The FAM104 proteins VCF1/2 promote the nuclear localization of p97/VCP
Figures

VCF1/2 bind to p97.
(A) Schematic overview of human VCF1 and VCF2 isoforms isolated in a yeast two-hybrid screen. Relevant amino acid residue numbers are shown, and conserved N- and C-terminal sequence motifs are indicated by purple and orange boxes, respectively. Sequence identity outside these boxes is indicated by different shades of gray (light, medium, dark). Internal deletions in VCF1 isoform 2 and VCF2 isoform 4 are indicated by thin lines. (B) Multiple-sequence alignment showing representative members of the FAM104 family. Regions with predicted alpha-helical secondary structure are indicated at the top by 'H'. The most highly conserved residues in the C-terminal sequence motif are boxed in red. Other conserved residues are boxed in black or gray, according to the degree of conservation. Numbers in square brackets indicate the length of insertions. For human VCF1 and VCF2, the sequences of isoforms 2 and 4, respectively, are shown, and numbers in round brackets indicate a 21-residue insertion present in isoforms 1 and 5 of VCF1 and a 1-residue insertion present in isoform 3 of VCF2, respectively. (C) Yeast two-hybrid analysis. Yeast PJ69-4a reporter cells transformed with the indicated combinations of bait (BD-) and prey (AD-) plasmids were spotted onto agar plates containing synthetic complete medium lacking uracil and leucine (control) or uracil, leucine, and histidine (-His). Growth was monitored after 3 d. (D) Glutathione sepharose pulldown assay using wild-type p97 and GST fusions of the indicated VCF1/2 proteins. Binding of p97 was analyzed by SDS-PAGE followed by Coomassie brilliant blue staining. (E, F) Glutathione sepharose pulldown assays as in (D), using the indicated p97 variants and GST fusions of VCF1 isoform 5 (E) and VCF2 isoform 3 (F), respectively. Arrowheads mark the position of the p97 N domain.
-
Figure 1—source data 1
Related to Figure 1D–F.
- https://cdn.elifesciences.org/articles/92409/elife-92409-fig1-data1-v2.zip
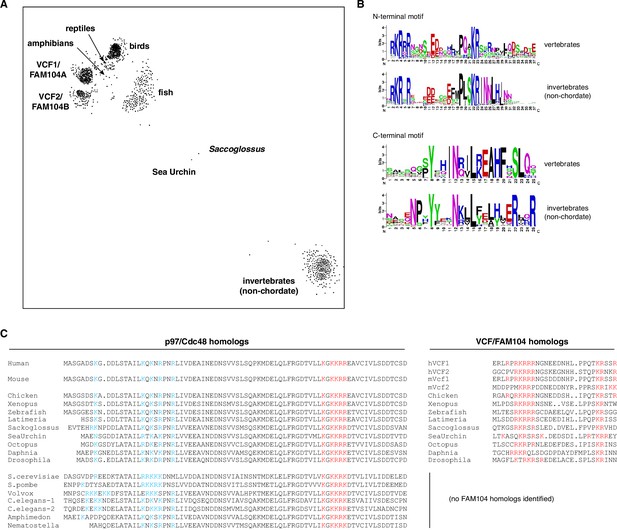
Evolutionary conservation of FAM104 proteins.
(A) CLANS diagram of the FAM104 family. The sequence relationship between different FAM104-related proteins is visualized by similarity-based clustering using the CLANS software. Each point represents one protein; point-to point distances are approximations of sequence divergence. Selected clusters are annotated. (B) Consensus sequences of conserved motifs. Separate sequence logos for the N- and C-terminal motifs were generated for the vertebrate and invertebrate clusters, respectively. (C) Comparison of the evolutionary conservation of predicted classical nuclear localization signals (cNLSs) in p97/Cdc48 (left multiple-sequence alignment) with the occurrence of VCF/FAM104 homologs (right multiple-sequence alignment). Positively charged residues in the consistently predicted cNLSs of p97/Cdc48 and FAM104 proteins are shown in red, whereas those in the region of the functional bipartite cNLS of S. cerevisiae Cdc48 are shown in blue. Note the lack of conservation of this bipartite cNLS in organisms possessing FAM104 homologs.

FAM104 proteins bind to p97 via their C-terminal helix.
(A) Schematic overview of the C-terminal truncations of VCF1 isoform 5 used for yeast two-hybrid analysis. Labeling as in Figure 1A. (B) Yeast two-hybrid analysis of p97 binding to the C-terminally truncated VCF1 isoform 5 variants shown in (A). (C) Expression levels of the two-hybrid fusion proteins in (B) were analyzed by western blot (WB) using antibodies against p97 and the Gal4 transactivation domain (Gal4-TA). The asterisk in the p97 blot marks a cross-reactivity with endogenous Cdc48. (D) Glutathione sepharose pulldown assay using wild-type p97 and GST fusions of the indicated full-length or C-terminally truncated VCF1/2 proteins. (E) Streptavidin sepharose pulldown assay using the biotinylated peptide CQGLYFHINQTLREAHFHSLQHRG spanning the conserved C-terminal alpha-helix and flanking residues of VCF1 (residues C180–G203 in isoform 1) and the indicated p97 variants. p97 binding to the immobilized peptide was analyzed by SDS-PAGE, followed by Coomassie brilliant blue staining. Arrowheads mark the position of the p97 N domain. (F, G) AlphaFold Multimer model of the C-terminal alpha-helix of VCF1 (turquoise) bound to the N domain of p97. (F) Overview showing binding to the subdomain cleft of the N domain. (G) Close-up view showing the interaction of the four most highly conserved residues with the N domain (green). Residue numbers refer to isoform 1 of VCF1. (H) Glutathione sepharose pulldown assay using wild-type p97 and GST fusions of the indicated full-length (wildtype, NL->AA, NL->RR) or C-terminally truncated (Cdel26) variants of VCF1 isoform 5. NL->AA, N188A/L191A double mutant; NL->RR, N188R/L191R double mutant.
-
Figure 2—source data 1
Related to Figure 2C.
- https://cdn.elifesciences.org/articles/92409/elife-92409-fig2-data1-v2.zip
-
Figure 2—source data 2
Related to Figure 2D, E, and H.
