Author Response
On behalf of my co-authors, I thank you very much for sending our manuscript (# eLifeRP-RA-2023-91223) entitled “Elimination of subtelomeric repeat sequences exerts little effect on telomere functions in Saccharomyces cerevisiae” for review and providing us an opportunity for revision. We also thank the reviewers for their critical and constructive comments and suggestions which have helped us to strengthen our study. We have performed more experiments to address the concerns the reviewers raised, and we have also revised or corrected some of our statements as the reviewers suggested.
Reviewer #1
1) The author’s data indicate that cells with many chromosomes are more dependent on possibly homologous recombination than SY12 cells with three chromosomes. Telomerase-deficient cells exhibit the type I and type II telomere structures, whereas telomerase-deficient SY12 cells often generate different telomere structures (named Type X survivors or atypical survivors). Type I survivor depends on Rad51 possessing tandem Y' elements whereas Type II survivor depends on Rad59 carrying long TG sequences (line 60-70). Both types require Rad52 (line 66-70). At the moment, it is not determined how Type X or atypical survivors are generated in telomerase-deficient SY12 cells.
The authors need to determine whether Type X or atypical survivors depend on other repair pathways from Type I and Type II, and what DNA sequences are retained adjacent to telomeres in Type X or atypical survivors by sequencing analysis (Fig. 2).
We thank the reviewer’s valuable comments and suggestions. Atypical survivor is a subtype of survivor that exhibits non-uniform telomere patterns, distinct from those observed in Type I, Type II, Type X, or circular survivors. To further determine its genetic requirements, we deleted RAD52 in SY12 tlc1Δ, SY12YΔ tlc1Δ, SY12XYΔ tlc1Δ, and SY12XYΔ+Y tlc1Δ strains. Southern blotting results showed that neither Type I nor Type II survivors were found in the series of strains; circular survivor was in the predomination; beside circular survivor, some survivors exhibiting non-uniform telomere patterns suggested they were atypical survivor. These results have been presented as Figure 2—figure supplement 6B, Figure 5—figure supplement 2B and Figure 6—figure supplement 4B in the revised version. The results showed that atypical survivors still emerged when Rad52 pathway was repressed, indicating that the formation of atypical survivors does not strictly rely on the homologous recombination.
Given that "atypical" clones exhibit non-uniform telomere patterns, it’s not surprising that their chromosome structures are variable and tanglesome. Consequently, it is hard for us to amplify and sequence the DNA sequences retained adjacent to telomeres.
Since no Type X survivor was detected in SY12 tlc1Δ rad52Δ strain (Author response image 1A), we deleted RAD50 or RAD51 in SY12 tlc1Δ strain to investigate on which pathway the formation of the Type X survivor relied. Results showed that Type X survivor emerged in the absence of Rad51 but not Rad50, suggesting that the formation of Type X survivor depended on Rad50 pathway. These results have been presented as Figure 2—figure supplement 7.
To determine the chromosomal end structure of the Type X survivor, we randomly selected a typical Type X survivor, and performed PCR-sequencing analysis. The results revealed the intact chromosome ends for I-L, X-R, XIII-L, XI-R, and XIV-R, albeit with some mismatches compared with the S. cerevisiae S288C genome, which possibly arising from recombination events that occurred during survivor formation. Notably, the sequence of the Y’-element in XVI-L could not be detected, while the X-element remained intact. Figure 2—figure supplement 5 in the revised manuscript.
2) Survivor generation of each type (Type I, Type II, Type X or atypical and circularization) needs to be accurately quantitated. The authors concluded that X or Y' elements are not strictly necessary for survivor formation (Fig. 5 and Fig. 6). However, their removal appears to increase atypical survivor and chromosome circularization (Fig. 2 vs Fig. 5 and 6).
