogeny of Mygalomorphae spider toxin superfamilies, where branches with high (BPP >0.95) and low (BPP <0.95) node supports are shown in thick black and thin grey lines, respectively. Coloured spheres alongside tree tips represent the spider genera, while the coloured outer circle indicates the spider family in which the respective toxin superfamily has been identified (Atracidae [red], Barychelidae [orange], and Theraphosidae [blue]).
-
Figure 2—source data 1
This zip archive contains sequence alignment used to perform phylogenetic analyses and the Bayesian phylogeny of Mygalomorphae DRP toxin superfamilies.
- https://cdn.elifesciences.org/articles/83761/elife-83761-fig2-data1-v2.zip
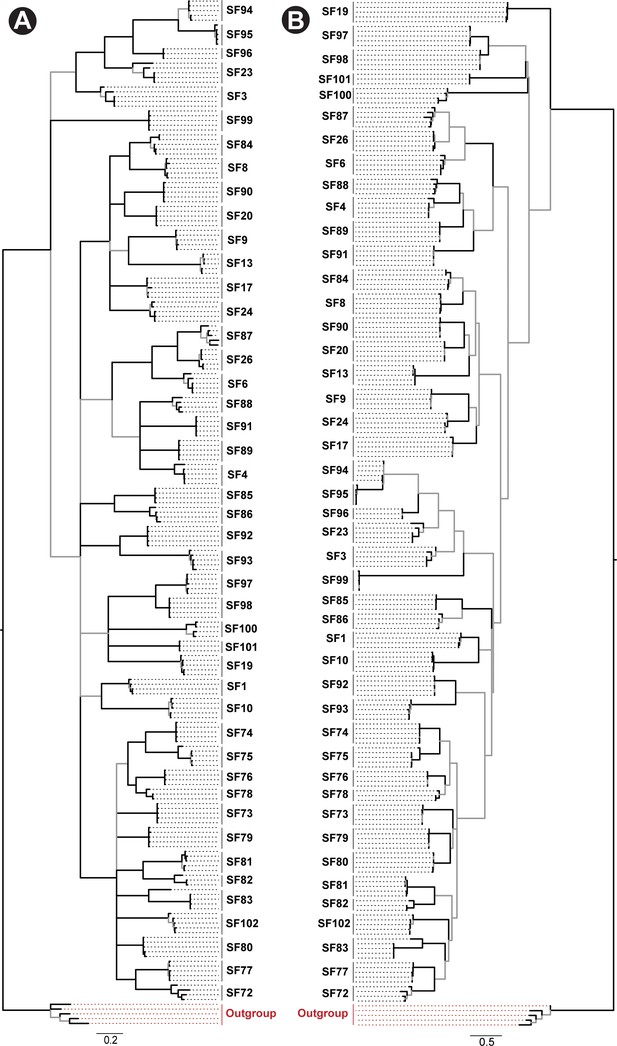
Phylogeny of Mygalomorphae spider toxin superfamilies.
Phylogenetic relationships of Mygalomorphae spider toxin superfamilies, assessed using Bayesian (BI; panel A) and maximum likelihood (ML; panel B) inferences, are shown in this figure. Node supports were estimated using Bayesian posterior probability (BPP) for the BI tree and bootstrapping replication (BS) for the ML tree. Branches with BPP lower than 0.95 in BI tree and BS lower than 90 in the ML tree are shown in grey. Cysteine-rich non-toxin outgroup sequences are coloured red.
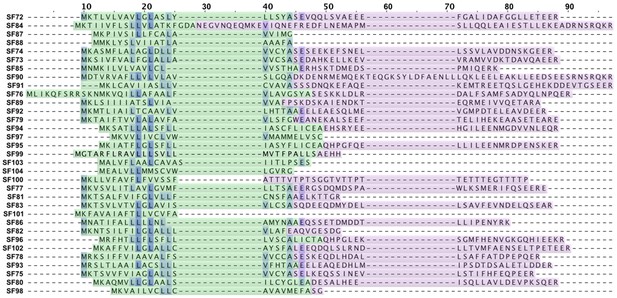
Signal peptide and propeptide alignment of novel mygalomorph superfamilies.
