Genome editing of an African elite rice variety confers resistance against endemic and emerging Xanthomonas oryzae pv. oryzae strains
Figures
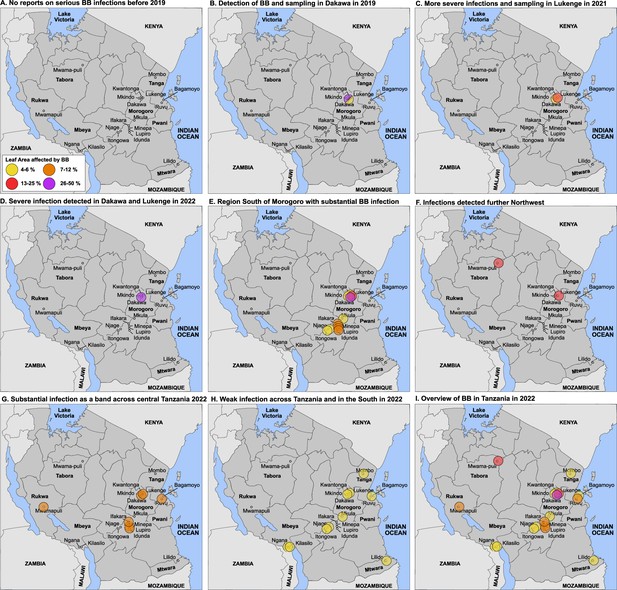
Detection of an outbreak and survey of BB in Tanzania.
(A) No reports of serious BB infections before 2019. (B) Detection in Dakawa; in 2019, (C) in Lukenge in 2021, (D–I) survey results from 2022 at different degrees of severity. Note that sampling and scoring occurred initially in the Morogoro region in Dakawa in 2019 and in Lukenge 2021, (no info on 2020 due to Covid-19); scoring and sampling was expanded to additional fields across Tanzania in 2022. Due to sampling bias early in the outbreak, data should be interpreted with caution. One may project that BB caused by the introduced strains will manifest in Tanzania and could spread to neighboring countries. Leaf area affected: yellow 4–6%, orange 7–12%, red 13–25% and purple 26–50%.

Photos of infected fields showing severe infection of the field in Dakawa, Morogoro, Tanzania, in 2019.

Photos of infected fields showing infection of the field in Lukenge, Morogoro, Tanzania, in 2021.
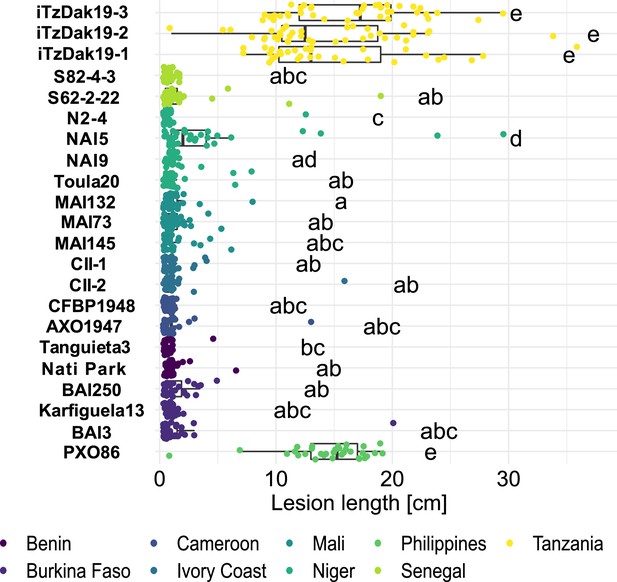
Resistance spectrum of wild-type Komboka rice against African Xoo strains.
Leaf-clip inoculation of wild-type Komboka rice plants with a panel of 21 Xoo strains originating from 8 African countries along with Asian reference strain PXO86 from the Philippines. Lesion length (in cm) was measured 14 days after Xoo inoculation. Data from three independent experiments are represented. Letters in the boxplot represent significant differences. As a representative of the isolates from Lukenge iTzLuk21-3 was tested for virulence on Komboka and shown to be fully susceptible (Figure 2—figure supplement 1).

Comparison of the susceptibility of Komboka to Tanzanian strains iTzDak19-1and iTzLuk21-3.