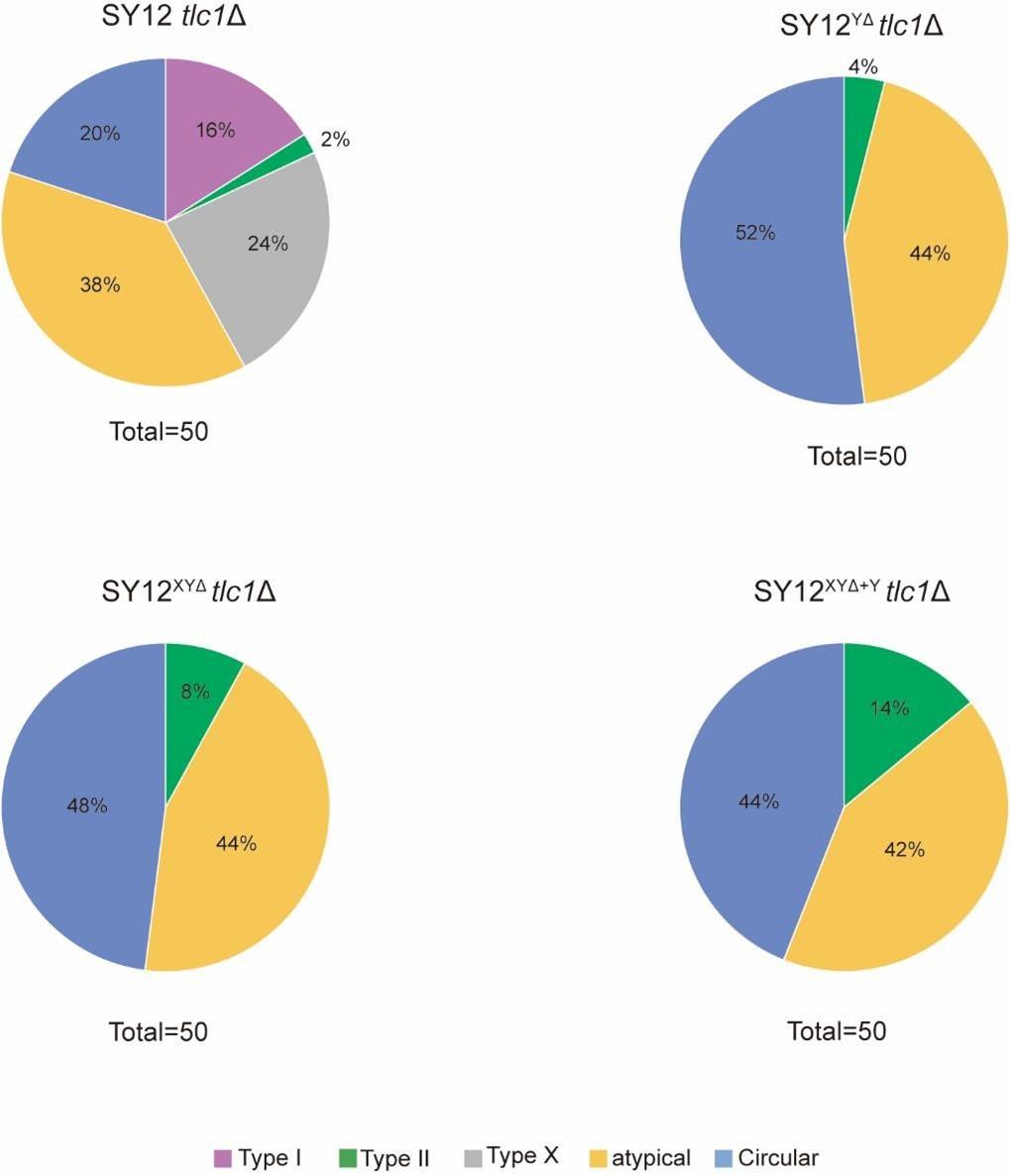
We are grateful for the reviewer’s critical and constructive suggestions. According to the reviewer’s requirement, we quantified each type of survivors in SY12 tlc1Δ, SY12YΔ tlc1Δ, SY12XYΔ tlc1Δ and SY12XYΔ+Y tlc1Δ strains (Figure 2D, 5C, 6A and 6B). In SY12 tlc1Δ strain, Type I survivors accounted for 16%, Type II survivors for 2%, Type X survivors for 24%, circular survivors for 20% and atypical survivors for 38%. In SY12YΔ tlc1Δ strain, 4% were Type II survivors, 52% were circular survivors and 44% were atypical survivors.
