Depletion of SMN protein in mesenchymal progenitors impairs the development of bone and neuromuscular junction in spinal muscular atrophy
Figures
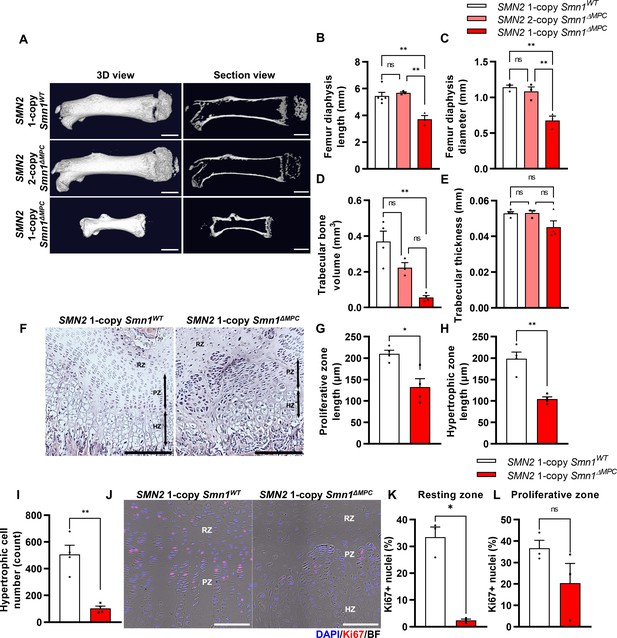
Skeletal growth abnormalities and altered growth plate homeostasis in SMN2 1-copy Smn1ΔMPC mice.
(A) Representative 3D images and longitudinal section view of the ossified femur bone. Scale bars, 1 mm. (B, C) SMN2 1-copy mutant’s femurs showed reduced growth in diaphysis length and diameter, and (D) decreased trabecular bone volume. (E) Trabecular bone thicknesses were not significantly different between the control and mutant groups. The micro-computed tomography (micro-CT) analysis was performed in femur diaphysis and metaphysis from SMN2 1-copy Smn1ΔWT, SMN2 2-copy, and 1-copy Smn1ΔMPC mice at P14. One-way analysis of variance (ANOVA) with Tukey’s post hoc test, n = 3–5 mice in each genotype (B–E). (F) Representative images of hematoxylin and eosin (H&E) staining in the distal femur growth plate of control and mutant mice with 1 copy of SMN2 at P14. Scale bars, 100 μm. Resting zone (RZ), hypertrophic zone (HZ), and proliferative zone (PZ). (G–I) Indicated by black arrows, the HZ and PZ lengths were reduced in SMN2 1-copy Smn1ΔMPC mice, and the hypertrophic cell number in a section of the 1-copy mutant was decreased (n = 4 mice in each genotype; unpaired t-test with Welch’s correction). (J) Representative images of Ki67 immunostaining in the distal femur growth plate of control and mutant mice with 1 copy of SMN2 at P14. Scale bars, 100 μm, and (K) decreased Ki67+ percentage in resting zone chondrocytes. (L) Ki67+ percentage in the proliferative zone was not significantly different between the control and mutant groups. n = 3 mice in each genotype; unpaired t-test with Welch’s correction (K–L). ns: not significantly different. *p < 0.05; **p < 0.01. Error bars show standard error of the mean (SEM).
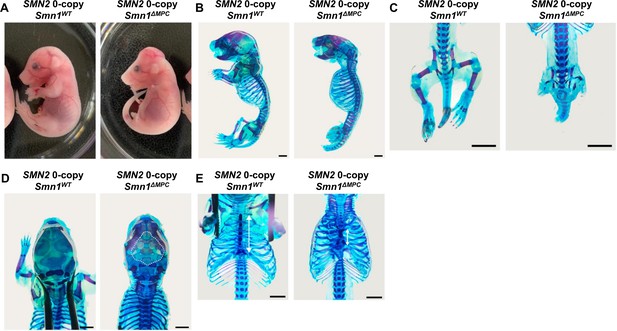
Growth defects in the Prrx1-lineage bone of SMN2 0-copy Smn1ΔMPC mice.
