Genome rearrangements and pervasive meiotic drive cause hybrid infertility in fission yeast
Figures
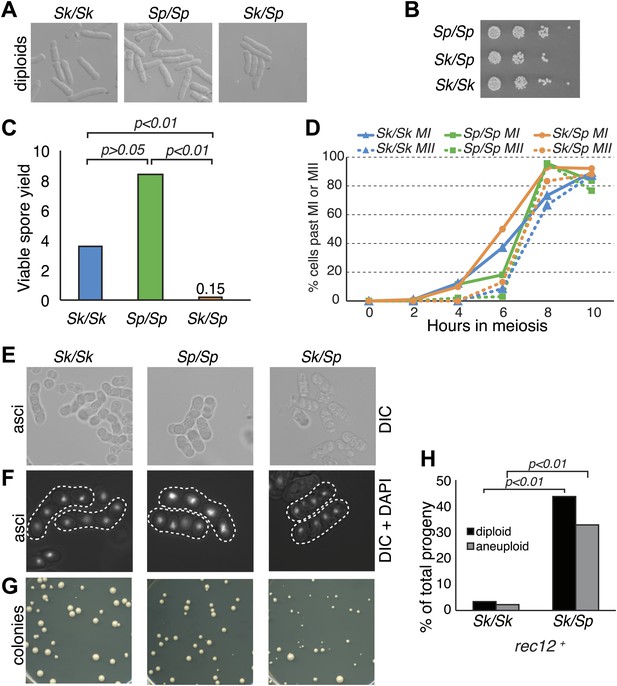
Sk/Sp hybrids are healthy but exhibit low fertility.
(A) Sk/Sp hybrid diploids are morphologically similar to pure species diploids. (B) Sk/Sp hybrid diploids show no gross growth defects relative to pure species controls. (C) Viable spore yield tests show that Sk/Sp fertility is low relative to pure species controls (averages of n ≥ 5 experiments, p-values obtained using t test). This assay does not directly measure viable spores per meiosis, so values can exceed 4. (D) Sk/Sp hybrids complete both meiotic divisions with timing similar to that of pure species controls (representative experiment of 3, n ≥200 cells for each data point). (E and F) The asci produced by Sk/Sp hybrids contain spores that are more irregular and transparent than pure species asci. (G) The viable spores produced by Sk/Sp hybrids often grow into small irregularly sized and shaped colonies. (H) The majority of the viable spores produced by Sk/Sp hybrids are aneuploid or diploid (p-values obtained using G-test, n >200 for each). These data are also shown in Figure 5A.
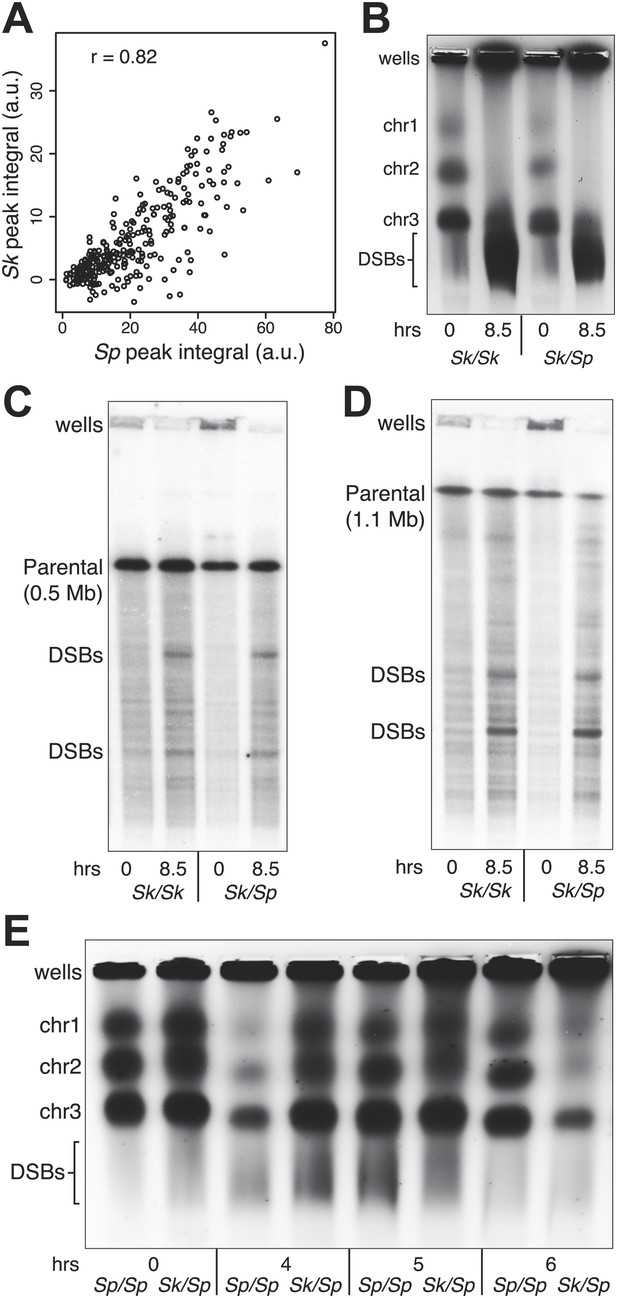
DSB hotspot divergence and repair in Sk/Sp hybrids.