- https://cdn.elifesciences.org/articles/92409/elife-92409-fig2-data2-v2.zip

The p97 binding mode is conserved in invertebrate FAM104 proteins.
AlphaFold Multimer model of the C-terminal alpha-helix of Drosophila melanogaster CG14229 (purple) superimposed on the model of VCF1 (turquoise) bound to the N domain of p97 (green) shown in Figure 2G. The modeled N domain of D. melanogaster TER94 was almost identical to that of p97 and was omitted for clarity. The four highly conserved residues contacting the N domain are labeled.
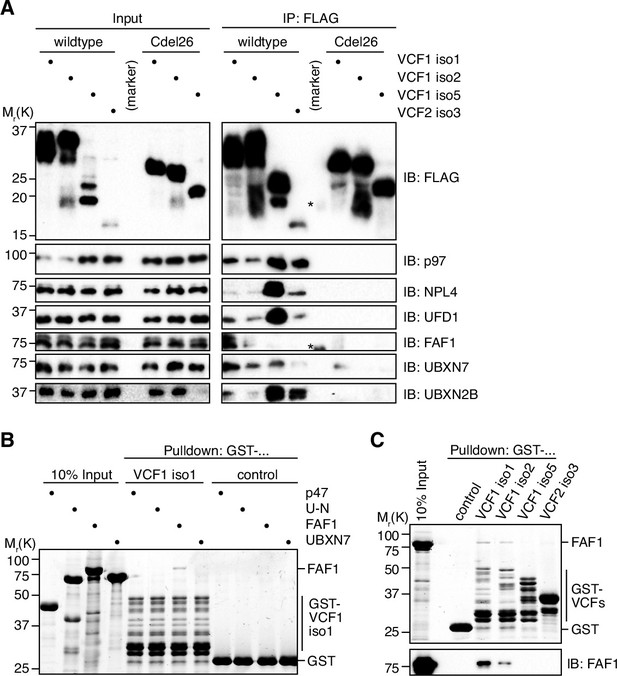
VCF1/2 form complexes with p97 and several p97 cofactors in cells.
(A) HEK293T cells ectopically expressing the indicated N-terminally FLAG-epitope-tagged wildtype or C-terminally truncated (Cdel26) VCF1/2 proteins were subjected to anti-FLAG immunoprecipitation (IP). Input and IP samples were immunoblotted for the FLAG epitope tag, p97, and the indicated p97 cofactors. The central empty lane had been loaded with a marker. The asterisks label marker bands cross-reactive with the FLAG and FAF1 antibody, respectively. (B) Glutathione sepharose pulldown assay, using the indicated p97 cofactors and GST-VCF1 isoform 1; U-N, UFD1-NPL4. (C) Glutathione sepharose pulldown assay, using FAF1 and GST fusions of the indicated VCF1/2 proteins. Upper panel: 2,2,2-trichloroethanol-stained gel; lower panel: immunoblot with FAF1 antibody.
-
Figure 3—source data 1
Related to Figure 3A.
- https://cdn.elifesciences.org/articles/92409/elife-92409-fig3-data1-v2.zip
-
Figure 3—source data 2
Related to Figure 3B.
- https://cdn.elifesciences.org/articles/92409/elife-92409-fig3-data2-v2.zip
-
Figure 3—source data 3
Related to Figure 3C.
- https://cdn.elifesciences.org/articles/92409/elife-92409-fig3-data3-v2.zip
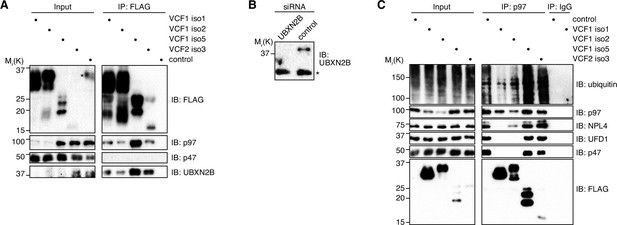
VCF1/2 associate with several p97-cofactor complexes.
(A) HEK293T cells ectopically expressing the indicated N-terminally FLAG-epitope-tagged VCF1/2 proteins or control cells were subjected to anti-FLAG immunoprecipitation (IP). Input and IP samples were immunoblotted for the FLAG epitope tag, p97, p47, and UBXN2B. In the input lane of the control sample, an asterisk in the FLAG immunoblot marks a spill-over from the strong signal in the neighboring IP lane. (B) Immunoblot of lysates from HeLa cells transfected with siRNAs targeting UBXN2B or with non-targeting control siRNAs, demonstrating the specificity of the UBXN2B antibody used in panel (A) and in Figure 3A. The asterisk indicates a nonspecific, cross-reactive band. (C) Lysates from HEK293T cells ectopically expressing the indicated N-terminally FLAG-epitope-tagged VCF1/2 proteins or control cells were subjected to IP of endogenous p97 or control IP (IgG). Input and IP samples were immunoblotted using the indicated antibodies.
-
Figure 3—figure supplement 1—source data 1
Related to Figure 3—figure supplement 1A–C.
- https://cdn.elifesciences.org/articles/92409/elife-92409-fig3-figsupp1-data1-v2.zip
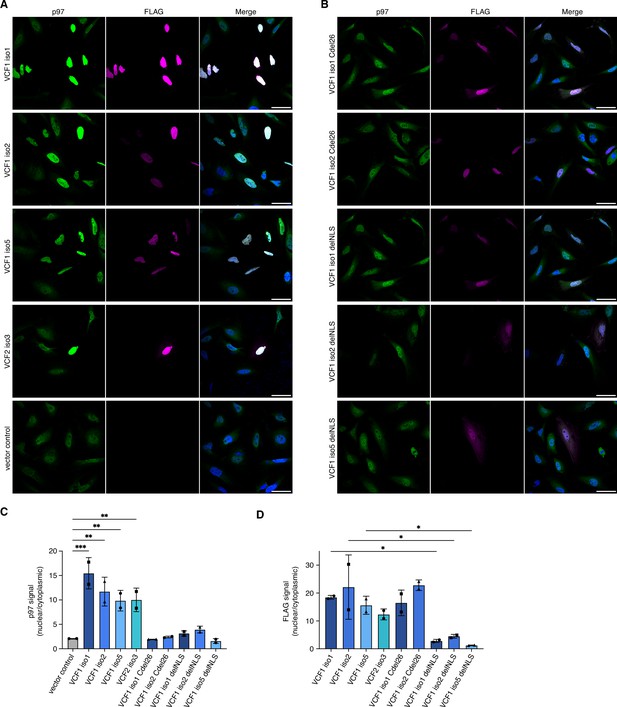
Ectopic expression of VCF1/2 increases nuclear p97 levels.
