Author Response
The following is the authors’ response to the original reviews.
We sincerely thank the editor and reviewers for their constructive feedback on our manuscript. Based on their recommendations, we've conducted additional experiments, made revisions to the text and figures, and provide a point-by-point response below.
Reviewer #1 (Recommendations for the authors):
1) The lack of behavioral/physiological measures of the depth of anesthesia (ventilation, heart rate, blood pressure, temperature, O2, pain reflexes, etc...) combined with the lack of dose-response and the use of different routes of administration makes the data difficult to interpret. Sure, there is a clear difference in network activation between KET and ISO, but are those effects due to the depth of the anesthesia, the route of administration, and the dose used? The lack of behavioral/physiological measures prevents the identification of brain regions responsible for some of the physiological effects and different effects of anesthetics.
We greatly appreciate the insightful feedback you have provided.
In response to the concerns about anesthesia depth:
a. We recorded EEG and EMG data both before and after drug administration. Supplementary Figure 1 showcases the changes in EEG and EMG power observed 30 minutes post-drug administration, normalized to a 5-minute baseline taken prior to the drug's administration. Notably, no significant differences were detected in the normalized EEG and EMG power between the ISO and KET groups. Given the marked statistical differences observed between the EEG power in the KET and saline groups, and the EMG power in the home cage and ISO groups, we infer that both anesthetics effectively induced a loss of consciousness.
b. We used standard methods and doses for inducing c-Fos expression with anesthetics, as documented in prior studies (Hua, T, et al., Nat Neurosci, 2020; 23(7): 854-868; Jiang-Xie, L F, et al., Neuron, 2019; 102(5): 1053-1065.e4; Lu, J, et al., J Comp Neurol, 2008; 508(4): 648-62). In future research, it might be more optimal to adopt continuous intraperitoneal or intravenous administration of ketamine.
c. Within the scope of our study, while disparities in anesthesia duration might potentially influence the direct statistical comparison of ISO and KET, such disparities wouldn't compromise the identification of brain regions activated by KET or ISO when assessed as distinct stimuli (ISO vs. home cage; KET vs. saline) or in relation to their individual functional network hub node results.
We hope these additions and clarifications adequately address your concerns and enhance the comprehensibility of our data.
2) Under anesthesia there should be an overall reduction of activity, is that the case? There is no mention of significantly downregulated regions. The authors use multiple transformations of the data to interpret the results (%, PC1 values, logarithm) without much explanation or showing the full raw data in Fig 1. It would be helpful to interpret the data to compare the average fos+ neurons in each region between treatment and control for each drug.
Absence of Significantly Downregulated Regions Under Anesthesia: There are two primary reasons for this observation:
a. Our study's sampling time for the home cage, ISO, saline, and KET groups was during Zeitgeber Time (ZT) 6-7.5. During this period, mice in both the home cage and saline groups typically showed reduced spontaneous activity or were in a sleep state. Our Supplementary Figure 1 EEG and EMG data corroborate this, revealing no significant statistical variations in EEG power between the home cage and ISO groups, nor in EMG power between the saline and KET groups.
b. Our immunohistochemical data showed that the total number of c-Fos positive cells in the two control groups was notably lower than in the experimental groups (Saline group vs KET group: 11808±2386 versus 308705±106131, P = 0.006; Home cage vs ISO group: 3371±840 vs 12326±1879, P = 0.001). This is in line with previous studies, like the one by Cirelli C and team, which found minimal c-Fos expression throughout the mouse brain during physiological sleep (Cirelli, C, and G Tononi, Sleep, 2000; 23(4): 453-69). Thus, in our analysis, we did not detect regions with significant downregulation when comparing anesthetized mice with controls.
Interpreting Raw Data from Figure 1: Regarding the average Fos+ neurons:
In Figures 4 and 5, we utilized raw data (c-Fos cell count) to assess cell expression differences across 201 brain regions within each group. Only brain regions that had significant statistical differences after multiple comparison corrections are shown in the figures.
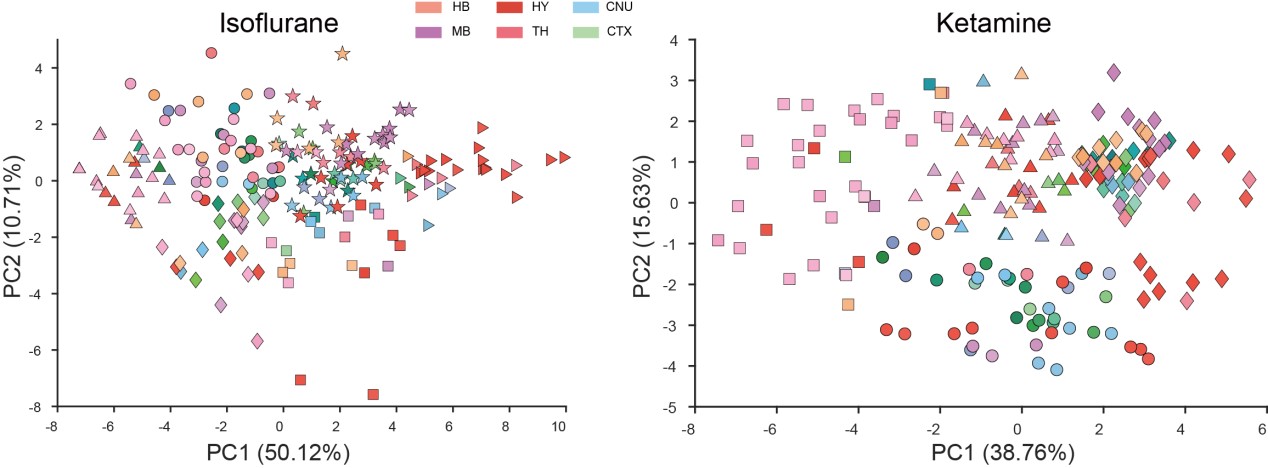
3) I do not understand their interpretation of the PCA analyses. For instance, in Fig 2 they claim that KET is associated with PC1 while ISO is associated with PC2. Looking at the distribution of points it's clear that the KET animals are all grouped at around +2.5 on PC1 and -2.0 on PC2, this means that KET is associated with both PC1 and PC2 to a similar degree (2 to 2.5). Moreover, I'm confused about why they use PCA to represent the animals/group. PCA is a powerful technique to reduce dimensionality and identify groups of variables that may represent the same underlying construct; however, it is not the best way to identify clusters of individuals or groups.
