Calaxin stabilizes the docking of outer arm dyneins onto ciliary doublet microtubule in vertebrates
Figures
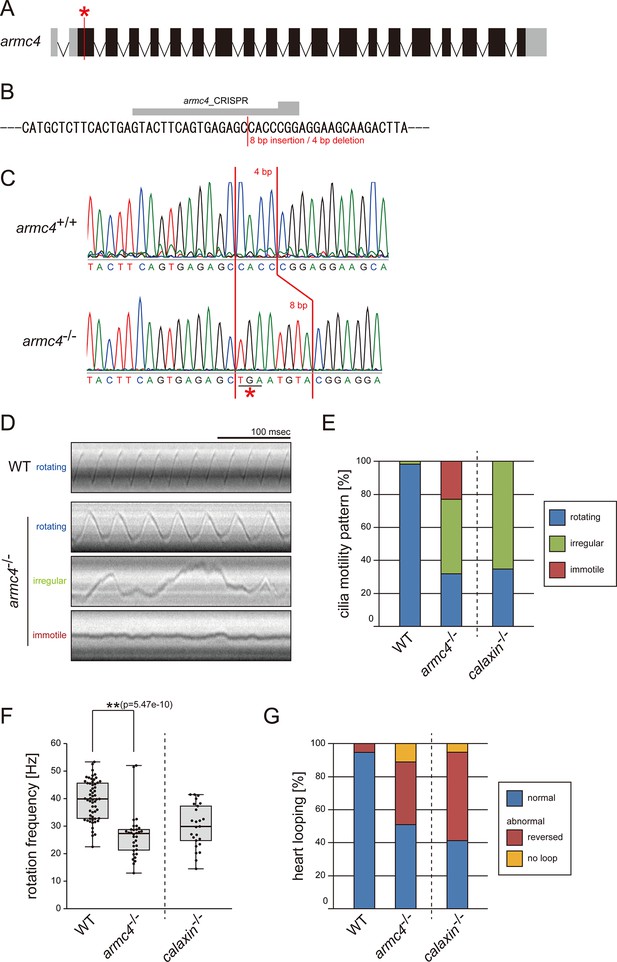
Mutation of armc4 causes abnormal motility of Kupffer’s vesicle cilia.
(A) Genomic organization of zebrafish armc4 gene. Black boxes: exons. Gray boxes: untranslated regions. Red asterisk: the genome-editing target site. (B) CRISPR/Cas9 target sequence. (C) Sanger sequencing of armc4+/+ and armc4−/− fish around the genome-editing target site. The 8 bp-insertion in armc4−/− includes a stop codon (red asterisk). (D) Typical kymographs of Kupffer’s vesicle cilia in WT and armc4−/− embryos. Kymograph patterns were categorized into three classes: rotating (blue), irregular (green), and immotile (red). Scale bar: 100 ms. (E) Ratios of each motility class. Number of cilia: 58 (WT) and 100 (armc4−/−). (F) Rotational frequencies of Kupffer’s vesicle cilia. Number of cilia: 58 (WT) and 32 (armc4−/−). Boxes correspond to the first and third quartiles, lines inside the boxes indicate the medians, and whiskers extend to the full range of the data. p-Value was calculated with Welch’s t-test. (G) Directions of heart looping. Number of embryos: 110 (WT) and 63 (armc4−/−). For comparison, calaxin-/- data from Sasaki et al., 2019 are displayed in (E, F, and G).
-
Figure 1—source data 1
Numerical data of Figure 1E.
Motility patterns of Kupffer’s vesicle cilia.
- https://cdn.elifesciences.org/articles/84860/elife-84860-fig1-data1-v2.zip
-
Figure 1—source data 2
Numerical data of Figure 1F.
Rotation frequencies of Kupffer’s vesicle cilia.
- https://cdn.elifesciences.org/articles/84860/elife-84860-fig1-data2-v2.zip
-
Figure 1—source data 3
Numerical data of Figure 1G.
Heart looping of embryos.
- https://cdn.elifesciences.org/articles/84860/elife-84860-fig1-data3-v2.zip
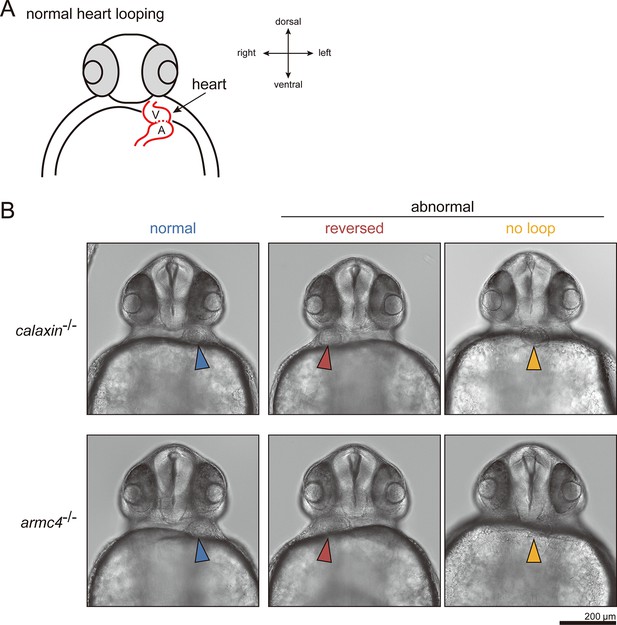
Heart looping of mutant embryos at 36 hpf.
(A) Diagram represents normal heart looping of the 36 hpf embryo. V, ventricle; A, atrium. (B) Typical images of the heart looping of mutant embryos. Arrowheads indicate the looping of the hearts. Scale bar: 200 μm.