(A) HeLa cells ectopically expressing full-length, N-terminally FLAG epitope-tagged VCF isoforms 1, 2, and 5, VCF2 isoform 3, and empty vector control, respectively, were analyzed by confocal immunofluorescence microscopy using antibodies against endogenous p97 and the FLAG epitope. Scale bars, 50 µm. All images were taken with identical acquisition settings and processed identically. (B) HeLa cells ectopically expressing N-terminally FLAG epitope-tagged VCF1 isoforms 1 and 2 lacking the C-terminal conserved helix (Cdel26) or the classical nuclear localization signal (cNLS) (delNLS), and VCF1 isoform 5 lacking the cNLS, were analyzed as in (A). Scale bars, 50 µm. (C) Quantification of the ratio of nuclear to cytoplasmic p97 signals in panels (A) and (B). Except for the vector control where all imaged cells were included (>80 cells per replicate and condition), only transfected cells (as judged by the FLAG channel) were included in the quantification. Shown is the mean ± SD; n = 2 biological replicates with 15–30 transfected cells per replicate and condition; one-way ANOVA. *p<0.05; **p<0.01; ***p<0.001; the differences between the Cdel26 and delNLS constructs and the vector control are all not significant (p>0.87). (D) Quantification of the ratio of nuclear to cytoplasmic FLAG signals in panels (A) and (B) was performed as described in (C).
-
Figure 4—source data 1
Related to Figure 4C and D.
- https://cdn.elifesciences.org/articles/92409/elife-92409-fig4-data1-v2.zip
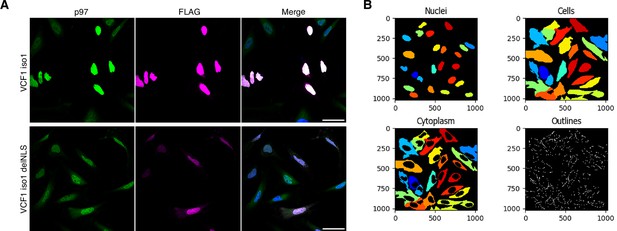
VCF1/2 promote the nuclear localization of p97.
(A) Comparison of the images from Figure 4A and B for full-length VCF1 isoform 1 and its classical nuclear localization signal (cNLS)-deleted variant. The images were processed for higher signal intensities in order to facilitate the visual inspection of cytoplasmic signals. Scale bars, 50 µm. (B) Example of the cell segmentation results obtained with CellProfiler for nuclear and cytoplasmic signals. See ‘Materials and methods’ section for details.

Ectopic expression of VCF1 isoforms 1 and 2 promotes the association of p97 with chromatin.
(A) HEK293T cells were transfected with plasmids encoding the indicated N-terminally FLAG-epitope-tagged VCF1 proteins or with empty vector (control). Total protein extracts were prepared by direct boiling part of the cells in SDS-PAGE sample buffer (SDS). The remaining cells were processed to cytoplasmic, soluble nuclear (nucleoplasmic), and chromatin fractions as indicated. Tubulin and ubiquitylated histone H2B (Ub-H2B) served as markers for the cytoplasmic and chromatin fractions, respectively, whereas the Coomassie staining of the membrane (total protein stain) served as loading control. (B) Fractionation of lysates from HEK293T cells ectopically expressing VCF1 isoform 1 or control cells using an alternative protocol including benzonase treatment that allows to distinguish solubilized chromatin-associated proteins (chromatin) from insoluble proteins (insoluble).
-
Figure 5—source data 1
Related to Figure 5A and B.
- https://cdn.elifesciences.org/articles/92409/elife-92409-fig5-data1-v2.zip
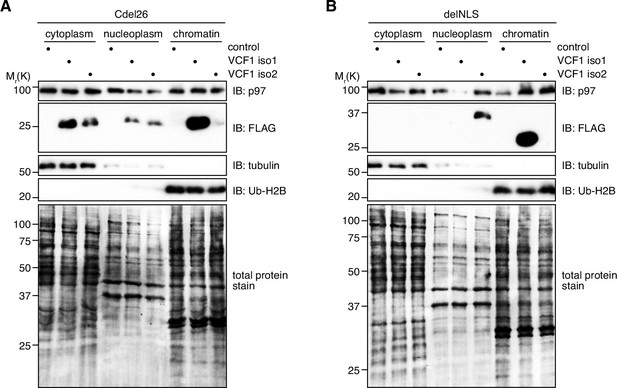
Control fractionations of cells ectopically expressing Cdel26 and delNLS variants of VCF1 isoforms 1 and 2.
(A) Fractionation into cytoplasmic, nucleoplasmic, and chromatin fractions of lysates from HEK293T cells ectopically expressing the FLAG-tagged, p97 binding-deficient Cdel26 variants of VCF1 isoforms 1 and 2 or control cells. Tubulin and ubiquitylated histone H2B (Ub-H2B) served as markers for the cytoplasmic and chromatin fractions, respectively, whereas the Coomassie staining of the membrane (total protein stain) served as loading control. (B) Same as (C), but using HEK293T cells ectopically expressing the delNLS variants of VCF1 isoforms 1 and 2.
-
Figure 5—figure supplement 1—source data 1
Related to Figure 5—figure supplement 1A and B.
- https://cdn.elifesciences.org/articles/92409/elife-92409-fig5-figsupp1-data1-v2.zip

Deletion of VCF1/2 reduces nuclear p97 levels.
