SEC24A deficiency lowers plasma cholesterol through reduced PCSK9 secretion
Figures
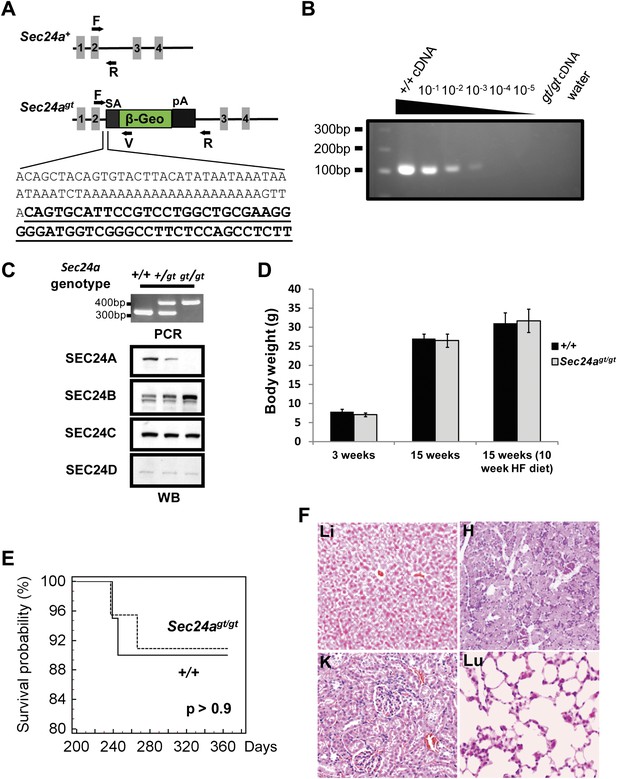
SEC24A null mice are viable and exhibit normal survival and development.
(A) Schematic of the first Sec24a mutant allele (Sec24agt). Gray blocks represent exons with specific numbers indicated. SA, splice acceptor cassette; β-Geo, β-galactosidase-neo fusion; pA, poly-adenylation sequence. F, R, and V represent genotyping primers. Bottom, sequence of Sec24agt gene trap insertion junction; sequence of the gene trap cassette is underlined. (B) RT-PCR detection of splicing between exons 2 and 3 in Sec24agt/gt mice. Liver cDNA of wild type mice was serially diluted into liver cDNA of Sec24agt/gt mice as indicated and used as template for PCR with primers Sec24a-Exon2 and Sec24a-Exon3 (see primer sequences). (C) Loss of SEC24A protein in Sec24agt/gt mice. Upper panel, PCR genotyping; lower panel, immunoblotting of brain protein extracts from mice with the genotypes indicated at the top, using the indicated SEC24A-D antibodies. (D) Body weights of SEC24A-deficient and wild type control mice. HF, high fat diet. Error bars represent SEM (standard error of the mean). At least six mice were included in each group at each time point. (E) Kaplan Meier plot for survival of SEC24A-deficient mice (n = 20) and littermate controls (n = 15). (F) Histology of several tissues from Sec24agt/gt mice. Li, liver; H, heart; K, kidney; Lu, lung.
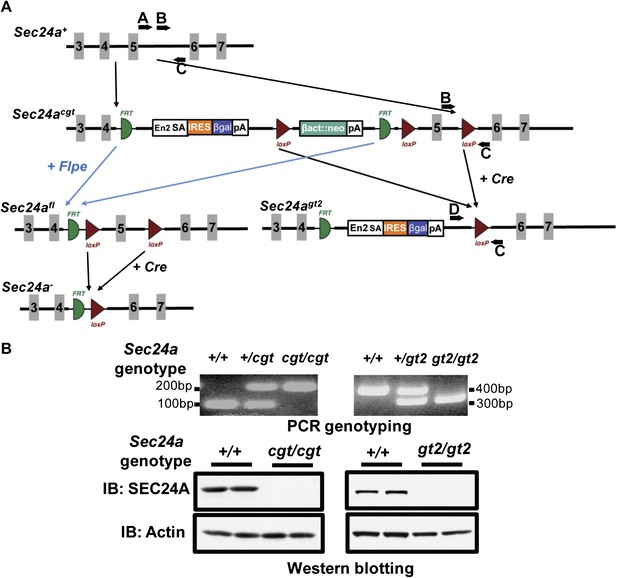
Additional targeted alleles of Sec24a.