Leaf-clip inoculation of wild-type Komboka rice plants with a panel of two Xoo strains from the Philippines (PXO99A and PXO61), one from Burkina Faso (BAI3) , one collected in 2019 in Dakawa, and one from Lukenge in 2021. Lesion length (in cm) was measured 14 days after Xoo inoculation. Data from three independent experiments are represented. Letters in the boxplot represent significant differences. Note that the wild type data were extracted from infection experiments shown in Figure 7 to demonstrate that the iTz strains isolated in the two locations in Morogoro in 2019 and 2021 behave similarly and are equally virulent on Komboka as PXO99A.

Virulence of the African Xoo diversity panel on the susceptible rice line Azucena.
Leaf-clip inoculation of wildtype Azucena with a panel of 21 Xoo strains originating from 8 African countries along with the Asian reference strain PXO86. Lesion lengths were measured 14 days after inoculation. Data from three independent experiments are represented with n=17–37 leaves per strain.

Presence of bacterial blight R genes in select rice varieties.
(A) QTL profiling of Komboka and other selected varieties from IRRI QTL database (https://rbi.irri.org/resources-and-tools/qtl-profiles). +: presence, -: absence, n.d., not determined. (B-D). Presence of Xa1 and Xa4 R genes in Komboka using their associated markers and sequencing. (B). Genotyping of O. sativa cultivars for an Xa4-associated marker gene (M: GeneRuler 1 kb Plus DNA Ladder, ThermoFisher Scientific). (C) Neighbor-joining phylogenetic analysis of Xa4 from O. sativa cultivars. Xa4 from Komboka shares 99.98% identity with Xa4 from IR64, Teqing, and IRBB4 (Hu et al., 2017). (D) Genotyping of O. sativa cultivars for presence of functional Xa1-like genes with gene-specific markers. (E) Neighbor-joining phylogenetic analysis of Xa1 from O. sativa cultivars. Komboka Xa1 is 99.88% homologous to the publishef Xa45(t) from Oryza nivara (Ji et al., 2020).
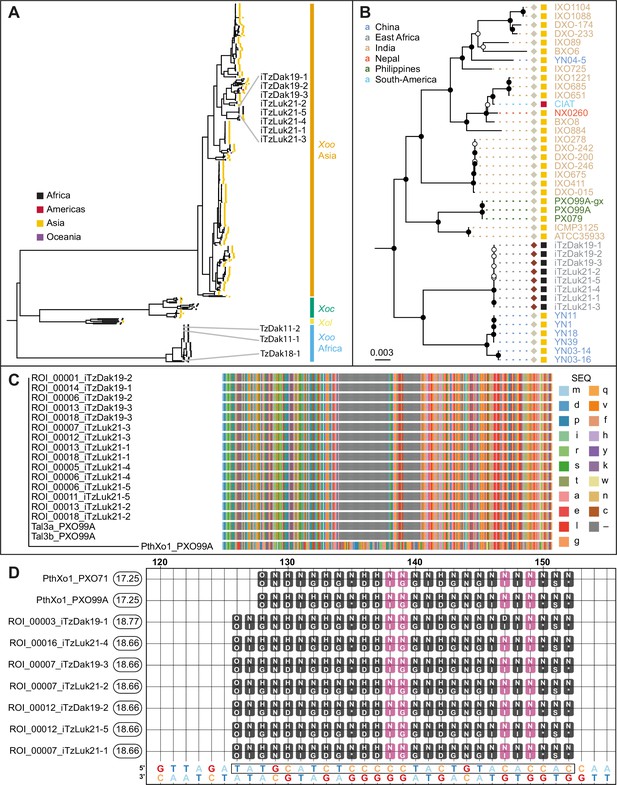
Analysis of the genomes of Tanzanian Xoo isolates.
(A) Core genome Xanthomonas oryzae phylogenetic tree. Only the names of Tanzanian isolates are indicated. Abbreviations: Xoo, X. oryzae pv. oryzae; Xoc, X. oryzae pv. oryzicola; Xol, X. oryzae pv. leersiae. (B) Close-up view of the branches of the tree in A, including the newly isolated Tanzanian strains and neighboring clades. Black-filled nodes have a bootstrap support value equal to or above 80%. The scale bar reflects branch length in mean number of nucleotide substitutions per site. Colored squares reflect the continent of origin (as in A). Text color refers to the subregion of origin. (C) Multiple alignment of the N- terminal domain sequences of PXO99A iTALes and the putative iTALes from the eight newly isolated Tanzanian iTz strains. The -terminal domain of PthXo1 from strainPXO99A was used as canonical TALe. TALe references; lowercase letters represent amino acids. Gaps are colored gray. (D) Talvez EBE predictions on the SWEET11a promoter. Repeat Variable Di-residue (RVD) sequences (rounded boxes) are aligned along their predicted matching nucleotide along the promoter sequence (the lowest row). Black-filled RVDs match their target nucleotide in the Talvez RVD-nucleotide association matrix with the best possible score for this RVD. Those in violet match with an intermediate score. Values in the rounded boxes near the TALe names correspond to Talvez prediction scores. Sequences are provided in Figure 3—source data 1.