Clarification on PCA Analyses in Figure 2: Thank you for pointing out the ambiguities in our initial presentation of the PCA analyses. We are grateful for the opportunity to address these concerns.
KET and ISO Associations with PC1 and PC2: You rightly observed that KET samples manifest both a positive value on PC1 (around +2.5) and a negative one on PC2 (around -2.0), suggesting that KET has a substantial influence on both principal components. In PCA, a positive score implies a positive association with that component, whereas a negative score suggests a negative association. Contrarily, ISO samples predominantly exhibit values around +2.5 on PC2, with nearly neutral values for PC1, underlining its stronger association with PC2 and lack of significant correlation with PC1. To ensure transparency and clarity, we've adjusted the corresponding descriptions in our manuscript, which can be found on Line 100.
Rationale Behind Using PCA to Represent Animals/Groups: Our initial step was to conduct PCA clustering analysis on the 201 brain regions within both the ISO and KET groups. In the accompanying chart, varying colors denote different brain regions, while distinct shapes represent separate clusters. There wasn't a pronounced distribution pattern within the ISO and KET groups, which led us to adopt the current computational method presented in the paper. This approach was chosen to directly contrast the relative differential expressions between ISO and KET.
We deeply value your feedback, which has steered us toward a clearer and more accurate presentation of our data. We genuinely appreciate your meticulous review.
Author response image 1.
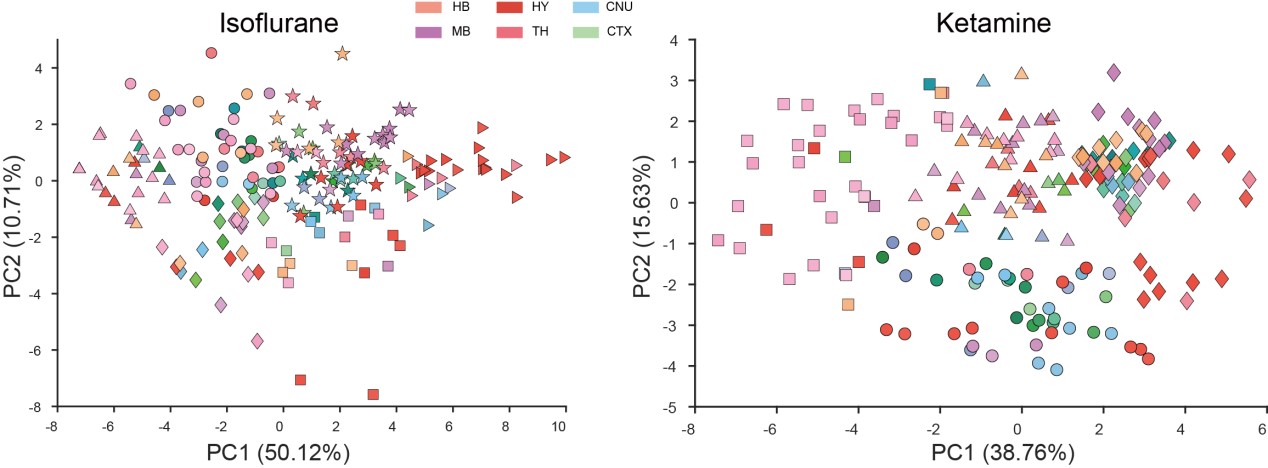
4) The actual metric used for the first PCA is unclear, is it the FOS density in each of the regions (some of those regions are large and consist of many subregions, how does that affect the analysis) is it the %-fos, or normalized cells? The wording describing this is variable causing some confusion. How would looking at these different metrics influence the analysis?
Thank you for raising concerns about the metrics used in our PCA analysis. We recognize the need for clearer exposition and appreciate the opportunity to clarify.
PCA Metrics: The metric for our PCA is calculated by obtaining the ratio of the Fos density within a specific brain region to the global Fos density across the brain. Briefly, this entails dividing the number of Fos-positive cells in a given region by its volume, and then comparing this to the Fos density of the whole brain. The logarithm of this ratio provides our PCA metric. We've elaborated on this in the Materials and Methods section (Lines 401) and enhanced clarity in our revised manuscript, particularly at Line 96.
In Figure 2A, we employed 53 larger, mutually exclusive brain regions based on the reference from the study by Do et al. (eLife, 2016;5:e13214). However, in Figure 3A, we used a more detailed segmentation, incorporating 201 distinct brain areas that are more granular than those in Figure 2A. Notably, the PCA results from both representations were consistent. The rationale behind selecting either the 53 or 201 brain regions can be found in our response to Question 10.
Rationale for Metric Choice: The log ratio of regional c-Fos densities relative to the global brain density was chosen due to:
a. Notable disparities in c-Fos cell expression across the groups.
b. A significant non-normal distribution of density values across animals within the group.
Employing the log ratio effectively mitigates the impact of extreme values and outliers, achieving a more standardized data distribution.
We've added PCA plots based on c-Fos densities, depicted in Author response image 2. However, the data dispersion has resulted in a significantly spread-out horizontal scale for these visuals.
