Trabid patient mutations impede the axonal trafficking of adenomatous polyposis coli to disrupt neurite growth
Figures
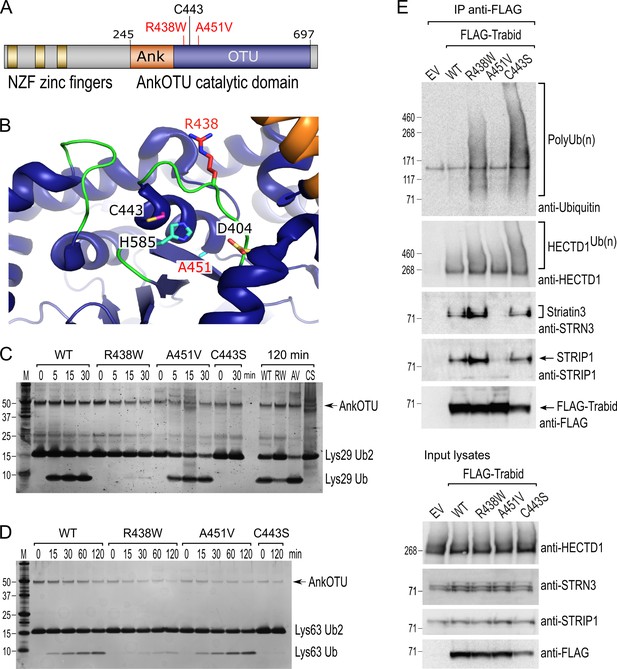
Trabid patient mutants are impaired in deubiquitylating (DUB) activity and Striatin-interacting phosphatase and kinase (STRIPAK) binding.
(A) Trabid patient mutations R438W and A451V flank on either side of the catalytic cysteine C443 in Trabid’s AnkOTU domain. (B) Residue R438 projects prominently into the catalytic cleft formed by the Ank and ovarian tumor (OTU) domain, whereas residue A451 resides at the back of the active site. The positions of the catalytic triad residues C443, H585, and D404 are shown. Crystal structure of AnkOTU domain: PDB 3ZRH. (C and D) In vitro DUB assays. Hydrolysis of Lys29-linked (C) and Lys63-linked (D) di-ubiquitin chains by purified wild-type and mutant Trabid AnkOTU proteins. The Trabid R438W AnkOTU domain exhibits strongly reduced DUB activity. M, molecular weight markers in kilodaltons. (E) Immunoprecipitation of FLAG-Trabid from the lysates of transfected HEK293T cells showed that FLAG-Trabid R438W and C443S mutants co-precipitated high amounts of endogenous polyubiquitin (PolyUb(n) smear), indicating impaired DUB activity. FLAG-Trabid A451V failed to efficiently co-precipitate Striatin3 and STRIP1, indicating loss of binding to the STRIPAK complex. All FLAG-Trabid proteins co-precipitated similar amounts of the E3 ubiquitin ligase HECTD1. EV, empty vector control.
-
Figure 1—source data 1
Uncropped images of the silver-stained gels depicted in Figure 1C and D.
- https://cdn.elifesciences.org/articles/90796/elife-90796-fig1-data1-v1.zip
-
Figure 1—source data 2
Uncropped images of the western blots depicted in Figure 1E.
Chemiluminescent signals on nitrocellulose blots were acquired using the ChemiDoc Imaging system (Bio-Rad). Included for each of the indicated antibody probe is the uncropped chemiluminescence image with a green box demarcating the region presented in Figure 1E. Also included is an image of the chemiluminescence signal merged with an image of the blotting membrane to visualize the positions of pre-stained molecular weight markers (M; HiMark, Invitrogen) relative to the chemiluminescent bands of interest.