-
Figure 3—source data 1
Compressed archive with Fasta Sequence files of Xoo strains CIX4462, CIX4506 and CIX4462.
CIX4505, CIX4507, CIX4508, CIX4457, CIX4458, Xoo3-1.
- https://cdn.elifesciences.org/articles/84864/elife-84864-fig3-data1-v3.docx

Pairwise counts of SNPs in the core genome of the newly isolated X. oryzae pv. oryzae strains from Tanzania and the closest neighboring clade.
Counts were obtained by applying the dist.dna function from the ape package (with the value 'N' to the model parameter) to pairs of genomes in the multiple SNP alignment file generated by parsnp and that included 99,626 SNPs. Rows and columns are in the same order as in the RAxML tree of Figure 3B.
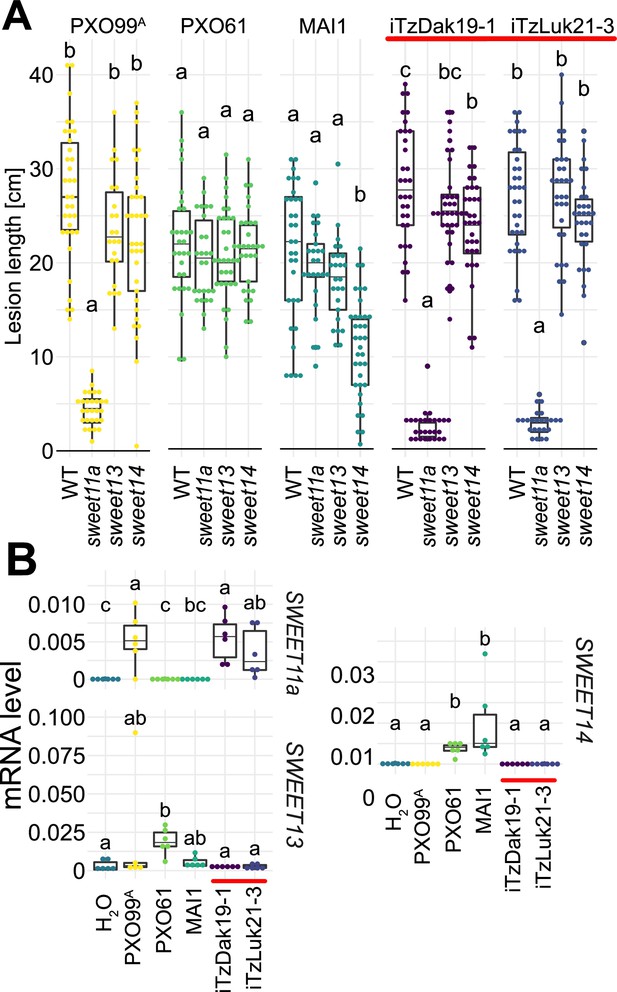
Virulence of the new Tanzanian Xoo strains depends on the induction of SWEET11a.
(A) Lesion lengths were measured 14 days after leaf-clipping inoculation of Kitaake individual sweet knock out lines (ossweet11a, ossweet13 and ossweet14) in the cultivar Kitaake (Eom et al., 2019) with PXO99A (PthXo1), PXO61 (PthXo2B/PthXo3), MAI1 (TalC/TalF), and Tanzanian strains (iTzDak19-1 and iTzLuk21-3) from the recent outbreaks (highlighted by a red bar). Results from two independent experiments are represented. (B) Relative mRNA levels (2-ΔCt) of SWEET11a, SWEET13 and SWEET14 in wild-type Kitaake upon infection by PXO99A (PthXo1), BAI3 (TalC), and Tanzanian pthXo1B dependent Xoo strains. Samples were collected 48 hr post infiltration. Data from three independent experiments were pooled and are represented together. Ct values were normalized to the rice EF1α elongation factor (∆Ct).