(A) Schematic of additional Sec24a alleles, Sec24acgt, Sec24agt2, Sec24afl and Sec24a-(adapted from the Knockout Mouse Project; general conditional gene targeting scheme: https://www.komp.org/alleles.php#conditional-promoter-csd, Sec24a targeting: http://www.knockoutmouse.org/martsearch/project/24915). Gray blocks represent exons. A, B, C, and D, genotyping primers. (B) PCR genotyping and immuno-blot analysis of brain extracts in tissues from Sec24acgt/cgt and Sec24agt2/gt2 mice. IB indicates immunoblotting antibody.
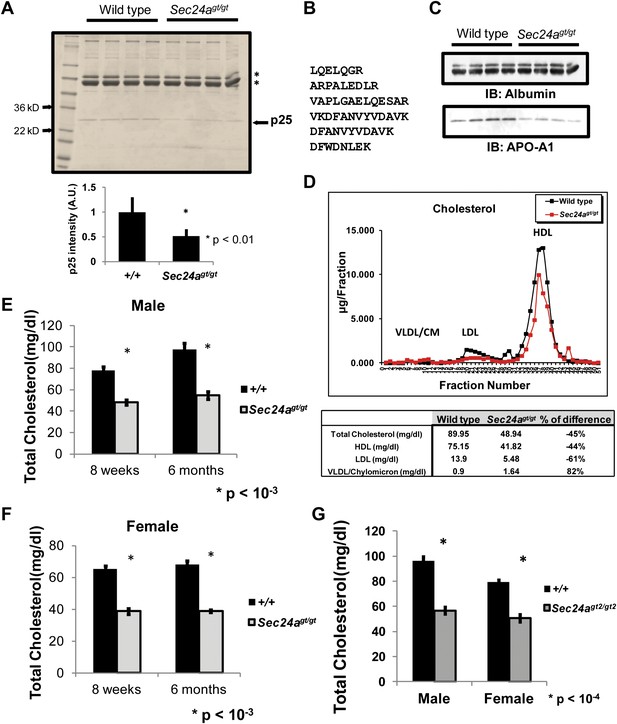
SEC24A-deficient mice develop hypocholesterolemia.
(A) Non-reduced plasma protein samples from four wild type and four Sec24agt/gt mice were separated on SDS-PAGE and stained with coomassie brilliant blue. The first lane contains size markers. Asterisks indicate transferrin and albumin. An ∼25kD protein (p25) is under-represented in the plasma of Sec24agt/gt mice. Lower panel, quantification of the intensity for the band labeled ‘p25' in the upper panel; error bars represent SEM. Asterisk, p<0.01 by Student's t-test. (B) Identification of p25 as APO-A1 by mass spectrometry. Peptide sequences detected in HPLC-ESI-MS/MS analysis; all six peptides exhibit 100% match with mouse APO-A1 sequence. (C) Non-reduced plasma protein samples from four wild type or four SEC24A-deficient mice were analyzed by immunoblotting with antibodies to albumin or APO-A1. (D) Pooled plasma samples from seven wild type and eight Sec24agt/gt mice were fractionated by FPLC and cholesterol in each fraction quantified with a colorimetric assay; total cholesterol for the fractions containing HDL, LDL, and VLDL/Chylomicrons are indicated in the table at the bottom. (E) Total plasma cholesterol in male wild type control (n = 10 for 8 weeks of age, n = 4 for 6 months of age) and Sec24agt/gt (n = 5 for 8 weeks of age, n = 6 for 6 months of age). Error bars represent SEM. Asterisk, p<0.001 by Student's t-test. (F) Total plasma cholesterol in female wild type control (n = 7 for 8 weeks of age, n = 4 for 6 months of age) and Sec24agt/gt (n = 5 for 8 weeks of age, n = 5 for 6 months of age). Error bars represent SEM. Asterisk, p<0.001 by Student's t-test. (G) Total plasma cholesterol in wild type control (n = 7 for male, n = 5 for female) and Sec24agt2/gt2 (n = 4 for male, n = 4 for female). Error bars represent SEM. *p<0.0001 by Student's t-test.
