Peer review process
Revised: This Reviewed Preprint has been revised by the authors in response to the previous round of peer review; the eLife assessment and the public reviews have been updated where necessary by the editors and peer reviewers.
Read more about eLife’s peer review process.Editors
- Reviewing EditorLibor MacurekInstitute of Molecular Genetics, Academy of Sciences of the Czech Republic, Prague, Czechia
- Senior EditorRichard WhiteUniversity of Oxford, Oxford, United Kingdom
Reviewer #1 (Public Review):
Summary:
Gain-of-function mutations and amplifications of PPM1D are fond across several human cancers and are associated with advanced tumor stage, worse prognosis, and increased lymph node metastasis. This manuscript presents important findings that SOD1 inhibition is a potential strategy to achieve therapeutic synergism for PPM1D-mutant leukemia; and demonstrates the redox landscape of PPM1D-mutant cells.
Strengths:
In this manuscript, Zhang and colleagues investigate the synthetic-lethal dependencies of PPM1D (protein phosphatase, Mg2+/Mn2+ dependent 1D) in leukemia cells using CRISPR/Cas9 screening. They identified that SOD1 (superoxide dismutase-1) as the top hit, whose loss reduces cellular growth in PPM1D-mutant cells, but not wildtype (WT) cells. Consistently, the authors demonstrate that PPM1D-mutant cells are more sensitive to SOD1 inhibitor treatment. By performing different in vitro studies, they show that PPM1D-mutant leukemia cells have elevated level of reactive oxygen species (ROS), decreased basal respiration, increased genomic instability, and impaired non-homologous end-joining repair. These data highlight the potential of SOD1 inhibition as a strategy to achieve therapeutic synergism for PPM1D-mutant leukemia; and demonstrates the redox landscape of PPM1D-mutant cells.
Weaknesses:
While the current study has identified synthetic lethality of PPM1D-mutant leukemia cells upon SOD1 inhibition, the underlying mechanism remains elusive. Although ROS levels have been assessed between wild-type (WT) and PPM1D-mutant leukemia cells, the specific redox alterations induced by SOD1 inhibition in PPM1D mutant versus WT cells have not been elucidated. To address this gap, direct comparisons of ROS levels using various probes should be conducted between PPM1D mutant and WT cells under conditions of SOD1 inhibition.
Reviewer #2 (Public Review):
The authors used a whole genome CRISPR screen to identify targetable synthetic lethalities associated with PPM1D mutations, known poor prognosis and currently undruggable factors in leukemia. The authors identified the cytosolic superoxide dismutase (SOD1, Cu/Zn SOD) as a major protective factor in PPMD1 mutant vs. wt cells, and their study investigates associated mechanisms of this protection. Using both genetic depletion and small molecule inhibitors of SOD1, the authors conclude that SOD1 loss exacerbates mitochondrial dysfunction, ROS levels and DNA damage phenotypes in PPM1D mutant cells, decreasing cell growth in AML cells. The data strongly support that PPMD1 mutant cells have high levels of total peroxides and elevated DNA breaks, and that genetic depletion of SOD1 decreases cell growth in two AML cell lines. However, the authors don't explain how superoxide radical (which is not damaging by itself) induces such damage, the on-target effects of the SOD1 inhibitors at the concentrations is not clear, the increase in total hydroperoxides is not supported by loss of SOD1, the changes in mitochondrial function are small, and there is no assessment of how the mitochondrial SOD2 expression or function, which dismutates mitochondrial superoxide, is altered. Overall these studies do not distinguish between signal vs. damaging aspects of ROS in their models and do not rule out an alternate hypothesis that loss of SOD1 increases superoxide production by cytosolic NADPH activity which would significantly alter ROS-driven regulation of kinase/phosphatase signal modulation, affecting cell growth and proliferation as well as DNA repair. Additionally, with the exception of growth defects demonstrated with sgSOD1, the majority of data are acquired using two chemical inhibitors, LCS1 and ATN-224, without supporting evidence that these inhibitors are acting in an on-target manner.
Overall, the authors address an important problem by seeking targetable vulnerabilities in PPM1D mutant AML cells, it is clear SOD1 deletion induces strong growth defects in the AML cell lines tested, most of the approaches are appropriate for the outcomes being evaluated, and the data are technically solid and well-presented. The major weakness lies in which redox pathways and ROS species are evaluated, how the resulting data are interpreted, and gaps in the follow-up experiments. Due to these omissions, as currently presented, the broader impact of these findings are unclear.