This figure shows the alignment of signal peptide and propeptide sequences from novel mygalomorph spider toxin superfamilies identified in this study. The signal peptide region is highlighted in green, while the propeptide region is represented in purple colour. Conserved sites are shaded in blue.
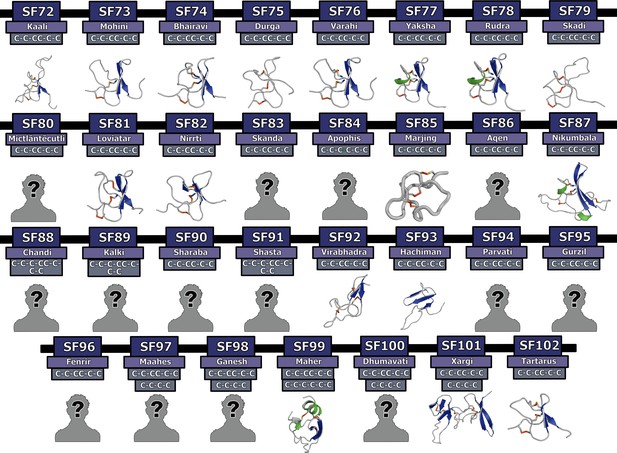
Homology models of novel Mygalomorphae toxin superfamilies.
This figure depicts the 3D homology models of disulfide-rich peptide Mygalomorphae toxin superfamilies. Here, helices are shown in green, β-strands in blue and disulfide bonds in orange. Cysteine arrangements in scaffolds are also provided above the model. Toxin SFs that lack structural data are indicated with a ‘?’ symbol.
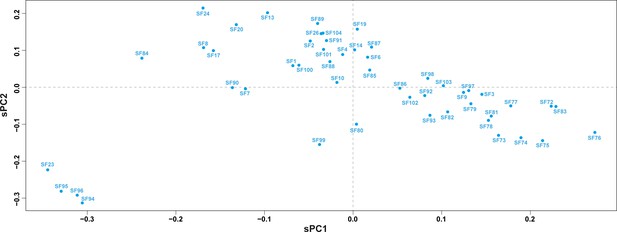
Principal component analyses for Mygalomorphae toxin superfamilies.
A scatter plot of scaled principal components, sPC1 and sPC2, for the signal peptide sequences of novel mygalomorph toxin superfamilies identified in this study is shown here. Signal peptide sequences belonging to a superfamily are represented as a single dot.
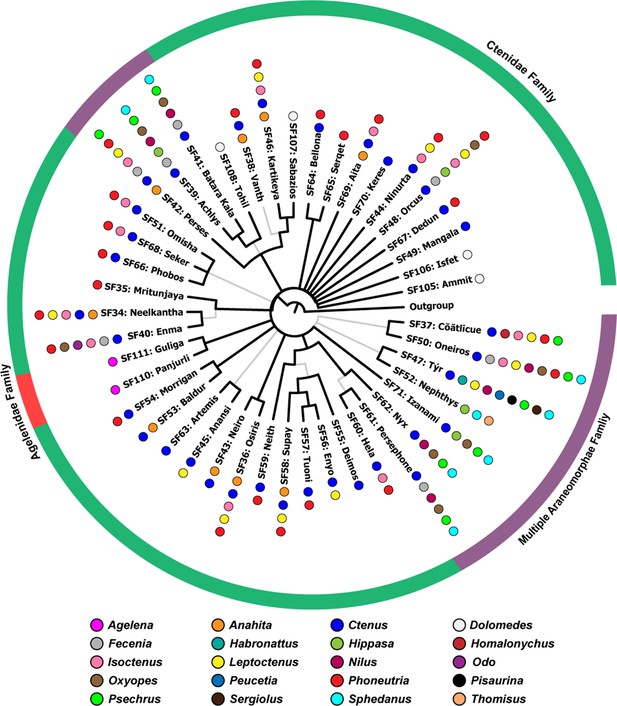
The Bayesian phylogeny of araneomorph spider venom toxin superfamilies.