For the SY12XYΔ tlc1Δ strain, 8% were Type II survivors, 48% were circular survivors and 44% were atypical survivors. In SY12XYΔ+Y tlc1Δ strain, the proportions of Type II, circular and atypical survivors were 14%, 44%, and 42%, respectively (Author response image 1).
In comparing SY12YΔ with SY12XYΔ, we observed a similar ratio of circular and atypical survivors. This result indicates that the remove of X-elements exert little effect on the formation of circular and atypical survivors. Similarly, in SY12XYΔ+Y strain, the proportions of circular and atypical survivors were comparable to those in SY12XYΔ strain, indicating that Y’-elements also have little effect on the formation of circular and atypical survivors. However, due to the unknown frequency of survivor formation, alternative explanations of these data are possible. For example, subtelomeric elements previously suggested to have no impact on the formation of any survivor types might influence every type to similar extents, leading to similar ratios across all survivor types. With our present data, it is still unclear whether the absence of X and Y'-elements enhances the formation of circular and atypical survivors. Therefore, we did not present these results in the revised manuscript.
Author response image 1.
Quantitation of each survivor type in SY12 subtelomerice engineered strains. The ratio of survivor types in SY12 tlc1Δ, SY12YΔ tlc1Δ, SY12XYΔ tlc1Δ and SY12XYΔ+Y tlc1Δ strains. Type
I, pulper; Type II, green; Type X, gray; atypical survivor, orange; circular survivor, blue.
3)The authors asked whether X and Y' elements are required for cell proliferation, stress response, telomere length control and telomere silencing (Fig. 4). Similar studies have been previously carried out by using synthetic chromosomes (see PMID: 28300123). The authors need to discuss this point.
Thanks for your suggestion, we have added the information in the revised version.
(p.24 line 449-453)
4) The Fig. 7 data support that circular chromosomes do not require Ku-dependent
DNA end protection. This is consistent with the current view that Ku binds and protects DNA ends. This finding by itself does not contribute significantly to our understanding of telomere maintenance. The authors need to more extensively discuss the significance of their findings in SY12 cells compared to wild-type cells with 16 chromosomes.
We agree with the logic that this reviewer has pointed out. Our results demonstrate that combinatorial deletion of YKU70 and TLC1 caused synthetic lethality in SY12 cells, which possess three linear chromosomes, However, it did not affect the viability of "circular survivors", supporting the notion that telomere deprotection leads to the synthetic lethality in yku70Δ tlc1Δ double mutants. Nevertheless, this conclusion merely confirms the current view observed in wild-type cells that Ku binds and protects DNA ends.
To avoid confusing readers and maintain the logical flow of the manuscript, we have deleted this section in the revised version.
Minor issues:
1) Line 112-113: " for SY13, which contains two chromosomes, could also have a high probability of circularizing all chromosomes for survival": The reference or the supplemental data are required.
Thank this reviewer for the suggestion. According to the reviewer’s comments, we performed a Southern blotting assay to examine the types of survivors in SY13 tlc1Δ strain. We found that the majority of SY13 tlc1Δ clones exhibited hybridization signal similar to SY14 tlc1Δ circular survivors, pointing to the possibility that two chromosomes in these survivors may undergo intra-chromosomal fusions. This result has been added to figure 1D in the revised version.
2) Line 349-350: The BY4742 mre11Δ haploid strain serves as a negative control. The authors need to explain why mre11 cells serve as a negative control.
Thank this reviewer for the comment. We employed mre11Δ as negative control because Mre11 is a member of the RAD52 epistasis group, which is involved in the repair of double-stranded breaks in DNA, and mutants in MRE11 exhibit defects in the repair of DNA damages caused by DNA damage drugs (Krogh and Symington, 2004; Lewis et al., 2004;
Symington, 2002). (p.23 line 420-422)
Reviewer #2
1) The qualification of survivor types mostly relies on molecular patterns in Southern blots. While this is a valid method for a standard strain, it might be more difficult to apply to the strains used in this study. For example, in SY8, SY11 and SY12, the telomere signal at 1-1.2 kb can be very faint due to the small number of terminal Y' elements left. As another example, for the Y'-less strain, it might seem obvious that no Type I survivor can emerge given that Y' amplification is a signature of Type I, but maybe Type-I-specific molecular mechanisms might still be used. To reinforce the characterization of survivor types, an analysis of the genetic requirements for Type I and Type II survivors (e.g. RAD51, RAD54, RAD59, RAD50) could complement the molecular characterization in specific result sections.