(A) Representative control and mutant mice with 0 copies of SMN2 were photographed at E18.5. (B, C) Alcian blue and alizarine red staining in the SMN2 0-copy Smn1ΔMPC mutant showed abnormal skeletal development of the limbs, (D) calvaria (shown by the dashed line), and (E) sternum (indicated by a white arrow). Scale bars, 5 mm (B, C) and 2 mm (D, E).
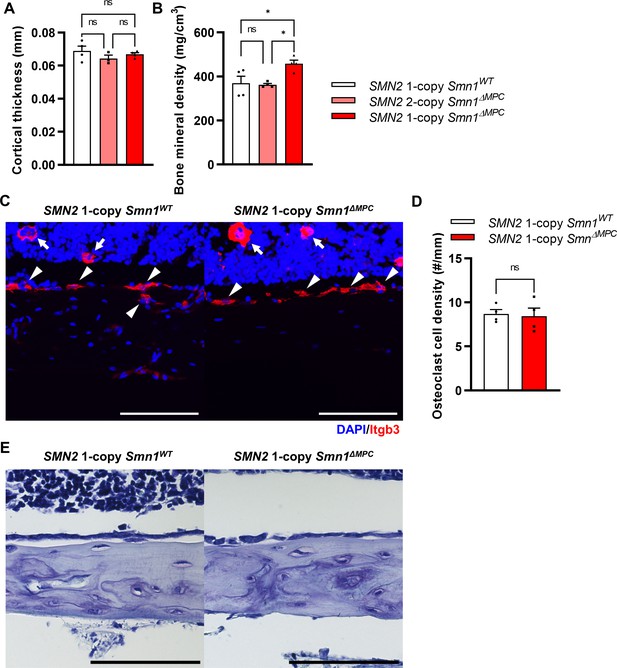
Osteoclasts and osteoblasts were undisturbed in SMN2 1-copy Smn1ΔMPC mice.
(A) Cortical bone thicknesses were not significantly different between the control and mutant groups. (B) Bone mineral density was slightly increased in the diaphysis of SMN2 1-copy Smn1ΔMPC mice. The micro-computed tomography (micro-CT) analysis was performed in femur diaphysis and metaphysis from SMN2 1-copy Smn1WT, SMN2 2-copy, and 1-copy Smn1ΔMPC mice at P14. One-way analysis of variance (ANOVA) with Tukey’s post hoc test, n = 3–5 mice in each genotype (A, B). (C) Representative images of osteoclast marker Itgb3 immunostaining in femur diaphysis cortical bone from mice at P14. Scale bars, 100 μm. Itgb3+ hematopoietic cell (indicated by arrow) and osteoclast (indicated by arrowhead) were imaged. (D) It showed similar osteoclast density. n = 3 mice in each genotype; unpaired t-test with Welch’s correction. (E) Representative images of toluidine blue staining in femur diaphysis cortical bone for osteoblast evaluation. Scale bars, 100 μm. ns: not significantly different. *p < 0.05. Error bars show standard error of the mean (SEM).

Decreased chondrocyte-derived IGF–AKT axis by limb mesenchymal cell-specific survival motor neuron (SMN) depletion in SMN2 1-copy Smn1ΔMPC mice.
(A) Representative images of p-AKT immunostaining in distal femur growth plate from mice at P14. Scale bars, 100 μm. (B, C) The p-AKT-positive percentage was decreased in the resting zone chondrocytes, not in the proliferative zone (n = 3–4 mice in each genotype; unpaired t-test with Welch’s correction). (D) Relative IGF axis mRNA expression in the livers of SMN2 1-copy control and mutant mice. The IGF pathway genes showed no difference when comparing controls to SMN2 1-copy Smn1ΔMPC mutants. (E, F) Relative IGF axis and chondrocyte differentiation marker mRNA expression in the chondrocytes of SMN2 1-copy control and mutant mice. The Igf1, Igfbp3, and hypertrophic marker Col10a1 expression were decreased in SMN2 1-copy Smn1ΔMPC mutants. n = 3 mice in each genotype; unpaired t-test with Welch’s correction (D–F). (G) Representative images of genomic PCR analysis from SMN2 1-copy Smn1ΔMPC mice tissues at P21. (H) Quantitative reverse transcription polymerase chain reaction (qRT-PCR) analysis from tissues of SMN2 1-copy Smn1WT and Smn1ΔMPC mice at P21 (n = 3 mice in each genotype; unpaired t-test with Welch’s correction). Deletion of Smn1 exon 7 was detected only in limb mesenchymal cells using genomic PCR (G) and full-length Smn1 mRNA expression (H). (I) Representative images of western blot analysis in cultured fibro-adipogenic progenitors (FAPs). SMN protein in FAPs of SMN2 1-copy Smn1ΔMPC mice exhibited a decrease comparable to that observed in the SMAΔ7 mice. (J) Relative SMN levels in cultured FAPs of the controls and mutants (n = 3 mice in each genotype; one-way analysis of variance (ANOVA) with Tukey’s post hoc test). ns: not significantly different. *p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.001. Error bars show standard error of the mean (SEM).