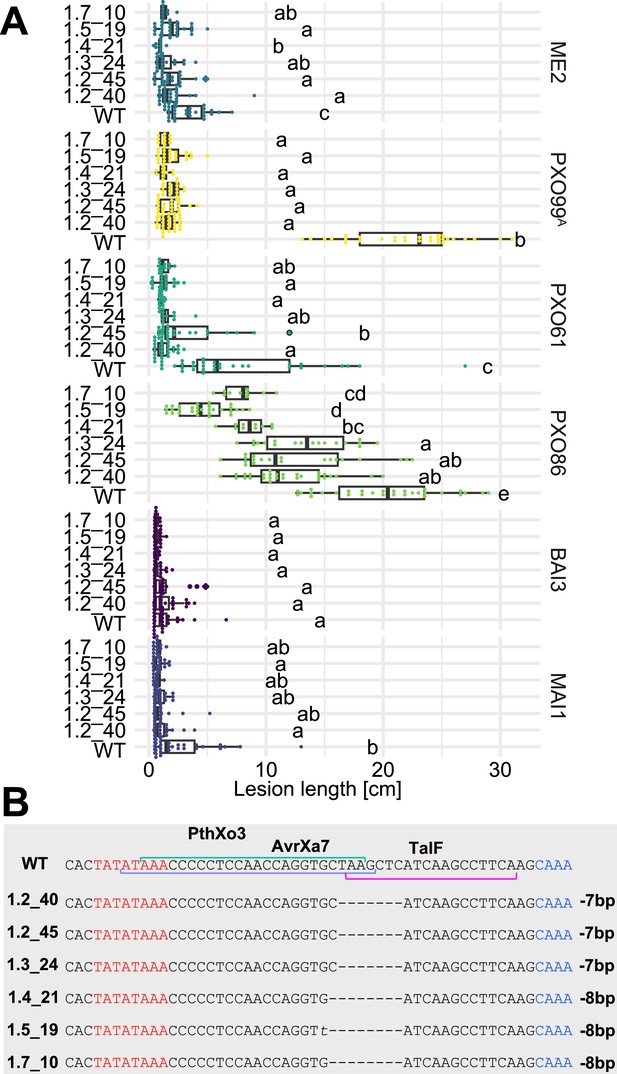
Resistance of EBE-edited Komboka lines against six representative Xoo strains.
(A) Reactions of WT and six edited Komboka lines to infection by Xoo strains (PXO99A, PXO61, PXO86, BAI3, and MAI1) harboring PthXo1, PthXo2 PthXo3, AvrXa7, TalC and/or TalF. ME2 is a PXO99A mutant strain with mutant deficient in PthXo1 and served as a negative control. The Komboka lines were carried edits in all six EBE sites validated by DNA sequencing.(B). Details of the EBEs targeted by PthXo3, AvrXa7 and TalF, and respective mutations in the six edited lines.

Comparison of EBE sequences for the SWEET11a, 13 and 14 promoters between Komboka and Kitaake.
Kitaake and Komboka have the same EBE sequences for PthXo1, TalC, TalF, AvrXa7, PthXo3. The promoter sequence of SWEET13 in wild type Komboka corresponds to the EBE for PthXo2A, while for Kitaake the sequence corresponds to the EBE for PthXo2B.

Hybrid CRISPR-Cas9/Cpf1 system.