(A) Control, VCF1 single-knockout (KO) and VCF1/2 double-knockout (DKO) HeLa cell pools were analyzed by confocal immunofluorescence microscopy using an antibody against endogenous p97. Scale bars, 50 µm. (B) HeLa cells were transfected with non-targeting or VCF1-specific siRNAs at the indicated final concentrations for 72 hr and analyzed as in (A). Scale bars, 50 µm. (C, D) Quantification of the ratios of nuclear to cytoplasmic p97 signals in (A) and (B), respectively. Shown is the mean ± SD; n = 3 biological with ≥80 cells per replicate and condition; *p<0.05; **p<0.01; unpaired, two-tailed Student´s t-test. (E) HeLa cell pools expressing wild-type p97 (FLAG-p97) or p97 carrying an N-terminal fusion of the SV40 cNLS (NLS-FLAG-p97) under the control of a doxycycline-inducible promoter, or empty vector control cells, were transfected with VCF1-targeting or non-targeting siRNAs, induced with doxycycline (Dox) for 40 hr where indicated, stained with antibodies detecting endogenous p97 or FLAG, and subjected to high-content microscopy, followed by automated image analysis of nuclear to cytoplasmic p97 signals. (F) Quantification of the ratios of nuclear to cytoplasmic p97 signals in (E); n = 2 biological replicates (performed in three technical replicates each) with ≥3000 cells per biological replicate and cell pool, shown are mean ± SD; two-way ANOVA; *p<0.05; **p<0.01; ns, not significant.
-
Figure 6—source data 1
Related to Figure 6C, D, and F.
- https://cdn.elifesciences.org/articles/92409/elife-92409-fig6-data1-v2.zip
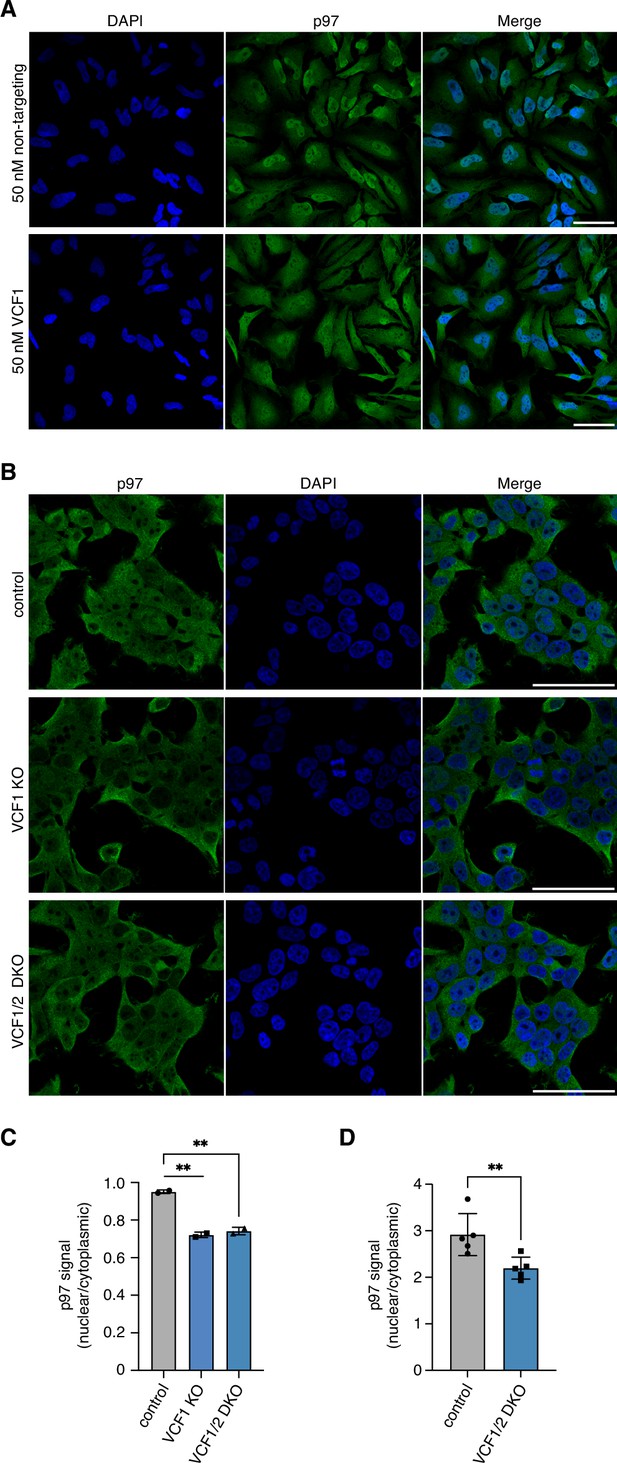
De(p)letion of VCF1/2 reduces nuclear p97 levels.
(A) Z stack images of control- or VCF1-depleted HeLa cells stained for endogenous p97. Shown are maximum intensity projections of stacks consisting of eight z planes with 2.5 µm distance. Scale bars, 50 µm. (B) Control, VCF1 single-knockout (KO) and VCF1/2 double-knockout (DKO) HEK293T cell pools were analyzed by confocal immunofluorescence microscopy using an antibody against endogenous p97. Scale bars, 50 µm. (C) Quantification of nuclear to cytoplasmic p97 signals detected in (B). Shown is the mean ± SD; n = 2 biological replicates with ≥100 cells per replicate and condition; unpaired, two-tailed Student´s t-test; **p<0.01. (D) High-content microscopy of control and VCF1/2 DKO HeLa cell pools stained with an antibody detecting endogenous p97, followed by automated image analysis of nuclear to cytoplasmic signals. n = 5 biological replicates with ≥3000 cells per replicate and cell pool; paired, two-tailed Student´s t-test; **p<0.01.
-
Figure 6—figure supplement 1—source data 1
Related to Figure 6—figure supplement 1C and D.
- https://cdn.elifesciences.org/articles/92409/elife-92409-fig6-figsupp1-data1-v2.zip

Deletion of VCF1/2 causes reduced growth and hypersensitivity to p97 inhibition.
