Fetal liver macrophages contribute to the hematopoietic stem cell niche by controlling granulopoiesis
Figures
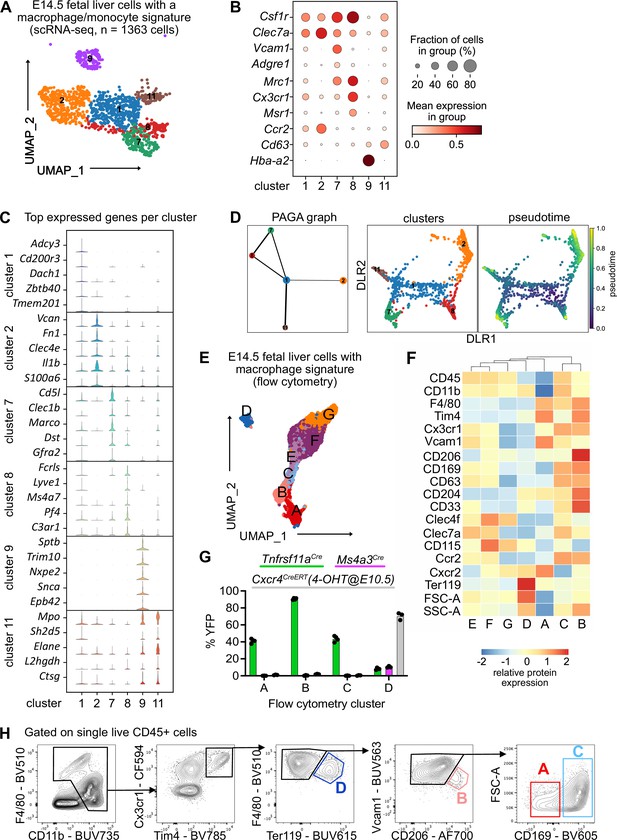
Characterization of fetal liver macrophage populations.
(A) Single-cell RNA-sequencing (scRNA-seq) analysis of wildtype CD11blow/+ cells isolated from a fetal liver at developmental day (E)14.5. Clusters of possible macrophage subsets identified by a monocyte/macrophage signature (see Figure 1—figure supplement 1) are visualized through Uniform Manifold Approximation and Projection (UMAP). (B) Expression of selected macrophage- and macrophage progenitor-specific genes in clusters from (A). (C) Violin plots of highly expressed genes within the clusters from (A). (D) Developmental trajectory analysis using Partition-based graph abstraction (PAGA) method (left) and pseudotime analysis (right) of the identified clusters in (A), excluding cluster 9. (E) Flow-cytometry analysis of CD11blow/+ cells with a macrophage signature, isolated from a fetal liver at developmental day E14.5. Cell surface marker expression was used to generate unbiased clusters using UMAP. (F) Heatmap of relative protein expression and cell size parameters used in the flow-cytometry analysis in (E). (G) Flow-cytometry analysis of E14.5 fetal liver macrophages using three different fate-mapping mouse models. CD11blow/+ cells with a macrophage signature were analyzed as shown in (E, F), resulting in similar cluster distribution (see Figure 1—figure supplement 2). YFP+ cells from the Tnfrsf11aCre; Rosa26YFP model (green) indicate a yolk sac origin. Ms4a3Cre; Rosa26YFP (pink) and Cxcr4CreERT; Rosa26YFP with 4-hydroxytamoxifen (4-OHT) injection at E10.5 (gray) indicate a monocytic and hematopoietic stem cell origin, respectively. Circles represent individual mice. (H) Simplified gating strategy to identify macrophage clusters in E14.5 livers using flow cytometry.
-
Figure 1—source data 1
Quantification of fate-mapped macrophages populations for Figure 1.
- https://cdn.elifesciences.org/articles/86493/elife-86493-fig1-data1-v2.xlsx
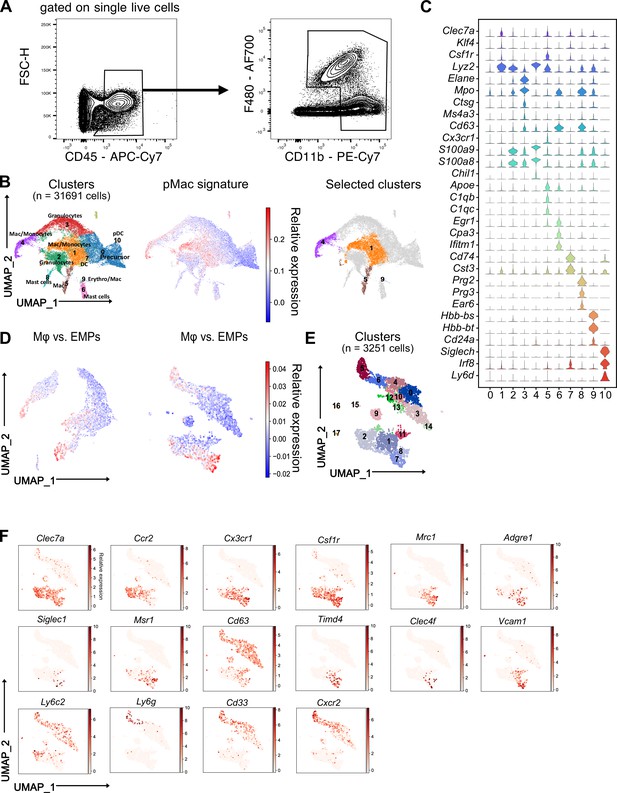
Sorting and characterization of fetal liver myeloid cells.