(A) Guide RNA design. Four guide RNAs were designed to target six known EBEs within OsSWEET11a, 13 and 14 promoters. EBE sequences are bolded, and the respective TALEs are indicated. Arrowheads indicate guide nuclease cleavages sites. Cas9 guide RNA is colored in pink (gTalC), and Cpf1 guide RNAs are colored in blue (cXo1, cXo2c, cXo2d, cTalF) (cX02d is for a different SWEET13 promoter variant, as present in Nipponbare SWEET13 (5´-ATAAAGCACCACAACTCCCTT-3´) that is targeted by the PthXo2B TALe variant). (B). Duplex CRISPR-Cas9 guide RNA system. Single Cas9 gRNA were cloned into the pENTR:gRNA. (C). Multiplex CRISPR-Cpf1 guide RNA system. Single Cpf1 cRNA were shuttled into the unit vectors and assembled into pENTR:cRNAs via GoldenGate assembling. (D). Three types of destination vectors: pDEST:Cas9 is the acceptance vector of pENTR:gRNAs, pDEST:Cpf1 is the acceptance vector of pENTR:cRNAs and hybrid pDEST:Cpf1&Cas9 is the acceptance vector for both Cas9 and Cpf1 guide RNAs. (E) pMUGW5 map that contains 4 Cpf1 cRNAs and 1 Cas9 gRNA targeting six EBEs on OsSWEET11a, 13, and 14. (F) Cloning scheme. The PCR fragment containing a rice-codon-optimized SpCas9-coding sequence driven by the maize Ubiquitin 1 (ZmUbi) promoter from pDEST:Cas9 was amplified and inserted into pDEST:Cpf1 to create a hybrid destination vector pDEST:Cpf1&Cas9. Four gBlock fragments were inserted into the vector pTLN using XbaI and XhoI to generate four intermediate pUNIT vectors: pTL-OsU6.1-CpfRNA1, pTL-OsU3-CpfRNA2, pTL-TaU3-CpfRNA3, pTL-ZmU3-CpfRNA4. All four pUNITs, after cloning of respective guide RNAs, were transferred into the donor vector pENTR4-ccdB to generate pENTR:cRNAs containing the Cpf1 cRNAs cXo1, cXo2c, cTalF, and cXo2d. Simultaneously, gTalC double-stranded oligonucleotides was inserted into the BtgZI-digested pENTR-gRNA. The cRNA expression cassette was PCR-amplified and inserted into pENTR:gRNAs at the XbaI site through Gibson cloning to generate the hybrid donor plasmid pENTR:gRNA-cRNAs. The gRNA and cRNAs cassettes were mobilized to the hybrid destination vector pDEST:Cpf1&Cas9 by Gateway to create a pCam1300-CRISPR plasmid named pCpf1&Cas9:gRNAs-cRNAs (pMUGW5).

Genotypes of T0 lines with biallelic mutations at six targeted EBEs.
TATAA box is labeled in red. LbCpf1 and SpCas9 PAM sequences are labeled in blue and purple, respectively. EBE sequences are indicated (e.g. PthXo1).
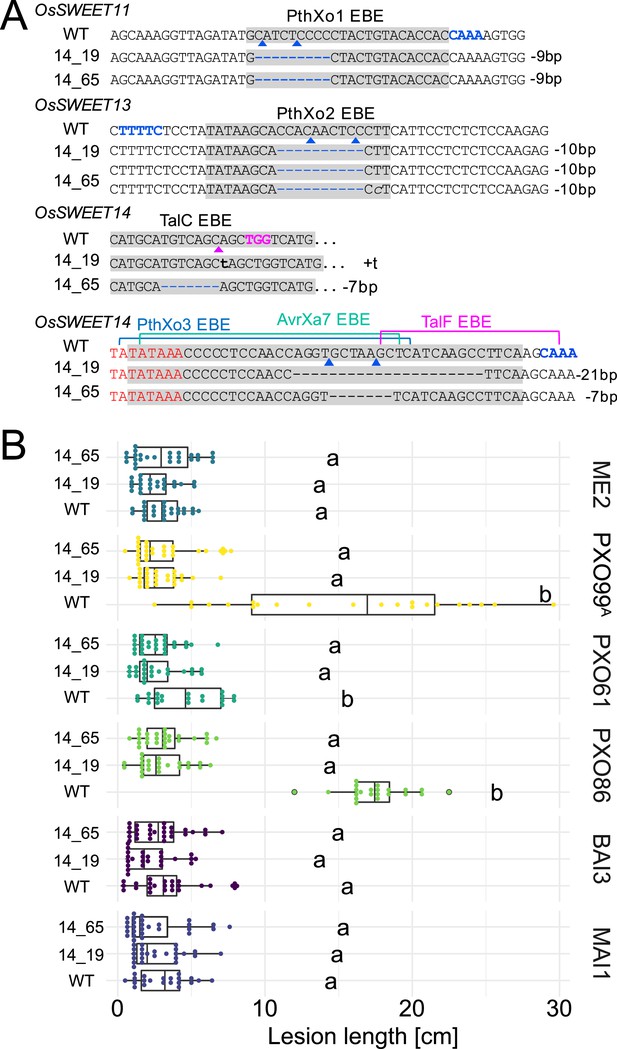
Genotypes and phenotypes of two EBE-edited lines generated in a second transformation experiment.
