Peer review process
Not revised: This Reviewed Preprint includes the authors’ original preprint (without revision), an eLife assessment, public reviews, and a provisional response from the authors.
Read more about eLife’s peer review process.Editors
- Reviewing EditorAnne-Florence BitbolEcole Polytechnique Federale de Lausanne (EPFL), Lausanne, Switzerland
- Senior EditorAleksandra WalczakCNRS, Paris, France
Reviewer #1 (Public Review):
This manuscript presents a pipeline incorporating a deep generative model and peptide property predictors for the de novo design of peptide sequences with dual antimicrobial/antiviral functions. The authors synthesized and experimentally validated three peptides designed by the pipeline, demonstrating antimicrobial and antiviral activities, with one leading peptide exhibiting antimicrobial efficacy in animal models. However, the manuscript as it stands, has several major limitations on the computational side.
Major issues:
(1) The choice of GAN as the generative model. There are multiple deep generative frameworks (e.g., language models, VAEs, and diffusion models), and GANs are known for their training difficulty and mode collapse. Could the authors elaborate on the specific rationale behind choosing GANs for this task?
(2) The pipeline is supposed to generate peptides showing dual properties. Why were antiviral peptides not used to train the GAN? Would adding antiviral peptides into the training lead to a higher chance of getting antiviral generations?
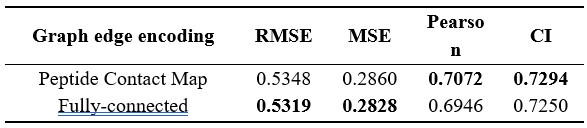
(3) For the antimicrobial peptide predictor, where were the contact maps of peptides sourced from?
(4) Morgan fingerprint can be used to generate amino acid features. Would it be better to concatenate ESM features with amino acid-level fingerprints and use them as node features of GNN?
(5) Although the number of labeled antiviral peptides may be limited, the input features (ESM embeddings) should be predictive enough when coupled with shallow neural networks. Have the authors tried simple GNNs on antiviral prediction and compared the prediction performance to those of existing tools?
(6) Instead of using global alignment to get match scores, the authors should use local alignment.
(7) How novel are the validated peptides? The authors should run a sequence alignment to get the most similar known AMP for each validated peptide, and analyze whether they are similar.
(8) Only three peptides were synthesized and experimentally validated. This is too few and unacceptable in this field currently. The standard is to synthesize and characterize several dozens of peptides at the very least to have a robust study.
Reviewer #2 (Public Review):
Summary:
This study marks a noteworthy advance in the targeted design of AMPs, leveraging a pioneering deep-learning framework to generate potent bifunctional peptides with specificity against both bacteria and viruses. The introduction of a GAN for generation and a GCN-based AMPredictor for MIC predictions is methodologically robust and a major stride in computational biology. Experimental validation in vitro and in animal models, notably with the highly potent P076 against a multidrug-resistant bacterium and P002's broad-spectrum viral inhibition, underpins the strength of their evidence. The findings are significant, showcasing not just promising therapeutic candidates, but also demonstrating a replicable means to rapidly develop new antimicrobials against the threat of drug-resistant pathogens.
Strengths:
The de novo AMP design framework combines a generative adversarial network (GAN) with an AMP predictor (AMPredictor), which is a novel approach in the field. The integration of deep generative models and graph-encoding activity regressors for discovering bifunctional AMPs is cutting-edge and addresses the need for new antimicrobial agents against drug-resistant pathogens. The in vitro and in vivo experimental validations of the AMPs provide strong evidence to support the computational predictions. The successful inhibition of a spectrum of pathogens in vitro and in animal models gives credibility to the claims. The discovery of effective peptides, such as P076, which demonstrates potent bactericidal activity against multidrug-resistant A. baumannii with low cytotoxicity, is noteworthy. This could have far-reaching implications for addressing antibiotic resistance. The demonstrated activity of the peptides against both bacterial and viral pathogens suggests that the discovered AMPs have a wide therapeutic potential and could be effective against a range of pathogens.
Reviewer #3 (Public Review):
Summary:
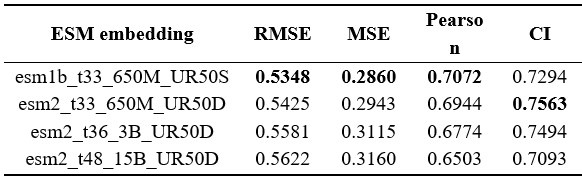
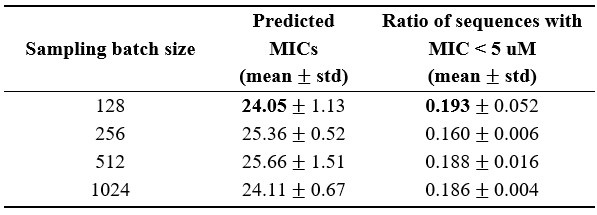
Dong et al. described a deep learning-based framework of antimicrobial (AMP) generator and regressor to design and rank de novo antimicrobial peptides (AMPs). For generated AMPs, they predicted their minimum inhibitory concentration (MIC) using a model that combines the Morgan fingerprint, contact map, and ESM language model. For their selected AMPs based on predicted MIC, they also use a combination of antiviral peptide (AVP) prediction models to select AMPs with potential antiviral activity. They experimentally validated 3 candidates for antimicrobial activity against S. aureus, A. baumannii, E. coli, and P. aeruginosa, and their toxicity on mouse blood and three human cell lines. The authors select their most promising AMP (P076) for in vivo experiments in A. baumannii-infected mice. They finally test the antiviral activity of their 3 AMPs against viruses.
Strengths:
-The development of de novo antimicrobial peptides (AMPs) with the novelty of being bifunctional (antimicrobial and antiviral activity).
-Novel, combined approach to AMP activity prediction from their amino acid sequence.
Weaknesses:
-I missed justification on why training AMPs without information of their antiviral activity would generate AMPs that could also have antiviral activity with such high frequency (32 out of 104).
-The justification for AMP predictor advantages over previous tools lacks rationale, comparison with previous tools (e.g., with the very successful AMP prediction approach described by Ma et al. 10.1038/s41587-022-01226-0), and proper referencing.
-Experimental validation of three de novo AMPs is a very low number compared to recent similar studies.
-I have concerns regarding the in vivo experiments including i) the short period of reported survival compared to recent studies (0.1038/s41587-022-01226-0, 10.1016/j.chom.2023.07.001, 0.1038/s41551-022-00991-2) and ii) although in Figure 2 f and g statistics have been provided, log scale y-axis would provide a better comparative representation of different conditions.
-I had difficulty reading the story because of the use of acronyms without referring to their full name for the first time, and incomplete annotation in figures and captions.