(A) Sorting strategy of CD11b+ cell for performing single-cell RNA-sequencing using wildtype E14.5 fetal livers. (B) Subclustering strategy and identification of cell types in the single-cell clusters from all cells passing quality control. Mac: macrophage, Erythro: erythroblast, DC: dendritic cell, pDC: plasmacytoid dendritic cell (left panel). The initial clusters of interest were identified by projecting the pre-macrophage (pMac) signature on the single cells (middle and right panels). (C) Top expressed genes from clusters in (B) to identify myeloid cell populations. (D) Subclustering strategy for the second and the third round of macrophage cluster selection. The initially selected clusters were analyzed for a second time, and a set of signature genes upregulated in macrophages (Mɸ) compared to erythro-myeloid progenitors (EMPs) were projected on them. The same procedure was repeated for a third time. (E) Clusters resulting from the third round of subsetting. Eighteen identified clusters were visualized using UMAP. (F) Expression of individual monocyte and macrophage-specific genes on the final identified clusters.
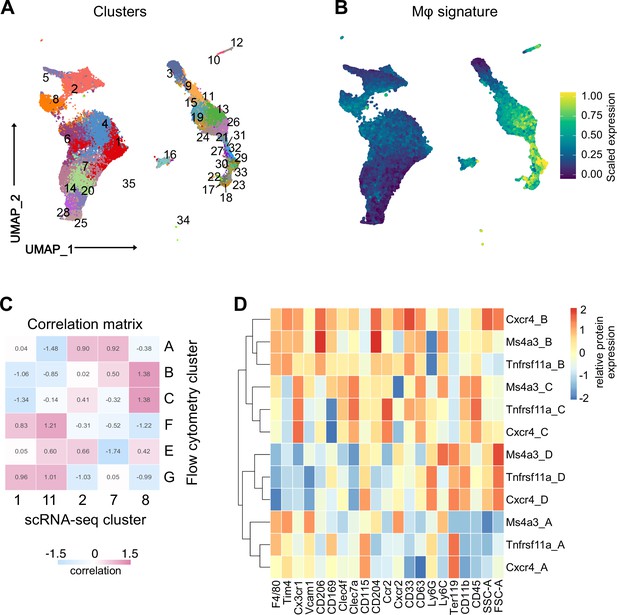
Flow-cytometry analysis of macrophage subpopulations and fate-mapping mouse models.
(A) Flow-cytometry analysis of CD11blow/+ cells with a macrophage signature, isolated from a fetal liver at developmental day E14.5. Cell surface marker expression was used to generate unbiased clusters using UMAP. (B) Projection of macrophage markers on the flow-cytometry data using UMAP visualization. (C) Correlation matrix between the flow cytometry (Figure 1E, F) and scRNA-seq data (Figure 1A–C) reveals highly correlating clusters between the two datasets. (D) Relative expression of markers used in the flow-cytometry analysis of three different fate-mapping mouse models. Tnfrsf11a: Tnfrsf11aCre; Rosa26YFP model. Ms4a3: Ms4a3Cre; Rosa26YFP; Cxcr4: Cxcr4CreERT; Rosa26YFP with 4-hydroxytamoxifen (4-OHT) injection at E10.5. The identified macrophage clusters (A–D) shown in Figure 1F could be identified in all models with similar expression patterns.
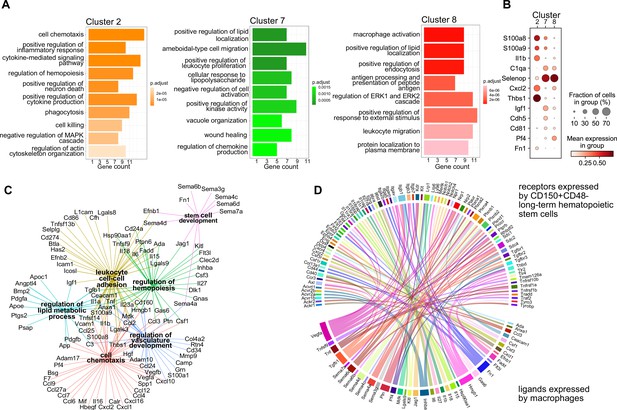
Transcriptional program and paracrine signaling of fetal liver macrophages.
(A) Gene set enrichment analysis (GSEA) of final macrophage clusters 2, 7, and 8 was performed on the differentially expressed genes of each cluster. (B) Expression of selected ligands in the final macrophage clusters. (C) Interaction and functional network of expressed ligands on the identified macrophage clusters 2, 7, and 8. Each hub with a color indicates the function of the ligands, while the edges show the interaction between them. (D) Potential ligand–receptor interactions between macrophages and long-term hematopoietic stem cells (LT-HSCs). The gene names on the bottom of the plot are expressed ligands in macrophage clusters 2, 7, and 8. Gene names on the top are expressed receptors on LT-HSCs at E14.5. Each ligand can target several receptors which are indicated with the same color.
