Diminishing neuronal acidification by channelrhodopsins with low proton conduction
Figures
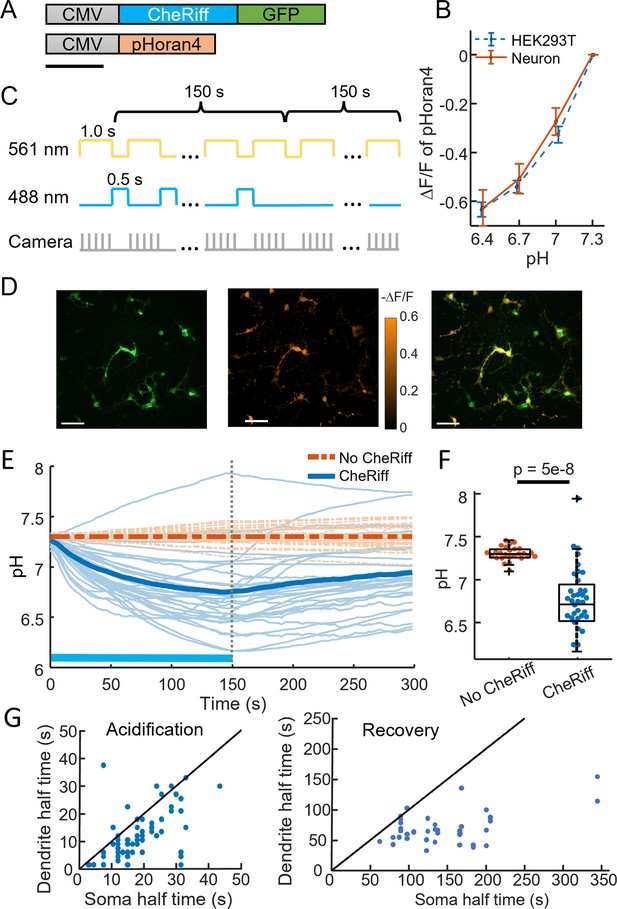
CheRiff acidifies polarized cells.
(A) Genetic constructs for simultaneous optogenetic stimulation and pH imaging. (B) Calibration of pHoran4 pH sensor in HEK cells and neurons. Error bars represent standard deviation (SD) of n = 8 measurements in HEK cells, 42 measurements in neurons. (C) Protocol for measuring pH responses to optogenetic stimulation. Stimulation (blue) and measurement (yellow) were interleaved for 150 s; then pH recovery was measured for 150 s without optogenetic stimulation. (D) Example images of cultured neurons showing (left) Green Fluorescent Protein (GFP) fluorescence, a marker for CheRiff expression, (middle) −ΔF/F in the pHoran4 channel after 150 s of the protocol shown in (C), (right) merge. Scale bars 100 μm. (E) Time-course of pH in cultured neurons. Cells expressing pHoran4 but not CheRiff did not acidify. Bold lines show population average. (F) CheRiff-expressing neurons acidified to a pH of 6.76 ± 0.35 (mean ± SD, n = 34 cells). Neurons not expressing CheRiff had significantly less acidification, pH = 7.3 ± 0.08 (mean ± SD, n = 26 cells, p = 5e−8 Wilcoxon rank sum test). Box plots show inter-quartile ranges, tick-marks show data range, + shows outlier. (G) Half-time of (left) acidification or (right) recovery for neuron somas vs dendrites stimulated with the protocol in (C). Black line shows equal kinetics.
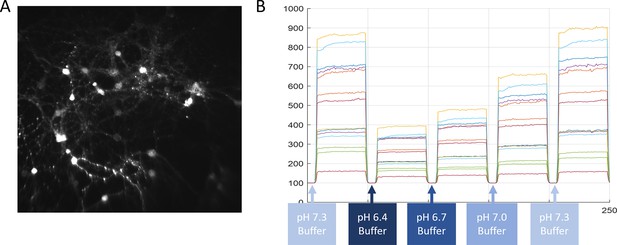
Procedure for calibrating pHoran4 pH measurements.
(A) Representative image of cultured neurons expressing pHoran4. The cells have been permeabilized with Nigericin and are in a high K+ extracellular medium (Methods). (B) Example fluorescence traces of individual cells as the dish is perfused with buffers of different pH values.