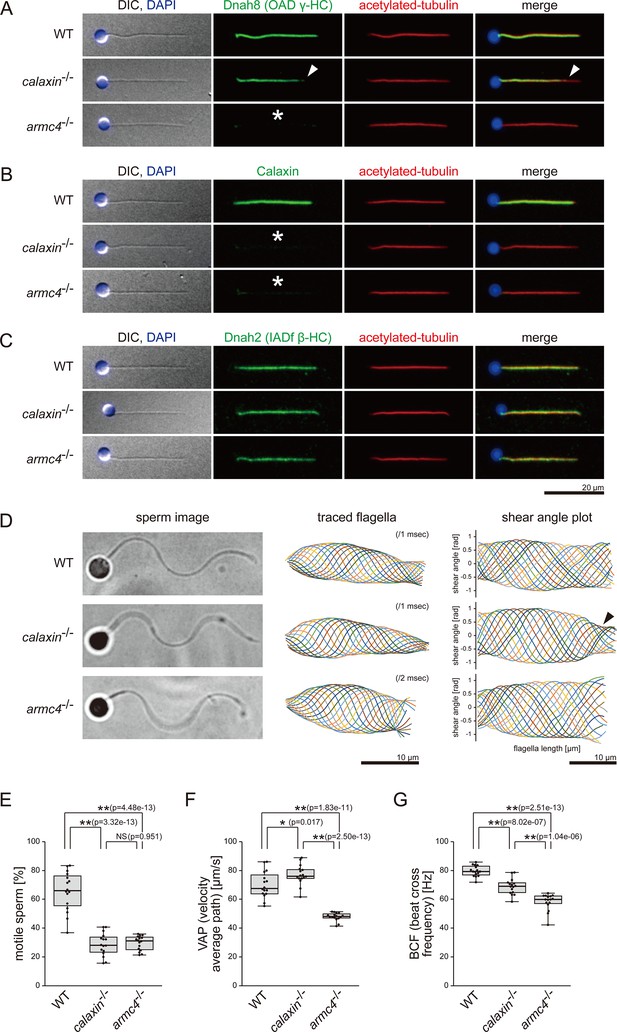
Mutations of calaxin and armc4 cause loss of OAD in sperm flagella.
(A–C) Immunofluorescence microscopy of zebrafish spermatozoa. Scale bar: 20 μm. (A) Dnah8 was localized along the entire length of sperm flagella in WT. In calaxin-/-, Dnah8 was lost at the distal region of sperm flagella (white arrowheads). In arm4-/-, Dnah8 was lost (white asterisk). (B) Calaxin was localized along the entire length of sperm flagella in WT. In calaxin-/- and arm4-/-, Calaxin was lost (white asterisks). (C) Dnah2 was localized along the entire length of sperm flagella in WT, calaxin-/-, and arm4-/-. (D) Phase-contrast microscopy images of swimming spermatozoa (left column), traces of beating flagella (middle column), and shear angle plots of traced flagella (right column). Swimming spermatozoa were filmed using a high-speed camera at 1000 fps (frames per second). Shear angles were plotted against the distance from the flagellar base. In calaxin-/-, shear angle plots lost their slopes at the distal region of the flagella (black arrowhead). Scale bars: 10 μm. (E–G) Motilities of swimming spermatozoa were filmed using a high-speed camera at 200 fps and analyzed by CASA modified for zebrafish. For each zebrafish line, more than 600 spermatozoa were analyzed in total with 16 technical replicates. Spermatozoa with less than 20 μm/s velocities were considered immotile. (H) Ratio of motile spermatozoa. (I) Velocity of spermatozoa on averaged paths. The averaged paths were drawn by connecting the points of averaged sperm positions of contiguous 33 frames. (J) Frequencies at which sperm heads crossed their averaged paths. Boxes correspond to the first and third quartiles, lines inside the boxes indicate the medians, and whiskers extend to the full range of the data. p-Values were calculated with Tukey-Kramer test.
-
Figure 2—source data 1
Numerical data of Figure 2E.
Ratios of motile spermatozoa in CASA.
- https://cdn.elifesciences.org/articles/84860/elife-84860-fig2-data1-v2.zip
-
Figure 2—source data 2
Numerical data of Figure 2F.
Velocity of spermatozoa on averaged paths in CASA.
- https://cdn.elifesciences.org/articles/84860/elife-84860-fig2-data2-v2.zip
-
Figure 2—source data 3
Numerical data of Figure 2G.
Frequencies at which sperm heads crossed their averaged paths in CASA.
- https://cdn.elifesciences.org/articles/84860/elife-84860-fig2-data3-v2.zip
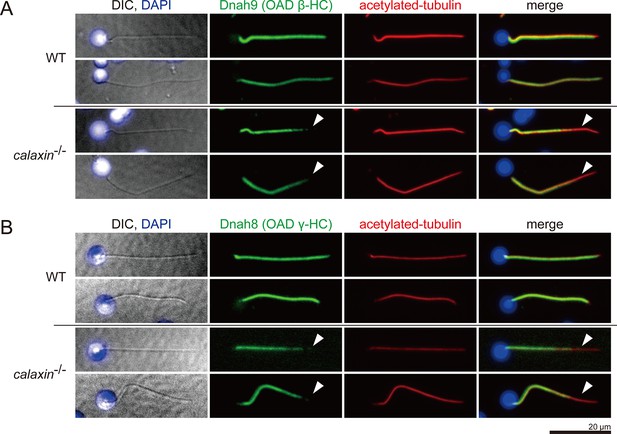
Immunofluorescence microscopy of OAD-HCs in zebrafish spermatozoa.
Two OAD HCs showed the same distribution patterns; (A) Dnah9, OAD β-HC and (B) Dnah8, OAD γ-HC. In WT, Dnah9 and Dnah8 were localized along the entire length of sperm flagella. In calaxin-/-, Dnah9 and Dnah8 were lost at the distal region of sperm flagella (white arrowheads). Scale bar: 20 μm.