-
Figure 2—source data 1
Original file for the gel electrophoresis of genomic PCR in Figure 2G (Cre, Smn1F7, Smn1Δ7) and western blot analysis in Figure 2I (anti-alpha-tubulin, anti-SMN).
- https://cdn.elifesciences.org/articles/92731/elife-92731-fig2-data1-v1.zip
-
Figure 2—source data 2
PDF containing Figure 2G, I and original scans of the PCR and western blot with highlighted bands and sample labels.
- https://cdn.elifesciences.org/articles/92731/elife-92731-fig2-data2-v1.zip
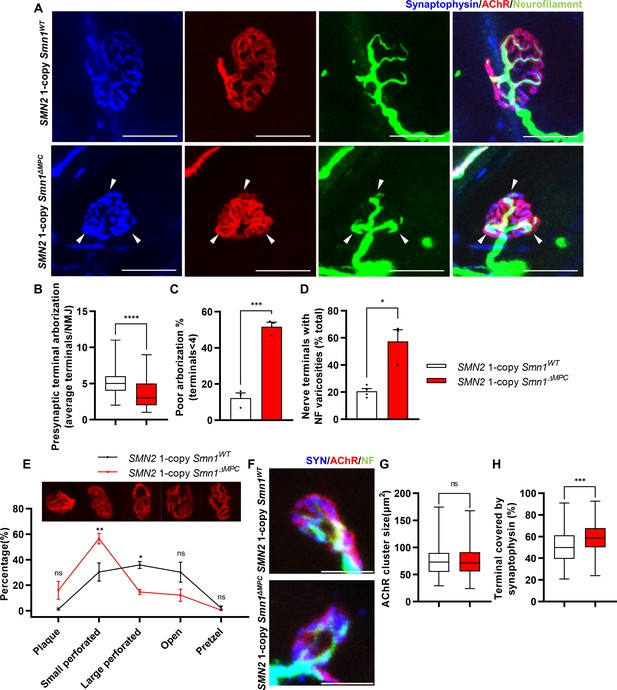
Aberrant postnatal neuromuscular junction (NMJ) maturation in SMN2 1-copy Smn1ΔMPC mice.