(A) HeLa VCF1/2 double-knockout (DKO) and control cells were seeded in 96-well plates, cultivated for 8 d, stained with Hoechst dye and automatically counted using a high-content microscopy platform. n = 6 (two biological replicates in three technical replicates each), shown are mean ± SD; unpaired two-tailed Student´s t-test, **p<0.01. (B) Relative growth after 8 d was determined at the indicated concentrations of the p97 inhibitor CB-5083. Values were normalized separately for control and knockout cells to the respective mean value at 0 nM CB-5083. A two-way ANOVA was performed using the normalized data. n = 6 (two biological replicates in three technical replicates each); *p<0.05; **p<0.01; ns, not significant. (C) Same as in (B), but cells were pretreated with 1 uM camptothecin for 70 min before starting the CB-5083 treatment. (D) VCF1 mRNA levels of HeLa cells grown in the presence or absence of 0.8 µM CB-5083 for 24 hr were determined by qRT-PCR using primer pairs specifically amplifying isoforms 1 and 2 or isoform 2 only, respectively. n = 3 biological replicates, shown are mean ± SD; two-tailed, one-sample t-test; *p<0.05. (E) Control and VCF1/2 DKO HeLa cell pools were treated with CB-5083 as indicated or left untreated, followed by confocal immunofluorescence microscopy, using an antibody against endogenous p97. Scale bars, 50 µm. (F) Quantification of the ratios of nuclear to cytoplasmic p97 signals in (E); n = 4, shown are mean ± SD; two-way repeated-measures ANOVA; *p<0.05; **p<0.01; ***p<0.001; ****p<0.0001.
-
Figure 7—source data 1
Related to Figure 7A–D and F.
- https://cdn.elifesciences.org/articles/92409/elife-92409-fig7-data1-v2.zip
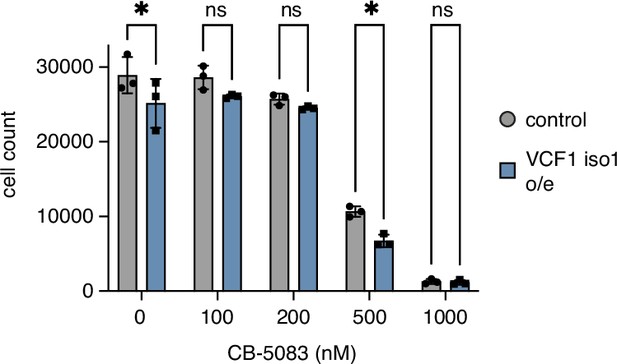
Overexpression of VCF1 isoform 1 causes moderate growth defects.
HEK293T cells ectopically expressing N-terminally FLAG epitope-tagged VCF1 isoform 1 (VCF1 iso1 o/e) or empty vector control, respectively, were seeded in 96-well plates, grown for 24 hr in the presence of the indicated concentrations of CB-5083, stained with Hoechst dye and automatically counted using a high-content microscopy platform. n = 3, shown are mean ± SD; unpaired, two-tailed Student´s t-test; *p<0.05; ns, not significant.
-
Figure 7—figure supplement 1—source data 1
Related to Figure 7—figure supplement 1.
- https://cdn.elifesciences.org/articles/92409/elife-92409-fig7-figsupp1-data1-v2.zip
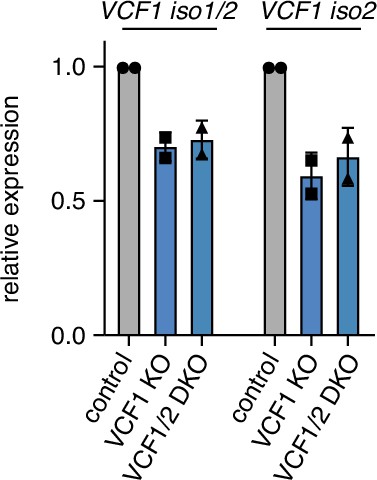
VCF1 expression levels in VCF1 and VCF1/2 knockout cell pools.
Steady-state VCF1 mRNA levels of control, VCF1 single-knockout (KO), and VCF1/2 double-knockout (DKO) HeLa cell pools were determined by qRT-PCR using primer pairs specifically amplifying isoforms 1 and 2 or isoform 2 only, respectively. n = 2, shown are mean ± SD.
-
Figure 7—figure supplement 2—source data 1
Related to Figure 7—figure supplement 2.
- https://cdn.elifesciences.org/articles/92409/elife-92409-fig7-figsupp2-data1-v2.zip
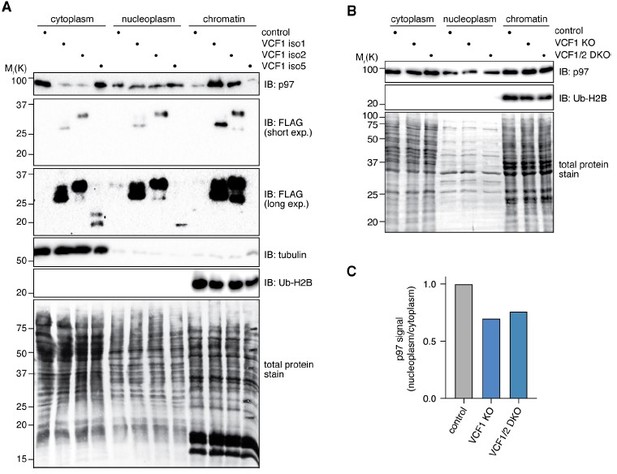
Fractionation of cell lysates after ectopic expression or deletion of VCF1/2.