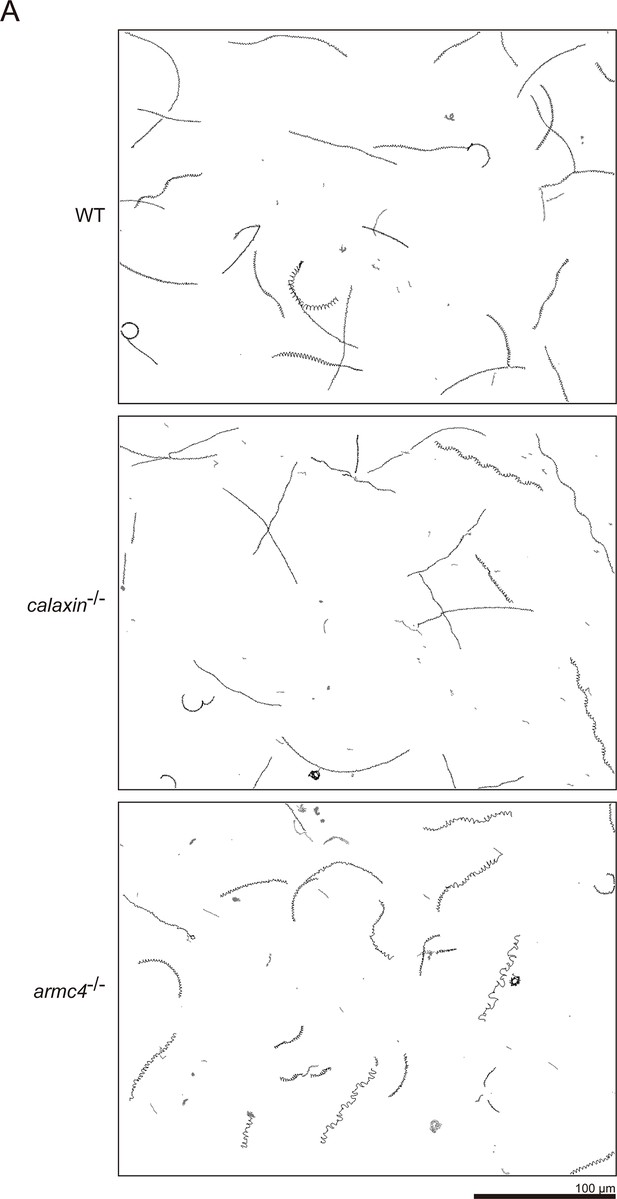
Sperm trajectories by CASA.
(A) Typical trajectories of sperm heads in WT, calaxin-/-, and armc4-/-. Swimming spermatozoa were filmed using a high-speed camera for 1 s at 200 fps. Scale bar: 100 μm.
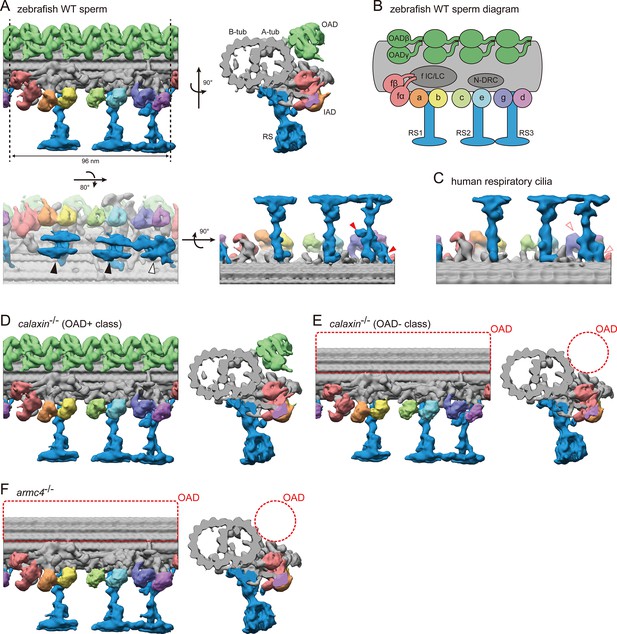
Cryo-ET structures of DMTs in WT and mutant sperm flagella.
(A) DMT structure of WT zebrafish sperm flagella. A-tub and B-tub: A- and B-tubule of DMT, respectively. OAD: outer arm dynein, IAD: inner arm dynein, RS: radial spoke. Upper left: side view, upper right: base-to-tip view, lower left: bottom view, and lower right: back view of RSs. Black arrowheads indicate the spoke heads of RS1 and RS2. White arrowhead indicates the spoke head of RS3. Red arrowheads indicate the RS3 structures which are not found in the RS3 in human respiratory cilia. (B) Diagram of DMT structure. N-DRC: nexin-dynein regulatory complex. f IC/LC: IAD-f intermediate chain and light chain complex. (C) Back view of RSs in human respiratory cilia (EMD-5950; Lin et al., 2014). (D–E) DMT structures of calaxin-/- sperm flagella. Structural classification sorted the subtomograms into two classes: (D) OAD+ class (70.7%, 6122 particles) and (E) OAD- class (29.3%, 2535 particles). (F) DMT structure of armc4-/- sperm flagella. Green: OAD, pale red: IAD-f, orange: IAD-a, yellow: IAD-b, light-green: IAD-c, cyan: IAD-e, indigo: IAD-g, violet: IAD-d, and blue: RSs. Red circles indicate the loss of OAD.
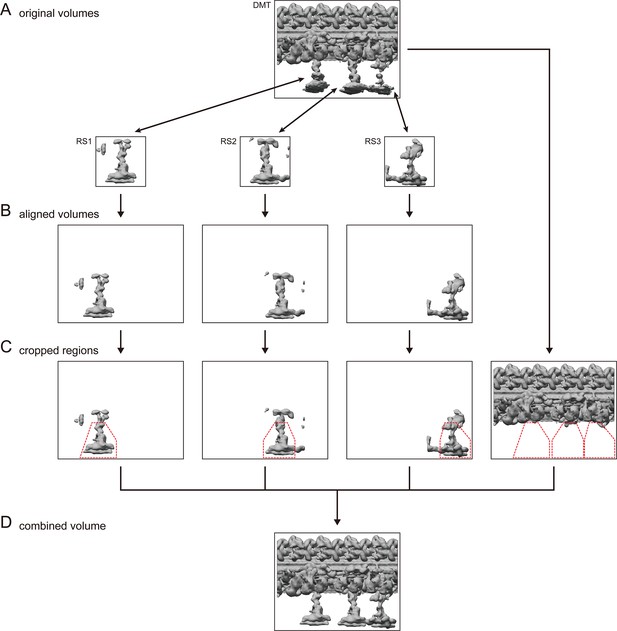
Computational process to generate combined volumes.