- https://cdn.elifesciences.org/articles/90796/elife-90796-fig1-data2-v1.zip
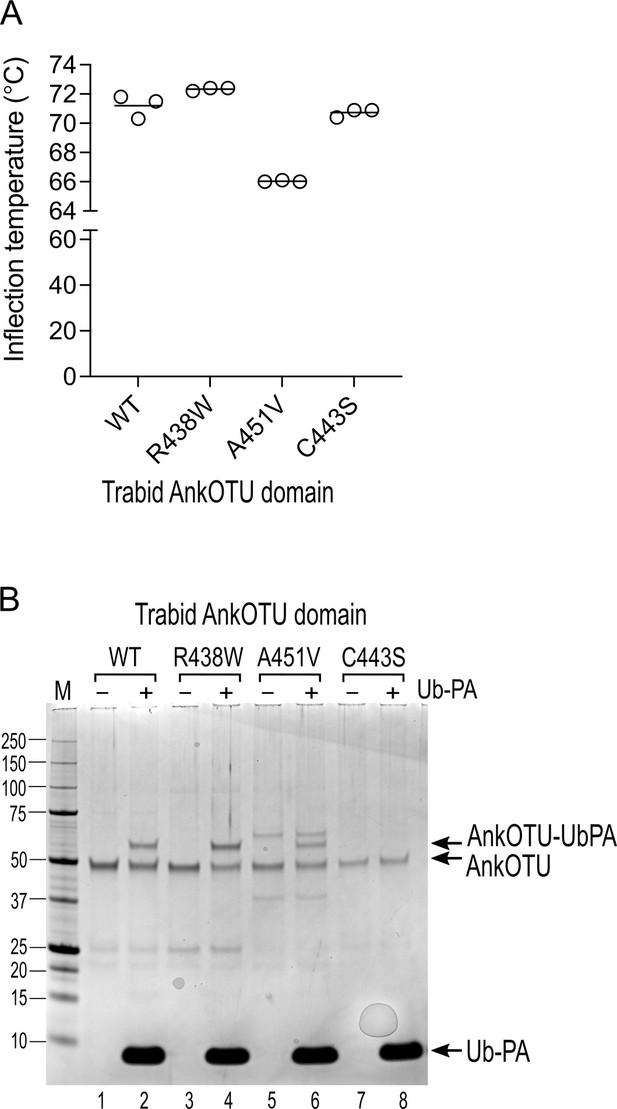
Thermal stability and catalytic cysteine reactivity of Trabid mutant AnkOTU domains.
(A) Purified Trabid AnkOTU domain proteins were subjected to thermal stability analysis using the Tycho system. The A451V mutant protein unfolded at a lower temperature than wild-type (WT) Trabid, indicating that it is a slightly less stable recombinant protein. (B) Purified Trabid AnkOTU proteins were incubated with the ubiquitin suicide probe ubiquitin-propargylamine (Ub-PA). WT, R438W, and A451V Trabid AnkOTU formed higher molecular weight species in the presence of Ub-PA, indicating that the Trabid patient variants retained a functional catalytic interaction with ubiquitin. M, molecular weight markers in kilodaltons.
-
Figure 1—figure supplement 1—source data 1
Uncropped images of the silver-stained gel depicted in Figure 1—figure supplement 1B.
- https://cdn.elifesciences.org/articles/90796/elife-90796-fig1-figsupp1-data1-v1.zip
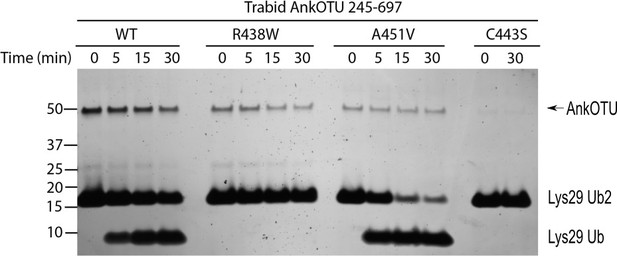
Replicate of the Lys29-linked di-ubiquitin deubiquitylating (DUB) assay using the same purified AnkOTU protein preparations used in Figure 1C.