(A) Immunostaining of NMJs in TA muscle of SMN2 1-copy Smn1WT and Smn1ΔMPC mice at P21 with anti-NF (green), anti-synaptophysin (blue), and α-Btx staining acetylcholine receptor (AChR; red). Scale bars, 20 μm. The confocal images of NMJs showed decreased presynaptic terminal branching and the existence of nerve terminal varicosities that were enlarged with neurofilament (NF; indicated by arrowheads) in the mutant. (B) The NMJs of the SMN2 1-copy mutant exhibited a significant decrease in presynaptic terminal arborization and (C) an increased percentage of poorly arborized NMJs (n = 3 mice in each genotype; unpaired t-test with Welch’s correction). (D) The percentage of NMJs exhibiting NF varicosities was higher in the SMN2 1-copy mutant group than in the control group (n = 3–4 mice in each genotype; unpaired t-test with Welch’s correction). (E) For quantification of the NMJ maturation stage, we classified NMJs into five distinct developmental stages (Plaque: plaque-shaped endplate without any perforation; Small perforated: plaque-shaped endplate with small perforations; Large perforated: plaque-shaped endplate with large perforations; Open: C-shaped endplate; Pretzel: pretzel-like shaped endplate) and then compared the frequency patterns of SMN2 1-copy control and mutant mice (n = 3 mice in each genotype; two-way analysis of variance (ANOVA) with Tukey’s post hoc test). The NMJs of SMN2 1-copy mutants displayed plaque-like shapes, indicating that they were in the immature stage. (F) Immunostaining of NMJs in TA muscle of SMN2 1-copy Smn1WT and Smn1ΔMPC mice at P3 with anti-NF (green), anti-synaptophysin (blue), and α-Btx staining AChR (red). Scale bars, 10 μm. (G) There were no significant differences in AChR cluster size between the SMN2 1-copy control and mutant at P3 (n = 3–4 mice in each genotype; unpaired t-test with Welch’s correction). (H) The ratio of the Synaptophysin area to the AChR area in NMJ was slightly higher in the SMN2 1-copy mutant at P3 (n = 3–4 mice in each genotype; unpaired t-test with Welch’s correction). ns: not significantly different. *p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001. All box-and-whisker plots show the median, interquartile range, minimum, and maximum. For the box-and-whisker plots, range bars show minimum and maximum (B, G, H). For the bar and line graph, error bars show standard error of the mean (SEM) (C–E).
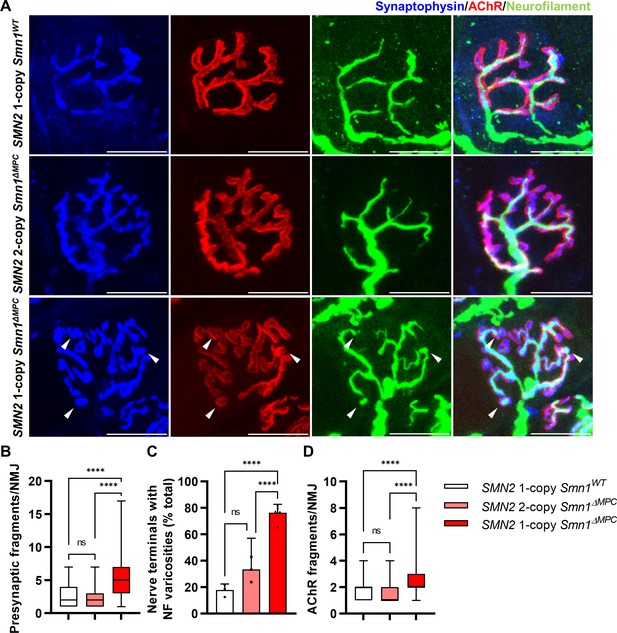
Morphological deterioration in neuromuscular junctions (NMJs) of adult SMN2 1-copy Smn1ΔMPC mice.
(A) Immunostaining of NMJs in TA muscle of SMN2 1-copy Smn1WT, SMN2 2-copy, and SMN2 1-copy Smn1ΔMPC mice at P56 with anti-NF (green), anti-synaptophysin (blue), and α-Btx staining acetylcholine receptor (AChR; red). Scale bars, 20 μm. The confocal images of NMJs showed fragmentation and bouton-like neurofilament (NF) varicosities (indicated by arrowheads) in the SMN2 1-copy Smn1ΔMPC mice. The NMJs of the SMN2 1-copy mutant displayed fragmented presynapse (B), endplate (D), and NF varicosities (C) compared to SMN2 1-copy Smn1WT and SMN2 2-copy Smn1ΔMPC mice (n = 3–5 mice in each genotype; Presynaptic fragments and AChR fragments: Brown–Forsythe and Welch analysis of variance (ANOVA) with Games–Howell’s test; NF varicosities: one-way ANOVA with Tukey’s post hoc test). ns; not significantly different. ****p < 0.0001. All box-and-whisker plots show the median, interquartile range, minimum, and maximum. For the box-and-whisker plots, range bars show minimum and maximum (B, D). For the bar graph, error bars show standard error of the mean (SEM) (C).

Reduced presynaptic neurotransmission ability in the neuromuscular junctions (NMJs) of SMN2 1-copy Smn1ΔMPC mice.