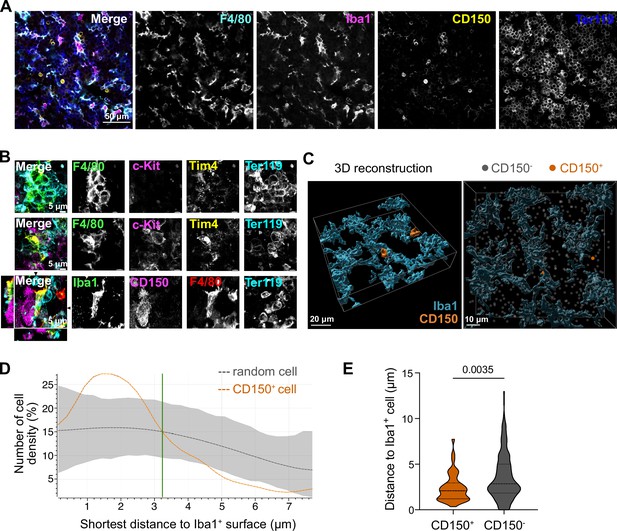
Macrophage interaction with CD150+ hematopoietic stem cells (HSCs).
(A) Immunostaining of E14.5 fetal liver sections with antibodies against F4/80, Iba1, CD150, and Ter119. Scale bar represents 50 µm. (B) Immunostaining of E14.5 fetal liver whole-mounts with antibodies against F4/80, c-Kit, Tim4, Ter119, Iba1, and CD150. The first row visualizes the interaction between EI macrophages surrounded by Ter119+ erythroblasts. The second and last rows highlight the interaction between macrophages and progenitor cells, including CD150+ long-term HSCs (LT-HSCs). The black arrows on the last row indicate the yz and xz dimensions which are shown on the left and bottom sides. Scale bar represents 5 µm. (C) 3D reconstruction of a E14.5 fetal liver whole mount stained with Iba1, CD150, and DAPI (shown as gray dots indicating CD150- cells). Scale bar represents 20 µm (left) or 10 µm (right). Measured distance of CD150+ and CD150− cells to the next Iba1+ macrophage (D) and average distance of cells to Iba1+ macrophages (n = 42 for CD150+ and n = 524 for CD150−) (E). Unpaired Student’s t-test.
-
Figure 3—source data 1
Distances measured between CD150+ or CD150− cells and Iba1 cells for Figure 3.
- https://cdn.elifesciences.org/articles/86493/elife-86493-fig3-data1-v2.xlsx
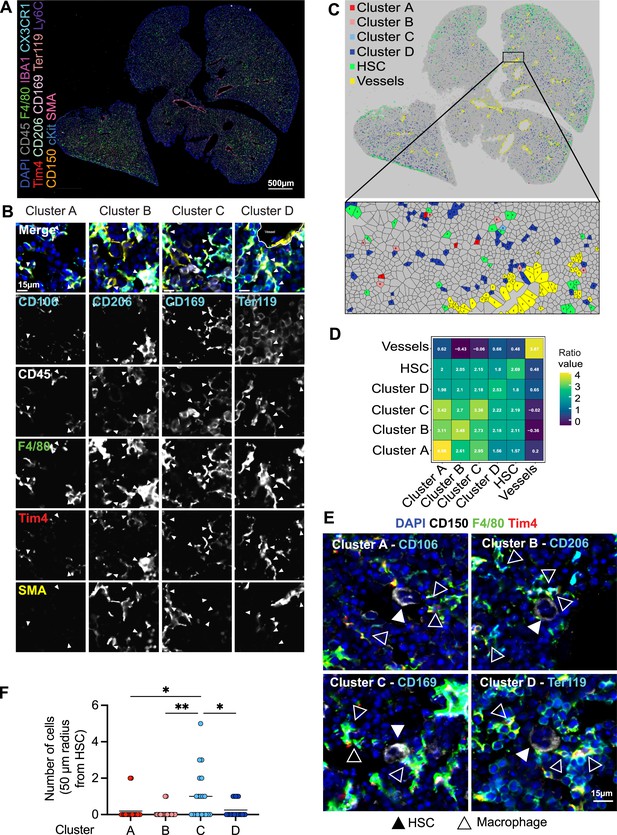
Macrophage heterogeneity and their interaction with hematopoietic stem cells (HSCs).
(A, B) A 5-µm frozen section of a fetal liver from an E14.5 wildtype embryo was stained with a 20-plex CODEX antibody panel. Representative image of the entire field of view is shown in (A) and enlargements showing representative images of the cells from clusters A–D are shown in (B). Arrows in the enlargements indicate macrophages from the corresponding cluster. Scale bars represent 500 µm in the overlay and 15 µm in the enlargements. (C) Voronoi diagram from (B) after manual cell classification using the HSCs, blood vessels, and cells from the four macrophage clusters as seeds. (D) Spatial analysis of interactions between cells from macrophage clusters, HSCs, and blood vessels in the fetal liver within a range of 5–50 µm. Values represent the calculated Log Odds Ratio. (E) Representative images of the interaction of HSCs with the macrophage clusters. Filled arrowheads indicate the HSC, and empty arrowheads indicate the macrophages from the corresponding cluster. CD150 staining is shown in white color. Scale bar represents 15 µm. (F) Absolute number of cells from the four macrophage clusters within a radius of 50 µm from one randomly selected HSC. Each dot represents one cell. Black lines in the plot represent the mean. One-way analysis of variance (ANOVA); *p < 0.05, **p < 0.01.