(A) 96 nm repeat unit of DMT was subdivided into four parts: DMT with axonemal dyneins, RS1, RS2, and RS3. Local refinements were performed individually. (B) Refined RSs were aligned on the volume of refined DMT. (C) For RSs, regions of interest were selected (red dotted lines) and segmented out. For DMT, regions corresponding to RS structures were cropped. (D) Four segmented volumes were joined together.
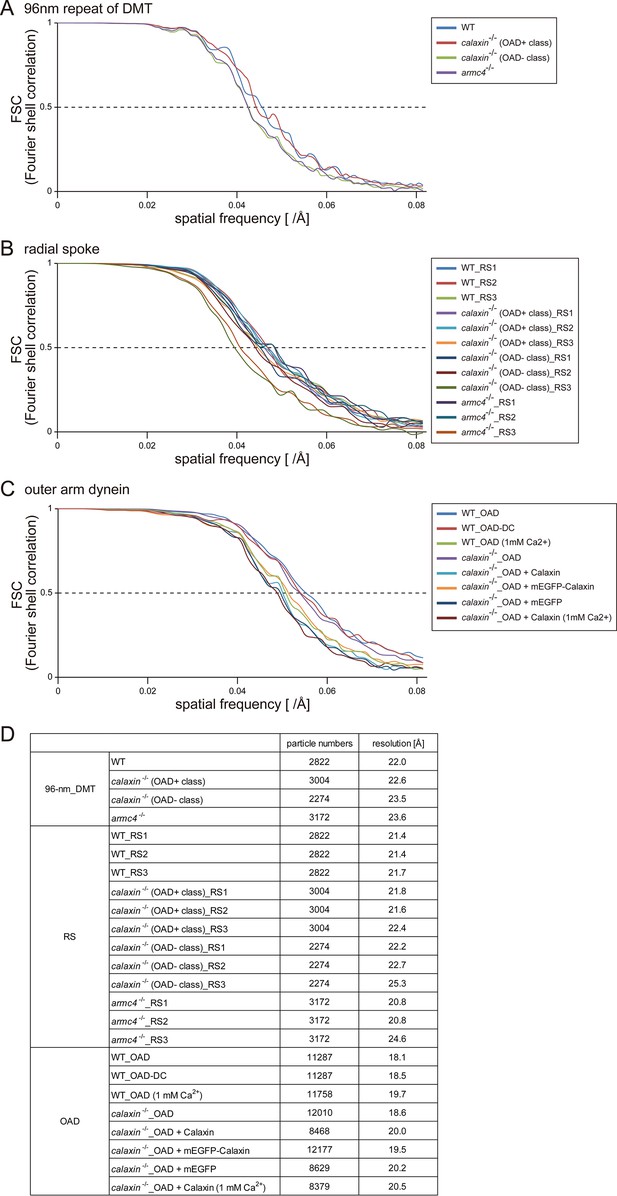
Fourier shell correlations of averaged subtomograms.
(A–C) Fourier shell correlation plots of each data set. Resolutions were determined with FSC 0.5 as a cutoff value. (A) 96 nm repeat units of DMT. (B) RSs. (C) OADs. (D) Particle numbers and resolutions of each data set.
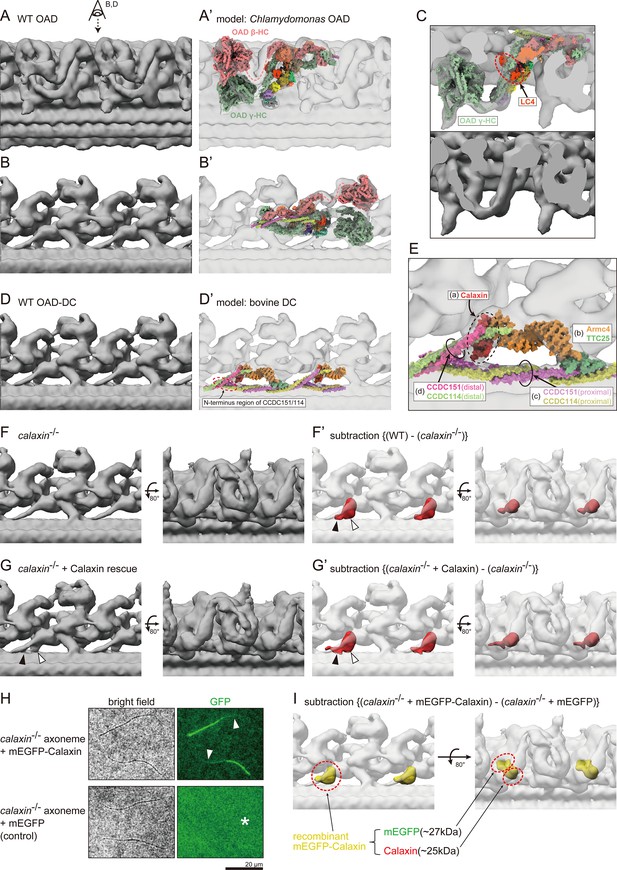
Cryo-ET structures of OADs in WT and calaxin-/- sperm flagella.