These specific concerns are outlined in detail below and I offer some suggestions regarding how to clarify the mechanisms underlying their initial observation of SOD1 synthetic lethality:
(1) Fig. 1 - SOD1 appears to be clustered with several other genes in the volcano plot (including FANC proteins). Did any other ROS-detoxifying enzymes show similar fitness scores? The effects of the SOD1 sgRNA are striking, however it would be useful to see qPCR or immunoblot data confirming robust depletion.
Does SOD1 co-expression in PPM1-mutant patient AML correspond to poorer disease outcomes? This can be evaluated in publicly available patient datasets and would support the idea of SOD1 synthetic lethality.
It would also be useful to know (given the subsequent results) whether expression of the SOD2, the mitochondrial superoxide dismutase, is altered in response to SOD1 loss.
(2) Fig. 2 - What are the relative SOD1 levels in the mutant PPM1D vs. wt. cell lines? The effects of the chemical inhibitors are stronger in MOLM-13 than the other two lines. These data could also point to whether LCS-1 and ATN-224 cytotoxicity is on-target or off-target at these concentrations, which is a key issue not currently addressed in these studies. This is a particular concern as the OCI-AML2 line shows a stronger growth defect with CRISPR SOD1 KO (in Fig 1) but the smallest effects with these chemical inhibitors.
While endogenous mitochondrial superoxide levels are elevated in PPM1D mutant lines, it is entirely unclear why SOD1 inhibition should affect mitochondrial superoxide as it detoxifies cytosolic superoxide. Also unclear why DCFDA signal (which measures total hydroperoxides) is *increased* under SOD1 inhibition - SOD1 dismutates superoxide radicals into hydrogen peroxide, therefore unless SOD2 is compensating for SOD1 loss, one might expect hydroperoxides to be lower (unless some entirely different oxidase is increasing their levels). None of these outcomes appear to be considered. Finally, it is not explained how lipid peroxidation, which requires production of hydroxyl or similarly high potency radicals, is being caused by increased superoxide or peroxides. One possibility is there is an increase in labile iron, in which case this phenotype would be rescued by the iron chelator desferal, and by the lipophilic antioxidant, ferrostatin.
Do the sgSOD1 cells also show similar increases in MitoSox green, DCFDA and BODIPY signal? These experiments would clarify whether the effects with the inhibitors are directly related directly to SOD1 loss or if they represent off-target effects from the inhibitors and/or compensatory changes in SOD2.
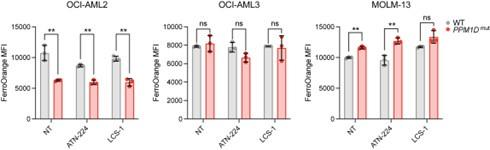
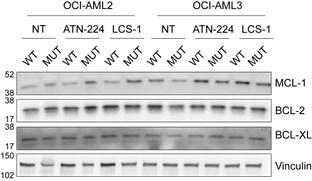
(3) Fig. 3 - the effects on mitochondrial respiratory parameters, while statistically significant, do not seem biologically striking. Also, these data are shown for OCI-AML2 cells which show the smallest cytotoxic effects with the SOD1 inhibitors among the 3 lines tested. They do however show the most robust growth defect with sgSOD1. This discrepancy could suggest that mitochondrial dysfunction does not underlie the observed growth defect and/or the inhibitor cytotoxicity is not on-target. Ideally mitochondrial profiling should also be carried out on this cell line with inducible SOD1 depletion. Have the authors assessed whether the mitochondrial Bcl family proteins are affected by the inhibitors?
(4) Fig. 4 - Currently the data in this figure do not support the authors claim that PPM1D-mutant cells have impaired antioxidant defense mechanisms, leading to an elevation in ROS levels and reliance on SOD1 for protection. It should be noted that oxidative stress specifically refers to adverse cellular effects of increasing ROS, not baseline levels of various redox parameters. Ideally levels of GSSG/GSH would be a better measure of potential redox stress tolerance than the total antioxidant capacity assay. Finally, oxidative stress can be assessed by challenging the wt and mutant PPM1D cell lines with oxidant stressors such as paraquat which elevates superoxide or drugs like erastin which elevate mitochondrial ROS. The immunoblot shows negligible changes in the antioxidant proteins assayed. Again, this blot should include SOD2 which is the most relevant antioxidant in the context of mitochondrial superoxide.