We thank this reviewer for his/her constructive comments and suggestions. To investigate whether Type-I-specific molecular mechanisms are still utilized in the survivor formation in Y'-less strain, we deleted RAD51 in SY12XYΔ tlc1Δ. SY12XYΔ tlc1Δ rad51Δ strain was able to generate three types of survivors, including Type II survivor, circular survivor and atypical survivor, similar to the observations in SY12XYΔ tlc1Δ strain. However, the ratios of circular and atypical survivors were 36% and 32%, respectively, lower than the 48% and 44% observed in SY12XYΔ tlc1Δ strain (supplementary file 5). This result indicates that Type-I-specific molecular mechanisms contribute to the survivor formation. Given that our work primarily focuses on the function of subtelomeric elements, we chose not to include this result in our revised manuscript to maintain a coherent logical flow.
To reinforce the characterization of survivor types, we deleted RAD50, RAD51 and RAD52 in SY12 tlc1Δ strain, respectively. Southern blotting assay revealed that in the absence of Rad51, no Type I survivor was detected; in the absence of Rad50, neither Type I nor Type X survivor was detected. However, circular and atypical survivors still emerged in the absence of Rad52, suggesting that the RAD52-mediated homologous recombination is not strictly necessary for the formation of circular and atypical survivors. These results have been presented as Figure 2—figure supplement 6 and Figure 2— figure supplement 7.
2) In the title, the abstract and throughout the discussion, the authors chose to focus on the effect of X- and Y'-element deletion on different phenotypes and on survivor formation, as the main message to convey. While it is a legitimate and interesting message, other important results of this work might benefit from more spotlight. Namely, the observation that strains with different chromosome numbers show different survivor patterns and that several survival strategies beyond Type I and II exist and can reach substantial frequencies depending on the chromosomal context.
Thanks for your valuable suggestion. While we value your suggestion to highlight additional aspects of our work, we would like to express our perspective on the current emphasis on the effect of X- and Y'-element deletion. We believe that by maintaining this focus, we can present a more coherent and impactful narrative for our readers. Additionally, we recognize that the relationship between chromosome numbers and survivor type frequencies is complex and warrants further experimental validation. We are considering exploring this aspect in more detail in our future projects. However, we fully acknowledge the importance of the observations you raised concerning strains with different chromosome numbers and the diversity of survival strategies.
3) In SY12 strain, while X- and Y'-elements are not essential for survivor emergence, they do modulate the frequency of each type of survivors, with more chromosome circularization events observed for SY12YΔ, SY12XYΔ and SY12XYΔ+Y strains. This result should be stated and discussed, maybe alongside the change in survivor patterns in the other SY strains, to more accurately assess the roles of these subtelomeric elements.
Following the reviewer’s suggestion, we compared the circular survivor ratios in SY12 tlc1Δ, SY12YΔ tlc1Δ, SY12XYΔ tlc1Δ and SY12XYΔ+Y tlc1Δ strains (supplementary file 5). It appears that the formation of circular survivors is less efficient in the SY12 tlc1Δ, with a ratio of 20%, much lower than that in SY12YΔ tlc1Δ, SY12XYΔ tlc1Δ or SY12XYΔ+Y tlc1Δ strains. However, it should be noted that SY12 tlc1Δ can generate Type I and Type X survivors, potentially decreasing the ratio of circular survivors.