-
Figure 1—figure supplement 2—source data 1
Uncropped images of the silver-stained gel depicted in Figure 1—figure supplement 2.
- https://cdn.elifesciences.org/articles/90796/elife-90796-fig1-figsupp2-data1-v1.zip
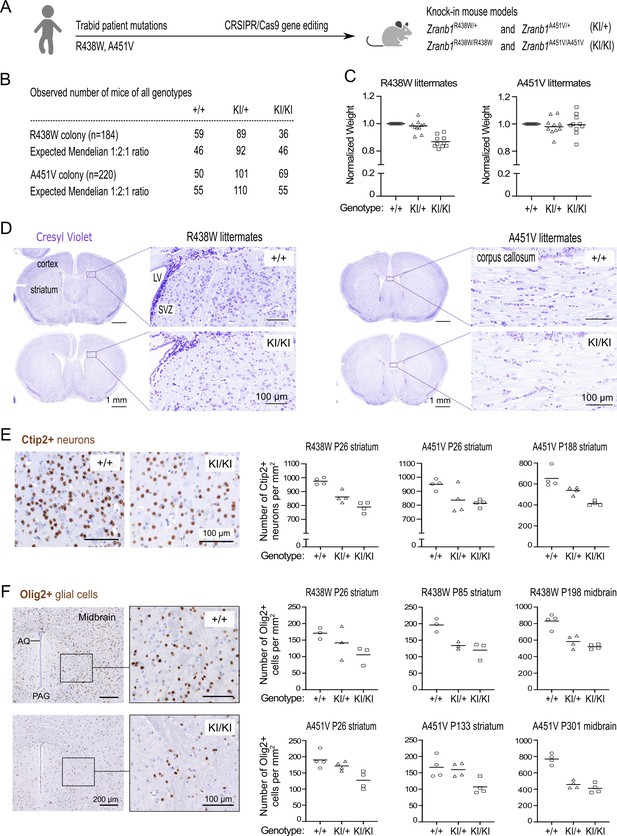
Decreased cell density in the brains of mice harboring Trabid patient mutations.
(A) Schematic of knock-in mouse strains harboring Zranb1/Trabid patient mutations generated by CRISPR/Cas9 genome editing. (B) Numbers of mice of all genotypes produced from heterozygous intercrosses in the R438W or A451V colony, respectively. (C) Normalized weight of 10 sets of weaned littermate mice (five males, five females; age range P26-P222) from each Trabid mutant colony. For each set of littermates, the weight of the wild-type (+/+) mouse was set at 1, and the weight of heterozygous (KI/+) or homozygous (KI/KI) mutant mice was expressed as a ratio of the wild-type mouse weight. (D) Coronal brain sections (Bregma ± 0.3 mm) of weaned (postnatal day P26) littermate males from the R438W and A451V mouse colonies were stained with Cresyl Violet to assess general anatomy and cellular organization in the cerebral cortex and striatum. LV, lateral ventricle; SVZ, subventricular zone. An overall decrease in Cresyl Violet staining intensity was evident in mutant homozygous (KI/KI) sections from both mouse strains compared to similar sections from the respective wild-type (+/+) littermate. (E) Ctip2 IHC revealed reduced numbers of medium spiny neurons in the striatum of mutant mice from the R438W and A451V colonies compared to wild-type littermate mice. Representative images shown are coronal brain sections of P26 male littermates from the R438W colony. Each data point represents the cell count in a randomly selected, non-overlapping 1 mm square area in the striatum from both brain hemispheres. The age and brain region of the cell count for three sets of littermate mice from the indicated mutant strain are specified. Cell numbers were quantified blinded to genotype using images processed in Fiji software. See Figure 2—figure supplement 1. (F) Olig2 IHC revealed reduced numbers of oligodendrocytes in the striatum and midbrain of homozygous mice from the R438W and A451V colonies compared to respective wild-type littermates. Representative images shown are coronal sections from the ventral midbrain of P301 female littermates from the A451V colony (AQ, cerebral aqueduct; PAQ, periaqueductal gray). Each data point represents the cell count in a randomly selected, non-overlapping 1 mm square area in the striatum or midbrain from both brain hemispheres. The age and brain region of the cell count for three sets of littermate mice from the indicated mutant strain are specified. Cell numbers were quantified blinded to genotype using images processed in Fiji. See Figure 2—figure supplement 1.