Author response image 2.
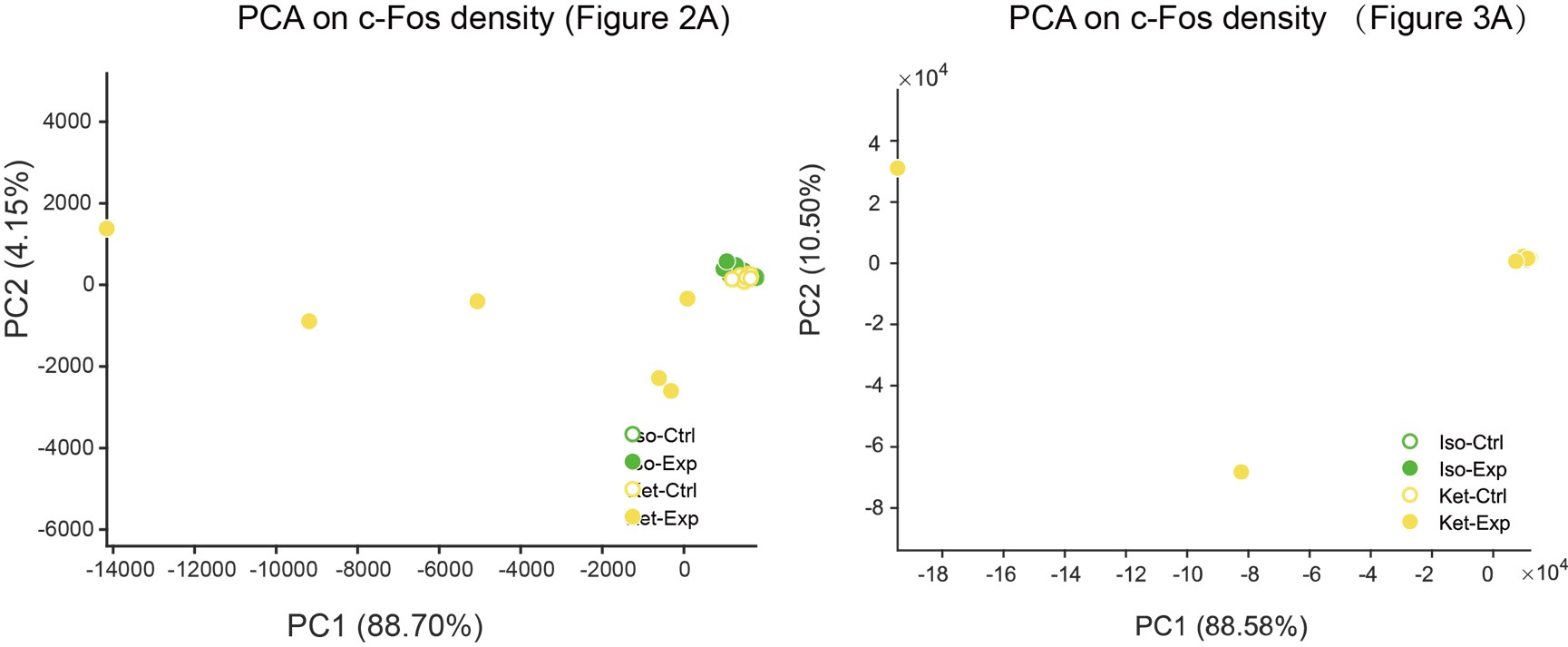
5) Based on Fig 3 the authors concludes that ISO activates the hypothalamic regions and inhibits the cortex, however, Fig 1 shows neither an activation of the hypothalamus in the ISO nor an inhibition of the cortex when compared to home cage control. If anything it suggests the opposite.
Thank you for your insightful observations regarding the discrepancies between Figures 2 and 3. We believe that when you refer to Figure 1, you are actually referencing Figure 2C.
ISO activation in Hypothalamus: In Figure 2C, we regret the oversight where we inadvertently interchanged the positions of ISO and Saline. When accurately represented, Figure 2C indeed shows that ISO notably activates the periventricular zone (PVZ) and the lateral zone (LZ) of the hypothalamus compared to the home cage group. Moreover, there's a discernible difference in the hypothalamic response between ISO and KET.
ISO's Effect on the Cortex: The main aim of Figure 3 was to highlight the differing responses between ISO and KET in the cortex. Notably, KET demonstrates a positive correlation with PC1 (+7 on PC1), whereas ISO shows a negative association (-3 on PC1). Given that the coefficient of PC1 for the cortical region is positive, it suggests that the cortical areas activated by KET are inhibited by ISO (with KET's distribution around 0 on PC2). However, the divergence between ISO and the home cage is most apparent in PC2, with ISO clusters at +4 and the home cage approximately at -2, suggesting that ISO activates a different set of cortical nuclei. In alignment with this, Figure 2C also illustrates that ISO activates specific cortical areas, such as ILA and PIR, in contrast to the home cage.
Thus, Figure 3 primarily employs PCA to delineate the contrasts between ISO and KET, whereas Figure 2C emphasizes the comparison of each against their respective controls.
6) Control for isoflurane should be air in the induction chamber rather than home cage. It is possible that Fos activation reflects handling/stress pre-anesthesia in the animals, which would increase Fos expression in the stress-related regions such as the BST, striatum (CeA), hypothalamus (PVH) and potentially the LC.
Thank you for emphasizing the importance of an appropriate control for Isoflurane.
In our efforts to minimize the potential impact of stress-induced c-Fos expression, we implemented several precautionary measures. Prior to the experiment, both groups of mice were subjected to handling and acclimatization within the induction chamber over four days. By the day of the experiment, for the mice in the experimental group, we ensured they were comfortable and exhibited no signs of distress or fear—such as cowering or evading. With care, we slowly relocated them to the nearby anesthesia induction chamber. Using 5% ISO, anesthesia was induced promptly, following a meticulously devised protocol to reduce stress impacts on c-Fos expression.