Therefore, we further compared the circular survivor ratios in SY12YΔ tlc1Δ, SY12XYΔ tlc1Δ and SY12XYΔ+Y tlc1Δ strains. In the SY12YΔ tlc1Δ strain, circular survivors accounted for 52% (26/50), comparable to 48% (24/50) in the SY12XYΔ tlc1Δ strain, indicating that X- elements exert little effect on the formation of circular survivor. Additionally, the ratio of circular survivors was 44% (22/50) in SY12XYΔ+Y tlc1Δ strain, also comparable to 48% (24/50) in the SY12XYΔ tlc1Δ strain, suggesting that Y’-element also has little effect on chromosome circularization. However, due to the unknown frequency of survivor formation, alternative explanations of these data are possible. For example, subtelomeric elements previously suggested to have no impact on the formation of any survivor types might influence every type to similar extents, resulting in similar ratios across all survivor types. With our current data, it is still uncertain whether X and Y'-elements modulate the frequency of each type of survivors. Therefore, we did not include these results in the revised manuscript.
4) The authors might want to update some general information about subtelomere structure and their diversity across yeast strain with the recent paper by O'Donnell et al. 2023 Nature Genetics, "Telomere-to-telomere assemblies of 142 strains characterize the genome structural landscape in Saccharomyces cerevisiae".
Thanks for your advice. We have added this information in the revised manuscript.
(p.3 line 51-54)
5) Although it is cited in the discussion, the recent work by the Malkova lab (Kockler et al. 2021 Mol Cell) could be mentioned in the introduction as it conceptually changes our views on survivor formation, its dynamics and the categorization into Type I and Type II.
Thanks for your advice. We have added this information in the revised manuscript.
(p.5 line 75-78)
6) p.7 line 128-130: rather than chromosome number, the ratio of survivor types might be controlled by the fraction of subtelomeres with Y'-elements and their relative configuration across chromosomes. A map of the structure of remaining subtelomeres in the SYn strains might be good to have.
We have added this information in supplementary file 2 in the revised manuscript.
7) Fig. 1C: in SY9 tlc1Δ, the lane with triangle mark looks like a type II.
The hybridization pattern of SY9 tlc1Δ clone 2 has both amplified Y’L-element and long heterogeneous TG1-3 repeats, it might be the “hybrid” survivor mentioned by Kockler’s work (Kockler et al., 2021). Therefore, we classify it as a no-classical survivor.
8) p.9 line 149: the title of this result section "Y'-element is not essential for the viability of cells carrying linear chromosomes" doesn't reflect well the content of the section, which is more about characterizing the survivor pattern in SY12.
Thanks for your advice. We have changed the title of this section into
“Characterizing the survivor pattern in SY12” in the revised manuscript. (p.9 line 155)
9) p.10 line 167: that type I can emerge in SY12 indicates that multiple Y'-elements in tandem are not required for type I recombination. I am not sure if this was already known, but it could be noted.
We appreciate the reviewer’s comment. We have added this information in the revised manuscript. (p.10 175-177)
10) p.18 line 318-320: the deletion of the Y' element also seems to remove the centromere-proximal telomere sequence adjacent to it. Maybe it should be stated as well. Even more importantly, in lines 327-329, the Y'-element that is reintroduced in the strain does not include the centromere-proximal short telomere sequence. This is important to interpret the Southern blots.
We thank the reviewer for this critical suggestion. The deletion of Y'-element including both Y’- and X- element sequence in XVI-L (supplementary file 4), and the Y’element in the XVI-L does not contain the centromere-proximal telomere sequence. The Y'-element reintroduced into the left arm of Chr 3 in SY12XYΔ strain was cloned from native left arm of XVI in SY12 strain which does not contain the centromere-proximal short telomere sequence. Besides listing these details in supplementary file 4, we also emphasize it in the revised manuscript (p.21 line 397-398).
11) p.29 lines 496-497: it seems that X and Y'-elements tend to inhibit formation of circular survivors either directly (by participating in end protection), or by promoting type I and type II, thus reducing the fraction of circular survivors. Maybe this could be added to the conclusion of this section.