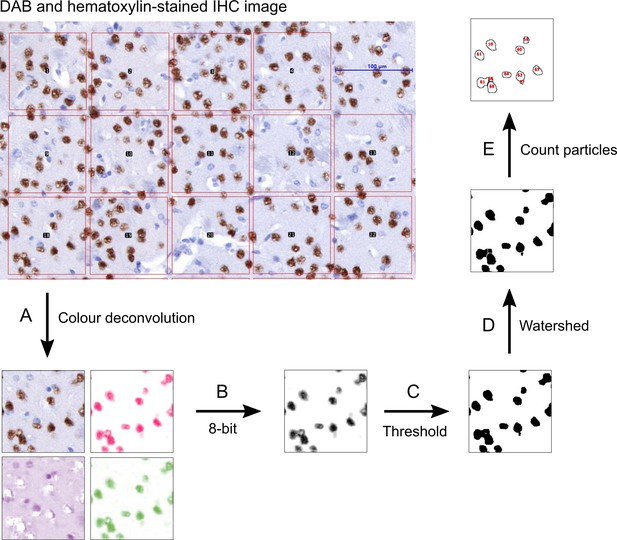
Workflow for quantification of cell numbers in Immunohistochemistry (IHC) images.
(A) A Fiji macro was generated to add squares of defined area in a DAB- and hematoxylin-stained IHC image. Color deconvolution was applied to each square to separate the color spectra. (B) The DAB-stained image (pink nuclei) is converted to 8-bit. (C) Thresholding was applied to demarcate nuclei. (D) To improve accuracy, further segmentation with the Watershed function was applied to separate partially connected nuclei. (E) Particle number was determined to provide the number of DAB-positive brown nuclei in each square. Nuclei located at the square edge are excluded from the count.
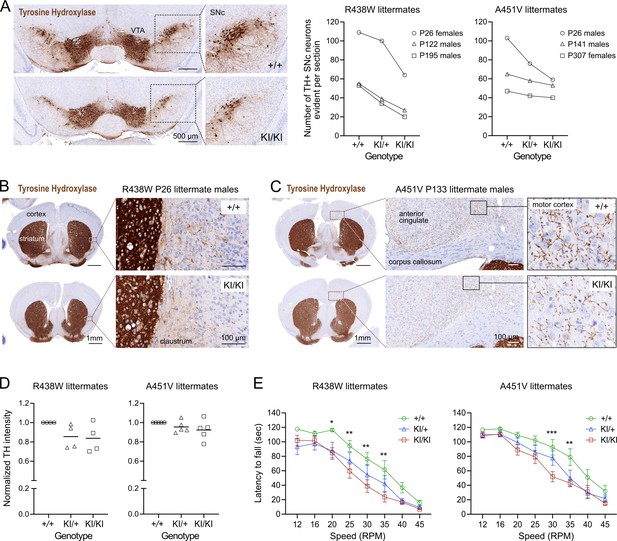
Trabid mutant mice exhibit a motor deficit consistent with reduced number of dopaminergic neurons and projections.