Moreover, existing studies have shown Isoflurane's activation of BST/CeA (Hua, T, et al., Nat Neurosci, 2020, 23: 854-868), PVH (Xu, Z, et al., British Journal of Anaesthesia, 2023, 130: 446-458), and LC (Lu, J, et al., J Comp Neurol, 2008, 508: 648-62), even when using oxygen controls. Such literature supports our findings, indicating that the activation we observed was indeed due to Isoflurane and not purely stress-related.
7) In the Ket network there are a few anticorrelated regions, most of which are amongst the list of the most activated regions, does this mean that the strong correlation results from an overall decreased activation? And if so, is it possible that the ketamine anesthesia was stronger than the isoflurane, causing a more general reduction in activity?
The pronounced correlations observed within the ketamine (KET) network do not signify a generalized decrease in activation. Instead, these correlations reflect significantly enhanced activity in specific regions under KET anesthesia. This amplified correlation is an indication of a more widespread increase in activity, rather than a decrease. These findings are consistent with previous research, which showed that anesthetic doses of ketamine produce patterns of Fos expression in the CNS similar to wakefulness (Lu, J, et al., J Comp Neurol, 2008; 508(4): 648-62).
Regarding the comparative strength of KET versus ISO anesthesia, our electroencephalographic evidence confirms that both agents induce a loss of consciousness. No significant differences were observed in EEG and EMG readings within the first 30 minutes post-administration. In future research, a continuous intravenous or intraperitoneal administration of KET might be a preferable method.
8) Since they have established networks it would be easy and useful to look at how the different regions identified (sleep, pain, neuroendocrine, motor-related, ...) work together to maintain analgesia, are they within the same module? Do they become functionally connected and is this core network of functional connections similar for KET and ISO?
Thank you for your suggestion. In response to your inquiry, we undertook analysis of the core functional networks for KET and ISO, using a set threshold at r>0.82 and P<0.05. For evaluating the modularity of each network, we utilized Newman's spectral community detection algorithm.
(A) The ISO’s core functional network (56 nodes, 372 edges) predominantly divides into two modules with a modularity quotient of 0.345. ISO-active regions include arousal-associated regions (PL, ILA, PVT), analgesia-related (CeA, LC, PB), neuroendocrine function nuclei (TU, PVi, ARH, PVH, SON) as detailed in Figure 5. Notably, ARH and SON weren't incorporated into the core network. Analgesia-associated regions, such as CeA, LC, and PB, reside within module 1, while neuroendocrine nuclei are spread between modules 1 and 2.
(B) In contrast, KET's core functional network (61 nodes, 1820 edges) splits into three distinct modules, but its low modularity quotient (0.06) indicates a lack of clear functional modularization, suggesting denser interconnections among brain regions. Furthermore, functionally-related regions such as arousal (PL, ILA, PVT, DR), analgesia-related (ACA, APN, PAG, LC), and neuroendocrine regulation (PVH, SON),etc., as seen in Figure 4, are distributed across different modules. This distribution may implies that functions like analgesia and neuroendocrine regulation are not governed by simple, linear processes, but arise from complex, overlapping pathways spanning various modules and functional zones.
In summary, the core functional networks of ISO and KET differ, with functionally-related regions spanning multiple modules, reflecting their diverse roles in varied physiological regulations.
Author response image 3.
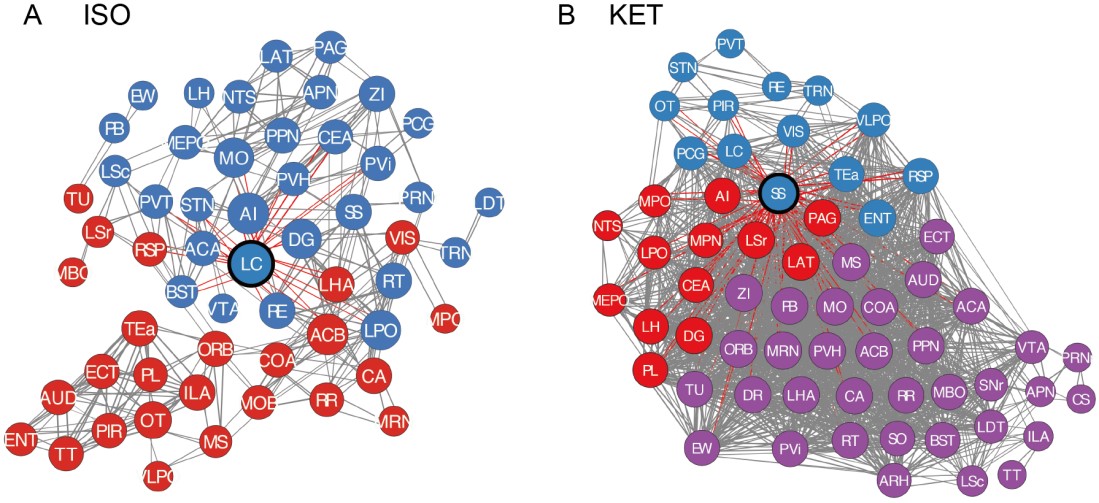
9) The naming of the function of some of the regions is very much debatable. For instance, PL/ILA are named "sleep-wakefulness regulation" regions in the paper. I can think of many more important functions of the PL/IL including executive functions, behavioral flexibility, and emotional control. It is unclear how the functions of all the regions were attributed. I am not sure that this biased labeling of structure-function is useful to the reports, it may instead suggest wrong conclusions.
Thank you for your thoughtful feedback regarding our classification of the functions of the PL/ILA regions in our manuscript.