This figure represents the Bayesian phylogeny of Araneomorphae spider toxin superfamilies, where branches with high (BPP >0.95) and low (BPP <0.95) node supports are shown in thick black and thin grey lines, respectively. Coloured spheres, alongside tree tips, represent the spider genera, while the coloured outer circle indicates the spider family (Agelenidae [red], Ctenidae [green], multiple araneomorph families [purple]: Ctenidae, Gnaphosidae, Homalonychidae, Lycosidae, Oxyopidae, Pisauridae, Psechridae, Salticidae, Thomisidae, Xenoctenidae) in which the respective toxin superfamily has been identified.
-
Figure 3—source data 1
This zip archive contains sequence alignment used to perform phylogenetic analyses and the Bayesian phylogeny of Araneomorphae DRP toxin superfamilies.
- https://cdn.elifesciences.org/articles/83761/elife-83761-fig3-data1-v2.zip
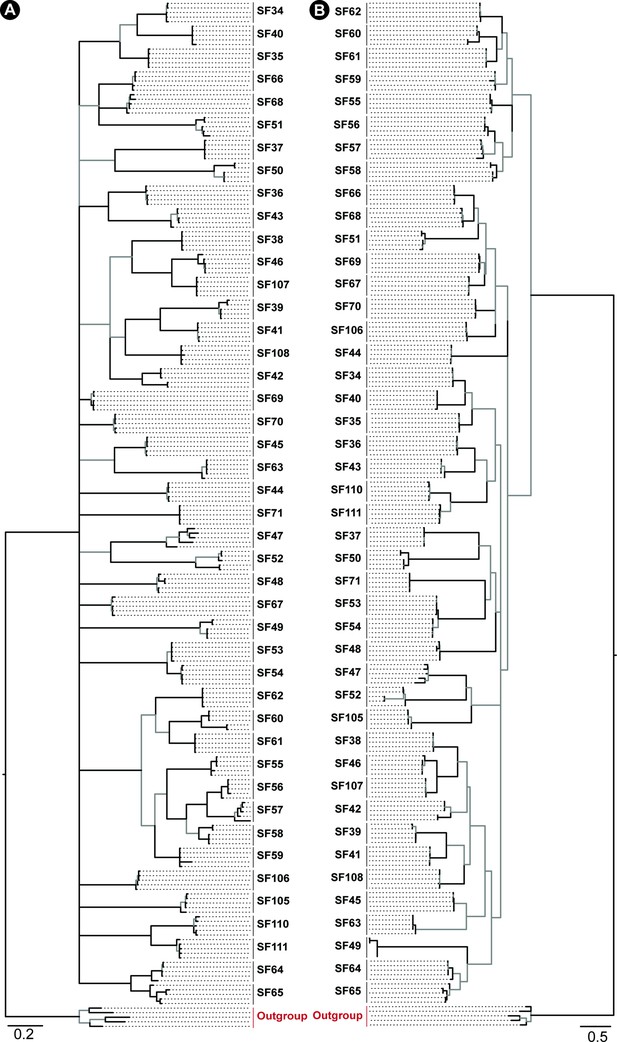

Phylogeny of Araneomorphae spider toxin superfamilies.
Phylogenetic relationships of Araneomorphae spider toxin superfamilies, assessed using Bayesian (BI; panel A) and maximum likelihood (ML; panel B) inferences, are shown in this figure. Node supports were estimated using Bayesian posterior probability (BPP) for the BI tree and bootstrapping replication (BS) for the ML tree. Branches with BPP lower than 0.95 in BI tree and BS lower than 90 in the ML tree are shown in grey. Cysteine-rich non-toxin outgroup sequences are coloured red.