(A) We used ChIP–chip of Rec12-FLAG from rad50S Sk meiotic cultures to assay DSB hotspots and compared the profile to the published DSB hotspot maps of Sp (Fowler et al., 2013). We then compared the Rec12-enrichment in Sk at 286 defined Sp hotspots and found a strong correlation between Rec12 enrichments between the two species at these sites. (B) Sk/Sp cells are proficient at inducing DSBs. Ethidium bromide stained pulsed-field gel of diploids at 0 and 8.5 hr after inducing meiosis in liquid cultures. These diploids are rad50S mutants, so DSBs form normally but are not repaired. As DSBs are formed, the three full-sized chromosome bands disappear and the DNA runs as smaller broken fragments on the gel. (C and D) We find that DSBs are formed at similar locations and similar frequencies in Sk/Sp and Sk/Sk. Southern blots of pulsed-field gels to obtain a closer view of DSB formation in rad50S diploids probed to visualize two NotI restriction fragments known as NotI J [shown in (C)] and NotI D [shown in (D)]. Prior to DSB formation, most of the DNA runs as a single large band. After all break formation (8.5 hr) smaller cut fragments become apparent at the same sites in Sk/Sp and Sk/Sk. (E) DSBs are efficiently repaired in Sk/Sp. Ethidium bromide stained pulsed-field gel of rad50+ diploids at the given times after the induction of meiosis show that DSBs do not accumulate more in Sk/Sp than the Sp/Sp control during meiotic prophase. Together with those in (B) these data demonstrate that Sk/Sp cells form and efficiently repair DSBs.
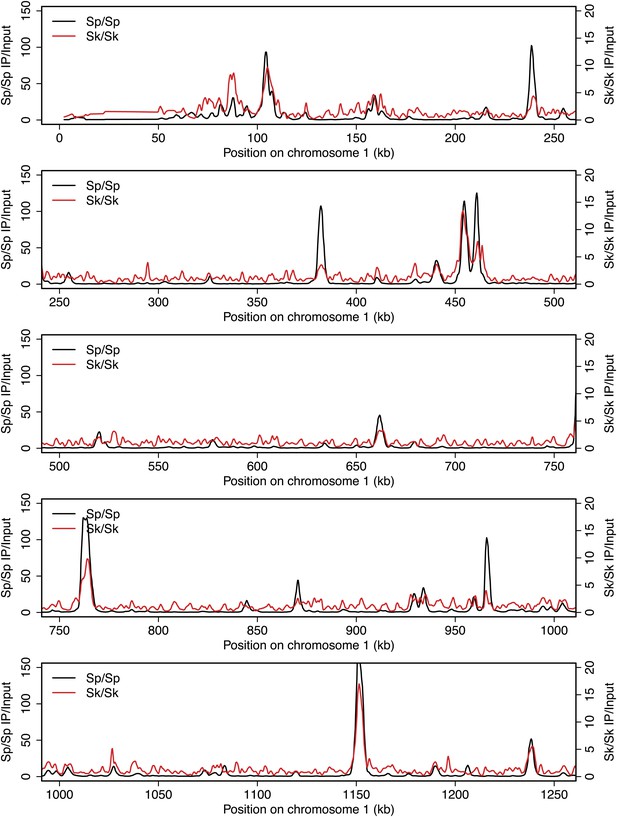
DSB hotspots in Sk and Sp.