-
Figure 4—source data 1
Number of CD150+ cells within each macrophage subpopulation measured in a 50-μm radius.
- https://cdn.elifesciences.org/articles/86493/elife-86493-fig4-data1-v2.xlsx
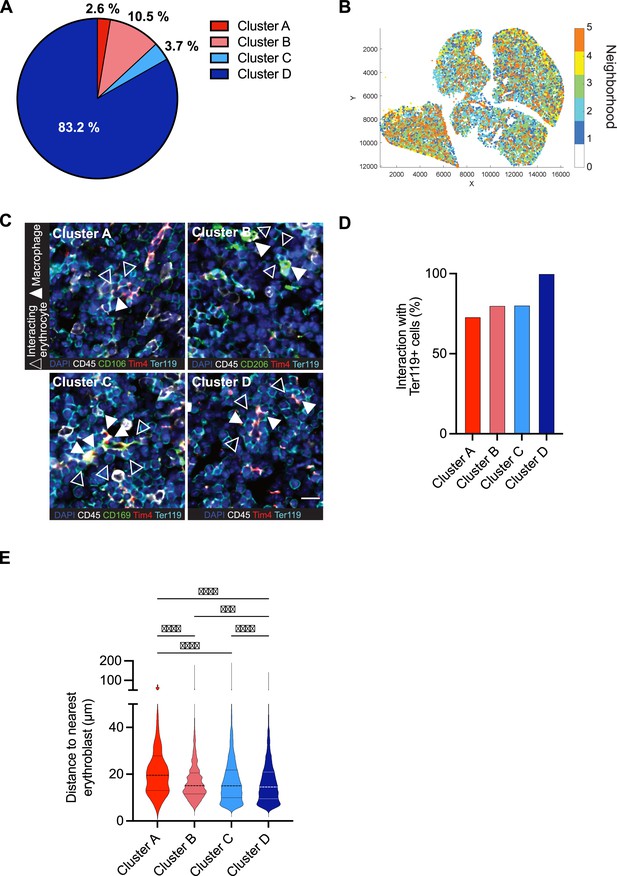
Spatial CODEX analyses of macrophage clusters in the fetal liver.
(A) Proportions of cells from the macrophage clusters within the total F4/80+ Iba1+ Cx3cr1+ Tim4+ macrophages from the CODEX image in Figure 3B. (B) Single objects detected in Figure 3B were spatially segmented into neighborhoods using a raster scan with a radius of 50 µm. Plot represents clustering of cellular neighborhoods based on their local composition of macrophages from the four clusters. Each color represents a neighborhood with a similar composition. (C) Representative images of interactions between macrophage subpopulations and Ter119+ erythroblasts. Filled arrowheads indicate macrophages from the corresponding cluster, whereas empty arrowheads indicate interacting erythrocytes. Scale bar represents 15 µm. (D) Interaction frequency between Ter119+ cells and distinct macrophage clusters, as indicated in (C). (E) The distance from the four macrophage clusters to their nearest erythrocyte in the entire tissue section was measured. One-way analysis of variance (ANOVA); ***p < 0.001, ****p < 0.0001.
-
Figure 4—figure supplement 1—source data 1
Quantification of macrophage subpopulations and their interaction with Ter119+ cells for Figure 4—figure supplement 1.
- https://cdn.elifesciences.org/articles/86493/elife-86493-fig4-figsupp1-data1-v2.xlsx
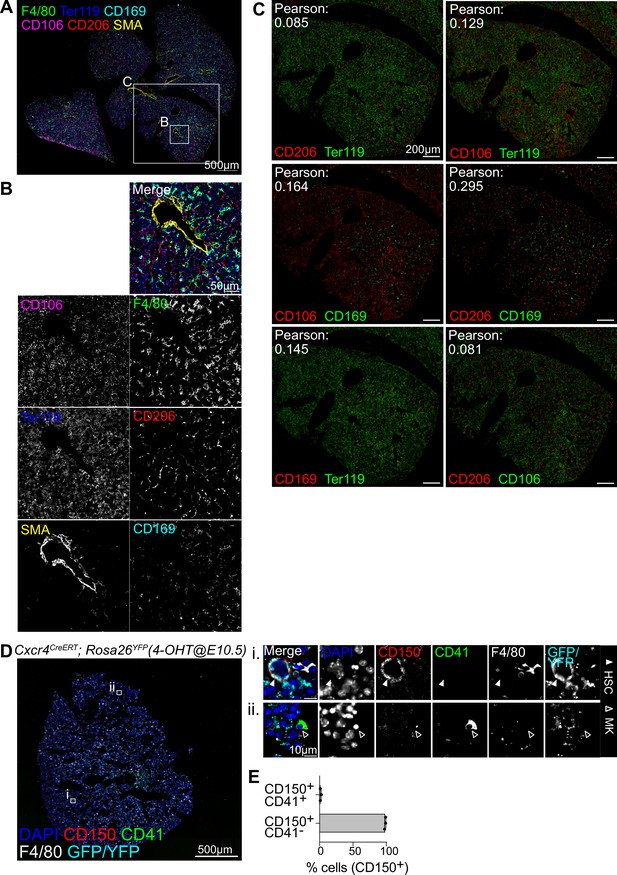
CODEX-based detection of macrophage heterogeneity and hematopoietic stem cells (HSCs).