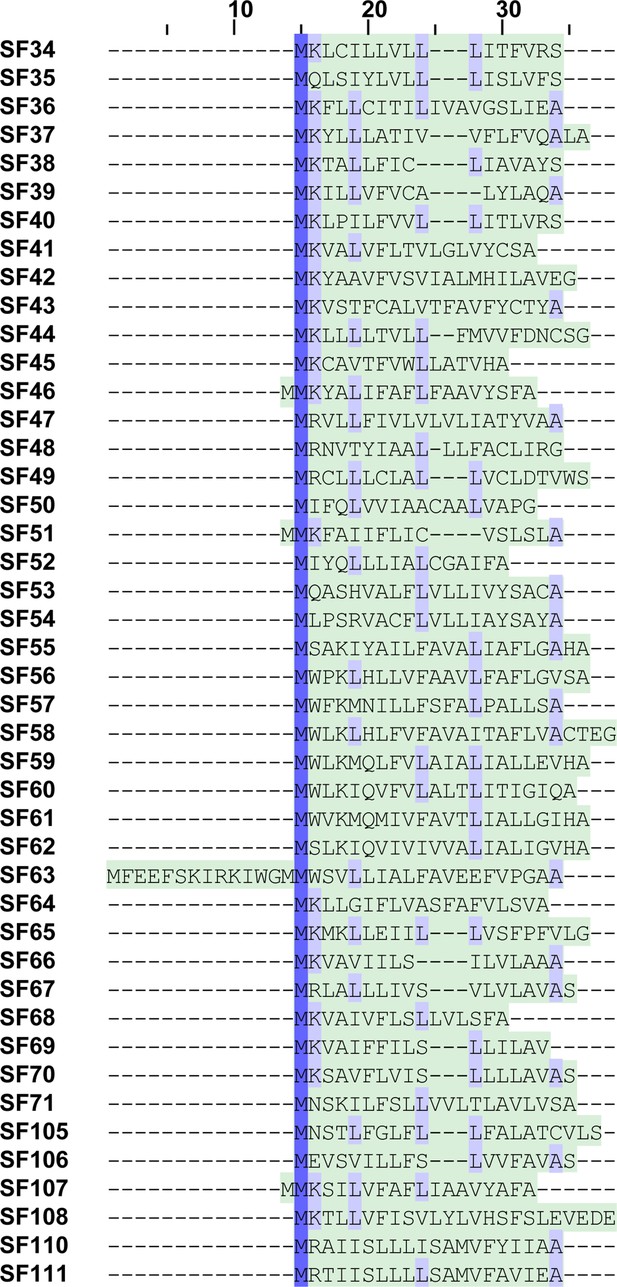

Signal peptide alignment of araneomorph superfamilies.
This figure shows the alignment of signal peptide sequences from novel araneomorph toxin superfamilies identified in this study. The signal peptide region is highlighted in green, while the conserved amino acid positions are shaded in blue. It should be noted that the propeptide region boundary could not be identified for all Araneomorphae toxin superfamilies.
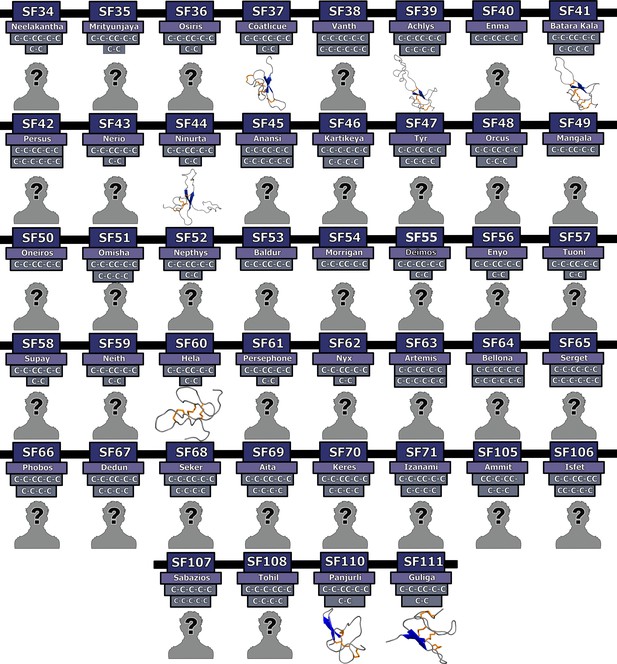
Homology models of novel Araneomorphae toxin superfamilies.