We used ChIP-chip of Rec12-FLAG from rad50S Sk meiotic cultures to assay DSB hotspots and compared the Sk profile (red) to the published DSB hotspot maps of Sp (black) (Fowler et al., 2013). Traces are shown in Figure 2—figure supplements 1–11.
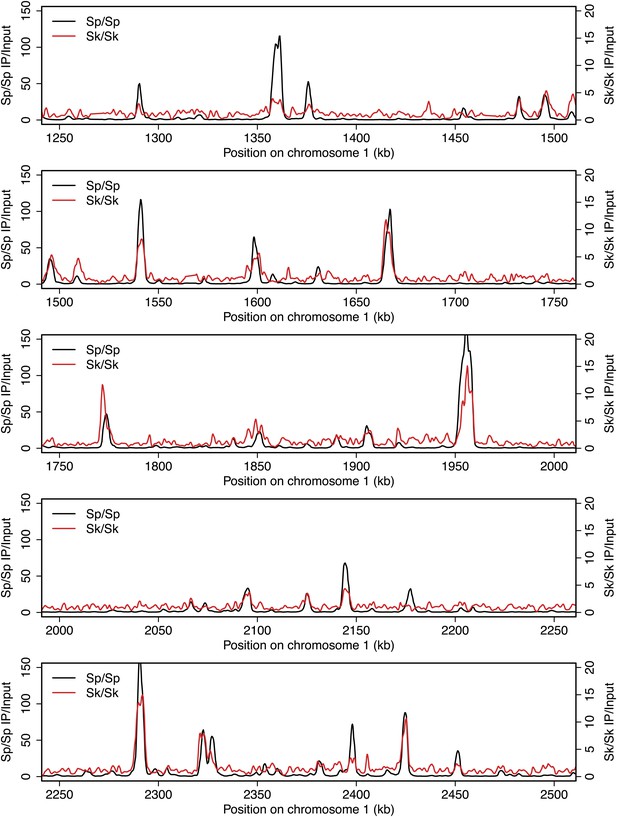
DSB hotspots in Sk and Sp.
We used ChIP-chip of Rec12-FLAG from rad50S Sk meiotic cultures to assay DSB hotspots and compared the Sk profile (red) to the published DSB hotspot maps of Sp (black) (Fowler et al., 2013). Traces are shown in Figure 2—figure supplements 1–11.
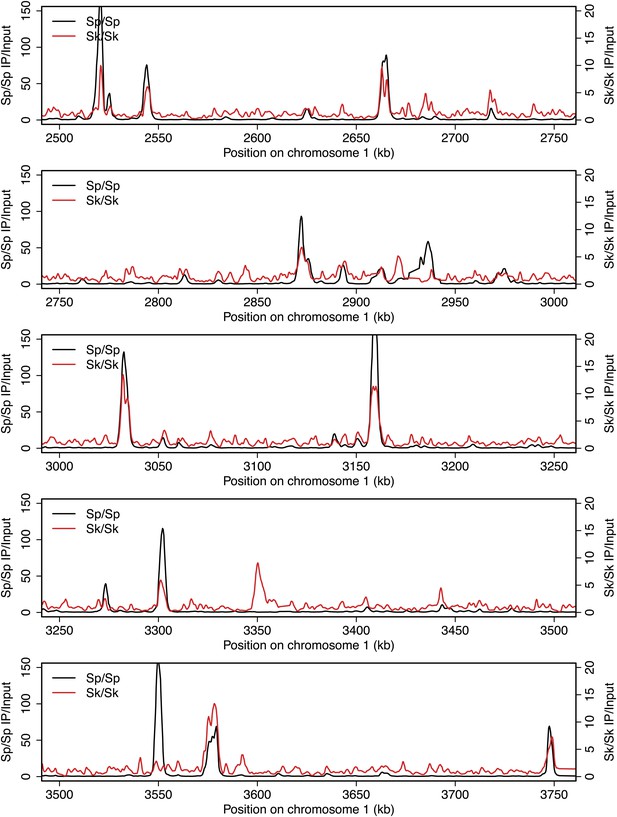
DSB hotspots in Sk and Sp.