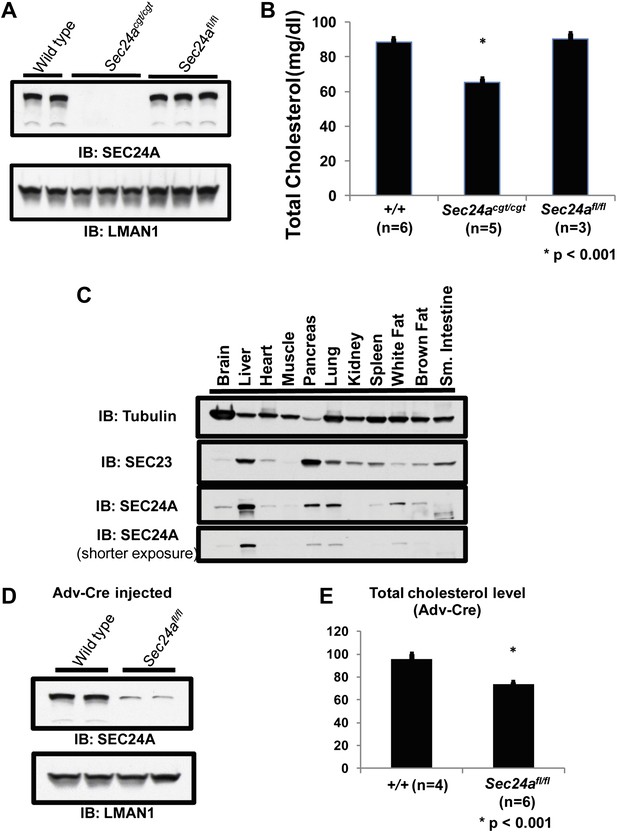
Loss of hepatic SEC24A expression leads to hypocholesterolemia.
(A) Immunoblotting of liver lysates from wild-type and Sec24acgt/cgt mice with an anti-SEC24A and control anti-LMAN1 antibody demonstrates loss of SEC24A expression in Sec24acgt/cgt mice. Removal of the gene trap from the Sec24acgt allele to generate Sec24afl restores wild type SEC24A expression. (B) Total plasma cholesterol levels from wild type, Sec24acgt/cgt , and Sec24afl/fl mice were quantified with a colorimetric assay; error bars represent SEM. Asterisk, p<0.001 by Student's t-test. (C) Protein extracts from the indicated tissues of a wild type mouse were subjected to immunoblotting with the indicated antibodies. (D) Hepatic inactivation of SEC24A was performed by intravenous injection of Sec24afl/fl mice with an adenovirus encoding Cre recombinase (Adv-Cre). Liver protein extracts from these mice and control Adv-Cre-injected wild type mice were subjected to immunoblotting with antibodies to SEC24A or a control (LMAN1). Hepatic SEC24A is reduced by 70–80% in the Adv-Cre-treated Sec24afl/fl mice. (E) Total plasma cholesterol was quantified in Adv-Cre-treated mice; error bars represent SEM. n = 4 for wild type mice; n = 6 for Sec24afl/fl mice. *p<0.001 by Student's t-test.
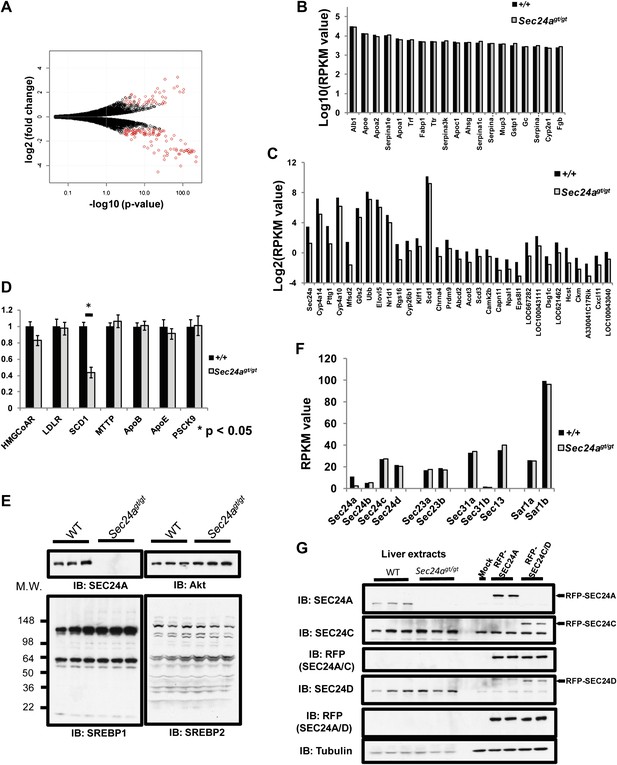
SEC24A deficiency does not alter SREBP signaling.