(A–C) A 5-µm frozen section of a fetal liver from an E14.5 wildtype embryo was stained with a 20-plex CODEX antibody panel. Representative image of the entire field of view is shown in (A) and enlargements showing all macrophage subcluster markers together (B) or in a pairwise comparison (C). Signal overlap in (C) was measured with the Pearson colocalization coefficient (related to Figure 4B). Scale bars represent 500 µm in the overview and 50 µm in the enlargements in (B) and 200 µm in the enlargements in (C). (D) A 5-µm frozen section of a fetal liver from an E14.5 Cxcr4CreERT; Rosa26YFP embryo where the pregnant mother was injected with 4-hydroxytamoxifen (4-OHT) at E10.5 was stained with a 22-plex CODEX antibody panel. Representative image of the entire field of view is shown on the left and enlargements showing a CD150+ YFP/GFP+ CD41− HSC in (i) and an example of a CD41 expressing megakaryocyte (Mg) cell in (ii). Scale bars represent 500 µm in the overview and 10 µm in the enlargements. (E) Quantification of CD150 and CD41 co-expressing cells in three sections as show in (D).
-
Figure 4—figure supplement 2—source data 1
Quantification of CD150 and CD41 expression for Figure 4—figure supplement 2.
- https://cdn.elifesciences.org/articles/86493/elife-86493-fig4-figsupp2-data1-v2.xlsx
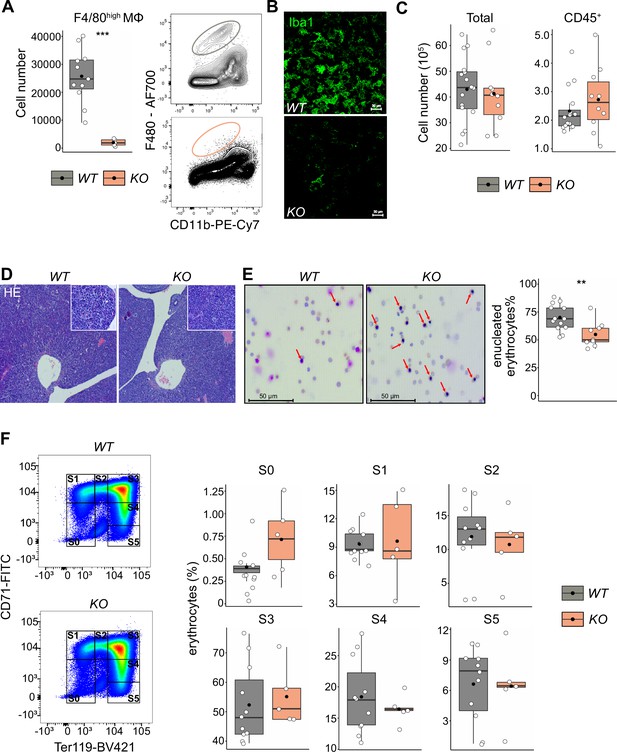
The effect of macrophage depletion on erythropoiesis.
(A) Quantification of F4/80+ macrophage cells in the E14.5 fetal liver of Tnfrsf11a+/+; Spi1f/f (WT) and Tnfrsf11aCre/+; Spi1f/f (KO) embryos using flow cytometry. n = 16 for WT, n = 10 for KO. (B) Immunofluorescent staining of macrophages using Iba1 antibody in E14.5 livers. Representative for n = 3. Scale bar represents 30 µm. (C) Quantification of single live and CD45+ cells in the E14.5 fetal liver of WT and KO knockout embryos using flow cytometry. n = 11 for WT, n = 5 for KO. (D) Hematoxylin and eosin stain (HE) of WT and KO fetal livers at E14.5. Representative for n = 5. Overviews were taken with a ×5 objective, insets with a ×20 objective. (E) On the left: representative pictures of blood smear using May–Grünwald–Giemsa staining. Arrows indicate nucleated erythroblasts. On the right; the percentage of enucleated erythrocytes in the blood of WT and KO embryos at E14.5. n = 13 for WT and n = 9 for KO. (F) On the left: representative gating strategy to capture differentiation stages of erythrocytes. On the right: comparison of the erythrocyte’s percentages in each of the differentiation stages between WT and KO embryos at E14.5. n = 11 for WT and n = 5 for KO. All statistical tests comparing WT and KO embryos: ***p < 0.001, **p < 0.01, Wilcoxon test.
-
Figure 5—source data 1
Quantified cell numbers of knockout and wildtype E14.5 embryos for Figure 5.