3D homology models of disulfide-rich peptide Araneomorphae toxin superfamilies are depicted in this figure. Here, helices are shown in green, β-strands in blue and disulfide bonds in orange. Cysteine arrangements in scaffolds are also provided above the model. Toxin SFs that lack structural data are indicated with a ‘?’ symbol.
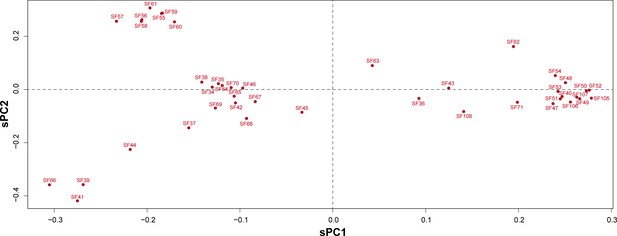
Principal component analyses for Araneomorphae toxin superfamily.
Scaled principal components, sPC1 and sPC2, for signal peptide sequences of novel araneomorph toxin superfamilies identified in this study are shown here in the form of a scatter plot. Signal peptide sequences belonging to a superfamily are represented as a single dot.
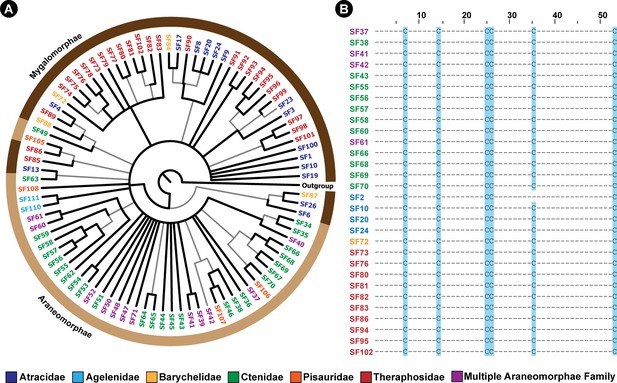
The Bayesian phylogeny and cysteine framework representation of spider venom DRPs.
This figure depicts the Bayesian phylogeny and alignment of representative sequences of Araneae DRP toxin superfamilies, where branches with high (BPP >0.95) and low (BPP <0.95) node supports are shown in thick black and thin grey lines, respectively. The coloured outer circle in panel A indicates the infraorder of spiders (Mygalomorphae and Araneomorphae shown in dark and light brown, respectively) in which the respective DRP superfamily was identified. In panel B, cysteine framework conserved across toxin SFs is highlighted in blue.
-
Figure 4—source data 1
This zip archive contains sequence alignment used to perform phylogenetic analyses and the Bayesian phylogeny of Araneae DRP toxin superfamilies.
A list of accession IDs for sequence data utilised in this study has also been provided.