(A) Tyrosine Hydroxylase (TH) Immunohistochemistry (IHC) of coronal midbrain sections revealed reduced numbers of dopaminergic neurons in the substantia nigra pars compacta (SNc) of homozygous mutant mice from the Trabid R438W and A451V colonies compared to similar midbrain sections of the respective wild-type littermates. Representative images shown are midbrain sections of P26 female littermates from the R438W colony. VTA, ventral tegmental area. Numbers of TH+ SNc neurons were quantified using IHC images processed in Fiji. See Figure 2—figure supplement 1. (B) Tyrosine Hydroxylase IHC of coronal brain sections (Bregma ± 0.3 mm) of P26 littermate mice from the Trabid R438W colony revealed reduced TH staining intensity in the striatum and fewer TH+ projections in the adjacent cortex of homozygous mutant mice compared to similar regions in the wild-type littermate (magnified area). (C) Tyrosine Hydroxylase IHC of coronal brain sections (Bregma ± 0.3 mm) of P133 littermate mice from the Trabid A451V colony revealed reduced abundance and intensity of TH+ projections in several cortical regions including the primary/secondary motor cortex of homozygous mutant mice compared to similar regions in the wild-type littermate (magnified area). (D) Tyrosine Hydroxylase staining intensity in the striatum of mutant mice from the R438W and A451V colonies normalized to the staining intensity of wild-type littermate sections. IHC signals were measured as the optical density of the region of interest demarcated manually using Fiji. Each symbol represents one mouse of the indicated genotype belonging to a set of littermate mice from the R438W colony (n=4 sets: P26 males, P93 females, P307 males, P480 females) or the A451V colony (n=5 sets: P26 males, P41 females, P133 males, P188 females, P303 females). (E) Rotarod performance of 3–4 month-old littermate mice from the R438W and A451V colonies. Each data point represents the average latencies of eight mice (four males, four females), where each mouse was subjected to six trials over 3 days. Wild-type (+/+), heterozygous (KI/+), and homozygous (KI/KI) littermate mice were tested together. The experimenter was blinded to genotype. A repeated measures two-way ANOVA and Dunnett’s multiple comparisons test were applied to the data, using wild-type mice as the control group. Asterisks denote statistically significant differences between wild-type (+/+) and homozygous (KI/KI) mutant mice. *p<0.05, **p<0.01, ***p<0.001. Error bars, ± SEM.
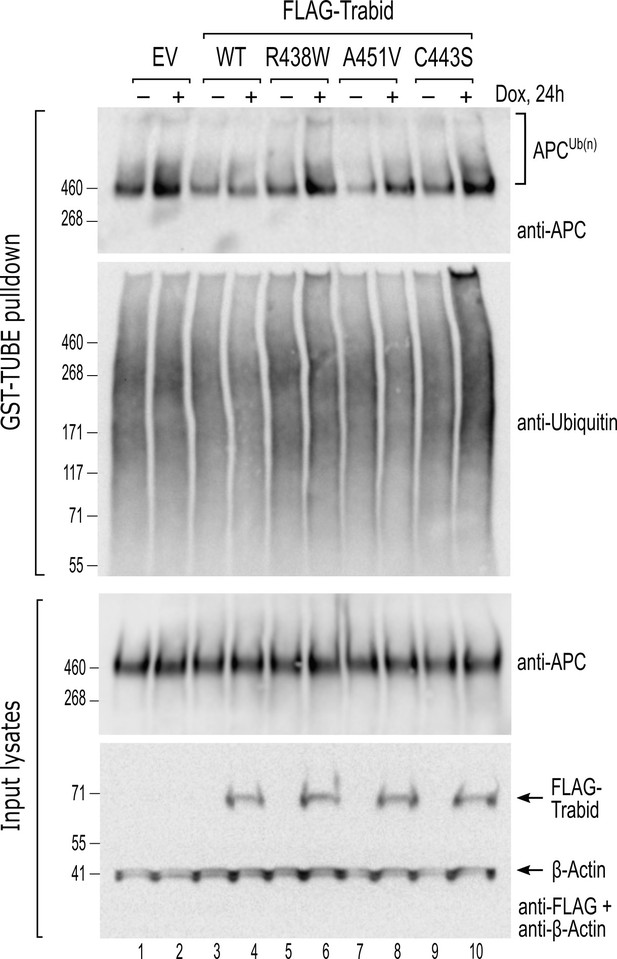
Trabid patient mutants fail to efficiently suppress adenomatous polyposis coli (APC) ubiquitylation in cells.