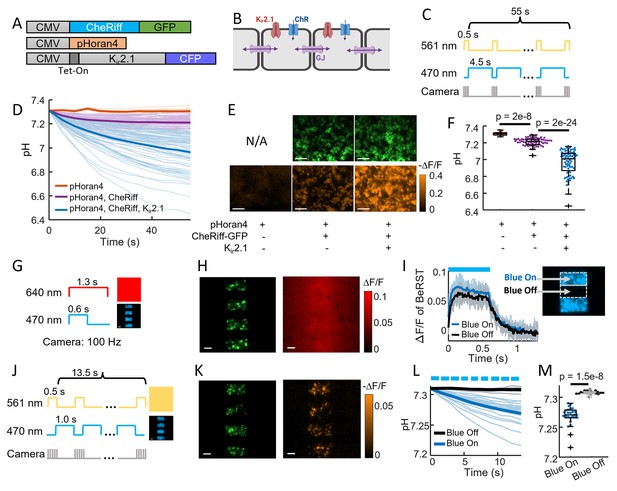
CheRiff exhibits high proton conductance.
(A) Genetic constructs for simultaneous optogenetic stimulation and pH imaging in polarized HEK293T cells. (B) Diagram of HEK cell monolayer connected by gap junctions. (C) Experimental paradigm for measuring pH responses to optogenetic stimulation. Stimulation (488 nm) and measurement (561 nm) were interleaved to avoid optical crosstalk. (D) Time-course of pH in HEK cells. Expression of Kir2.1 increased the driving force for proton influx, substantially enhancing the acidification. (E) Images of HEK cell monolayers showing (top) GFP fluorescence, a marker for CheRiff expression and (bottom) −ΔF/F in the pHoran4 channel after protocol shoin (C). Scale bars 100 μm. (F) Quantification of the data in (D–E). pHoran4 alone: pH = 7.31 ± 0.02 (mean ± standard deviation [SD], n = 13 cells); CheRiff and pHoran4 pH = 7.21 ± 0.05 (n = 70 cells); CheRiff, pHoran4, and Kir2.1: pH = 6.96 ± 0.15 (n = 75 cells). Statistical comparisons via Wilcoxon signed-rank test. (G) Protocol for mapping voltage responses to patterned optogenetic stimulation (488 nm) via fluorescence of BeRST1 (640 nm exc.). (H) Images of HEK cell monolayers showing (left) fluorescence of GFP with patterned blue illumination, (right) ΔF/F of BeRST1. Scale bars 100 μm. (I) Time-course of BeRST1 fluorescence in HEK cells inside (Blue On) and outside (Blue Off) the optogenetic stimulus regions. (J) Protocol for measuring pH responses to patterned optogenetic stimulation. Stimulation (488 nm) and measurement (561 nm) were interleaved to avoid optical crosstalk. (K) (Left) Fluorescence of GFP with patterned blue illumination, (right) ΔF/F in the pHoran4 channel after protocol shown in (J). Scale bars 100 μm. (L) Time-course of pH inside (Blue On) and outside (Blue Off) the optogenetic stimulus regions. (M) Quantification of the data in (L). Directly stimulated cells acidified to pH = 7.27 ± 0.016 (mean ± SD, n = 26 cells), indirectly depolarized cells (Blue Off) did not acidify: pH = 7.31 ± 0.003 (n = 19 cells, p = 1.5e−8 Wilcoxon signed-rank test).