- https://cdn.elifesciences.org/articles/83761/elife-83761-fig4-data1-v2.zip
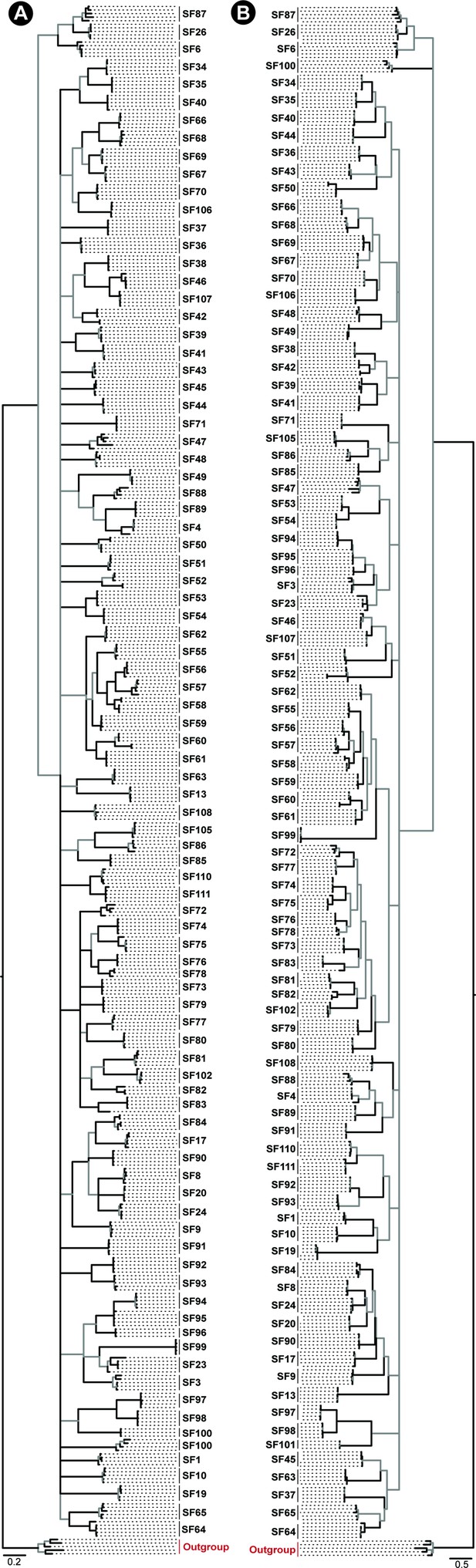

Phylogeny of Araneae spider toxin superfamilies.
Phylogenetic relationships of Araneae toxin superfamilies built using Bayesian (BI; panel A) and maximum likelihood (ML; panel B) inferences are shown in this figure. Node supports were estimated using Bayesian posterior probability (BPP) for the BI tree and bootstrapping replication (BS) for the ML tree. Branches with BPP lower than 0.95 in BI tree and BS lower than 90 in the ML tree are shown in grey. Cysteine-rich non-toxin outgroup sequences are coloured red.
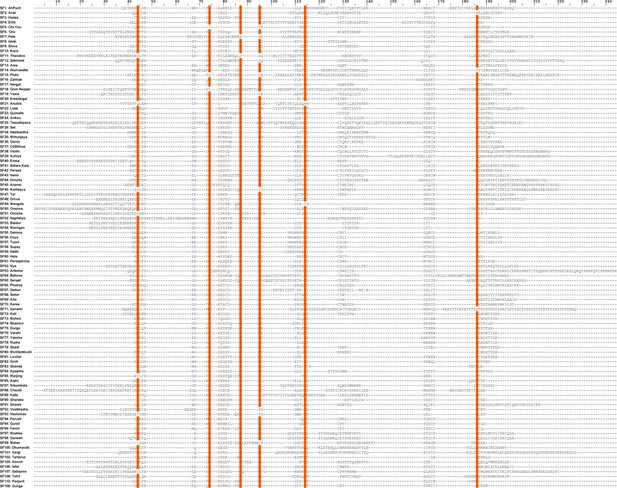
Mature peptide alignment of mygalomorph and araneomorph DRP superfamilies.
An alignment of mature peptide sequences from mygalomorph and araneomorph spider toxin superfamilies is shown here. Conserved amino acid positions (sequence identity ≥90%) are shaded orange.
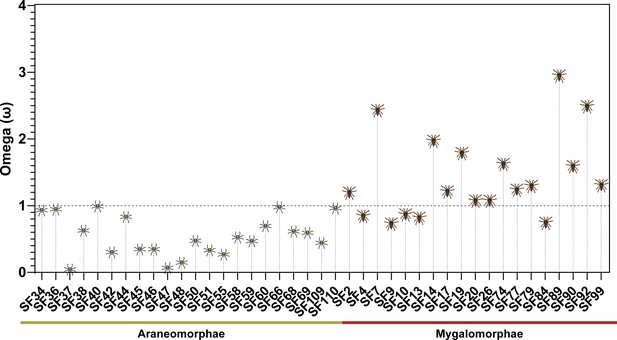
Molecular evolution of spider toxin superfamilies.