(A and B) OAD structure of WT zebrafish sperm flagella. Local refinement was performed focusing on OAD HCs. (B) shows the top view of A (eye and arrow). (A’ and B’) Comparison of zebrafish OAD structure with Chlamydomonas OAD model (PDB-7kzm; Walton et al., 2021). α-HC and DC linkers were omitted from the Chlamydomonas OAD model. (C) Detailed structure around OAD γ-HC. Upper: composite image of zebrafish OAD and Chlamydomonas OAD model. Lower: zebrafish OAD only. Chlamydomonas OAD has the LC4 protein (red circle) attached to the γ-HC tail. (D) OAD-DC structure of WT zebrafish sperm flagella. Local refinement was performed focusing on DC. (D’) Comparison of zebrafish DC structure with bovine DC model (PDB-7rro; Gui et al., 2021). Red dotted circle indicates the N-terminus region of the CCDC151/114. (E) Detailed structure of DC. DC is composed of four linker structures: (a) Calaxin, (b) the Armc4-TTC25 complex, (c) the proximal CCDC151/114, and (d) the distal CCDC151/114. (F) OAD-DC structure of calaxin-/- sperm flagella. (F’) Composite image of difference map (red; subtraction of F from WT) and calaxin-/- OAD-DC (translucent). Difference map shows the densities corresponding to the Calaxin (white arrowhead) and the adjacent CCDC151/114 (black arrowhead). (G) OAD-DC structure of calaxin-/- incubated with recombinant Calaxin protein. (G’) Composite image of difference map (red; subtraction of calaxin-/- from G) and calaxin-/- OAD-DC (translucent). (H) calaxin-/- sperm axoneme was incubated with recombinant proteins of mEGFP-Calaxin or mEGFP (control). Upper row: mEGFP-Calaxin binds to the limited region of calaxin-/- axoneme, with the partial loss of EGFP signals (white arrowheads). Lower row: mEGFP has no interaction with calaxin-/- axoneme (asterisk). Scale bar: 20 μm. (I) Composite image of difference map (yellow; subtraction of calaxin-/- incubated with mEGFP from calaxin-/- rescued by mEGFP-Calaxin) and calaxin-/- OAD-DC (translucent). Difference map shows the densities of mEGFP and Calaxin.
-
Figure 4—source data 1
Original SDS-PAGE image of recombinant proteins.
- https://cdn.elifesciences.org/articles/84860/elife-84860-fig4-data1-v2.zip
-
Figure 4—source data 2
Annotated SDS-PAGE images of recombinant proteins.
(A) Original SDS-PAGE image. (B) Contrast adjusted image of A, with annotations for each lane.
- https://cdn.elifesciences.org/articles/84860/elife-84860-fig4-data2-v2.zip
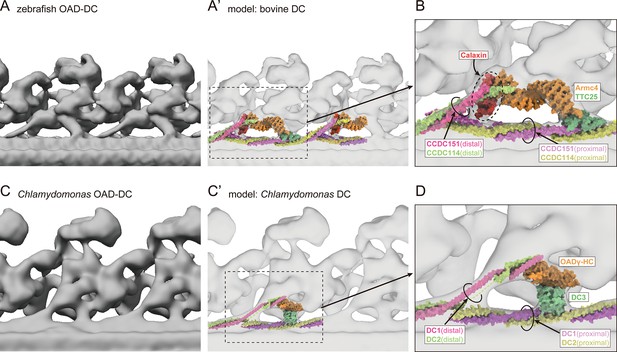
Comparison of OAD-DC structures in vertebrates and Chlamydomonas.
(A) OAD-DC structure of WT zebrafish sperm flagella. (A’) Fitting of bovine DC model (PDB-7rro; Gui et al., 2021) to zebrafish DC structure. (B) Detailed structure of vertebrate DC, composed of four structures: Calaxin, the Armc4-TTC25 complex, the proximal CCDC151/114, and the distal CCDC151/114. (C) OAD-DC structure of Chlamydomonas flagella (EMD-9768; Owa et al., 2019). (C’) Fitting of Chlamydomonas DC model (PDB-7kzo; Walton et al., 2021) to Chlamydomonas DC structure. (D) Detailed structure of Chlamydomonas DC, composed of three structures: DC3, the proximal DC1/2, and the distal DC1/2.
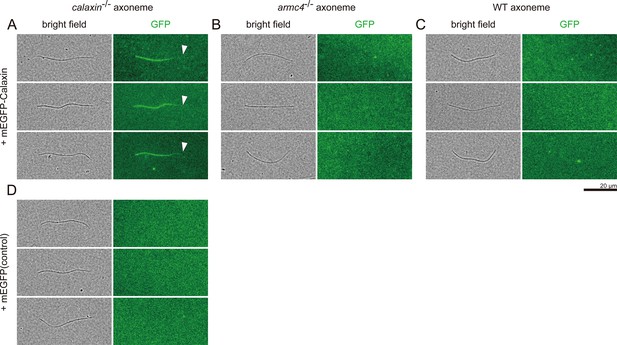
Interactions of recombinant Calaxin protein with WT and mutant sperm axonemes.
Sperm axonemes were incubated with recombinant proteins of mEGFP-Calaxin (A-C) or mEGFP (D, control). (A) mEGFP-Calaxin binds to the limited region of calaxin-/- axoneme, with the partial loss of EGFP signals (white arrowheads). (B-C) mEGFP-Calaxin showed no significant interaction with (B) armc4-/- axoneme or (C) WT axoneme. (D) mEGFP has no interaction with calaxin-/- axoneme (asterisk). Scale bar: 20 μm.
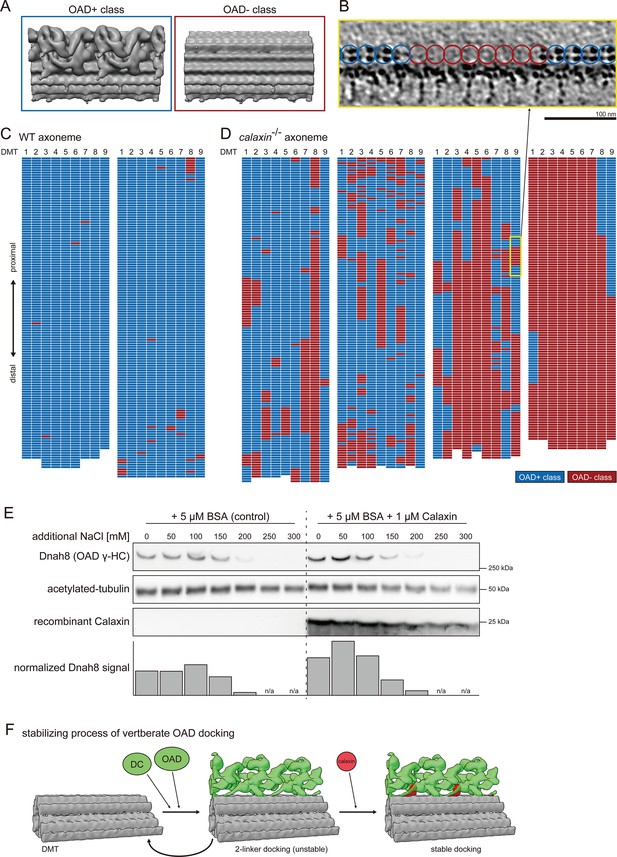
Calaxin is required for the stable docking of OAD onto DMT.