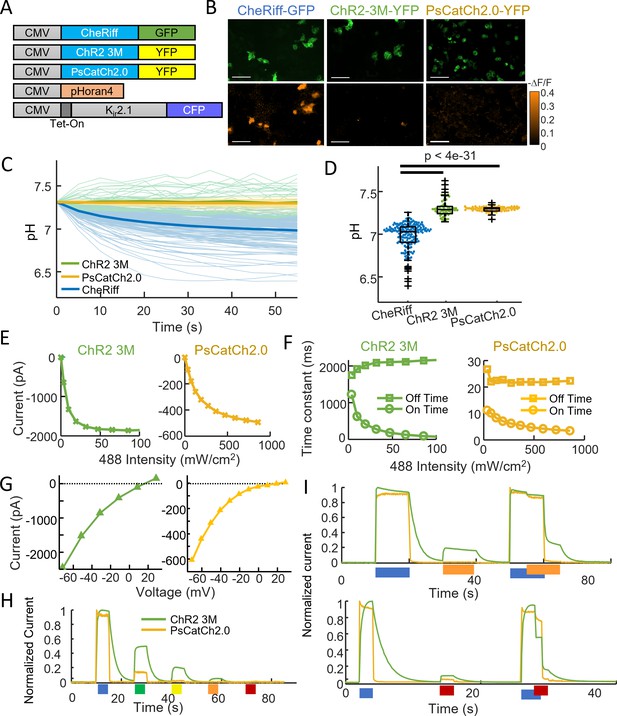
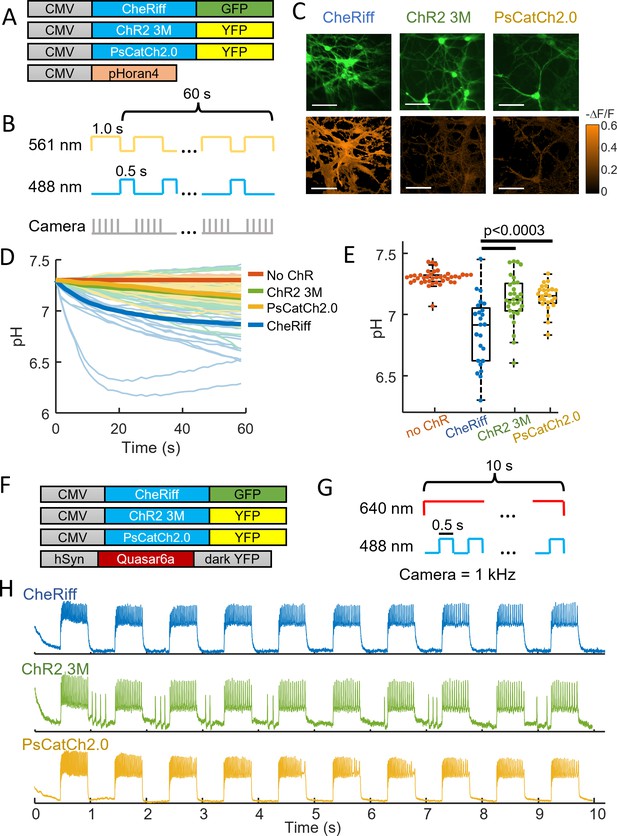
ChR2-3M and PsCatCh2.0 are potent non-acidifying channelrhodopsins.
(A) Genetic constructs for simultaneous optogenetic stimulation using channelrhodopsin variants and pH imaging in polarized HEK cells. (B) Images of HEK cells showing (top) GFP or YFP fluorescence, a marker for channelrhodopsin expression and (bottom) −ΔF/F in the pHoran4 channel, measured after protocol shown in Figure 2C. Scale bars 100 μm. (C) Time-course of pH in HEK cells expressing the three opsins. (D) Quantification of theta in (C). CheRiff: pH = 6.98 ± 0.15 (mean ± standard deviation [SD], n = 170 cells); ChR2-3M: pH = 7.31 ± 0.10 (n = 63 cells); PsCatCh2.0: pH = 7.30 ± 0.03 (n = 74 cells); p = 4e−31, p = 4e−35, Wilcoxon signed-rank test. (E–I) Whole-cell voltage clamp measurements on HEK cells expressing channelrhodopsins. (E) Steady-state photocurrents as a function of blue illumination intensity. (F) Opening and closing kinetics as a function of blue light intensity. (G) Steady-state photocurrents as a function of holding voltage. (H) Normalized photocurrents from stimulation with light at 488, 532, 561, 594, and 640 nm (50 mW cm−2 in all cases). (I) Normalized photocurrents from combinations of blue (488 nm, 240 mW cm−2) and orange (594 nm, 1 W cm−2) or red (640 nm, 8 W cm−2) light corresponding to intensities typical for all-optical electrophysiology.