We thank the reviewer for his/her comments and have analyzed survivor types in SY12 tlc1Δ, SY12YΔ tlc1Δ, SY12XYΔ tlc1Δ and SY12XYΔ+Y tlc1Δ strains (supplementary file 5). Circular survivor formation appears less efficient in the SY12 tlc1Δ, with a ratio of 20%, significantly lower than SY12YΔ tlc1Δ, SY12XYΔ tlc1Δ or SY12XYΔ+Y tlc1Δ strains. However, it is noteworthy that SY12 tlc1Δ can generate Type I and Type X survivors, potentially impacting the circular survivor ratio.
We further compared circular survivor ratios in SY12YΔ tlc1Δ, SY12XYΔ tlc1Δ and SY12XYΔ+Y tlc1Δ strains. SY12YΔ tlc1Δ had 52% circular survivors, similar to SY12XYΔ tlc1Δ with 48%, indicating minimal impact of X- elements. Additionally, SY12XYΔ+Y tlc1Δ had 44% circular survivors, also similar to SY12XYΔ tlc1Δ, suggesting that Y’-element has little effect on chromosome circularization. However, due to unknown frequency of survivor formation, alternative explanations, like subtelomeric elements affecting all the type of survivor similarly, are possible. With our current data, it remains unclear whether X and Y'-elements are involved in end protection and consequently inhibit the formation of circular survivors.
Therefore, these results were not included in the revised manuscript.
12) p.32 line 533: this result section doesn't really fit with the rest of the paper, does it?
Thanks for your valuable advice. To avoid confusing readers and to keep the fluency of logic flow of the manuscript we have deleted this section in the revised version.
13) The methods section does not describe the experiments sufficiently and it often lacks specific details such as the manufacturer or references. Some sections of the methods are more exhaustive than others. They should all be written with the same level of detail in my opinion.
Thanks for your advice. We have described the experiments more sufficiently and added the manufacturer or references in the ‘materials and methods’ part in the revised manuscript. (p.41 line741-745, p.42 line 755-756, p.42 line 762-770, p.43 line 788 and p.45 line 812-813)
Minor comments, typos and grammatical errors:
p.3 line 33: "INTROUDUCTION" should be "INTRODUCTION".
We have corrected it in the revised manuscript. (p.3 line 33) p.4 line 54: "S, cerevisiae", use dot instead of comma. R15: We have corrected it in the revised manuscript. (p.4 line 57)
p.4 line 55: I believe TLC1 as the RNA moiety should be in (non-italicized) capital
letters and not written as a protein.
We have corrected it in the revised manuscript. (p.4 line 58)
p.7 line 115: please indicate that pRS316 uses URA3 as a marker, otherwise the counterselection with 5'-FOA is not obvious.
Thank this reviewer for the comment. We have added this statement in the revised manuscript. (p.7 line 121-122)
p.12 line 206: tlc1Δ should be in italic.
We have corrected it in the revised manuscript. (p.10 line 184)
p.13 lines 227-229: "where only one hybridization signal", a verb seems to be missing.
We thank the reviewer’s kind reminder and have corrected the mentioned errors in the revised manuscript. (p.14 line 254-255)
Reviewer #3
1) A weakness of the manuscript is the analysis of telomere transcriptional silencing. They state: "The results demonstrated a significant increase in the expression of the MPH3 and HSP32 upon Sir2 deletion, indicating that telomere silencing remains effective in the absence of X and Y'-elements". However, there are no statistical analyses performed as far as I can see. For some of the strains, the significance of the increased expression in sir2 (especially for MPH3) looks questionable. In addition, a striking observation is that the SY12 strain (with only three chromosomes) express much less of both MPH3 and HSP32 than the parental strain BY4742 (16 chromosomes), both in the presence and absence of Sir2. In fact, the expression of both MPH3 and HSP32 in the SY12 sir2 strain is lower than in the BY4742 SIR2+ strain. In addition, relating this work to previous studies of subtelomeric sequences in other organisms would make the discussion more interesting.
First, I enjoyed reading your manuscript. It would be great if you performed the statistical analysis on the RT-qPCR data in figure 4B and addressed the issue of the difference of the BY4742 and SY12 strains. A model could be that this is a titration effect of silencing proteins due to fewer telomeres, which could be investigated by performing the analyses on more SY-strains with variable numbers of telomeres.