(A–D) Structural classification sorted the 24 nm repeat units of DMT into OAD+ class (blue) and OAD- class (red). (A) Averaged structures of each class. (B) Tomographic slice shows the side view of DMT. Blue or red circles indicate the class of each 24 nm repeat unit of DMT. Scale bar: 100 nm. (C–D) Distribution patterns of each class. 24 nm repeat units on each DMT are schematically displayed as individual colored grids. (C) Two typical tomograms of WT axoneme. (D) Four typical tomograms of calaxin-/- axoneme. Yellow square indicates the region displayed in B. (E) Immunoblot of calaxin-/- sperm axonemes incubated with or without recombinant Calaxin protein in different salt concentration buffers. Bottom row shows the bar graph of Dnah8 signals, normalized by the amount of acetylated tubulin signals. (F) Model of the stabilizing process of vertebrate OAD docking onto DMT.
-
Figure 5—source data 1
Original blot images of Figure 5E, Dnah8.
Chemiluminescence and epi-illumination images of the blot membrane.
- https://cdn.elifesciences.org/articles/84860/elife-84860-fig5-data1-v2.zip
-
Figure 5—source data 2
Original blot images of Figure 5E, acetylated tubulin and recombinant Calaxin.
Chemiluminescence and epi-illumination images of the blot membrane.
- https://cdn.elifesciences.org/articles/84860/elife-84860-fig5-data2-v2.zip
-
Figure 5—source data 3
Annotated blot images of Figure 5E, Dnah8.
(A) Original epi-illumination image. (B) Contrast adjusted image of A. (C) Original chemiluminescence image. (D) Contrast adjusted image of C, with annotations for each lane. (E) Annotations for lanes in D.
- https://cdn.elifesciences.org/articles/84860/elife-84860-fig5-data3-v2.zip
-
Figure 5—source data 4
Annotated blot images of Figure 5E, acetylated tubulin and recombinant Calaxin.
(A) Original epi-illumination image. (B) Contrast adjusted image of A. (C) Original chemiluminescence image, with annotations for each lane. (D) Annotations for lanes in C.
- https://cdn.elifesciences.org/articles/84860/elife-84860-fig5-data4-v2.zip
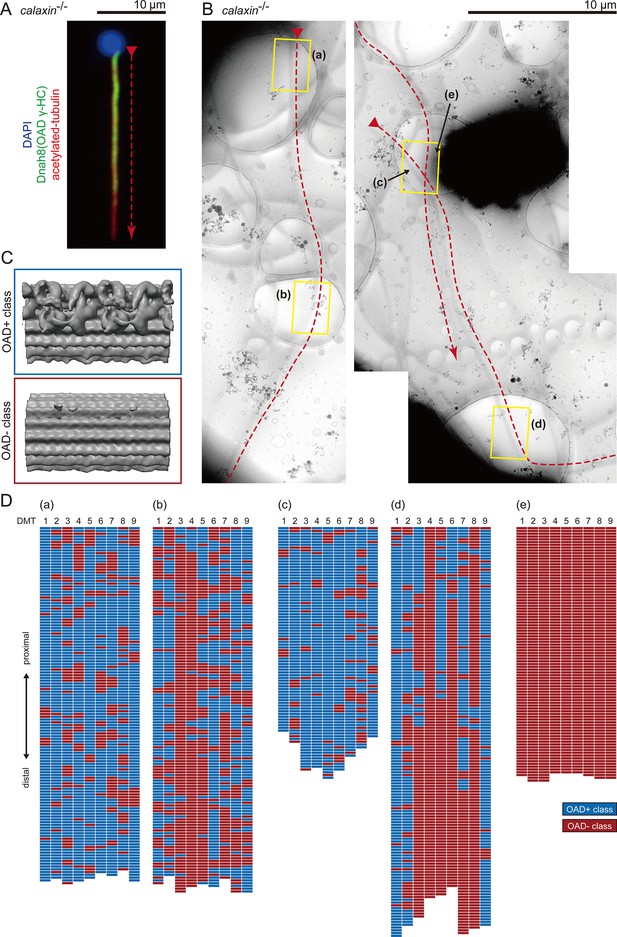
Detailed distribution of OADs in calaxin-/- sperm axoneme.
(A) Immunofluorescence microscopy shows OAD distribution in calaxin-/- spermatozoa. (B) Cryo-TEM images of calaxin-/- axonemes, traced with red dotted lines. Yellow squares indicate the cryo-ET observation areas. Proximal-distal directions of the axonemes were judged from the 3D-reconstructed structures of the axonemes. Scale bars: 10 μm. (C–D) Structural classification sorted the 24 nm repeat units of DMT into OAD+ class (blue) and OAD- class (red). (C) Averaged structures of each class. (D) Distribution patterns of each class.
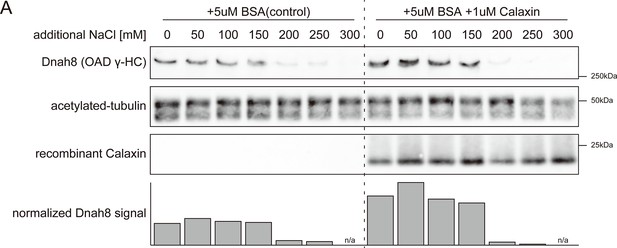
Experimental replication confirmed the reproducibility of data shown in Figure 5E.
(A) The same experiment as Figure 5E was performed with independently collected calaxin-/- sperm axonemes. Axonemes were incubated with or without recombinant Calaxin protein in different salt concentration buffers. Bottom row shows the bar graph of Dnah8 signals, normalized by the amount of acetylated tubulin signals.
-
Figure 5—figure supplement 2—source data 1
Original blot images of Figure 5—figure supplement 2, Dnah8.
Chemiluminescence and epi-illumination images of the blot membrane.