(A) HEK293T cells ectopically expressing the indicated VCF1 isoforms were processed to cytoplasmic, soluble nuclear (nucleoplasmic) and chromatin fractions and analyzed by immunoblotting as indicated. (B) As in (A), but using VCF1 single and VCF1/2 double knockout cell pools transfected with siRNA targeting VCF1. (C) Quantification of the ratio of nucleoplasmic to cytoplasmic p97 levels in (B). p97 band intensities were normalized to the whole lane intensity of the total protein stain (Coomassie) of the respective lane. The p97 intensities in the nucleoplasmic fraction were divided by the intensity in the cytoplasmic fraction, and this ratio was set to 1 for the control cells.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Strain, strain background (Escherichia coli) | XL1 Blue | Stratagene | Cat# 200249 | |
| Strain, strain background (E. coli) | XL10 Gold | Agilent | Cat# 200516-4 | |
| Strain, strain background (E. coli) | BL21(DE3) pRIL | Agilent | Cat# 230245 | |
| Strain, strain background (Saccharomyces cerevisiae) | PJ69-4A | James et al., 1996 | N/A | |
| Cell line (Homo sapiens) | HeLa | ATCC | CCL-2 | |
| Cell line (H. sapiens) | HEK293T | ATCC | CRL-3216 | |
| Cell line (H. sapiens) | HeLa VCF1 knockout cell pool | This study | N/A | See ‘Materials and methods,’ section ‘Mammalian cell culture’ |
| Cell line (H. sapiens) | HeLa VCF1/2 double-knockout cell pool | This study | N/A | |
| Cell line (H. sapiens) | HeLa non-human-target control cell pool | This study | N/A | |
| Cell line (H. sapiens) | HeLa pINDUCER20 control cell pool | This study | N/A | |
| Cell line (H. sapiens) | HeLa pINDUCER20 p97 wildtype cell pool | This study | N/A | |
| Cell line (H. sapiens) | HeLa pINDUCER20 SV40NLS-p97 cell pool | This study | N/A | |
| Cell line (H. sapiens) | HEK293T VCF1 knockout cell pool | This study | N/A | |
| Cell line (H. sapiens) | HEK293T VCF1/2 double-knockout cell pool | This study | N/A | |
| Cell line (H. sapiens) | HEK293T non-human-target control cell pool | This study | N/A | |
| Antibody | Anti-alpha-tubulin (mouse monoclonal) | Sigma-Aldrich | Cat# T5168, RRID:AB_477579 | WB 1:2500 |
| Antibody | Anti-GAL4-TA (mouse monoclonal) | Santa Cruz | Sc-1663 | WB 1:500 |
| Antibody | Anti-FLAG (rabbit polyclonal) | Thermo Fisher Scientific | Cat# PA1-984B, RRID:AB_347227 | WB 1:2000 |
| Antibody | Anti-FLAG (mouse monoclonal) | Sigma-Aldrich | Cat# F1804 | WB 1:2000 IF 1:200 |
| Antibody | Anti-VCP (rabbit polyclonal) | Bethyl Laboratories | Cat# A300-589A, RRID:AB_495512 | WB 1:5000 IF 1:300 |
| Antibody | Anti-VCP (mouse monoclonal) | Santa Cruz | Cat# sc-57492, RRID:AB_793927 | IF 1:50 |
| Antibody | Anti-FAF1 (rabbit polyclonal) | Max Planck Institute of Biochemistry, animal house | AB65 | WB 1:10,000 |
| Antibody | Anti-UBXN7 (rabbit polyclonal) | Sigma-Aldrich | Cat# HPA049442 | WB 1:1000 |
| Antibody | Anti-NPL4 (rabbit polyclonal) | Sigma-Aldrich | Cat# HPA021560 | WB 1:1000 |
| Antibody | Anti-UFD1 (rabbit polyclonal) | Proteintech | Cat# 10615 | WB 1:2000 |
| Antibody | Anti-Ubiquityl-Histone H2B (rabbit monoclonal) | Cell Signaling | Cat# 5546 | WB 1:1000 |
| Antibody | Anti-UBXN2B (rabbit polyclonal) | Lee et al., 2018 | N/A | WB 1:1000 |
| Antibody | Anti-UBXD1 (rabbit polyclonal) | Novus | NBP2-57653 | WB 1:1000 |
| Antibody | Anti-MCM7 (rabbit polyclonal) | Proteintech | 11225-1-AP | WB 1:1000 |
| Antibody | Anti-MYC (rabbit monoclonal) | Biocenter, University of Würzburg, made in-house | N/A | WB 1:2000 |
| Antibody | Anti-mouse IgG HRP (goat polyclonal) | Dianova (Jackson ImmunoResearch) | Cat# 115-035-003, RRID:AB_10015289 | 1:7500 |
| Antibody | Anti-rabbit IgG HRP (goat polyclonal) | Dianova (Jackson ImmunoResearch) | Cat# 111-035-045, RRID:AB_2337938 | 1:7500 |
| Antibody | Alexa Fluor 488 goat anti-rabbit IgG (H+L) | Thermo Fisher Scientific | Cat# A-11070, RRID:AB_142134 | 1:500 |
| Antibody | Alexa Fluor 488 goat anti-mouse IgG (H+L) | Thermo Fisher Scientific | Cat# A-11017, RRID:AB_143160 | 1:500 |
| Antibody | Alexa Fluor 594 goat anti-rabbit IgG (H+L) | Thermo Fisher Scientific | Cat# A-11072, RRID:AB_142057 | 1:500 |
| Antibody | Alexa Fluor 594 goat anti-mouse IgG (H+L) | Thermo Fisher Scientific | Cat# A-11020, RRID:AB_141974 | 1:500 |
| Recombinant DNA reagent | psPAX2 | Addgene | Cat# 12260 | |
| Recombinant DNA reagent | pMD2.G | Addgene | Cat# 12259 | |
| Recombinant DNA reagent | pCMV-Tag2B | Agilent | Cat# 211172 | |
| Recombinant DNA reagent | pCMV VCF1 isoform 1 | This study | pAB2225 | See ‘Materials and methods,’ section ‘Plasmids’ |
| Recombinant DNA reagent | pCMV VCF1 isoform 2 | This study | pAB2208 | |