HEK293 cells stably transfected with doxycycline-inducible constructs for the expression of wild-type or mutant FLAG-Trabid were untreated (-) or treated with dox (+; 100 ng/mL) for 24 h. Cell lysates were subjected to a GST-TUBE pulldown to enrich the ubiquitylated proteome. Precipitated material and input lysates were processed for Western blotting using the indicated antibodies. Endogenous APC protein levels remained unchanged irrespective of FLAG-Trabid expression (input lysates), but ubiquitylated APC species were only efficiently suppressed by wild-type FLAG-Trabid expression (APCUb(n); WT +). EV, empty vector control.
-
Figure 4—source data 1
Uncropped images of western blots depicted in Figure 4.
Chemiluminescent signals on nitrocellulose blots were acquired using the ChemiDoc Imaging system (Bio-Rad). Included for each of the indicated antibody probe is the uncropped chemiluminescence image with a green box demarcating the region presented in Figure 4. Also included is an image of the chemiluminescence signal merged with an image of the blotting membrane to visualise the positions of pre-stained molecular weight markers (M; HiMark, Invitrogen) relative to the chemiluminescent bands of interest.
- https://cdn.elifesciences.org/articles/90796/elife-90796-fig4-data1-v1.zip
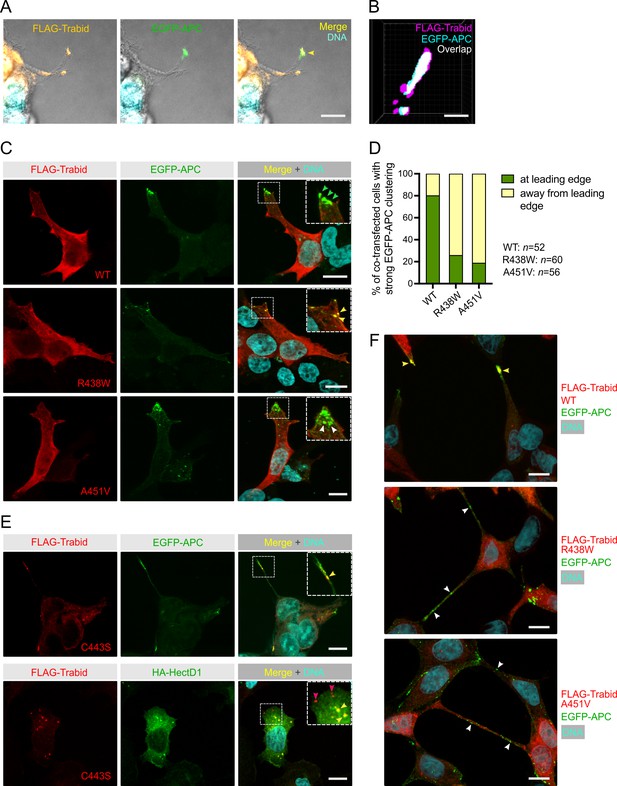
Trabid patient mutants impede the transport of EGFP-APC to the leading edge of migrating cells.
