Peer review process
Not revised: This Reviewed Preprint includes the authors’ original preprint (without revision), an eLife assessment, public reviews, and a provisional response from the authors.
Read more about eLife’s peer review process.Editors
- Reviewing EditorDipyaman GangulyAshoka University, Sonepat, India
- Senior EditorRichard WhiteUniversity of Oxford, Oxford, United Kingdom
Reviewer #1 (Public review):
In this manuscript, Rishiq et al. investigate whether natural killer (NK) cells can interact with Fusobacterium nucleatum and identify the molecular mediators involved in this interaction. The authors propose that the bacterial adhesin RadD may bind to the activating NK cell receptor NKp46 (NCR1 in mice), leading to NK cell activation and tumor control. While the topic is of significant interest and the hypothesis intriguing, the manuscript lacks critical experimental evidence, contains several technical concerns, and requires substantial revisions.
Major Concerns:
(1) Lack of Direct Evidence for RadD-NKp46 Interaction
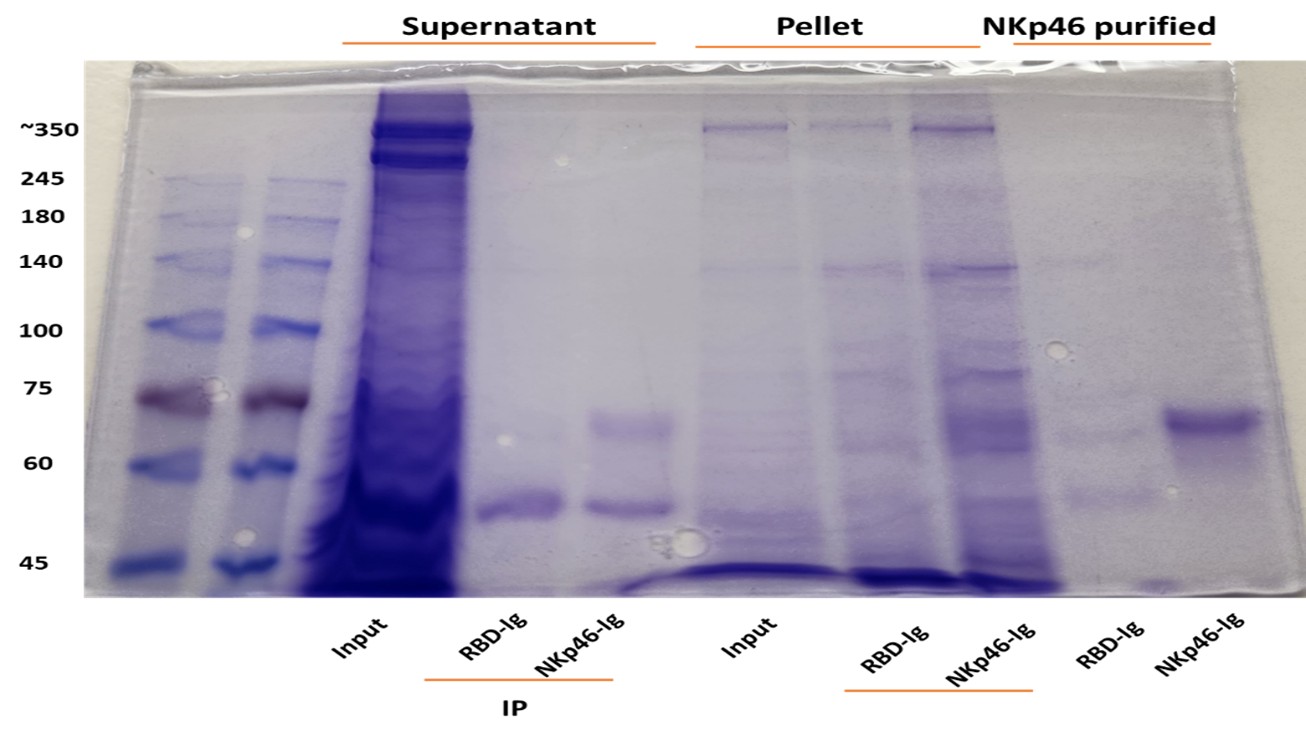
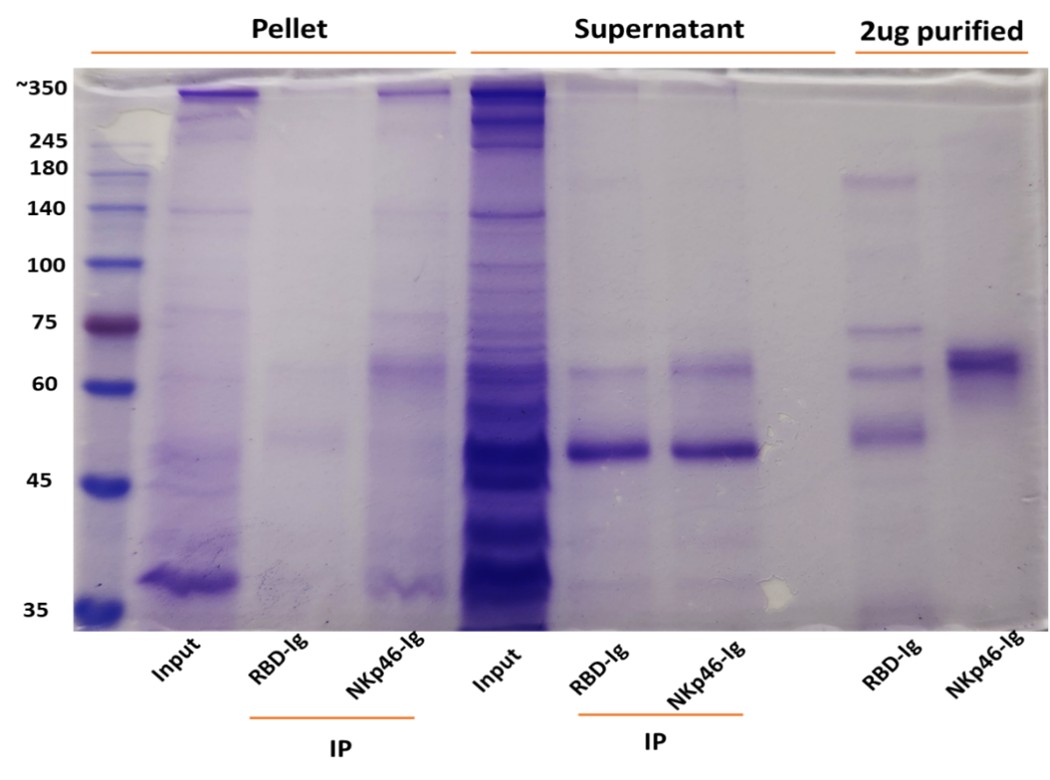
The central claim that RadD interacts with NKp46 is not formally demonstrated. A direct binding assay (e.g., Biacore, ELISA, or pull-down with purified proteins) is essential to support this assertion. The absence of this fundamental experiment weakens the mechanistic conclusions of the study.