| Recombinant DNA reagent | pCMV VCF1 isoform 5 | This study | pAB2223 | |
| Recombinant DNA reagent | pCMV VCF2 isoform 3 | This study | pAB2277 | |
| Recombinant DNA reagent | pCMV VCF1 isoform 1 Cdel26 | This study | pAB2280 | |
| Recombinant DNA reagent | pCMV VCF1 isoform 2 Cdel26 | This study | pAB2275 | |
| Recombinant DNA reagent | pCMV VCF1 isoform 5 Cdel26 | This study | pAB2276 | |
| Recombinant DNA reagent | pCMV VCF1 isoform 1 delNLS | This study | pAB2230 | |
| Recombinant DNA reagent | pCMV VCF1 isoform 2 delNLS | This study | pAB2231 | |
| Recombinant DNA reagent | pCMV VCF1 isoform 5 delNLS | This study | pAB2227 | |
| Recombinant DNA reagent | pGEX-4T1 | Cytiva | Cat# 28954549 | |
| Recombinant DNA reagent | pGEX-4T1 VCF1 isoform 1 | This study | pAB2212 | See ‘Materials and methods,’ section ‘Plasmids’ |
| Recombinant DNA reagent | pGEX-4T1 VCF1 isoform 2 | This study | pAB2195 | |
| Recombinant DNA reagent | pGEX-4T1 VCF1 isoform 5 | This study | pAB2213 | |
| Recombinant DNA reagent | pGEX-4T1 VCF2 isoform 3 | This study | pAB2298 | |
| Recombinant DNA reagent | pGEX-4T1 VCF1 isoform 1 Cdel26 | This study | pAB2295 | |
| Recombinant DNA reagent | pGEX-4T1 VCF1 isoform 2 Cdel26 | This study | pAB2296 | |
| Recombinant DNA reagent | pGEX-4T1 VCF1 isoform 5 Cdel26 | This study | pAB2297 | |
| Recombinant DNA reagent | pGEX-4T1 VCF2 isoform 3 Cdel26 | This study | pAB2299 | |
| Recombinant DNA reagent | pGEX-4T1 VCF1 isof. 5 NL->AA | This study | pAB3163 | |
| Recombinant DNA reagent | pGEX-4T1 VCF1 isof. 5 NL->RR | This study | pAB3162 | |
| Recombinant DNA reagent | mini-pRSETA | Perrett et al., 1999 | N/A | |
| Recombinant DNA reagent | mini-pRSETA UBXN7 | This study | pAB2008 | See ‘Materials and methods,’ section ‘Plasmids’ |
| Recombinant DNA reagent | mini-pRSETA p47 | Allen et al., 2006 | pAB356 | |
| Recombinant DNA reagent | pQE30 FAF1 | Jensen et al., 2001 | N/A | |
| Recombinant DNA reagent | pET28a(+) UFD1 | Fernández-Sáiz and Buchberger, 2010 | pAB425 | |
| Recombinant DNA reagent | pET21d NPL4 | Fernández-Sáiz and Buchberger, 2010 | pAB1340 | |
| Recombinant DNA reagent | pProExHT p97 | Fernández-Sáiz and Buchberger, 2010 | pAB1312 | |
| Recombinant DNA reagent | pProExHT p97 N domain | Fernández-Sáiz and Buchberger, 2010 | pAB1342 | |
| Recombinant DNA reagent | pProExHT p97 ND1 | Fernández-Sáiz and Buchberger, 2010 | pAB1343 | |
| Recombinant DNA reagent | pProExHT p97 ΔN | Rothballer et al., 2007 | pAB749 | |
| Recombinant DNA reagent | pGAD-C1 | James et al., 1996 | N/A | |
| Recombinant DNA reagent | pGAD-C1 VCF1 isoform 1 | This study | pAB2217 | See ‘Materials and methods,’ section ‘Plasmids’ |
| Recombinant DNA reagent | pGAD-C1 VCF1 isoform 2 | This study | pAB2205 | |
| Recombinant DNA reagent | pGAD-C1 VCF1 isoform 5 | This study | pAB2215 | |
| Recombinant DNA reagent | pGAD-C1 VCF2 isoform 3 | This study | pAB2233 | |
| Recombinant DNA reagent | pGBDU | James et al., 1996 | N/A | |
| Recombinant DNA reagent | pGBDU p97 | This study | pAB1184 | See ‘Materials and methods,’ section ‘Plasmids’ |
| Recombinant DNA reagent | pGAD-C1 VCF1 isoform 5 Cdel4 | This study | pAB2312 | |
| Recombinant DNA reagent | pGAD-C1 VCF1 isoform 5 Cdel7 | This study | pAB2313 | |
| Recombinant DNA reagent | pGAD-C1 VCF1 isoform 5 Cdel13 | This study | pAB2314 | |
| Recombinant DNA reagent | pGAD-C1 VCF1 isoform 5 Cdel18 | This study | pAB2315 | |
| Recombinant DNA reagent | pGAD-C1 VCF1 isoform 5 Cdel26 | This study | pAB2235 | |
| Recombinant DNA reagent | pINDUCER20 | Meerbrey et al., 2011 | pAB3012 | |
| Recombinant DNA reagent | pINDUCER20 p97 wildtype | This study | pAB3041 | See ‘Materials and methods,’ section ‘Plasmids’ |
| Recombinant DNA reagent | pINDUCER20 SV40NLS-p97 | This study | pAB3261 | |
| Recombinant DNA reagent | pLentiCRISPRv2 | Addgene | Cat# 52961 | |
| Recombinant DNA reagent | Non-human control sgRNAs in pLentiCRISPRv2 | Manuel Kaulich, University of Frankfurt | N/A | |
| Recombinant DNA reagent | VCF1 sgRNAs in pLentiCRISPRv2 | Manuel Kaulich, University of Frankfurt | N/A | |
| Recombinant DNA reagent | VCF2 sgRNAs in pLentiCRISPRv2 | Manuel Kaulich, University of Frankfurt | N/A | |
| Sequence-based reagent | ON-TARGETplus human FAM104A siRNA-SMARTpool | Dharmacon | Cat# L-015015-02-0005 | |
| Sequence-based reagent | ON-TARGETplus human UBXN2B siRNA-SMARTpool | Dharmacon | Cat# L-025945-01-0005 | |
| Sequence-based reagent | ON-TARGETplus non-targeting pool | Dharmacon | Cat# D-001810-10-05 | |
| Sequence-based reagent | FAM104A_1_fwd | This paper | qPCR primer | CTCCGTCCCAGGAAAAGGAG |