(A) Volcano plot of liver transcriptome analysis by mRNA-Seq. X-axis, -Log(p-value); y-axis, Log2(fold difference WT/Sec24agt/gt). Significantly altered genes (fold change > 2, RPKM > 0.1, and FDR < 0.05) are colored in red. (B) Twenty most abundant hepatic transcripts detected in wild type and Sec24agt/gt mice by mRNA-Seq. X-axis, gene name; y-axis: Log(RPKM value). (C) Hepatic transcripts significantly down-regulated by SEC24A deficiency. X-axis, gene name; y-axis: Log2(RPKM value). (D) Liver mRNA samples from wild type (n = 4) or SEC24A-deficient mice (n = 4) were subjected to quantitative-PCR (q-PCR) with primers for the indicated SREBP-regulated transcripts. Error bars represent SEM. Asterisk, p<0.01 by Student's t-test. (E) Liver protein extracts from wild type and Sec24agt/gt mice were subjected to immunoblotting with the indicated antibodies. (F) Transcript abundance detected by mRNA-Seq for COPII genes in the liver of wild type and Sec24agt/gt mice. (G) SEC24A and SEC24C/D are expressed at comparable levels in the liver. Liver protein extracts from three wild type and three Sec24agt/gt mice and 293T cells expressing RFP-tagged SEC24A, SEC24C or SEC24D as references were analyzed with the indicated antibodies.
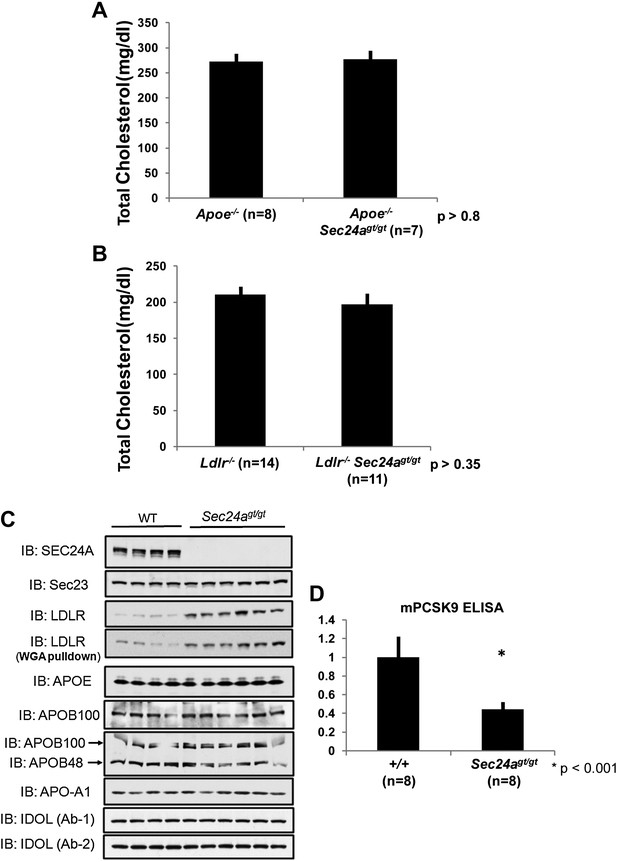
SEC24A deficiency up-regulates LDLR protein levels by decreasing circulating PCSK9.