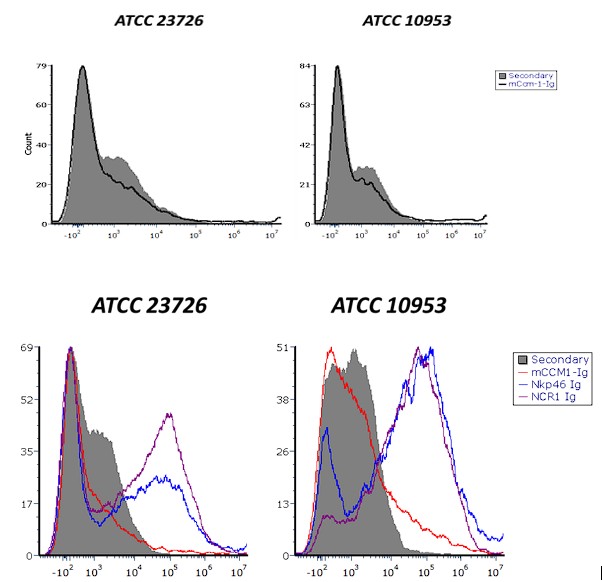
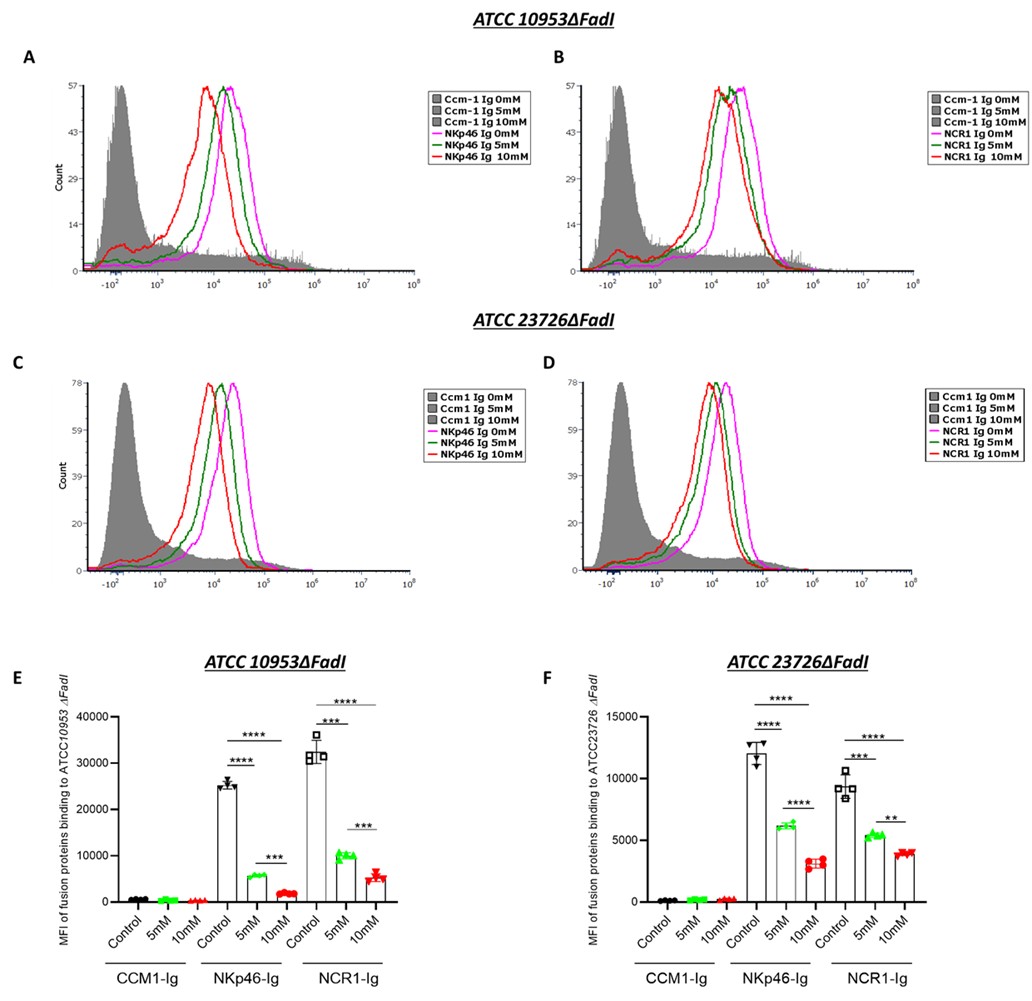
(2) Figure 2: Binding Specificity and Bacterial Strains
A CEACAM1-Ig control should be included in all binding experiments to distinguish between specific and non-specific Ig interactions. There is differential Ig binding between strains ATCC 23726 and 10953. The authors should quantify RadD expression in each strain to determine if the difference in binding is due to variation in RadD levels.