| Sequence-based reagent | FAM104A_1_rev | This paper | qPCR primer | AGGGTTTCTGCTACTTCTTTTGG |
| Sequence-based reagent | FAM104A_2_fwd | This paper | qPCR primer | TGGCAACGAAGAAGACAACC |
| Sequence-based reagent | FAM104A_2_rev | This paper | qPCR primer | TCACTGCCTGAAGACTCTGTG |
| Sequence-based reagent | HPRT_fwd | This paper | qPCR primer | TGGACAGGACTGAACGTCTTG |
| Sequence-based reagent | HPRT_rev | This paper | qPCR primer | CAGTCATAGGAATGGATCTATCAC |
| Sequence-based reagent | PBGD_fwd | This paper | qPCR primer | CCCTGGAGAAGAATGAAGTGG |
| Sequence-based reagent | PBGD_rev | This paper | qPCR primer | TTCTCTGGCAGGGTTTCTAGG |
| Sequence-based reagent | Fam104A1-KO-2-R_79 | This paper | gRNA | CGTAGCTTCCATCCGCCAGC |
| Sequence-based reagent | Fam104A1-KO-3-R_38 | This paper | gRNA | CCTCGGGCCTTGGCTCTCGC |
| Sequence-based reagent | Fam104A2-KO-1-R_190 | This paper | gRNA | TGTCCGGGCTATTGATGCTG |
| Sequence-based reagent | Fam104A2-KO-2-R_170 | This paper | gRNA | ACCGCGCAGAACGCTTTGTT |
| Sequence-based reagent | Fam104A3-KO-1-R_172 | This paper | gRNA | CTCCGCGAAGAGAGGGAACA |
| Sequence-based reagent | Fam104A3-KO-3-R_70 | This paper | gRNA | CCGAAACACAACCCCCTCTG |
| Sequence-based reagent | Fam104B1-KO-1-R_187 | This paper | gRNA | CTGTATCTTGAGAATCCTGA |
| Sequence-based reagent | Fam104B1-KO-2-R_18 | This paper | gRNA | CTTGCTCTCTCTGGGATATT |
| Sequence-based reagent | Fam104B1-KO-3-R_139 | This paper | gRNA | CATTAATATCCCAGAGAGAG |
| Sequence-based reagent | Fam104B2-KO-1-R_19 | This paper | gRNA | GATTGTTACTGAACCCGATG |
| Sequence-based reagent | Fam104B2-KO-2-R_85 | This paper | gRNA | GTTTTCATGGAGTGATAATG |
| Sequence-based reagent | Non-human-target-309-KO-1-R_156 | This paper | gRNA | AACATGACGTTCAAGATTGG |
| Sequence-based reagent | Non-human-target-365-KO-5-R_5 | This paper | gRNA | ACCACTGTTCTACGCGCAGG |
| Sequence-based reagent | Non-human-target-415-KO-2-R_24 | This paper | gRNA | TTGAACGGGCCGCGGAAGCG |
| Sequence-based reagent | Non-human-target-42-KO-15-R_115 | This paper | gRNA | CCCGCATGACACCGTCACTT |
| Peptide, recombinant protein | Biotin- CQGLYFHINQTLREAHFHSLQHRG-COOH | PANATecs GmbH | N/A | See ‘Materials and methods,’ section ‘In vitro binding assays’ |
| Commercial assay or kit | Pierce BCA Protein Assay Kit | Thermo Fisher Scientific | Cat# 23225 | |
| Commercial assay or kit | QuikChange XLII Mutagenesis Kit | Agilent | Cat# 200521 | |
| Commercial assay or kit | Transcriptor High Fidelity cDNA synthesis kit | Roche | Cat# 5081963001 | |
| Commercial assay or kit | NucleoSpin RNA kit | Macherey-Nagel | Cat# REF 740955 | |
| Chemical compound, drug | Camptothecin | Selleckchem | NSC-100880 | |
| Chemical compound, drug | CB-5083 | Selleckchem | Cat# S8101 | |
| Commercial assay or kit | Clarity Western ECL Substrate | Bio-Rad | Cat# 1705061 | |
| Chemical compound, drug | cOmplete, EDTA-free Protease Inhibitor Cocktail | Roche | Cat# 04693132001 | |
| Chemical compound, drug | Glutathione Sepharose 4 Fast Flow | Cytiva | Cat# 17513202 | |
| Chemical compound, drug | Ni-NTA Agarose | QIAGEN | Cat# 30230 | |
| Chemical compound, drug | Opti-MEM | Thermo Fisher Scientific | Cat# 31985602 | |
| Chemical compound, drug | Polybrene | Santa Cruz | Cat# sc-134220 | |
| Chemical compound, drug | Polyethylenimine (PEI) | Polysciences | Cat# 23966-1 | |
| Chemical compound, drug | ProLong Glass Antifade Mountant | Thermo Fisher Scientific | Cat# P36980 | |
| Chemical compound, drug | Anti FLAG-M2 affinity agarose beads | Sigma-Aldrich | Cat# A2220 | |
| Chemical compound, drug | Oligofectamine | Thermo Fisher Scientific | Cat# 12252011 | |
| Chemical compound, drug | Benzonase Nuclease | Sigma-Aldrich | Cat# 70664 | |
| Chemical compound, drug | PowerUp SYBR Green Master Mix | Thermo Fisher | Cat# A25741 | |
| Chemical compound, drug | Protein G Sepharose 4 fast flow | Cytiva | Cat# 17-5132-01 | |
| Software, algorithm | Fiji | http://fiji.sc | RRID:SCR_002285 | |
| Software, algorithm | Image Lab Software | Bio-Rad | RRID:SCR_014210 | |
| Software, algorithm | CellProfiler | https://cellprofiler.org/ | RRID:SCR_007358 | |
| Software, algorithm | GraphPad Prism | https://www.graphpad.com/scientific-software/prism/ | RRID:SCR_002798 | |
| Software, algorithm | QuantStudio Design & Analysis Software | https://www.thermofisher.com/de/de/home/global/forms/life-science/quantstudio-3-5-software.html |
Additional files
-
Supplementary file 1
Summary of VCF1/2 yeast two-hybrid hits.
- https://cdn.elifesciences.org/articles/92409/elife-92409-supp1-v2.xlsx
-
MDAR checklist
- https://cdn.elifesciences.org/articles/92409/elife-92409-mdarchecklist1-v2.pdf