- https://cdn.elifesciences.org/articles/84860/elife-84860-fig5-figsupp2-data1-v2.zip
-
Figure 5—figure supplement 2—source data 2
Original blot images of Figure 5—figure supplement 2, acetylated tubulin.
Chemiluminescence and epi-illumination images of the blot membrane.
- https://cdn.elifesciences.org/articles/84860/elife-84860-fig5-figsupp2-data2-v2.zip
-
Figure 5—figure supplement 2—source data 3
Original blot images of Figure 5—figure supplement 2, recombinant Calaxin.
Chemiluminescence and epi-illumination images of the blot membrane.
- https://cdn.elifesciences.org/articles/84860/elife-84860-fig5-figsupp2-data3-v2.zip
-
Figure 5—figure supplement 2—source data 4
Annotated blot images of Figure 5—figure supplement 2.
(A, C, E) Epi-illumination images (contrast adjusted). (B, D, F) Chemiluminescence images (contrast adjusted), with annotations for each lane. (G) Annotations for lanes in B, D, and F.
- https://cdn.elifesciences.org/articles/84860/elife-84860-fig5-figsupp2-data4-v2.zip
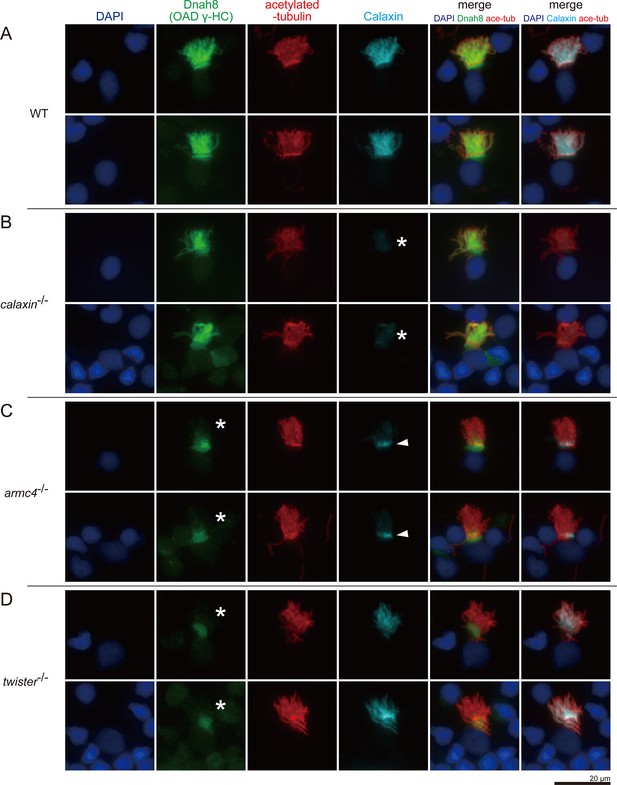
Calaxin requires Armc4 to be localized to cilia.
(A–D) Immunofluorescence microscopy of multiciliated cells of zebrafish olfactory epithelium. (A) WT. (B) calaxin-/-. Calaxin signal was lost (white asterisks). (C) armc4-/-. Ciliary localization of Dnah8 was lost (white asterisks). Calaxin was accumulated at the ciliary base (white arrowheads). (D) twister-/-. Ciliary localization of Dnah8 was lost (white asterisks). Scale bar: 20 μm.
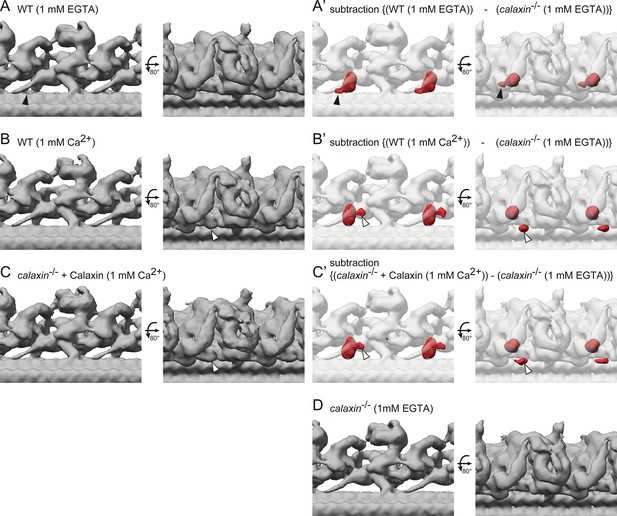
Cryo-ET structures of OADs in different Ca2+ conditions.
(A–B) OAD-DC structures of WT sperm flagella in different Ca2+ conditions: (A) 1 mM EGTA condition (for Ca2+-free) and (B) 1 mM Ca2+ condition. (C) OAD-DC structure of calaxin-/- sperm flagella incubated with recombinant Calaxin protein in 1 mM Ca2+ condition. (A’-C’) Composite images of difference map (red; subtraction of calaxin-/- in 1 mM EGTA condition from A, B, and C, respectively) and calaxin-/- OAD-DC (translucent). Black arrowheads in A indicate the CCDC151/114 structure adjacent to Calaxin, which is not observed in B and C. White arrowheads in B and C indicate the additional density around DC, which is not observed in A. (D) OAD-DC structure of calaxin-/- sperm in 1 mM EGTA condition, which was used to generate (A’-C’).
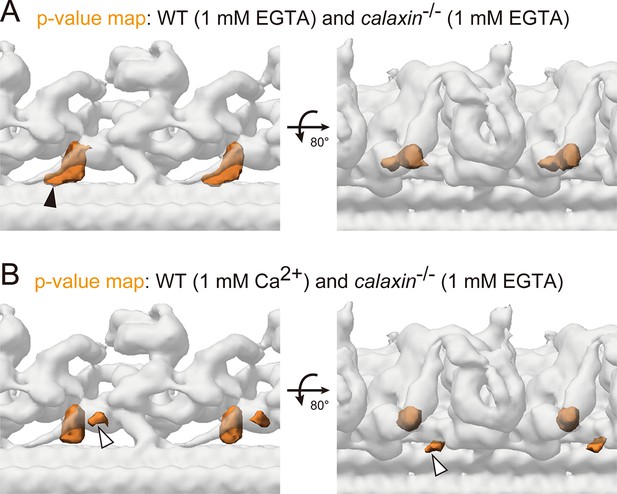
Student’s t-test to compare the WT and calaxin-/- OAD-DC structures.