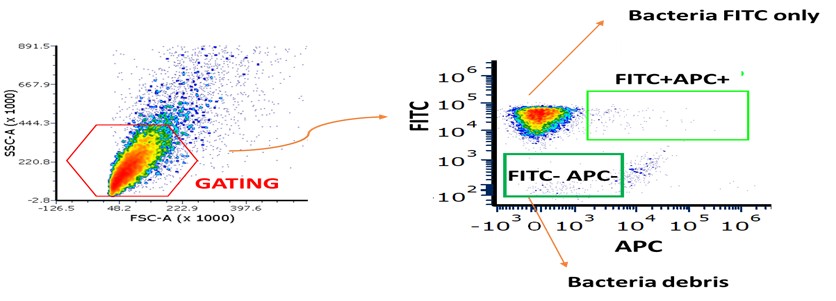
(3) Figure 3: Flow Cytometry Inconsistencies and Missing Controls
What do the FITC-negative, Ig-negative events represent? The authors should clarify whether these are background signals, bacterial aggregates, or debris.
Panel B, CEACAM1-Ig binding appears markedly increased compared to WT bacteria. The reason for this enhancement should be discussed-does it reflect upregulation of the bacterial ligand or an artifact of overexpression? Fluorescence compensation should be carefully reviewed for the NKp46/NCR1-Ig binding assays to ensure that the signals are not due to spectral overlap or nonspecific binding. Importantly, binding experiments using the FadI/RadD double knockout strain are missing and should be included. This control is essential.
In Panel E, the basis for calculating fold-change in MFI is unclear. Please indicate the reference condition to which the change is normalized.
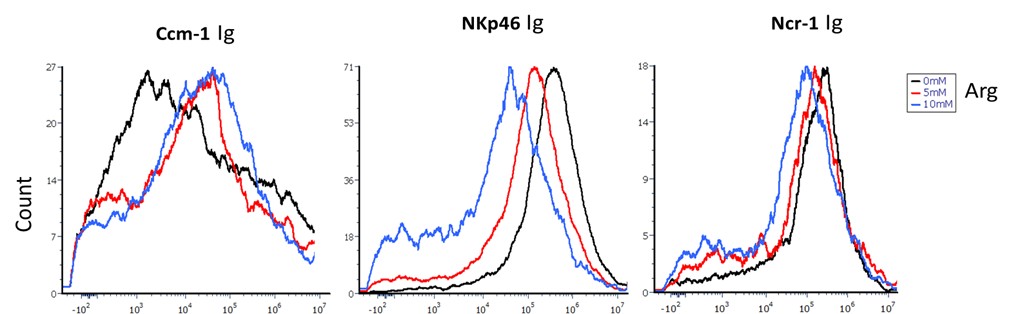
(4) Figure 4: Binding Inhibition and Receptor Sensitivity
Panel A lacks representative FACS plots and is currently difficult to interpret. Differences in the sensitivity of human vs. mouse NKp46 to arginine inhibition should be discussed, given species differences in receptor-ligand interactions. What are the inhibition results using F. nucleatum strains deficient in FadI?
In Panel B, CEACAM1-Ig and RadD-deficient bacteria must be included as negative controls for binding specificity upon anti-NKp46 blocking.