This figure shows the distribution of ω values (Y-axis) for araneomorph and mygalomorph spider venom toxin superfamilies (X-axis). The horizontal dotted black line represents neutral evolution (ω=1), with ω values above and below it indicating positive (ω>1) and negative (ω<1) selection, respectively.
-
Figure 5—source data 1
Molecular evolution of toxin superfamilies.
(a): Positively selected sites detected by the Bayes Empirical Bayes approach implemented in M8; (b): Fast Unconstrained Bayesian AppRoximation (c): Sites detected as experiencing episodic diversifying selection (0.05 significance) by the Mixed Effects Model Evolution (MEME). Sites detected at 0.99 and 0.95 significance are indicated in the parenthesis; (d): number of sites under pervasive diversifying selection at the posterior probability ≥0.9 (FUBAR); (e): Number of sites under pervasive purifying selection at the posterior probability ≥0.9 (FUBAR); ω: mean dN/dS. Biochemical properties evaluated: Equilibrium Const. – ionisation, COOH (pK); Hydropathy (h); Long-range n.b. energy (El); Polarity (p); Total n.b. energy (Et). Structural properties: α−helical tendencies (Pα); β-structure tendencies (Pβ); Average # surrounding residues (Ns); Bulkiness (Bl); Chromatographic index (RF); Coil tendencies (Pc); Compressibility (K°); Helical contact energy (Ca); Mean r.m.s. fluctuation displacement. (F); Molecular volume (Mv); Molecular weight (Mw); Partial specific volume (V°); Polar requirement (Pr); Power to be – C-term. α-helix (αc); Power to be – middle, α-helix (αm); Power to be – N-term., α-helix (αn); Refractive index (µ); Solvent accessible reduct. ratio (Ra); Thermodynamics transfer hydrophobicity (Ht).
- https://cdn.elifesciences.org/articles/83761/elife-83761-fig5-data1-v2.zip
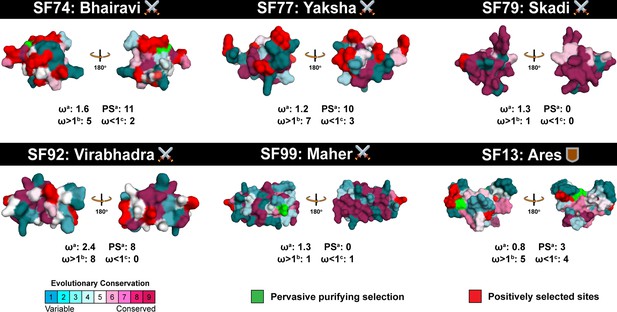
Deployment strategies dictate the evolution of spider venom.
This figure highlights the distinct regimes of evolutionary selection pressures acting on defensive and offensive spider venom toxin superfamilies. Positively selected sites detected by PAML (M8) and FUBAR are highlighted in red, while sites under the effect of pervasive purifying selection (FUBAR) are shown in green. A colour code indicating strength of selection is also provided. Here, ω: ratio of non-synonymous to synonymous substitutions; (a): ω and positively selected sites (Bayes Empirical Bayes) detected by model 8 of PAML. (b): sites experiencing pervasive influence of positive selection identified by FUBAR (ω>1); and c: sites experiencing pervasive influence of negative selection identified by FUBAR (ω<1).