(A) SEC24A deficiency does not cause hypocholesterolemia in the setting of APOE deficiency. Total plasma cholesterols in Apoe-/- (n = 8) and Apoe-/-Sec24agt/gt (n = 7) mice. Error bars represent SEM. p=∼0.8 by Student's t-test. (B). Total plasma cholesterol levels from Ldlr-/- (n = 14) and Ldlr-/-Sec24agt/gt (n = 11) mice. Error bars represent SEM. p=∼0.35 by Student's t-test. (C) Liver protein extracts from wild type and Sec24agt/gt mice were subjected to immunoblotting with the indicated antibodies. (D) Plasma PCSK9 levels from wild type (n = 8) and Sec24agt/gt (n = 8) mice were quantified by ELISA. Error bars represent SEM. *p<0.001 by Student's t-test.
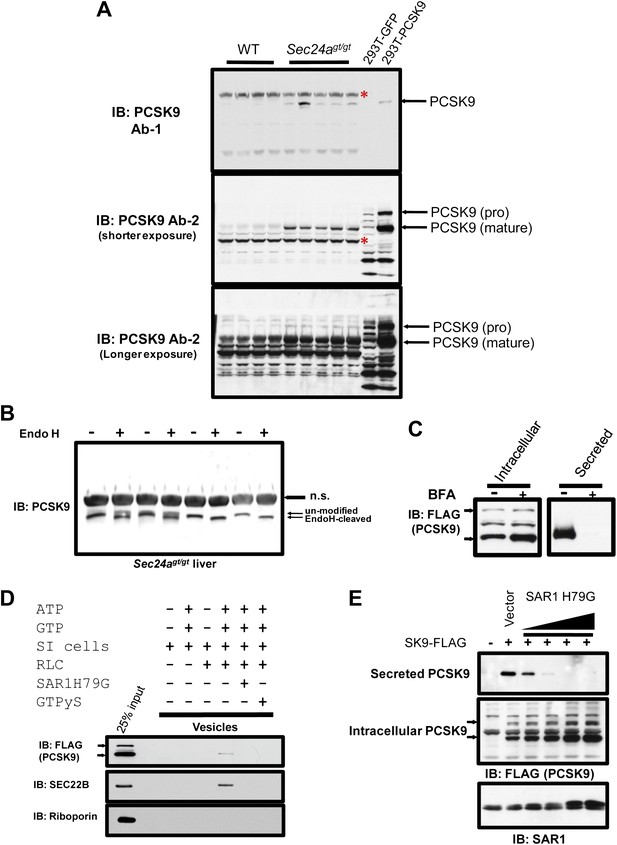
PCSK9 is a soluble COPII cargo.
(A) Liver protein extracts from wild type and Sec24agt/gt mice and cell lysates from 293T cells expressing PCSK9 were subjected to immunoblotting with two different anti-PCSK9 antibodies. Red asterisks indicate non-specific bands. (B) Liver protein extracts from SEC24A-deficient mice were subjected to Endo H treatment before SDS-PAGE and immunoblotting with an anti-PCSK9 antibody. n.s., non-specific band. (C) Cell lysates and conditioned medium from 293T cells stably expressing PCSK9-FLAG were analyzed by immunoblotting with an anti-FLAG antibody following treatment with or without BFA. The arrows indicate un-cleaved (upper) and auto-cleaved (lower) forms of PCSK9. (D) Permeabilized 293T cells stably expressing PCSK9-FLAG were employed in an in vitro COPII budding assay; the resulting vesicle fractions and permeabilized cell inputs were separated by SDS-PAGE and visualized by immunoblotting. The arrows indicate un-cleaved (upper) and auto-cleaved (lower) forms of PCSK9. (E) Cell lysates and conditioned medium from 293T cells stably expressing PCSK9-FLAG transfected with a vector control or a plasmid expressing a dominant-negative mutant SAR1 (H79G) and analyzed by immunoblotting with an anti-FLAG antibody. The arrows indicate un-cleaved (upper) and auto-cleaved (lower) forms of PCSK9.