- https://cdn.elifesciences.org/articles/86493/elife-86493-fig5-data1-v2.xlsx
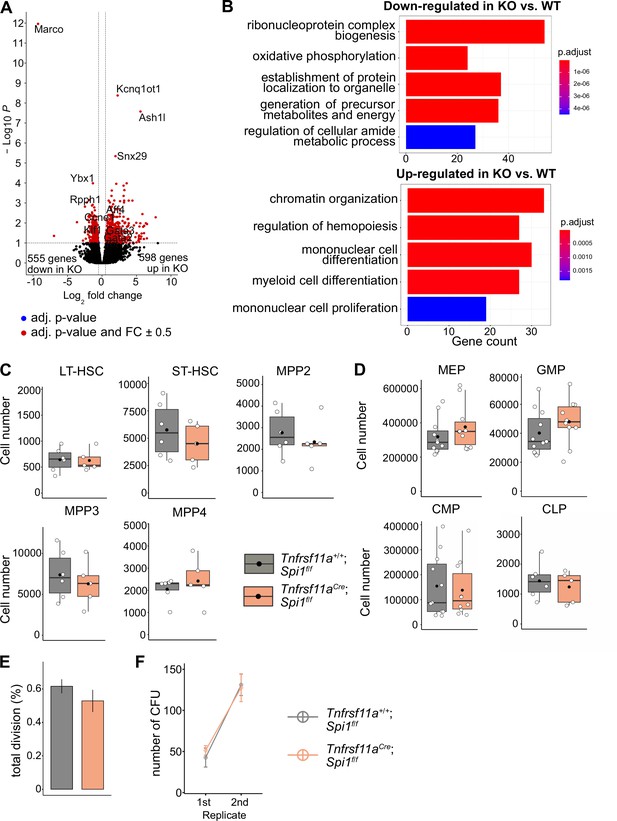
The effect of macrophage depletion on hematopoiesis.
(A) Volcano plot of differentially expressed genes between CD150+ LT-HSCs sorted from Tnfrsf11a+/+; Spi1f/f (WT, n = 4) and Tnfrsf11aCre/+; Spi1f/f (KO, n = 6) embryos. Blue dots are significant (adjusted p-value <0.1), red dots are significant with a fold-change of ±0.5. (B) Gene set enrichment analysis of significant up- and downregulated genes from (A) between the control and knockout embryos. (C) Quantification of the stem- and progenitor cells from WT and KO fetal livers at E14.5. LT-HSC: long-term hematopoietic stem cells; ST-HSC: short-term hematopoietic stem cells; MPP: multipotent progenitors. n = 6 for WT and n = 5 for KO. (D) Quantification of progenitors from WT and KO fetal livers at E14.5. n = 6–10 for WT and n = 5–10 for KO. CLP: common lymphoid progenitor; CMP: common myeloid progenitor; GMP: granulocyte–macrophage progenitor; MEP: megakaryocyte–erythrocyte progenitor. (E) Cell proliferation assay. LT-HSCs were harvested from the fetal liver at E14.5 using FACS for performing a single-cell colony assay. n = 77 LT-HSCs from n = 5 fetal livers for WT and n = 96 LT-HSCs from n = 5 fetal livers for KO. (F) Serial transfer colony-forming assay showing the number of observed colonies after seeding E14.5 fetal liver cells into media (1st replicate) and re-seeding the cultured colonies (2nd replicate). n = 7 for WT and n = 8 for KO.
-
Figure 6—source data 1
Quantified cell numbers of knockout and wildtype E14.5 livers and cells from serial colony-forming unit (CFU) and proliferation assays for Figure 6.
- https://cdn.elifesciences.org/articles/86493/elife-86493-fig6-data1-v2.xlsx
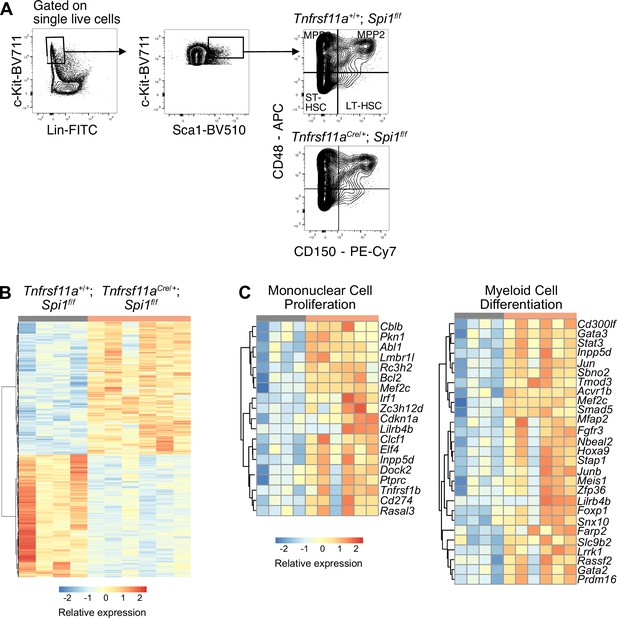
Bulk RNA-sequencing and analysis of long-term hematopoietic stem cell (LT-HSC).