(5) Fig. 5 - These data support that DNA breaks are elevated in PPM1D mutant vs. wt cells. However, the data with the chemical SOD1 inhibitor again do not convince that the enhanced levels are due to on-target effects on SOD1. Use of the alkaline comet assay is appropriate for these studies and the 8-oxoguanine data do indicate contributions from oxidative DNA base damage. But these are unlikely to result directly from altered superoxide levels, as this species cannot directly oxidize DNA bases or cause DNA strand breaks.
The following points summarize my specific experimental and textual recommendations:
(1) These studies require an assessment of on-target efficacy of the inhibitors at the relevant concentration ranges. Ideally, they should have minimal effects against SOD1 knockout cell lines (acute challenge at a time point before the growth defects become apparent) and show better efficacy in SOD1-overexpressing lines. Key experiments (changes in superoxide, OCR profiling, DNA alkaline comet assay) would be more convincing if they are carried out with SOD1 knockout lines to compare against the inhibitor effects (3-4 days after introducing sgSOD1 when growth defects are not apparent).
(2) Instead of using NAC, which elevates glutathione synthesis but also has several known side-effects, the authors may want to determine whether Tempol, a SOD mimetic can rescue the effects of SOD1 knockout or inhibition. This would directly prove that SOD1 functional loss underlies the observed growth defect and cytotoxicity from genetic SOD1 knockdown or chemical inhibition.
(3) The complete lack of consideration of SOD2 in these studies is a missed opportunity as it reduces mitochondrial superoxide levels but elevates hydrogen peroxide levels. It would be very interesting to see whether SOD1 inhibition leads to compensatory increases in SOD2. SOD2 can be easily measured by immunoblot. Furthermore, measuring total superoxide via hydroethidium in a flow cytometric assay vs. mitochondrial ROS in PPM1D mut vs. wt cells and under SOD1 knockout would enable a determination of which species dominates (cytosolic or mitochondrial). These experiments are required to fill some logical gaps in interpretation of their redox data.
(4) Given the DNA breaks observed in PPM1D mutant cells, it is highly recommended the authors assess whether iron levels are elevated in mut vs. wt cells and whether desferal can rescue observed SOD1 inhibition defects.
(5) The authors may want to assess whether Rac1 or NADPH oxidase activity is altered in the SOD1 KO in wt vs. PPM1D cells. Their results may be the consequence of compromised ROS-driven survival signaling or DNA repair rather than direct ROS-induced damage, which is not caused directly by superoxide (or hydrogen peroxide).
(6) It is recommended the discussion focus more strongly on how the signaling function of superoxide vs. its reactions with other molecular entities to induce genotoxic outcomes could be contributing to the observed phenotypes. The discussion of FANC proteins, which were targets with similar fitness scores but not experimentally investigated at all, is an unwarranted digression.
Reviewer #3 (Public Review):
Summary:
Authors performed a genome-wide CRISPR-based screen for synthetic lethal interactions in leukemic cells expressing a mutant form of PPM1D and identified SOD1. Loss of SOD1 or its inhibition with small molecule compounds reduced survival of the cells containing truncated PPM1D. Further analysis revealed that mitochondria are functionally deficient in PPM1D mutant cells resulting in increased levels of ROS. Surprisingly, expression profiling and reverse phase protein arrays revealed that PPM1D mutant cells did not respond appropriately to the increased levels of ROS. The precise molecular mechanism underlying this phenotype remains currently unclear, nevertheless the study convincingly shows that PPM1D mutant cells are vulnerable to oxidative stress.
Strengths:
Experimental procedures used in the study are appropriate and overall the presented data are very convincing. The study identified an important vulnerability of leukemic cells that carry PPM1D mutation and provides a fundamental background for testing SOD1 inhibitors in preclinical research. In the revised version of the manuscript, authors provide several new experiments that support their former conclusions. In particular, they showed that deletion of SOD1 in AML cells improved survival of the transplanted mice and this effect was more prominent when using cells carrying the mutant PPM1D. Further, they included an important control experiment that showed decreased SOD1 activity after treatment with ATN-224 inhibitor.
Weaknesses:
In the opinion of reviewer, there are no obvious weaknesses in this study. In broader view, the findings presented here using in vitro cultures will need to be validated in vivo by future research. Cell lines used in the study were generated by CRSIPR approaches in AML cells that have already been transformed. In addition, genome editing is inheritably connected with a risk of off target effects. It would therefore be great to identify AML samples carrying the PPM1D mutation that has been naturally selected during the transformation process.