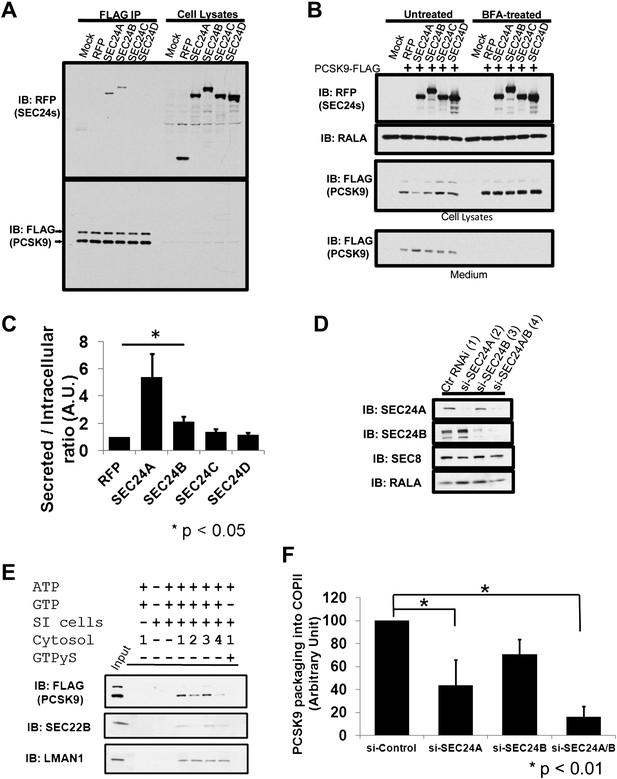
SEC24A regulates PCSK9 secretion.
(A) Cell lysates from 293T cells stably expressing PCSK9-FLAG transfected with plasmids expressing RFP-tagged SEC24A-D or a control RFP vector together with GFP-tagged SEC23A, subjected to immune-precipitation; and immune-complexes and cell lysates then examined by immunoblotting with anti-RFP or anti-FLAG antibodies. The arrows indicate un-cleaved (upper) and auto-cleaved (lower) forms of PCSK9. (B) Cell lysates and conditioned medium from 293T cells co-transfected with PCSK9-FLAG and SEC23/24 plasmids as in (A) were subjected to immunoblotting with the indicated antibodies. Cells treated with BFA were employed as controls. (C) Ratio of secreted PCSK9/intracellular PCSK9 for each transfected RFP-tagged SEC24 and control. Quantification was performed from five independent experiments. Error bars represent SEM. Asterisk, p<0.05 by Student's t-test. (D) Deficient McA-RH777 cells treated with the indicated siRNAs were subjected to immunoblotting to determine SEC24A and SEC24B levels. Numbers indicate the cells receiving different siRNAs. (E) Deficient McA-RH777 cells treated with the indicated siRNAs used as the source of cytosol for in vitro COPII budding assay as in (7d). (F) Quantification of PCSK9 packaging into COPII vesicles from four different experiments. Error bars represent standard deviation. * p<0.01 by Student's t-test.
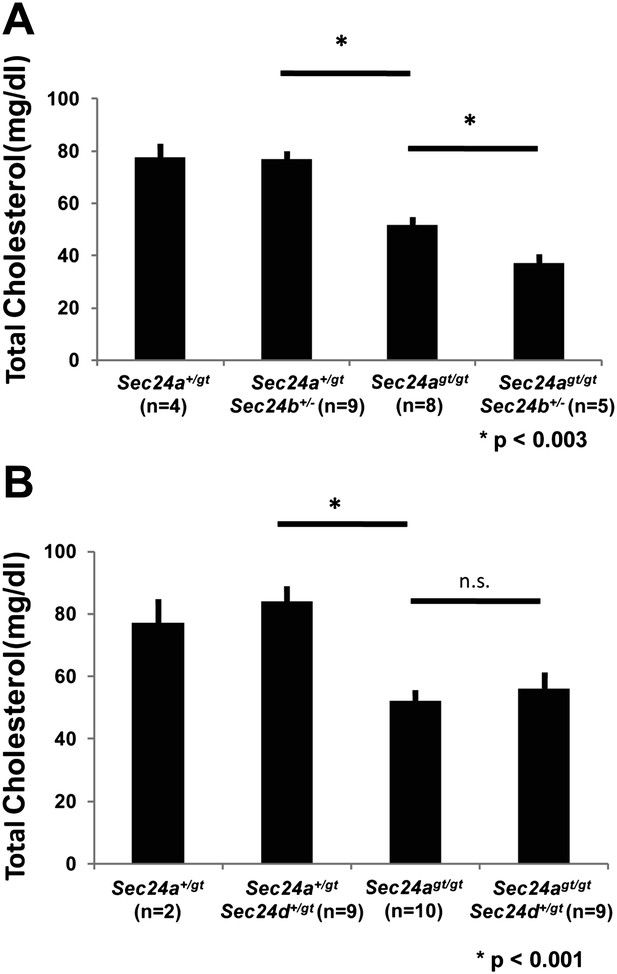
SEC24B but not SEC24D exhibit partial overlap in function with SEC24A in vivo.