(A) Representative traces of Miniature endplate potential (mEPP) from SMN2 1-copy Smn1ΔWT (top) and SMN2 1-copy Smn1ΔMPC (bottom) mice. (B, C) SMN2 1-copy mutant’s NMJs showed an increase in mEPP amplitude and no differences in mEPP frequency (1-copy control, n = 25, 9 mice; 1-copy mutant, n = 21, 8 mice; unpaired t-test with Welch’s correction). (D) Representative traces of evoked endplate potential (eEPP) from SMN2 1-copy Smn1ΔWT (top) and SMN2 1-copy Smn1ΔMPC (bottom) mice. (E) The mutant’s NMJs showed a stronger amplitude of eEPPs (1-copy control, n = 12, 4 mice; 1-copy mutant, n = 12, 3 mice; unpaired t-test with Welch’s correction). (F) Representative traces of paired-pulse response from SMN2 1-copy Smn1ΔWT (top) and SMN2 1-copy Smn1ΔMPC (bottom) mice. (G) Paired-pulse response was not different between SMN2 1-copy control and mutant NMJs, indicating a comparable neurotransmitter release probability (1-copy control, n = 8, 3 mice; 1-copy mutant, n = 6, 3 mice; unpaired t-test with Welch’s correction). The electrophysiological recording was performed in the extensor digitorum longus (EDL) muscle at P56. ns: not significantly different. *p < 0.05. All box-and-whisker plots show the median, interquartile range, minimum, and maximum. For the box-and-whisker plots, range bars show minimum and maximum (B, C, E, G).
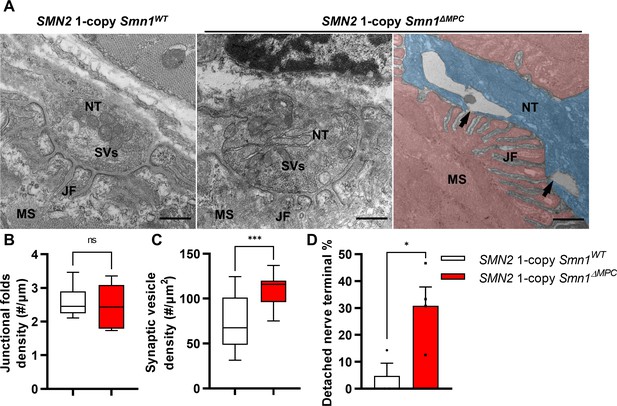
Abnormal nerve terminal ultrastructure in SMN2 1-copy mutant.
(A) Representative transmission electron microscopy (TEM) images from neuromuscular junctions (NMJs) of SMN2 1-copy Smn1ΔWT and SMN2 1-copy Smn1ΔMPC mice at P56. Scale bars, 500 nm. Nerve terminal (NT; indicated by the blue zone). Synaptic vesicles (SVs). Muscle fiber (MS; indicated by the red zone). Endplate junctional folds (JF). Nerve terminal detachment (indicated by arrow) was observed in SMN2 1-copy Smn1ΔMPC mice. (B) The density of junctional folds in the NMJ of SMN2 1-copy Smn1ΔMPC mice no significant change compared to the control, whereas (C) the density of synaptic vesicles was increased (n = 3–4 mice in each genotype; unpaired t-test with Welch’s correction). (D) The detachment of the nerve terminal occurs more frequently at the NMJ of mutants (n = 3–4 mice in each genotype; unpaired t-test with Welch’s correction). ns: not significantly different. *p < 0.05; ***p < 0.001. All box-and-whisker plots show the median, interquartile range, minimum, and maximum. For the box-and-whisker plots, range bars show minimum and maximum (B, C). For the bar graph, error bars show standard error of the mean (SEM) (D).