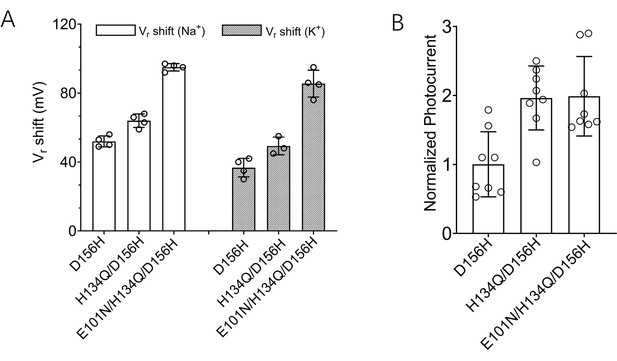
Engineering of ChR2-3M, a channelrhodopsin with high Na+ and K+ selectivity and high photocurrent amplitude.
(A) Shifts in reversal potential (Vr) of ChR2 variants upon changing extracellular Na+ or K+ concentration from 120 to 1 mM (mean ± standard deviation [SD], n = 3–4 cells). (B) Photocurrent amplitudes of ChR2 variants (mean ± SD, n = 8 cells). The triple mutant E101N/H134Q/D156H was modified with trafficking, ER export, and signal peptides and designated ChR2-3M. Experiments were performed with Xenopus oocytes expressing different channelrhodopsin variants.

Depolarization of HEK cell monolayers via patterned stimulation of channelrhodopsins.
(A) Blue light stimulation patterns. (B) Fluorescence of GFP or YFP tags on the opsins expressed in confluent monolayers of HEK cells, under patterned blue light excitation. (C) BeRST1 ΔF/F showing depolarization from patterned stimulation. Scale bars 100 μm.
Chr2-3M and PsCatCh2.0 acidify neurons less than CheRiff.
(A) Genetic constructs for simultaneous optogenetic stimulation and pH imaging. (B) Experimental paradigm for measuring pH responses to optogenetic stimulation. Stimulation (blue) and measurement (yellow) are interleaved for 60 s to avoid optical crosstalk. (C) Images of cultured neurons showing (top) GFP or YFP fluorescence, a marker for channelrhodopsin expression, (bottom) ΔF/F in the pHoran4 channel after the protocol shown in (B). (left) CheRiff-GFP, (middle) ChR2-3M-YFP, (right) PsCatCh2.0. Scale bars 100 μm. (D) Time-course of pH dynamics in cultured neurons. Cells expressing ChR2-3M and PsCatCh2.0 acidify less than CheRiff. (E) Neurons expressing ChR2-3M, pH = 7.13 ± 0.19 (mean ± standard deviation [SD], n = 31 cells), and PsCatCh2.0, pH = 7.14 ± 0.11 (mean ± SD, n = 25 cells) had significantly less acidification (p = 2.5e−4, p = 4e−5), respectively, (Wilcoxon signed-rank test) than CheRiff-expressing neurons, pH of 6.87 ± 0.27 (mean ± SD, n = 24 cells). (F) Genetic constructs for simultaneous optogenetic stimulation and voltage imaging. (G) Experimental paradigm for measuring voltage responses to optogenetic stimulation. Stimulation (blue) and measurement (red). (H) Time-course of optogenetically activated spiking in cultured neuron expressing (top) CheRiff, (middle) ChR2-3M, or (bottom) PsCatCh2.0.
Tables
Comparison of channelrhodopsin gating properties.
EPD50 is the effective power density for 50% activation. CheRiff data are from Fig. S9 and Table S4 of Hochbaum et al., 2014. CheRiff reversal potential is from Zhang et al., 2016.
| Reversal potential (mV) | ton fastest (ms) | ton at EPD50 (ms) | toff (ms) | EPD50 (mW cm−2) | Steady-state photocurrent (pA) | Ipeak/ISS | |
|---|---|---|---|---|---|---|---|
| CheRiff | 4 | 4.5 ± 0.3 | - | 16 ± 0.8 | 22 ± 4 | 1300 ± 80 | 0.65 |
| ChR2-3M (n = 4) | 16.6 ± 3.6 | 57 ± 21 | 800 ± 550 | 1950 ± 500 | 11.6 ± 8.7 | 1378 ± 618 | 1.00 |
| PsCatCh2.0 (n = 6) | 12.3 ± 4.7 | 4.2 ± 3.5 | 9.3 ± 1.2 | 17.6 ± 3.4 | 116 ±1 3 | 847 ± 359 | 0.92 |