(A) Total plasma cholesterol levels from mice generated from a Sec24a+/gtSec24b+/- X Sec24agt/gt cross. Error bars represent SEM. Asterisk, p<0.003 by Student's t-test. (B) Total plasma cholesterol levels from mice generated from a Sec24a+/gtSec24d+/gt X Sec24agt/gt cross. Error bars represent SEM. * p<0.003; n.s., p=∼0.6, by Student's t-test.
Tables
Distributions of offspring from intercross
| Crosses | Genotype distribution in % | p value (χ2) | |||
| +/+ | +/− | −/− | |||
| Expected % | 25% | 50% | 25% | ||
| Sec24a+/gt X Sec24a+/gt | 25% (36) | 48.6% (70) | 26.4% (38) | > 0.9 | |
| Sec24a+/cgt X Sec24a+/cgt | 23.3% (7) | 43.3% (13) | 33.3% (10) | > 0.5 | |
| Sec24a+/gt2 X Sec24a+/gt2 | 27.9% (17) | 49.2% (30) | 23% (14) | > 0.8 | |
| Sec24a+/gtSec24b+/- X Sec24agt/gt | Sec24a+/gt 25% | Sec24a+/gtSec24b+/- 25% | Sec24agt/gt 25% | Sec24agt/gtSec24b+/- 25% | |
| Observed | 20.7% (19) | 32.6% (30) | 25% (23) | 21.7% (20) | > 0.35 |
| Sec24a+/gtSec24d+/gt X Sec24agt/gt | Sec24a+/gt 25% | Sec24a+/gtSec24d+/gt 25% | Sec24agt/gt 25% | Sec24agt/gtSec24d+/gt 25% | |
| Observed | 22.1% (21) | 29.5% (28) | 27.4% (26) | 20.1% (20) | > 0.6 |
-
Observed numbers are listed in parentheses.
Complete blood count survey
| WT (n = 10) | Sec24agt/gt (n = 6) | p value | |
| WBC (X103) | 11.77 ± 2.67 | 11.53 ± 1.60 | 0.85 |
| RBC (X106) | 9.89 ± 0.41 | 9.28 ± 0.46 | 0.02 * |
| HGB (g/dl) | 12.9 ± 0.7 | 12.7 ± 0.5 | 0.51 |
| HCT (%) | 50.2 ± 1.5 | 49.2 ± 2.7 | 0.34 |
| MCV (fl) | 50.78 ± 0.97 | 52.72 ± 0.66 | 0.001 * |
| MCH (pg) | 13.06 ±0.35 | 13.48 ± 0.23 | 0.02 * |
| MCHC (%) | 25.75 ± 0.55 | 25.60 ± 0.28 | 0.55 |
| CHCM (g/dl) | 26.39 ± 0.21 | 26.28 ± 0.17 | 0.31 |
| RDW (%) | 12.92 ± 0.61 | 12.62 ± 0.70 | 0.38 |
| HDW (%) | 1.692 ± 0.086 | 1.630 ± 0.071 | 0.16 |
| PLT (X104) | 123.6 ± 19.1 | 118.8 ± 8.5 | 0.58 |
| MPV (fl) | 5.2 ± 1.5 | 5.73 ± 0.2 | 0.46 |
-
*
p < 0.05 by Student's t-test.
Additional files
-
Supplementary file 1
RNA-Seq ENRAGE results. (A) Genes. (B) Exons.
- https://doi.org/10.7554/eLife.00444.014
-
Supplementary file 2
Gene-ontology analysis of potentially biological pathways in Sec24agt/gt mice. (A) Down-regulated. (B) Up-regulated.
- https://doi.org/10.7554/eLife.00444.015