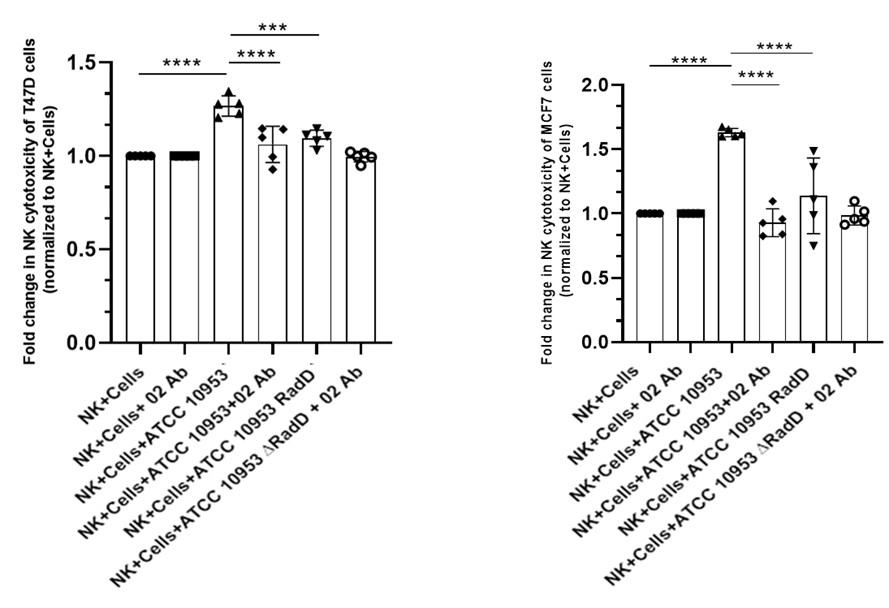
(5) Figure 5: Functional NK Activation and Tumor Killing
In Panels B and C, the key control condition (NK cells + anti-NKp46, without bacteria) is missing. This is needed to evaluate if NKp46 recognition is involved in tumor killing. The authors should explicitly test whether pre-incubation of NK cells with bacteria enhances their anti-tumor activity. Alternatively, could bacteria induce stress signals in tumor cells that sensitize them to NK killing? This distinction is critical.
(6) Figure 5D: Mechanism of Peripheral Activation
It is suggested that contact between bacteria and NK cells in the periphery leads to their activation. Can the authors confirm whether this pre-activation leads to enhanced killing of tumor targets, or if bacteria-tumor co-localization is required? The literature indicates that F. nucleatum localizes intracellularly within tumor cells. If so, how is RadD accessible to NKp46 on infiltrating NK cells?
(8) Figure 5E and In Vivo Relevance
Surprisingly, F. nucleatum infection is associated with increased tumor burden. Does this reflect an immunosuppressive effect? Are NK cells inhibited or exhausted in infected mice (TGIT, SIGLEC7...)? If NK cell activation leads to reduced tumor control in the infected context, the role of RadD-induced activation needs further explanation. RadD-deficient bacteria, which do not activate NK cells, result in even poorer tumor control. This paradox needs to be addressed: how can NK activation impair tumor control while its absence also reduces tumor control?
(9) NKp46-Deficient Mice: Inconsistencies
In Ncr1⁻/⁻ mice, infection with WT or RadD-deficient F. nucleatum has no impact on tumor burden. This suggests that NKp46 is dispensable in this context and casts doubt on the physiological relevance of the proposed mechanism. This contradiction should be discussed more thoroughly.
Reviewer #2 (Public review):
Summary:
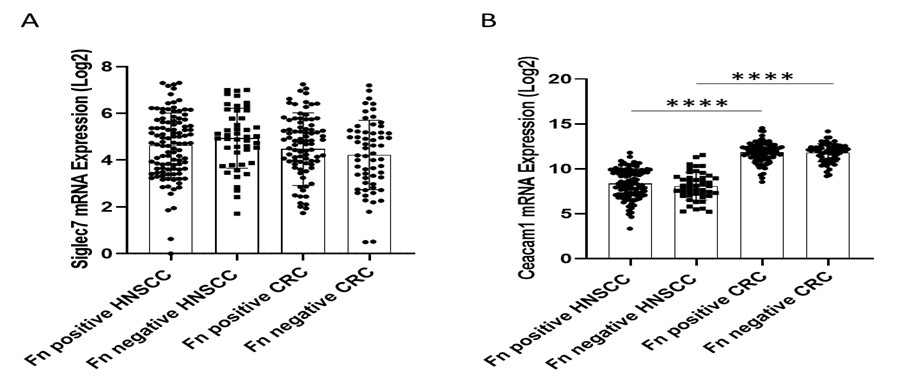
In the present study, Rishiq et al. investigated whether the RadD protein expressed by Fusobacterium nucleatum subsp. Nucleatum serves as a natural ligand for the NK-activating receptor NKp46, and whether RadD-NKp46 interaction enhances NK cell cytotoxicity against tumor cells. To address this, the authors first performed an association analysis of F. nucleatum abundance and NKp46 expression in head and neck squamous cell carcinoma (HNSC) and colorectal cancer (CRC) using the TCMA and TCGA databases, respectively. While a positive association between NKp46⁺ and F. nucleatum⁺ status with improved overall survival was observed in HNSC patients, no such correlation was found in CRC.