We recognize the challenge in accurately defining the functions of brain regions. While there is evidence highlighting the role of PL/ILA in arousal pathways, we also acknowledge their documented roles in executive functions, behavioral flexibility, and emotional control. In response to your comments, we have refined our description, changing "sleep-wakefulness regulation" to "wake-promoting pathways" (see Line: 159, 164).
It's worth noting that many brain regions, including the PL/ILA, have multiple functions. We agree that a single label might not capture the entirety of their roles. To provide a broader perspective, we will add a section in our manuscript that sheds light on the varied functions of these regions (Line: 181).
10) A point of concern and confusion is the number of brain regions analyzed. In the introduction, it is mentioned that 987 brain regions are considered, but this is reduced to 53 selected brain regions in Figure 2, then 201 brain regions in Figure 3, and reduced again to 63 for the network analysis. The rationale for selecting different brain regions is not clear.
For the 987 brain regions: Using the standard mouse atlas available at http://atlas.brain-map.org/, the mouse brain is organized into nine levels. The broadest category is the grey matter, which then progresses to more specific subdivisions, totaling 987 unique regions.
For the 53 brain regions: To effectively understand the activation patterns of ISO and KET, we started with a broad approach, looking at larger brain areas like the thalamus and hypothalamus. This broad view, presented in Figure 2, focuses on the 5th-level brain regions, encompassing 53 primary areas. This methodology is also employed in the study by Do et al. (Elife, 2016; 5: e13214). We have added the rationale for selecting these brain regions in the main text (Line: 92).
Regarding the 201 brain regions in Figures 3, 4, and 5: We delved deeper, examining the 6th-level brain regions, a common granularity in neuroscience research. This detailed view allowed us to highlight specific areas, like the CeA and PVH (Line:129).
Finally, for Figures 6 and 7, we selected 63 regions that were activated by both ISO and KET, as well as regions previously reported to be related to the mechanism of general anesthesia(Leung, L, et al., Progress in neurobiology, 2014; 122: 24-44) (Line: 220). Using these regions, we analyzed the correlation of c-Fos expression, aiming to construct a functional brain network with strong positive connections.
We hope this clarifies our approach and the rationale behind our region selection at each stage of the study. Thank you for your attention to this detail.
11) The statistical analysis does not seem appropriate considering the high number of comparisons. They use simple t-tests without correction for multiple comparisons.
Thank you for pointing out the concern regarding our statistical analysis. In the revised manuscript, we addressed the issue of multiple comparisons correction in our t-tests. We adopted the statistical methods detailed in the papers by Renier, N, et al., Cell, 2016; and Benjamini, Y, and Y Hochberg, 1995. P-values were adjusted for multiple comparisons using the two-stage linear step-up procedure of Benjamini, Krieger, and Yekutieli, with a false discovery rate (FDR) threshold (Q) of 0.05. This approach is now explained in the Materials and Methods section (Line: 434). After this adjustment, the brain regions we initially identified remained statistically significant. Furthermore, we revisited the original immunohistochemical images to confirm the differences in c-Fos cell expression between the experimental and control groups, reinforcing our conclusions.
12) There is no statistical analysis in Fig 2C。
Thank you for bringing to our attention the lack of statistical analysis in Fig 2C. We have now added the relevant statistical data in Supplementary Table 1 and provided annotations in Fig 2C to reflect this.
Reviewer #2
1) The authors report 987 brain regions in the introduction, but I cannot find any analysis that incorporates these or even which regions they are. Very little rationale is provided for the regions included in any of the analyses and numbers range from 53 in Figure 1, to 201 in Figure 3, to 63 in Figure 6. It would help if the authors could first survey Fos+ counts across all regions to identify a subset that is of interest (significantly changed by either condition compared to control) for follow up analysis.
Thank you for your insightful comments on the number of brain regions analyzed in our study.
987 Brain Regions: The reference to 987 brain regions from the standard mouse atlas (http://atlas.brain-map.org/) represents the entire categorization of the mouse brain across nine levels. We recognize that a comprehensive analysis of all these regions would be valuable, but to ensure clarity and depth, we took a focused approach.
Region Selection Rationale:
Figure 2: Concentrated on 5th-level brain regions (53 areas), inspired by methods from Do et al. (eLife, 2016;5:e13214). This provided a broad overview of c-Fos expression differences.
Figures 4 and 5: Delved into 6th-level brain regions (201 areas), a common practice in neuroscience for more detailed study.
Figure 6: We focused on 63 regions, which encompass not only the regions activated by both ISO and KET but also those previously reported to be associated with the mechanisms of general anesthesia.
Methodological Approach: Our region selection was rooted in identifying areas with significant changes under anesthetic conditions compared to controls. This staged approach allowed a targeted analysis of the most affected regions, ensuring robust conclusions.
Enhancements: We've incorporated comparative analyses of activated brain regions at different hierarchical levels in Figures 4 and 5. For clearer comprehension, we’ve added clarifications in the manuscript at Lines: 92, 130, and 220.
2) Different data transformations are used for each analysis. One that is especially confusing is the 'normalization' of brain regions by % of total brain activation for each animal prior to PCA analysis in Figures 2 and 3. This would obscure any global differences in activation and make it unlikely to observe decreases in activation (which I think is likely here) that could be identified using the Fos+ counts after normalizing for region size (ie. Fos+ count / mm3) which is standard practice in such Fos-based activity mapping studies. While PCA can be powerful approach to identify global patterns, the purpose of the analysis in its current form is unclear. It would be more meaningful to show that regional activation patterns (measured as counts/mm3) are on separate PCs by group.