(A) Gating strategy for sorting LT-HSC from Tnfrsf11a+/+; Spi1f/f and Tnfrsf11aCre; Spi1f/f fetal livers at E14.5 for RNA-sequencing. (B) Heatmap of all differentially expressed genes between LT-HSCs from Tnfrsf11a+/+; Spi1f/f and Tnfrsf11aCre; Spi1f/f at E14.5 with adjusted p-value <0.1. (C) Heatmap of genes belonging to the GO terms ‘mononuclear cell proliferation’ and ‘myeloid cell differentiation’.
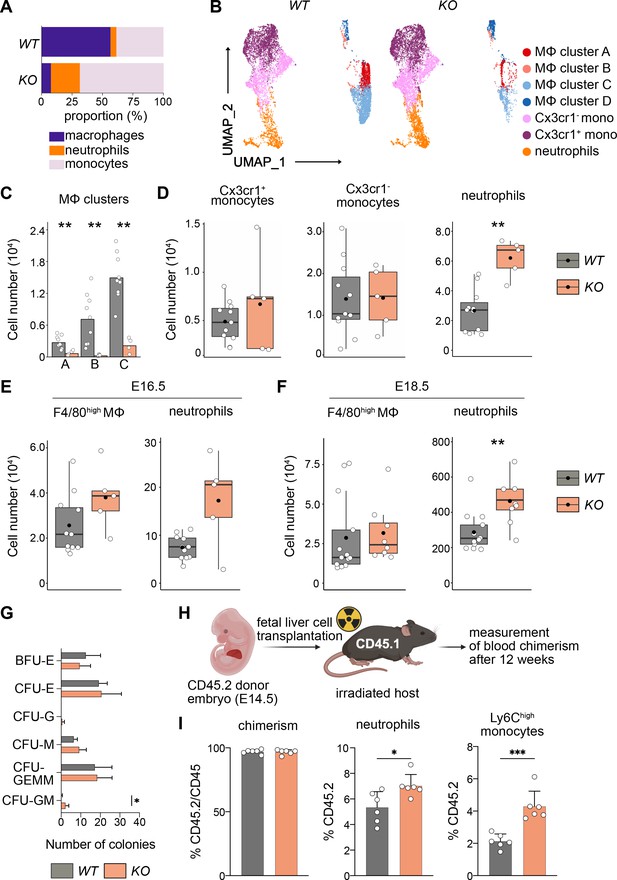
Macrophage depletion shifts hematopoiesis toward the granulocyte–macrophage progenitor (GMP) lineage.
(A) Relative proportions of myeloid cells cells isolated from Tnfrsf11a+/+; Spi1f/f (WT) and Tnfrsf11aCre/+; Spi1f/f (KO) fetal livers at E14.5. Average of n = 11 for WT and n = 5 for KO. (B) Flow-cytometry analysis of CD11blow/+ cells isolated from WT and KO fetal livers at E14.5. Cell surface marker expression was used to generate unbiased clusters using UMAP, which were subsequently used for a gating strategy to quantify respective macrophage (Mϕ), monocyte, and neutrophil populations. (C) Quantification of Mϕ subclusters from WT and KO fetal livers at E14.5. n = 10 for WT and n = 4 for KO. (D) Quantification of monocytes (Cx3cr1+ and Cx3cr1−) and neutrophils from WT and KO fetal livers at E14.5. n = 11 for WT and n = 5 for KO. Quantification of F4/80 Mϕ and neutrophils from WT and KO fetal livers at E16.5, n = 11 for WT and n = 5 for KO (E) and at E18.5, n = 13 for WT and n = 8 for KO (F). (G) Colony-forming unit assay from WT and KO fetal livers at E14.5. n = 4 for WT and n = 5 for KO. BFU: burst-forming erythroid, E: erythroid; G: granulocyte; GEMM; granulocyte, erythroid, macrophage, megakaryocyte; GM granulocyte, macrophage; M: macrophage. (H) Scheme of transplantation experimental setup. (I) Quantification of CD45.2 chimerism and % of neutrophils and Ly6Chigh monocytes in the blood of hosts 12 weeks after WT or KO fetal liver cells were transplanted. n = 6 for WT and KO. Wilcoxon test was performed comparing WT and KO cell number in (C), two-way analysis of variance (ANOVA) test was performed for the CFU assay in (G), unpaired Student’s t-test was performed for adoptive transfer experiment in (I). ***p < 0.001, **p < 0.01, and *p < 0.05.
-
Figure 7—source data 1
Quantified cell numbers of knockout and wildtype livers and cells from colony-forming unit (CFU) and adoptive transfer assays for Figure 7.