(A) Mutations at the EBEs for PthXo1, PthXo2, TalC, TalF, PthXo3, and AvrXa7 in two Komboka edited lines, 14_19 and 14_65. (B). Reactions of wild-type Komboka and the two edited lines to the infections by PXO99A (PthXo1), PXO61 (PthXo2B, PthXo3), PXO86 (AvrXa7), BAI3 (TalC) and MAI1 (TalC, TalF). PXO99AME2 (ME2) is a PXO99A mutant carrying a transposon insertion in pthXo1 and served as a negative control.
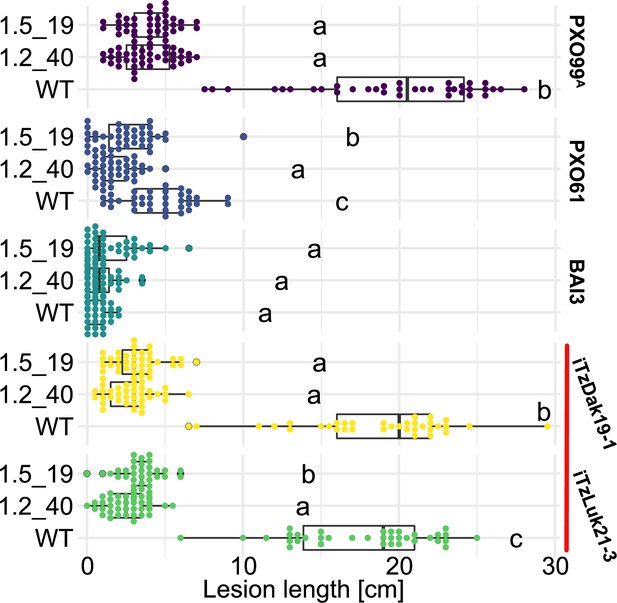
Komboka SWEET promoter edited lines are fully resistant against Tanzanian strains of Xoo.
Lesion lengths measured 14 days after leaf-clip inoculation of wild-type Komboka and the two multi-edited lines 1.2_40 and 1.5_19 with Xoo strains PXO99A (PthXo1), BAI3 (TalC), PXO61 (PthXo2B, PthXo3), and the newly isolated Tanzanian strains iTzLuk21-3 and iTzDak19-1 (both carrying PthXo1 homologs and iTAles, red bar). Data from four independent experiments are represented.

qRT-PCR analysis of SWEET11a mRNA accumulation of edited Komboka lines after infection with Tanzanian Xoo strains.
Relative mRNA levels (2-ΔCt) of SWEET11a in Komboka wild type and the multi-EBE-edited lines 1.2_40 and 1.5_19 upon infection by PXO99A (PthXo1), BAI3 (TalC) and Tanzanian PthXo1B-dependent Xoo strains (red line). Samples were collected 48 hr post infiltration. Three independent experiments are represented. Three technical replicates were performed for each sample. All Ct values were normalized to the rice EF1α elongation factor mRNA level (∆Ct). The number of infected plants (or leaves) was n=9 (except line 1.5_19 infected with iTzLuk21-3: n=7).
Tables
Efficiency of resistance genes in IRBB rice lines towards a diversity panel of 26 African Xoo strains.
Quantitative scores of lesion length produced upon leaf-clip inoculation of a diversity panel consisting of 26 Xanthomonas oryzae pv. oryzae strains from West and East Africa on IRBB near-isogenic-lines (NILs) carrying different Xanthomonas resistance genes (Xa) and on the variety Azucena used as susceptible control (IRBB1: Xa1; IRBB3: Xa3; IRBB4: Xa4; IRBB5: xa5; IRBB7: Xa7; IRBB23: Xa23; IR24: recipient parent). Resistance or susceptibility of rice plants to Xoo was determined based on lesion length measured 14 days after inoculation: Resistant (R) <5 cm; moderately resistant (MR)=5–10 cm; moderately susceptible (MS)=10–15 cm; and susceptible (S) >15 cm.