Thank you for your thoughtful comments. We regret any confusion caused by our initial presentation. For the PCA analysis in Figures 2A and 3A, we calculated the ratio of cell density in each brain region to the overall brain density, and then applied a logarithmic transformation to this ratio. Our approach in Figure 2C was to use the proportion of c-Fos cell counts in individual brain regions to the total cell counts throughout the brain. This methodology considers variations in overall c-Fos cell counts across animals, effectively mitigating potential biases due to differential global activation levels across subjects.
Furthermore, our direct comparison of differences in c-Fos cell counts between ISO, KET, and their respective control groups in Figures 4 and 5 addresses your concerns about potential decreases in activation. Notably, we did not identify any brain regions with significant suppression in these figures, which is consistent with the trends observed post-normalization in Figure 2C.
Given your feedback, we conducted another PCA using cell densities for each region (counts/mm3). However, we found significant variability and non-normal distribution of c-Fos density across the groups, leading to extensive data dispersion. Consequently, normalizing the cell counts across regions and then applying a logarithmic transformation before PCA might be more appropriate.
Author response image 4.
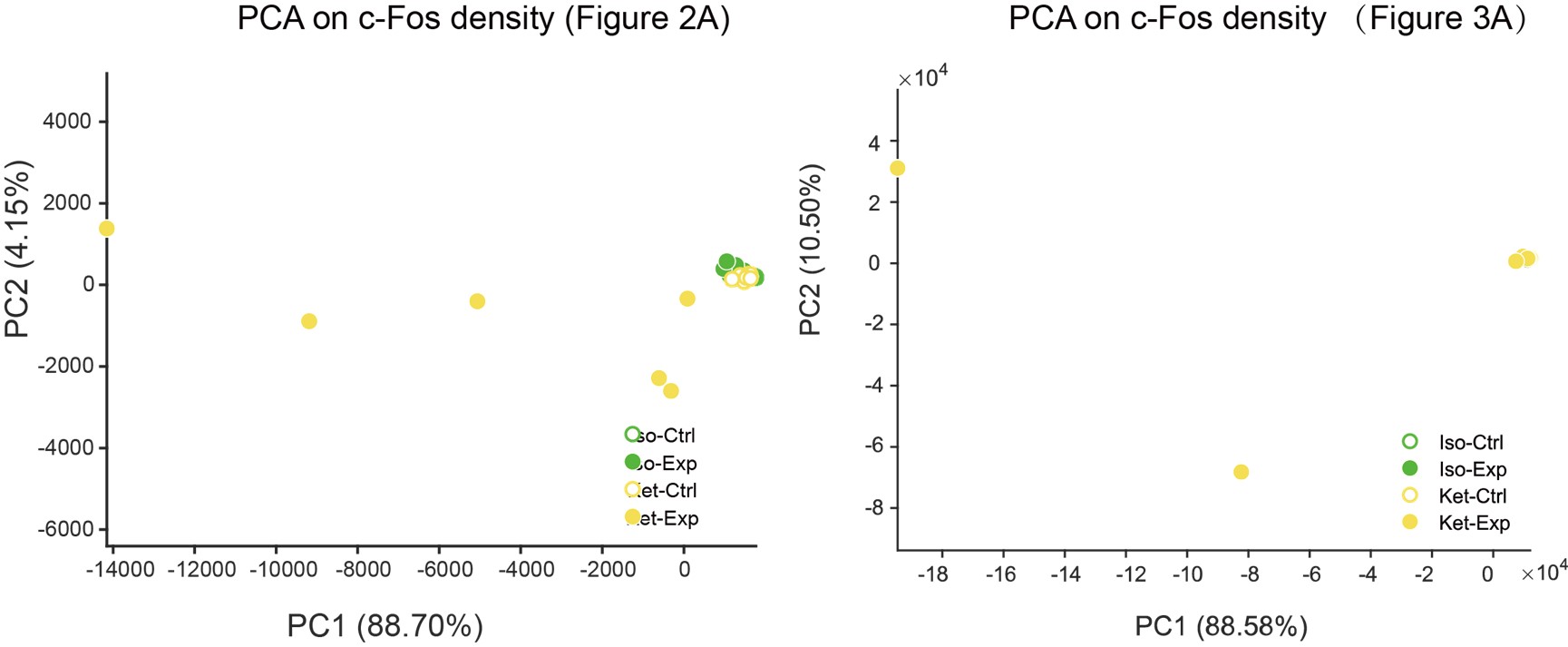
Additionally, our exploration of regional activation patterns using PCA analysis for ISO and KET separately, based on the logarithm ratio of the c-Fos density, revealed that there was no distinct clustering feature among the different brain regions (as illustrated in Author response image 5: colors represented distinct brain regions, while the shapes were indicative of different clusters). This observation further suggests that our original statistical approach might be more suitable.
Author response image 5.
3) Critical problem: The authors include a control group for each anesthetic (ketamine vs. saline, isofluorane vs. homecage) but most analyses do not make use of the control groups or directly compare Fos+ counts across the groups. Strictly speaking, they should have compared relative levels of induction by ketamine versus induction by isoflurane using ANOVAs. Instead, each type of induction was separate from the other. This does not account for increased variability in the ketamine versus isoflurane groups. There is no mention in the Statistics section or in Results section that any multiple comparison corrections were used. It appears that the authors only used Students t-test for each region and did not perform any corrections.
We appreciate the reviewer's insights and have addressed your concerns:
Given the pronounced difference in c-Fos cell count expression between the KET and ISO groups, a direct comparison of Fos+ counts may not effectively capture their inherent disparities. To better highlight these distinctions, we used the logarithm ratio of c-Fos density in our PCA analysis (Figure 3), mitigating potential disparities in overall cell counts between samples and emphasizing relative variations. However, in response to your feedback, we've included additional analyses. Author response image 6 depicts the c-Fos density (cells/mm^3) across different brain regions for the home cage, ISO, saline, and KET groups, with regions like the cerebral cortex, cerebral nuclei, thalamus, and others differentiated by shaded backgrounds. Data are represented as mean ± SEM. We performed a one-way ANOVA followed by Tukey’s post hoc test, marking significant differences between ISO and KET with asterisks: P < 0.001, P < 0.01, P < 0.05.