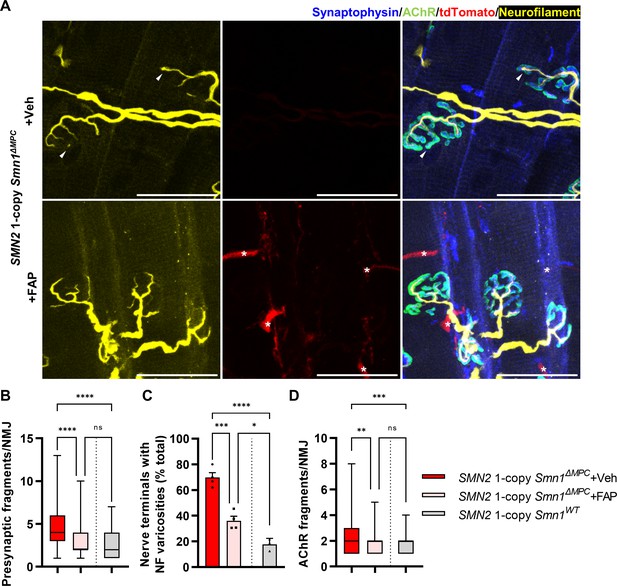
Improved postnatal neuromuscular junction (NMJ) development in the TA muscle of SMN2 1-copy Smn1ΔMPC mice following healthy fibro-adipogenic progenitors (FAPs) transplantation.
(A) Immunostaining of NMJs in tdTomato+ FAP-transplanted TA muscle (+FAP) and vehicle-treated contralateral muscle (+Veh) in SMN2 1-copy Smn1ΔMPC mice at P56 with anti-NF (yellow), anti-synaptophysin (blue), α-Btx staining acetylcholine receptor (AChR; green), and tdTomato fluorescence (red). Scale bars, 40 μm. The images revealed neurofilament (NF) varicosities (indicated by arrowheads) in the +Veh NMJs. The tdTomato+ FAPs (marked by asterisks) were transplanted into +FAP NMJs, which exhibited (B) decreased presynaptic fragmentation, (C) NF varicosities, and (D) AChR fragmentation compared to +Veh NMJs and similar to wild-type NMJs (n = 3–4 mice in each group; unpaired t-test with Welch’s correction). ns: not significantly different. *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001. All box-and-whisker plots show the median, interquartile range, minimum, and maximum. For the box-and-whisker plots, range bars show minimum and maximum (B, D). For the bar graph, error bars show standard error of the mean (SEM) (C).
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Genetic reagent (Mus musculus) | Prrx1Cre | The Jackson Laboratory | Strain #: 005584; RRID:IMSR_JAX:005584 | |
| Genetic reagent (M. musculus) | Smn1f7/+ | The Jackson Laboratory | Strain #: 006138; RRID:IMSR_JAX:006138 | |
| Genetic reagent (M. musculus) | Rosa26LSL-YFP/+ | The Jackson Laboratory | Strain #: 006148; RRID:IMSR_JAX:006148 | |
| Genetic reagent (M. musculus) | Rosa26LSL-tdTomato/+ | The Jackson Laboratory | Strain #: 007914; RRID:IMSR_JAX:007914 | |
| Genetic reagent (M. musculus) | SMN2+/+; Smn1+/− | The Jackson Laboratory | Strain #: 005024; RRID:IMSR_JAX:005024 | |
| Genetic reagent (M. musculus) | Smn1+/−; SMN2+/+; SMNΔ7+/+ | The Jackson Laboratory | Strain #: 005025; RRID:IMSR_JAX:005025 | |
| Antibody | anti-SMN (Mouse monoclonal) | BD Biosciences | Cat. #: 610646; RRID:AB_397973 | WB (1:1000) |
| Antibody | anti-alpha-tubulin (Rabbit monoclonal) | Abcam | Cat. #: ab176560; RRID:AB_2860019 | WB (1:1000) |
| Antibody | anti-Neurofilament M (Rabbit polyclonal) | Merckmillipore | Cat. #: ab1987; RRID:AB_91201 | IF (1:1000) |
| Antibody | anti-Synaptophysin 1 (Guinea pig polyclonal) | Synaptic Systems | Cat. #: 101 004; RRID:AB_1210382 | IF (1:500) |
| Antibody | anti-GFP (Chicken polyclonal) | Abcam | Cat. #: ab13970; RRID:AB_300798 | IF (1:500) |
| Antibody | anti-Ki67 (Rabbit polyclonal) | Abcam | Cat. #: ab15580; RRID:AB_443209 | IF (1:500) |
| Antibody | anti-Itgb3 (Rabbit monoclonal) | Cell Signaling | Cat. #: 13166; RRID:AB_2798136 | IF (1:100) |