| Strain name | CIX code | Country | Azucena | IR24 | IRBB1 | IRBB3 | IRBB4 | IRBB5 | IRBB7 | IRBB23 | Komboka |
|---|---|---|---|---|---|---|---|---|---|---|---|
| NatiPark | 607 | Benin | S | R | R | R | R | R | R | R | R |
| Karfiguela13 | 705 | Burkina Faso | S | R | R | R | R | R | R | R | R |
| N2-4 | 4482 | Niger | S | R | R | R | R | R | R | R | R |
| Tanguieta3 | 609 | Benin | S | MR | R | MR | R | R | MR | R | R |
| Toula20 | 629 | Niger | S | MR | R | MR | R | R | MR | MR | R |
| BAI250 | 4127 | Burkina Faso | S | MR | R | MR | MR | R | R | MR | R |
| S62-2-22 | 2374 | Senegal | S | MR | R | MS | R | R | MR | R | R |
| CII-1 | 4083 | Ivory Coast | S | MS | MR | MS | R | R | MR | MR | R |
| CII-2 | 1042 | Ivory Coast | S | MS | MR | MS | MR | R | MR | MR | R |
| NAI9 | 2787 | Niger | S | MS | MR | MS | MR | MR | MS | S | R |
| AXO1947 | 1917 | Cameroon | S | S | R | S | R | R | MS | MS | R |
| MAI145 | 894 | Mali | S | S | R | S | MR | MS | S | MR | R |
| BAI3 | 4092 | Burkina Faso | S | MS | MR | S | MR | MR | MS | S | R |
| CFBP1948 | 2801 | Cameroon | S | MS | MR | S | MR | MR | S | S | R |
| NAI5 | 4099 | Niger | S | S | MR | S | MR | MR | S | S | R |
| MAI73 | 4079 | Mali | S | S | R | S | MS | MR | S | MR | R |
| S82-4-3 | 2976 | Senegal | S | MS | MR | S | MS | MR | MS | S | R |
| MAI132 | 4517 | Mali | S | S | R | S | S | MS | S | MR | R |
| iTzDak19-1 | 4457 | Tanzania | S | S | S | S | S | S | S | R | S |
| iTzDak19-2 | 4458 | Tanzania | S | S | S | S | S | S | S | R | S |
| iTzDak19-3 | 4462 | Tanzania | S | S | S | S | S | S | S | R | S |
| iTzLuk21-1 | 4506 | Tanzania | nd | nd | nd | nd | nd | nd | nd | nd | nd |
| iTzLuk21-2 | 4509 | Tanzania | nd | nd | nd | nd | nd | nd | nd | nd | nd |
| iTzLuk21-3 | 4505 | Tanzania | nd | nd | nd | nd | nd | nd | nd | nd | S |
| iTzLuk21-4 | 4507 | Tanzania | nd | nd | nd | nd | nd | nd | nd | nd | S |
| iTzLuk21-5 | 4508 | Tanzania | nd | nd | nd | nd | nd | nd | nd | nd | nd |
-
n.d. not determined.
Additional files
-
Supplementary file 1
Supplementary file with additional data.
(a) List of Xoo strains used in this study (b) Disease survey in multiple rice growing areas in Tanzania in 2022. (c) Sequences of Xa1 and Xa4 genes from Oryza sativa cv. Komboka. (d) Characteristics of the TALome of Xanthomonas oryzae pv. oryzae Tanzanian strains. (e) OsSWEET promoter sequences for the rice varieties Komboka and Kitaake. (f) Guide RNA sequences. (g) EBE sequences in select OsSWEET promoters of Komboka wild-type and CRISPR-edited lines. (h) List of SNPs and Indels in OsSWEET promoters after editing of the rice varieties of Komboka and Kitaake (i) List of T0 lines with biallelic mutations at all targeted EBEs. (j) Screening of the T1 generation for homozygous mutations in all six SWEET EBEs. (k) List of T1 lines with homozygous mutations of all targeted EBEs. n.d. not determined. (l) List of primers used in this study (m) Alignment of OsSWEET11a promoters from the rice cv. Komboka and Kitaake.
- https://cdn.elifesciences.org/articles/84864/elife-84864-supp1-v3.docx
-
Supplementary file 2
Accession numbers and main features of genome assemblies used for strain phylogeny inference.
- https://cdn.elifesciences.org/articles/84864/elife-84864-supp2-v3.xlsx
-
Supplementary file 3
SWEET11a-targeting RVD sequences of the iTz strains.
- https://cdn.elifesciences.org/articles/84864/elife-84864-supp3-v3.xlsx
-
Source data 1
Summary of raw data files available at dryad https://doi.org/10.5061/dryad.xpnvx0kk3.
- https://cdn.elifesciences.org/articles/84864/elife-84864-data1-v3.xlsx
-
MDAR checklist
- https://cdn.elifesciences.org/articles/84864/elife-84864-mdarchecklist1-v3.pdf