Regarding multiple comparison corrections, we've conducted thorough analyses on the data in Figure 2C and Figures 4, 5, and 6, implementing multiple comparison corrections. The detailed methodology is provided in the “Statistical analysis” section.
Author response image 6.
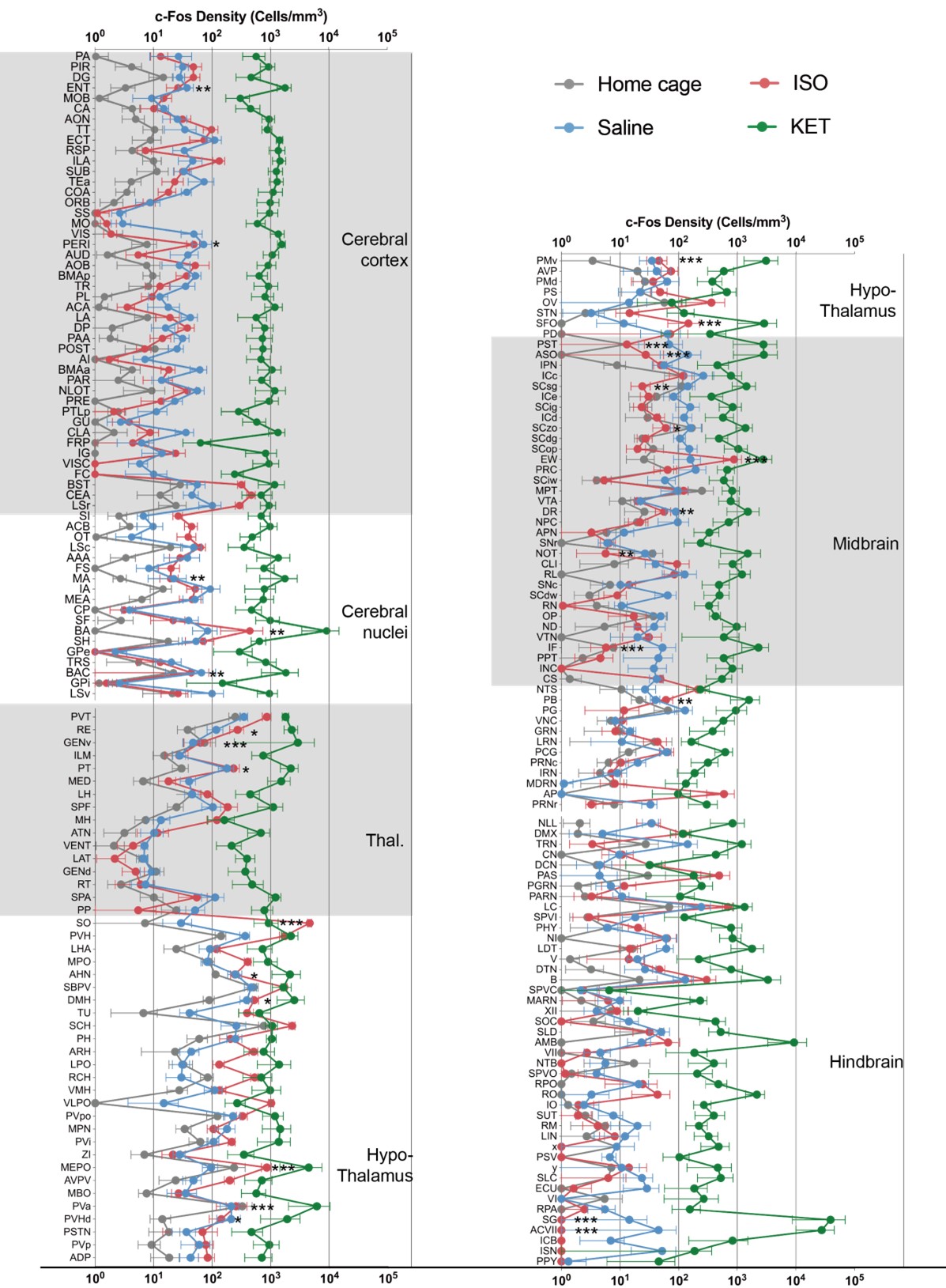
4) Figures 4 and 5 show brain regions 'significantly activated' following KET or ISO respectively, but again a subset of regions are shown and the stats seem to be t-tests with no multiple comparisons correction. It would help to show these two figures side by side, include the same regions, and keep the y axis ranges similar so the reader can easily compare the 'activation patterns' across the two treatments. Indeed, it looks like KET/Saline induced activation is an order or magnitude or two higher than ISO/Homecage. I would also recommend that this be the first data figure before any other analyses and maybe further analysis could be restricted to regions that are significantly changed in following KET or ISO here.
Thank you for your constructive feedback regarding Figures 4 and 5.
Comparison and Presentation of Figures 4 and 5: We acknowledge your suggestion to present these figures side by side for easier comparison. In the supplementary figure provided in the previous question, we've placed Figures 4 and 5 adjacent to each other, with consistent y-axis ranges, ensuring that readers can make direct comparisons between the activation patterns elicited by KET and ISO.
Statistical Concerns and Region Selection: As mentioned in our previous response, we have conducted multiple comparison corrections on the data presented in Figures 4 and 5. Detailed procedures are elaborated in the “Statistical analysis” section. We believe this approach addresses your concerns regarding the use of t-tests without corrections for multiple comparisons.