- https://cdn.elifesciences.org/articles/86493/elife-86493-fig7-data1-v2.xlsx
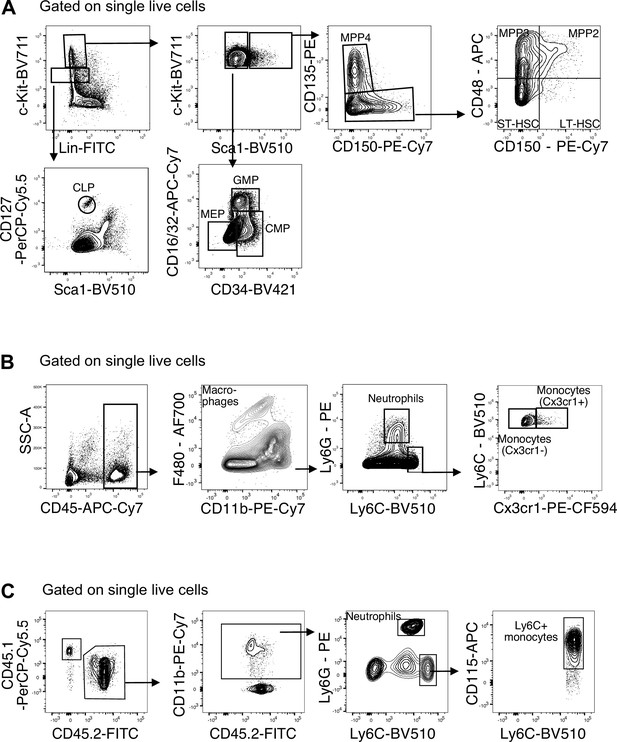
Gating strategies for flow-cytometry data.
(A) Gating strategy for quantification of stem and progenitor cells in fetal livers at E14.5. CLP: common lymphoid progenitor; CMP: common myeloid progenitor; GMP: granulocyte–macrophage progenitor; LT-HSC: long-term hematopoietic stem cells; MEP: megakaryocyte–erythrocyte progenitor; MPP: multipotent progenitors; ST-HSC: short-term hematopoietic stem cells. (B) Gating strategy for quantification of myeloid cells in fetal livers at E14.5. (C) Gating strategy for quantification of CD45.1/CD45.2 chimerism after adoptive transfer.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Commercial assay or kit | CODEX conjugation kit | Akoya Biosciences | 7000008 | |
| Commercial assay or kit | CODEX staining kit | Akoya Biosciences | 7000009 | |
| Commercial assay or kit | CODEX Buffer | Akoya Biosciences | 7000001 | |
| Chemical compound, drug | BS3 fixative | Thermo Fisher Scientific | 21580 | |
| Chemical compound, drug | CollagenaseD | Sigma-Aldrich | 11088882001 | |
| Chemical compound, drug | DAPI (4′,6-diamidino-2-phenylindole, dilactate) | BioLegend | 422801 | 1:10,000 |
| Chemical compound, drug | DNase | Sigma-Aldrich | DN25-1G | |
| Chemical compound, drug | DRAQ7 | BioLegend | 424001 | 1:1000 |
| Chemical compound, drug | Fetal calf serum | Bio&Sell | FBS. S 0615HI | |
| Chemical compound, drug | Giemsa solution | Merck | 109204 | |
| Chemical compound, drug | May–Grünwald solution | Merck | 101424 | |
| Chemical compound, drug | MethoCult | StemCell Technologies | 3434 | |
| Chemical compound, drug | Normal Goat Serum | VWR | ICNA 08642921 | |
| Chemical compound, drug | Paraformaldehyde (PFA) | Thermo Fisher Scientific | 11586711 | |
| Chemical compound, drug | Progesterone | Sigma-Aldrich | P3972-5G | |
| Chemical compound, drug | QIAzol Lysis Reagent | QIAGEN | 79306 | |
| Chemical compound, drug | Rat serum | Bio-Rad | C13SD | |
| Chemical compound, drug | Roswell Park Memorial Institute medium (Seahorse XF RPMI medium) | Agilent Technologies | 103576-100 | |
| Chemical compound, drug | Tamoxifen | Sigma-Aldrich | T5648 | |
| Chemical compound, drug | Weise buffer tablet | Merck | 109468 | |
| Software | CODEX instrument manager | Akoya Biosciences | ||
| Software | CODEX Processor | Akoya Biosciences | ||
| Software | CODEX MAV | Akoya Biosciences | ||
| Software | QuPath | Bankhead et al., 2017 | ||
| Software | ImageJ | Schindelin et al., 2012 | ||
| Software | FlowJo | BD | v.10.8.1 |
Additional files
-
Supplementary file 1
Macrophage subcluster-specific gene set enrichment analysis (GSEA).
- https://cdn.elifesciences.org/articles/86493/elife-86493-supp1-v2.xlsx
-
Supplementary file 2
Ligands expressed by macrophages at E14.5.
- https://cdn.elifesciences.org/articles/86493/elife-86493-supp2-v2.xlsx
-
Supplementary file 3
Differentially expressed genes in knockout versus wildtype long-term hematopoietic stem cells (LT-HSCs).
- https://cdn.elifesciences.org/articles/86493/elife-86493-supp3-v2.xlsx
-
Supplementary file 4
GO term analysis of knockout versus wildtype long-term hematopoietic stem cells (LT-HSCs).
- https://cdn.elifesciences.org/articles/86493/elife-86493-supp4-v2.xlsx
-
Supplementary file 5
Antibody list.
- https://cdn.elifesciences.org/articles/86493/elife-86493-supp5-v2.xlsx
-
Supplementary file 6
Barcodes and reporters for co-detection by indexing CODEX.
- https://cdn.elifesciences.org/articles/86493/elife-86493-supp6-v2.xlsx
-
MDAR checklist
- https://cdn.elifesciences.org/articles/86493/elife-86493-mdarchecklist1-v2.pdf