(A–B) Composite image of p-value maps (orange) and calaxin-/- OAD-DC (translucent). p-Values of each voxel were calculated as described in Oda and Kikkawa, 2013. The isosurface threshold of p-values corresponds to 0.05% probability in one-tailed test. p-value maps indicate not only Calaxin structures but also the adjacent CCDC151/114 (A, black arrowhead) and the additional density around DC (B, white arrowheads) as the statistically significant difference between WT and calaxin-/- OAD-DC.
Videos
Motilities of Kupffer’s vesicle cilia in WT and armc4-/-.
Typical movies of Kupffer’s vesicle cilia, filmed by a high-speed camera at 1000 fps and played at 30 fps. Scale bar: 5 μm.
Waveforms of swimming spermatozoa in WT, calaxin-/-, and armc4-/-.
Typical swimming spermatozoa, filmed by a high-speed camera at 1000 fps and played at 30 fps. Scale bar: 10 μm.
Motilities of swimming spermatozoa in WT, calaxin-/-, and armc4-/-.
Typical movies of swimming spermatozoa for CASA, filmed by a high-speed camera at 200 fps and played at 30 fps. Scale bar: 100 μm.
Cryo-ET structure of WT DMT.
Cryo-ET structure of calaxin-/- DMT (OAD+ class).
Cryo-ET structure of calaxin-/- DMT (OAD- class).
Cryo-ET structure of armc4-/- DMT.
Cryo-ET structure of WT OAD.
Left: OAD structure of WT zebrafish sperm flagella. Right: Comparison of zebrafish OAD structure with Chlamydomonas OAD model (PDB-7kzm; Walton et al., 2021). α-HC and DC linkers were omitted from the Chlamydomonas OAD model.
Cryo-ET structure of WT OAD-DC.
Left: OAD-DC structure of WT zebrafish sperm flagella. Right: Comparison of zebrafish DC structure with bovine DC model (PDB-7rro; Gui et al., 2021).
Cryo-ET structure of calaxin-/- OAD-DC.
Left: OAD-DC structure of calaxin-/- sperm flagella. Right: Composite of difference map (red; subtraction of calaxin-/- from WT) and calaxin-/- OAD-DC (translucent).
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Gene (Danio rerio) | calaxin | NA | ZFIN: ZDB-GENE-040914–40 | also known as efcab1; odad5 |
| Gene (Danio rerio) | armc4 | NA | ZFIN: ZDB-GENE-100316–7 | also known as odad2 |
| Antibody | Anti-acetylated tubulin (mouse monoclonal) | Sigma-Aldrich | Sigma-Aldrich: T6793 | 1:500 in immunofluorescence microscopy; 1:5000 in immunoblot analysis |
| Antibody | Anti-Dnah8 (rabbit polyclonal) | PMID:29741156 | against aa 895–1402; 1:50 in immunofluorescence microscopy; 1:400 in immunoblot analysis | |
| Antibody | Anti-Dnah2 (rabbit polyclonal) | PMID:29741156 | against aa 802–1378; 1:50 in immunofluorescence microscopy; 1:400 in immunoblot analysis | |
| Antibody | Anti-Calaxin (guinea pig polyclonal) | PMID:31240264 | against full-length; 1:50 in immunofluorescence microscopy; 1:400 in immunoblot analysis | |
| Antibody | Anti-Dnah9 (rabbit polyclonal) | This paper | against aa 535–1002; 1:50 in immunofluorescence microscopy; 1:400 in immunoblot analysis | |
| Software, algorithm | CASA modified for zebrafish | PMID:17137620 | ||
| Software, algorithm | SerialEM | PMID:16182563 | ||
| Software, algorithm | MotionCor2 | PMID:28250466 | ||
| Software, algorithm | IMOD | PMID:8742726 | ||
| Software, algorithm | PEET | PMID:16917055 | ||
| Software, algorithm | UCSF Chimera | PMID:15264254 | ||
| Software, algorithm | EMAN2 | PMID:16859925 |
Oligonucleotide sequences used in this study.
| Purpose | Name | Sequence |
|---|---|---|
| Mutant generation | calaxingRNA.oligoF | ATTTAGGTGACACTATAGCGTCGGTCATCCCGAA AGTGGTTTTAGAGCTAGAAATAGCAAG |
| armc4gRNA.oligoF | ATTTAGGTGACACTATAGTACTTCAGTGAGAGCC ACCGTTTTAGAGCTAGAAATAGCAAG | |
| constant.oligoR | AAAAGCACCGACTCGGTGCCACTTTTTCAAGTTG ATAACGGACTAGCCTTATTTTAACTTGCTATTTCT AGCTCTAAAAC | |
| calaxin_check.F | GGAGAGCAGGCAGAGAGAAAG | |
| calaxin_check.R | CTGCACTGCAAATTGTGATTG | |
| armc4_check.F | CTAGAGAACAGCCTCCTGAATA | |
| armc4_check.R | GTGAAATCAGACACTTCTAGAGAT | |
| Recombinant Calaxin protein | EcoRI_calaxin.F | GGGAATTCCCATGCTGAAAATGTCGGCGATG |
| EcoRI_calaxin.R | GGGAATTCTTATTCTTTGCAGTGTTCGTGTTTCTG | |
| mEGFP.F | ATGGTGAGCAAGGGCGAG | |
| mEGFPdel229.R | GATCCCGGCGGCGGTCAC | |
| pGEX6p2-mEGFP.R | GCCCTTGCTCACCATGGGAATTCCTGGGGATCC | |
| mEGFPdel229-calaxin.F | ACCGCCGCCGGGATCATGCTGAAAATGTCGGCGA | |
| Dnah9 antigen | BamHI_dnah9.F | CGGGATCCGAGCAGCCGCTGATAGCA |
| SalI_dnah9.R | CGGTCGACTTTGCGGTCGTCCACGTA |