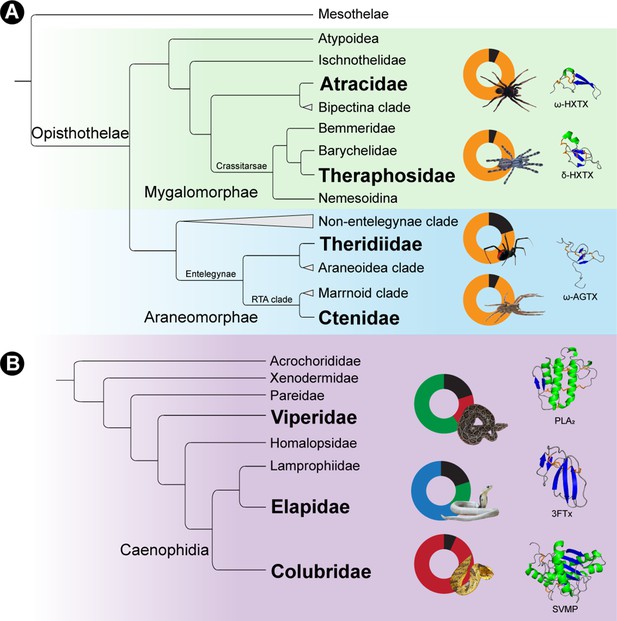
Distinct toxin scaffold recruitment strategies in spiders and snakes.
This figure depicts distinct toxin scaffold recruitment strategies in (A) spiders and (B) advanced snakes. The Araneae phylogeny highlights the domination of disulfide-rich peptide toxins in spiders [Atracidae: Atrax sp.; Theraphosidae: Poecilotheria formosa; Theridiidae: Latrodectus mactans; Ctenidae: Phoneutria nigriventer: e.g., Palagi et al., 2013; Oldrati et al., 2017; Diniz et al., 2018], whereas venoms of advanced snakes are constituted by diverse phylogenetically unrelated toxin superfamilies (Viperidae: Daboia russelii, Elapidae: Naja naja, Colubridae: Spilotes sulphureus: e.g., Senji Laxme et al., 2021a; Senji Laxme et al., 2021b; Modahl et al., 2018). Doughnut charts, portraying the major molecular scaffolds in venom are also shown disulfide-rich peptides (yellow), snake venom metalloproteinases (SVMP, red), phospholipase A2 (PLA2, green), three-finger toxins (3 FTx, blue) and other minor components (black). Structures of the major scaffolds are also shown, with helices coloured in green, β-strands in blue and disulfide bonds in orange.
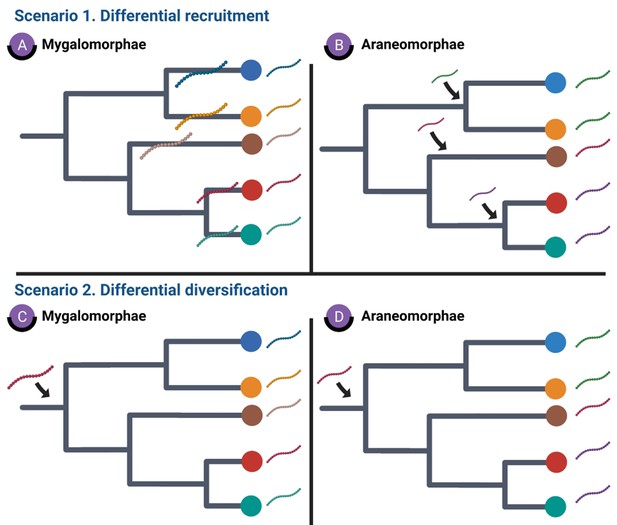
Hypotheses explaining the stark differences in recruitment and diversification of toxin SFs in Araneomorphae and Mygalomorphae.
This figure depicts various hypotheses that explain distinct toxin SF recruitment and diversification in spiders. Scenario 1 depicts genus- or family-specific recruitment of spider toxin SFs in Mygalomorphae and Araneomorphae, respectively, while scenario 2 highlights the implications of differential rates of diversification.
Additional files
-
Supplementary file 1
List of spider toxin superfamilies.
The table lists spider toxin superfamilies described till date with names conferred upon and their respective mythological references. A list of genera from which the superfamily was identified has also been provided.
- https://cdn.elifesciences.org/articles/83761/elife-83761-supp1-v2.xlsx
-
MDAR checklist
- https://cdn.elifesciences.org/articles/83761/elife-83761-mdarchecklist1-v2.docx