We highly appreciate the reviewer’s valuable comments and suggestions, which included a point that has also left us confused. We conducted statistical analyses on the
RT-qPCR data, and the t-test result revealed that upon the deletion of Sir2, SY12YΔ, SY12XYΔ and SY12XYΔ+Y strains exhibited a significant increase in MPH3 expression (located on the right arm of chr X) with a P value < 0.05. In the case of SY12, the deletion of Sir2 resulted in an increase in gene expression (P value < 0.1). Similar tendencies were observed in the BY4742 strain. The statistical analyses of RTqPCR results on XVI-L mirrored those of X-R.
The results demonstrated a significant increase in MPH3 and HSP32 expression upon SIR2 deletion in SY12YΔ, SY12XYΔ and SY12XYΔ+Y strains, leading to the conclusion that telomere silencing remains effective in the absence of X-and Y’-elements. However, as the reviewer has pointed out, no statistically significant differences in MPH3 and HSP32 expression were observed between the SY12 and SY12 sir2Δ strain. For HSP32, this lack of significance may be attributed to the greater distance between HSP32 and telomere XVI-L in SY12 compared to SY12YΔ, SY12XYΔ or SY12XYΔ+Y strains, resulting in a weaker telomere position effect on HSP32 and a non-significant increase in gene expression in SY12. However, this explanation does not apply to MPH3, as SY12YΔ, with a same distance between MPH3 and telomere X-R as in SY12, still exhibits an effective telomere position effect on MPH3. We cannot provide a compelling explanation at this moment, and we suspect that the lack of statistically significant differences may be due to random clonal variation.
Additionally, the SY12 strain (with three chromosomes) exhibited lower expression levels of both MPH3 and HSP32 compared to the parental strain BY4742 (with 16 chromosomes). Notably, it has been reported that the expression of genes coding silencing proteins in SY14 (with one chromosomes) were nearly identical to that of BY4742 (with 16 chromosomes)(Shao et al., 2018). Consequently, with respect to the reduced chromosome numbers, the silencing proteins appeared to be relatively overexpressed. Therefore, as pointed out by the reviewer, this observed phenomenon may be attributed to a titration effect of silencing proteins due to fewer telomeres. We have added the statistical analyses result in Figure 4B.
We have related our work with previous studies of subtelomeric sequences in fission yeast in the discussion part. (p.37 line 655-676)
Minor points are to correct the figure legend for Figure 6 supplement 1 (the strain designations) and line 55, RNAs are written with all caps, i.e. TLC1, and line 537 delete the "which" in the sentence.
Thanks for your advice. We have corrected them in the revised manuscript.
1) The strain has been replaced with SY12XYΔ+Y (p.35 line 617, 618 and 620)
2) “Tlc1” has been replaced with “TLC1” (p.4 line 58).
3) We have deleted the section of “Circular chromosome maintain stable when double knockout of yku70 and tlc1” according to the suggestions raised by reviewer 1 and 2, the deleted section contain the sentence in line 537 you mentioned.
Kockler, Z.W., Comeron, J.M., and Malkova, A. (2021). A unified alternative telomerelengthening pathway in yeast survivor cells. Molecular Cell 81, 1816-1829.e1815.
Krogh, B.O., and Symington, L.S. (2004). Recombination proteins in yeast. Annu Rev Genet 38, 233-271.
Lewis, L.K., Storici, F., Van Komen, S., Calero, S., Sung, P., and Resnick, M.A. (2004). Role of the nuclease activity of Saccharomyces cerevisiae Mre11 in repair of DNA double-strand breaks in mitotic cells. Genetics 166, 1701-1713.
Shao, Y., Lu, N., Wu, Z., Cai, C., Wang, S., Zhang, L.L., Zhou, F., Xiao, S., Liu, L., Zeng, X., et al. (2018). Creating a functional single-chromosome yeast. Nature 560, 331-335.
Symington, L.S. (2002). Role of RAD52 epistasis group genes in homologous recombination and double-strand break repair. Microbiol Mol Biol Rev 66, 630-670, table of contents.