Difference in Activation Levels: We observed that the c-Fos activation due to KET is significantly higher than that from ISO. When presented side-by-side using the same scale, ISO activations appear less prominent, potentially mask subtle differences in the activation patterns of ISO, particularly if both KET and ISO showed changes in the same direction in certain brain regions but differed in magnitude. To address this, we used the proportion of c-Fos cell counts in Figure 2C, the logarithm ratio of c-Fos density in Figure 2A and Figure 3. This method emphasizes the relative changes, rather than absolute values, giving a more balanced view of the effects of each treatment.
5) Analyses in Figure 6 and 7 are interesting but again the choice of regions to include is unclear and makes interpreting the results impossible. For example, in Figure 7 it is unclear why the list of regions in bar graphs showing Degree and Betweenness Centrality are not the same even within a single row?
Thank you for your pertinent observation. The choice of brain regions in Figures 6 and 7 was carefully determined based on two main criteria: regions that were significantly activated by ISO or KET within the scope of our study, and those previously reported to be associated with anesthesia mechanisms and sleep-wake regulation.
Regarding your second concern on Figure 7, the discrepancies observed in the x-axes of the bar graphs arise from our methodological approach. We prioritized presenting the top 20% of regions based on their Degree or Betweenness Centrality values. By separately ranking these regions from highest to lowest, the regions presented for each metric inherently differ. This approach was taken to elucidate nodes that consistently emerge as significant across both metrics, thereby highlighting core nodes in the functional network. Were we to use a consistent x-axis without this ranking, it would not only necessitate a more extensive presentation but might also dilute the emphasis on key information. To clarify this methodology and its rationale for our readers, we have expanded upon this in the manuscript at Line 243.
We hope these clarifications address your concerns and facilitate a clearer understanding of our findings.
Reviewer #1 (Recommendations For The Authors):
Minor points
1) In Table 1: the separation of which substructures belong to which brain structure is not clear
2) Line 132 on page 3 seems to repeat the sentence earlier in the paragraph "KET predominantly affects brain regions within the cerebral cortex (CTX), while significantly inhibiting the hypothalamus, midbrain, and hindbrain."
3) Typos
a) Line 99/100 and 130 Central nucleus (CNU) should be cerebral nucleus
b) Comma on line 166
c) Fig. 4D: KET instead of Keta
d) Line 263 "ep"
e) Line 332: 35" "ml (add space)
4) Will data and code be made available?
Thank you for your detailed feedback.
-
We have revised Table 1 to clarify which substructures belong to which brain structures.
-
We acknowledge the redundancy and have now edited line 139 on page 3 to remove the repeated sentence regarding the effects of KET on brain regions.
-
We have addressed the typos you pointed out:
a. The terms "Central nucleus (CNU)" have been corrected to "cerebral nucleus."
b. The comma issue on line 166 has been rectified.
c. In Fig. 4D, we have corrected "Keta" to "KET."
d. We have corrected the typo "ep" on line 263.
e. A space has been added between "35" and "ml" on line 332 as you indicated.
- Regarding the availability of data and code, we are currently conducting additional analyses related to this study. Once these analyses are completed, we will be more than happy to make the data and code available.
Thank you for assisting us in improving our manuscript.
Reviewer #2 (Recommendations For The Authors):
Minor comments:
6) The term 'whole-brain mapping' in the title suggests that the mapping was performed on 'intact brains' where in fact serial sections were used here. Maybe the authors could change to 'brain-wide mapping' to align better with the study.
Thank you for your insightful comments.
We have revised the title as suggested, changing "whole-brain mapping" to "brain-wide mapping".
7) It is unclear if the mice were kept under anesthesia for the 90-min duration and how the authors monitored the level of sedation. Additionally, if the KET mice were already sedated why were they further sedated with ISO before perfusions and tissue extraction? The methods should be clarified and any potential confounds discussed.
To maintain consistency in the experimental protocol and to reduce stress reactions in the mice, ISO was used before perfusion in all cases. However, this does not affect c-Fos expression as the expression of c-Fos protein starts 20-30 minutes after stimulation (Lara Aparicio, S Y, et al., NeuroSci, 2022; 3(4): 687-702).
We appreciate your guidance in enhancing the clarity of our manuscript.
Reviewer #3 (Recommendations For The Authors):
Recommendation: Minor corrections.
1) The authors should delve deeper into the molecular mechanisms underlying the observed effects, particularly the changes associated with NMDA and GABA receptors. Exploring these mechanisms would provide a more comprehensive understanding of how Ketamine and Isoflurane modulate neural activity and induce anesthesia.
2) The clinical relevance of these findings has not been sufficiently addressed. It would be valuable to elaborate on how the current research outcomes could potentially lead to changes in current anesthesia practices. For instance, identifying the distinct pathways of action for Ketamine and Isoflurane could aid anesthesiologists in selecting the most appropriate anesthetic based on the specific needs of individual patients or surgical procedures.
3) Both Ketamine and Isoflurane have been associated with neurotoxicity. It is important to discuss how the c-Fos activation induced by these anesthetics could contribute, at least partially, to anesthesia-related neurotoxicity. Examining the potential neurotoxic effects would provide a more comprehensive understanding of the risks associated with these anesthetics and aid in the development of safer anesthesia protocols.
Thank you for your valuable suggestions.
Regarding the three points (1, 2, and 3) you've raised, we fully recognize their significance. In the current study, our primary focus was on the differential impacts of Isoflurane and Ketamine on widespread c-Fos expression in the brain. However, we indeed acknowledge the importance of delving deeper into these mechanisms and their clinical relevance. Therefore, we intend to explore these critical issues in greater detail in our future research endeavors.
We appreciate your feedback, which provides constructive guidance for our subsequent research directions.