Next, they examined the binding of NKp46-Ig to various F. nucleatum strains. To confirm that this interaction was mediated specifically by RadD, they employed a RadD-deficient mutant strain. Finally, to establish the functional relevance of the RadD-NKp46 interaction in promoting NK cell cytotoxicity and anti-tumor responses, they utilized a syngeneic mouse breast cancer model. In this setup, AT3 cells were orthotopically implanted into the mammary fat pad of C57BL/6 wild-type (WT) or Ncr1-deficient (NCR1⁻/⁻; murine orthologue of human NKp46) mice, followed by intravenous inoculation with either WT F. nucleatum or the ∆RadD mutant strain.
Strengths:
A notable strength of the work is that it identifies a previously unrecognized activating interaction between F. nucleatum RadD and the NK cell receptor NKp46, demonstrating that the same bacterial protein can engage distinct NK cell receptors (activating or inhibitory) to exert context-dependent effects on anti-tumor immunity. This dual-receptor insight adds depth to our understanding of F. nucleatum-immune interactions and highlights the complexity of microbial modulation of the tumor microenvironment.
Weaknesses:
(1) A previous study by this group (PMID: 38952680) demonstrated that RadD of F. nucleatum binds to NK cells via Siglec-7, thereby diminishing their cytotoxic potential. They further proposed that the RadD-Siglec-7 interaction could act as an immune evasion mechanism exploited by tumor cells. In contrast, the present study reports that RadD of F. nucleatum can also bind to the activating receptor NKp46 on NK cells, thereby enhancing their cytotoxic function.
While F. nucleatum-mediated tumor progression has been documented in breast and colon cancers, the current study proposes an NK-activating role for F. nucleatum in HNSC. However, it remains unclear whether tumor-infiltrating NK cells in HNSC exhibit differential expression of NKp46 compared to Siglec-7. Furthermore, heterogeneity within the NK cell compartment, particularly in the relative abundance of NKp46⁺ versus Siglec-7⁺ subsets, may differ substantially among breast, colon, and HNSC tumors. Such differences could have been readily investigated using publicly available single-cell datasets. A deeper understanding of this subset heterogeneity in NK cells would better explain why F. nucleatum is passively associated with a favorable prognosis in HNSC but correlates with poor outcomes in breast and colon cancers.
(2) The in vivo tumor data (Figure 5D-F) appear to contradict the authors' claims. Specifically, Figure 5E suggests that WT mice engrafted with AT3 breast tumors and inoculated with WT F. nucleatum exhibited an even greater tumor burden compared to mice not inoculated with F. nucleatum, indicating a tumor-promoting effect. This finding conflicts with the interpretation presented in both the results and discussion sections.
(3) Although the authors acknowledge that F. nucleatum may have tumor context-specific roles in regulating NK cell responses, it is unclear why they chose a breast cancer model in which F. nucleatum has been reported to promote tumor growth. A more appropriate choice would have been the well-established preclinical oral cancer model, such as the 4-nitroquinoline 1-oxide (4NQO)-induced oral cancer model in C57BL/6 mice, which would more directly relate to HNSC biology.
(4) Since RadD of F. nucleatum can bind to both Siglec-7 and NKp46 on NK cells, exerting opposing functional effects, the expression profiles of both receptors on intratumoral NK cells should be evaluated. This would clarify the balance between activating and inhibitory signals in the tumor microenvironment and provide a more mechanistic explanation for the observed tumor context-dependent outcomes.