Peer review process
Not revised: This Reviewed Preprint includes the authors’ original preprint (without revision), an eLife assessment, public reviews, and a provisional response from the authors.
Read more about eLife’s peer review process.Editors
- Reviewing EditorAnne-Florence BitbolEcole Polytechnique Federale de Lausanne (EPFL), Lausanne, Switzerland
- Senior EditorAleksandra WalczakCNRS, Paris, France
Reviewer #1 (Public review):
Summary:
The authors developed a novel theoretical/computational procedure to count bacterial populations without introducing artificial randomness effects due to dilution. Surprisingly, this very important aspect of studies of bacterial systems has been overlooked. The proposed method provides a simple and transparent approach to eliminate the randomness of bacterial accounting procedures, allowing now to fully concentrate on the intrinsic effects of the studied systems.
Strengths:
A very simple and clear procedure is introduced and explained in full detail. This elegant approach finds an excellent compromise between mathematical rigor and computational efficiency, which is important for practical applications. The provided examples are convincing beyond a doubt, clearly indicating the potential strong impact of the proposed framework. Various complications and possible issues are also discussed and analyzed. This seems to be a very powerful novel method that should significantly advance the analysis of complex biological systems.
Weaknesses:
The only minor weakness that I found is the assumption of independence of bacterial species, which is expressed as the well-stirred approximation. One could imagine that bacterial species might cooperate, leading to non-uniform distributions that are real. How to distinguish such situations?
I believe that this method can be extended to determine if this is the case or not before the application. For example, if the bacteria species are independent of each other and one can use the binomial distributions, then the Fano factor would be proportional to the overall relative fraction of bacterial species. Maybe a simple test can be added to test it before the application of REPOP. However, I believe that this is a minor issue.
Reviewer #2 (Public review):
Summary:
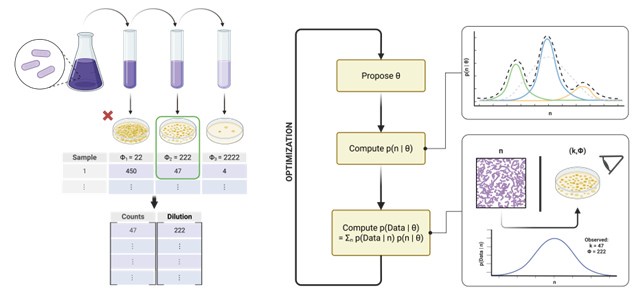
Microbial population abundances are regularly estimated by multiplying plate counts by dilution factors, with inferences made about sample heterogeneity without taking into account heterogeneity generated through dilution and plating methods. The authors have developed REPOP, a method for disentangling methodological stochasticity from ecological heterogeneity using a Bayesian framework. They present three models: a unimodal distribution, a multimodal distribution, and a multimodal distribution that incorporates a colony count cutoff. They use a combination of simulated and experimental data to show the effectiveness of the REPOP method in resolving true microbial population distributions.
Strengths:
Overall, this paper addresses a significant issue in microbial ecology and reliably demonstrates that the REPOP method improves upon current methods of estimating microbial population heterogeneity, particularly with simulation data. The three models presented build upon each other and are discussed in a way that is fairly accessible to a broad audience. The authors also show that leveraging the information provided by non-countable plates is important. Additionally, the authors address the potential for extending this method to other sources of methodological stochasticity that may occur in microbial plating. However, it does seem that they could extend this further by discussing ways that this method could be applied to non-microbial systems, allowing this work to appeal to a broader audience.
Weaknesses:
A more thorough discussion of when and by how much estimated microbial population abundance distributions differ from the ground truth would be helpful in determining the best practices for applying this method. Not only would this allow researchers to understand the sampling effort necessary to achieve the results presented here, but it would also contextualize the experimental results presented in the paper. Particularly, there is a disconnect between the discussion of the large sample sizes necessary to achieve accurate multimodal distribution estimates and the small sample sizes used in both experiments.
Reviewer #3 (Public review):
Summary:
In microbiology, accurately characterizing microbial populations and communities is essential. One widely used approach is to measure the absolute or relative abundance of microbial species. Recent research in microbial ecology, for instance, has shown that even genetically identical hosts exposed to the same microbial pool can develop very different gut microbiota, largely due to random colonization events. This study builds on that idea but adds a valuable layer: it suggests that some of the observed variability might actually result from experimental noise, specifically the randomness introduced by dilution and plate counting techniques. To address this, the authors introduce REPOP, a new tool designed to improve the quantification of microbial populations by explicitly accounting for the inherent stochasticity in these methods. They test REPOP using both simulated and experimental datasets, showing how it can help recover meaningful trends.
Strengths:
Overall, this paper is a good contribution to the field. The motivation is clear: improving our ability to quantify microbial populations is crucial for many research areas. The authors make a strong case that ignoring experimental noise is no longer acceptable, and they offer a well-argued solution. The manuscript is well-written and easy to follow, and the logic behind REPOP is convincingly laid out. The use of simulated data is especially valuable, as it allows the authors to test whether the method can recover known inputs, an important validation step. Even with experimental data, where true values are unknown, the method seems to behave in a reasonable and expected way, which is reassuring. All in all, this is an important step forward in how we quantify microbial populations.
Weaknesses:
While the study is promising, there are a few areas where the paper could be strengthened to increase its impact and usability. First, the extent to which dilution and plating introduce noise is not fully explored. Could this noise significantly affect experimental conclusions? And under what conditions does it matter most? Does it depend on experimental design or specific parameter values? Clarifying this would help readers appreciate when and why REPOP should be used. Second, more practical details about the tool itself would be very helpful. Simply stating that it is available on GitHub may not be enough. Readers will want to know what programming language it uses, what the input data should look like, and ideally, see a step-by-step diagram of the workflow. Packaging the tool as an easy-to-use resource, perhaps even submitting it to CRAN or including example scripts, would go a long way, especially since microbiologists tend to favor user-friendly, recipe-like solutions. Third, it would be great to see the method tested on existing datasets, such as those from Nic Vega and Jeff Gore (2017), which explore how colonization frequency impacts abundance fluctuation distributions. Even if the general conclusions remain unchanged, showing that REPOP can better match observed patterns would strengthen the paper's real-world relevance. Lastly, it would be helpful for the authors to briefly discuss the limitations of their method, as no approach is without its constraints. Acknowledging these would provide a more balanced and transparent perspective.