(A) Combined differential interference contrast and immunofluorescence images of HEK293T cells extending long cell processes that contain FLAG-Trabid and EGFP-APC in the cortical protrusions of co-transfected cells (arrowhead in merged image). Scale bar, 10 μm. (B) 3D-SIM super-resolution microscopy analysis of HEK293T cell tip clusters (n=5) revealed ~30% overlap of FLAG-Trabid and EGFP-APC signals at resolutions of ~100 nm in xy and 320 nm in z. Scale bar, 100 nm. (C) EGFP-APC membrane clusters at the leading edge of migrating HEK293T cells (green arrowheads) are readily detected with FLAG-Trabid wild-type (WT) co-expression. In cells expressing FLAG-Trabid patient mutants, EGFP-APC aggregate in punctate structures near the leading edge (white arrowheads), that are often sequestered together with the FLAG-Trabid R438W mutant (yellow arrowheads). Scale bar, 10 μm. (D) Quantification of strong EGFP-APC clustering at or away from the leading edge membrane of co-transfected HEK293T cells, as shown in (C). Numbers of cells analyzed from three independent transfections are shown. (E) In co-transfected HEK293T cells, DUB-dead FLAG-Trabid C443S form puncta that sequesters EGFP-APC near the tips of long cell protrusions or sequesters HA-HECTD1 in the cytoplasm (yellow arrowheads). Notably, HA-HECTD1 was not detected in FLAG-Trabid C443S punctae that form near the leading edge membrane of migrating cells (red arrowheads). Scale bar, 10 μm. (F) In near-confluent HEK293T cells 48 h post-transfection, FLAG-Trabid patient mutants induce long thin tubes that connect neigboring cells and extensive EGFP-APC aggregations are conspicuous along these tubular structures. This phenomenon was not observed in cells transfected with wild-type FLAG-Trabid. Scale bar, 10 μm.
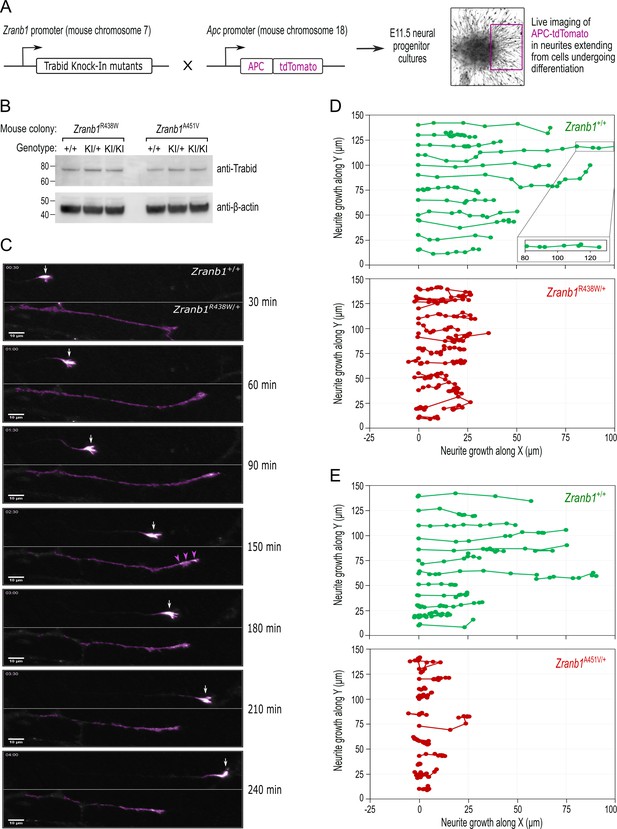
Trabid patient mutations perturb the trafficking of APC-tdTomato and the trajectory of neurite outgrowth.