| Antibody | anti-p-AKT(S473) (Rabbit monoclonal) | Cell Signaling | Cat. #: 4060; RRID:AB_2315049 | IF (1:100) |
| Antibody | anti-CD45-APC (Rat monoclonal) | Biolegend | Cat. #: 103111; RRID:AB_312976 | FACS (3 µl per test) |
| Antibody | anti-CD31-APC (Rat monoclonal) | Biolegend | Cat. #: 102409; RRID:AB_312904 | FACS (3 µl per test) |
| Antibody | anti-Sca-1(Ly6a)-FITC (Rat monoclonal) | Biolegend | Cat. #: 122507; RRID:AB_756192 | FACS (3 µl per test) |
| Antibody | anti-Sca-1(Ly6a)-Pacific blue (Rat monoclonal) | Biolegend | Cat. #: 108120; RRID:AB_493273 | FACS (3 µl per test) |
| Antibody | anti-Vcam1-Biotin (Rat monoclonal) | Biolegend | Cat. #: 105703; RRID:AB_313204 | FACS (3 µl per test) |
| Antibody | anti-Rabbit IgG-HRP (Goat monoclonal) | Promega | Cat. #: W4011; RRID:AB_430833 | WB (1:10,000) |
| Antibody | anti-Mouse IgG-HRP (Goat monoclonal) | Promega | Cat. #: W4021; RRID:AB_430834 | WB (1:10,000) |
| Antibody | anti-Rabbit IgG-Alexa fluor 488 (Goat monoclonal) | Invitrogen | Cat. #: A11034; RRID:AB_2576217 | IF (1:500) |
| Antibody | anti-Chicken IgY-Alexa fluor 488 (Goat monoclonal) | Invitrogen | Cat. #: A11039; RRID:AB_2534096 | IF (1:500) |
| Antibody | anti-Rabbit IgG-Alexa fluor Plus 647 (Goat monoclonal) | Invitrogen | Cat. #: A32733; RRID:AB_2633282 | IF (1:500) |
| Antibody | anti-Guine pig IgG-Alexa fluor 405 (Goat monoclonal) | Abcam | Cat. #: ab175678; RRID:AB_2827755 | IF (1:500) |
| Peptide, recombinant protein | PE-Cy7-Streptavidin | Biolegend | Cat. #: 405206 | FACS (3 µl per test) |
| Peptide, recombinant protein | Alpha-bungarotoxin-Alexa fluor 555 | Invitrogen | Cat. #: B35451 | IF (1:1000) |
| Peptide, recombinant protein | Alpha-bungarotoxin-Alexa fluor 488 | Invitrogen | Cat. #: B13422 | IF (1:1000) |
| Peptide, recombinant protein | μ-conotoxin GIIIB | Alomone | Cat. #: C-270 | Electrophysiology (2.5 µM) |
| Commercial assay or kit | Pierce BCA protein assay kits | Thermo Fisher Scientific | Cat. #: C-23225 | |
| Software | ImageJ | ImageJ | RRID:SCR_003070 | |
| Software | AccuCT | PerkinElmer | ||
| Software | Microsoft Excel | Microsoft | RRID:SCR_016137 | |
| Software | Leica Application Suite X | Leica | RRID:SCR_013673 | |
| Software | GraphPad Prism | Graphpad | RRID:SCR_002798 | |
| Software | pClamp | Molecular devices | RRID:SCR_011323 | |
| Software | QIAGEN | QIAGEN | RRID:SCR_008539 | |
| Other | Gill No. 3 formula hematoxylin | Sigma | Cat. #: GHS332 | Hematoxylin staining solution (1×) |
| Other | Eosin Y solution | Sigma | Cat. #: HT110116 | Eosin staining solution (1×) |
| Other | Toluidine blue | Sigma | Cat. #: 198161 | Toluidine blue staining (1%) |
Additional files
-
MDAR checklist
- https://cdn.elifesciences.org/articles/92731/elife-92731-mdarchecklist1-v1.docx
-
Supplementary file 1
A primer list of genomic PCR and qRT-PCR.
- https://cdn.elifesciences.org/articles/92731/elife-92731-supp1-v1.xlsx