(A) Schematic of outcross between knock-in mice harboring Zranb1/Trabid patient mutations and ApctdTomato mice, where the APC-tdTomato fluorescent protein is expressed under the control of the endogenous Apc gene promoter. Midbrain neural progenitors derived from E11.5 embryos of these outcrosses were used for live cell imaging of APC-tdTomato trafficking in growing neurites. (B) Western blot analysis of Trabid protein expression in E11.5 midbrain neural progenitors derived from heterozygous intercrosses in the Zranb1 R438W or A451V mouse colonies. Mutant Trabid protein levels are expressed at comparable levels to wild-type Trabid protein (compare KI/KI to +/+). (C) Live cell imaging of APC-tdTomato in neurites extending from E11.5 midbrain neural progenitors of a Zranb1R438W/+; ApctdTom/+ outcross. The frames shown are from a 4 h time-lapse portraying the movement of APC-tdTomato. APC-tdTomato accumulated in the tips of neurites with wild-type Trabid (Zranb1+/+) to generate growth cones that drive axon elongation (white arrows). In neurons with mutant Trabid (Zranb1R438W/+; Zranb1A451V/+ data not shown), APC-tdTomato intensities were broadly distributed along neurites (purple arrowheads) and the formation of growth cones was abrogated. Scale bar, 10 μm. (D and E) The movement of Apc-tdTomato in individual neurite tips was tracked to visualize the growth trajectory. Neurites extending from E11.5 midbrain neural progenitors of Zranb1R438W/+ ;ApctdTom/+ (D) or Zranb1A451V/+; ApctdTom/+ (E) outcrosses were analyzed. APC-tdTomato fluorescence synchronized with the movement of a membrane dye during neurite extension and retraction (data not shown) and was, therefore, used as a proxy for neurite growth along an arbitrary X and Y plane. Each data point represents the location of APC-tdTomato at 15 min intervals of a 4 h time-lapse. Neurite tracking data were compiled from the imaging of neurite outgrowth in three independent neurosphere cultures established from E11.5 embryos of three independent Zranb1R438W/+; ApctdTom/+, or Zranb1A451V/+; ApctdTom/+ outcrosses. All image acquisition, processing, and analyses were performed blinded to genotype.
-
Figure 6—source data 1
Uncropped images of western blots depicted in Figure 6B.
Chemiluminescent signals on polyvinylidene fluoride (PVDF) blots were acquired using the ChemiDoc Imaging system (Bio-Rad). Included is the uncropped chemiluminescence image with green boxes demarcating the regions presented in Figure 6B. Also included is the chemiluminescence image merged with an image of the blotting membrane to visualize the positions of pre-stained molecular weight protein standards (M; Novex Sharp, Invitrogen) relative to the chemiluminescent bands of interest.
- https://cdn.elifesciences.org/articles/90796/elife-90796-fig6-data1-v1.zip
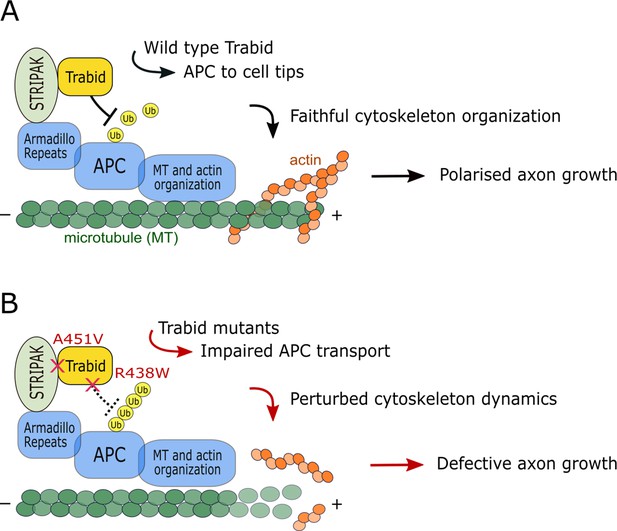
Model of Trabid’s mechanism of action in axon growth and guidance.
(A) Striatin-interacting phosphatase and kinase (STRIPAK) recruits Trabid to deubiquitylate adenomatous polyposis coli (APC) (and possibly other substrates). This promotes efficient APC accumulation at microtubule plus ends to coordinate the actin and microtubule cytoskeleton dynamics that drive directional cell migration and polarized axon growth. (B) In cells expressing Trabid mutants that are DUB-impaired (R438W) or STRIPAK-binding-deficient (A451V), APC becomes persistently modified with ubiquitin chains that retard its transport to cell tips. This impairs cytoskeleton organization which leads to defective axon elongation and cell migration. Future studies will aim to identify the mechanism of Trabid recruitment to STRIPAK, the ubiquitin-modified residue(s) on APC (and possibly other substrates), and the composition of polyubiquitin linkages on substrates including the Lys29- and Lys33-linked atypical chain types preferentially targeted by Trabid.