We used ChIP-chip of Rec12-FLAG from rad50S Sk meiotic cultures to assay DSB hotspots and compared the Sk profile (red) to the published DSB hotspot maps of Sp (black) (Fowler et al., 2013). Traces are shown in Figure 2—figure supplements 1–11.
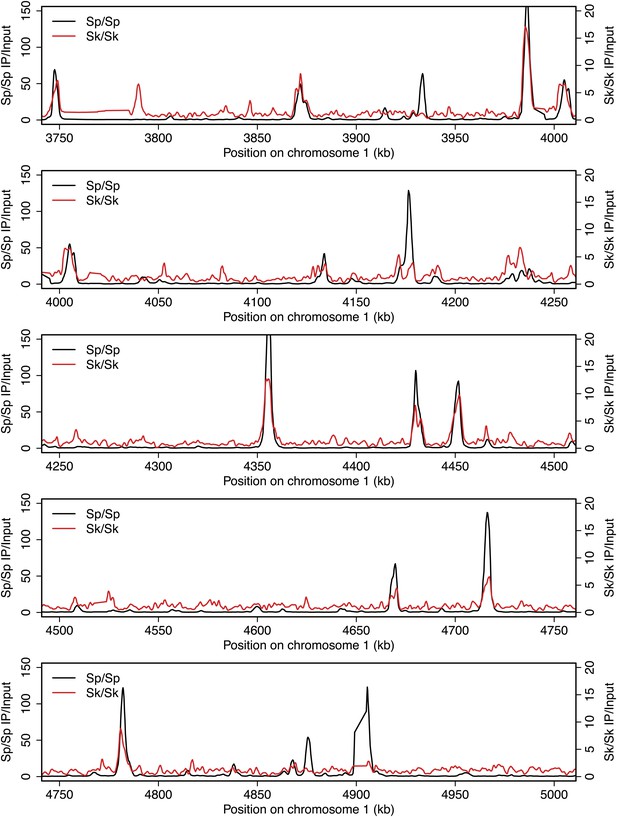
DSB hotspots in Sk and Sp.
We used ChIP-chip of Rec12-FLAG from rad50S Sk meiotic cultures to assay DSB hotspots and compared the Sk profile (red) to the published DSB hotspot maps of Sp (black) (Fowler et al., 2013). Traces are shown in Figure 2—figure supplements 1–11.
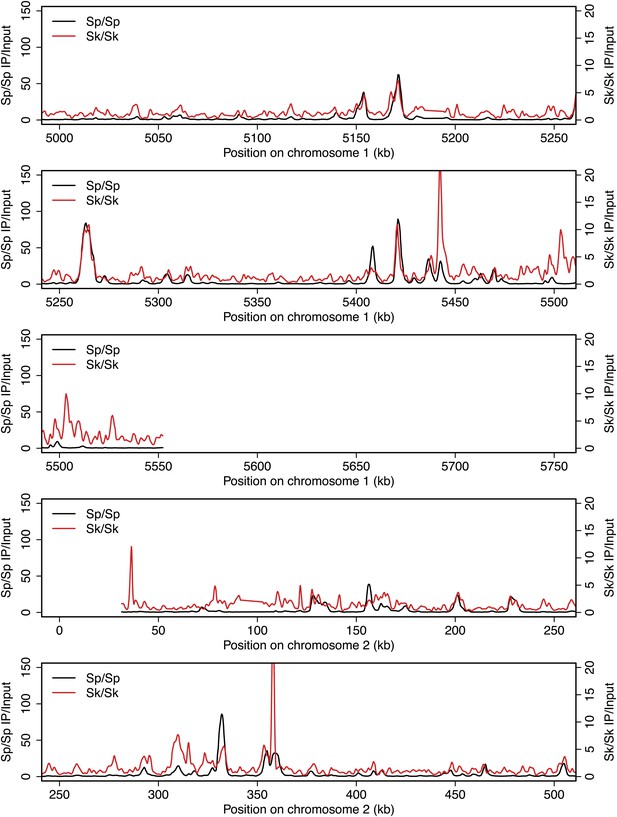
DSB hotspots in Sk and Sp.
We used ChIP-chip of Rec12-FLAG from rad50S Sk meiotic cultures to assay DSB hotspots and compared the Sk profile (red) to the published DSB hotspot maps of Sp (black) (Fowler et al., 2013). Traces are shown in Figure 2—figure supplements 1–11.
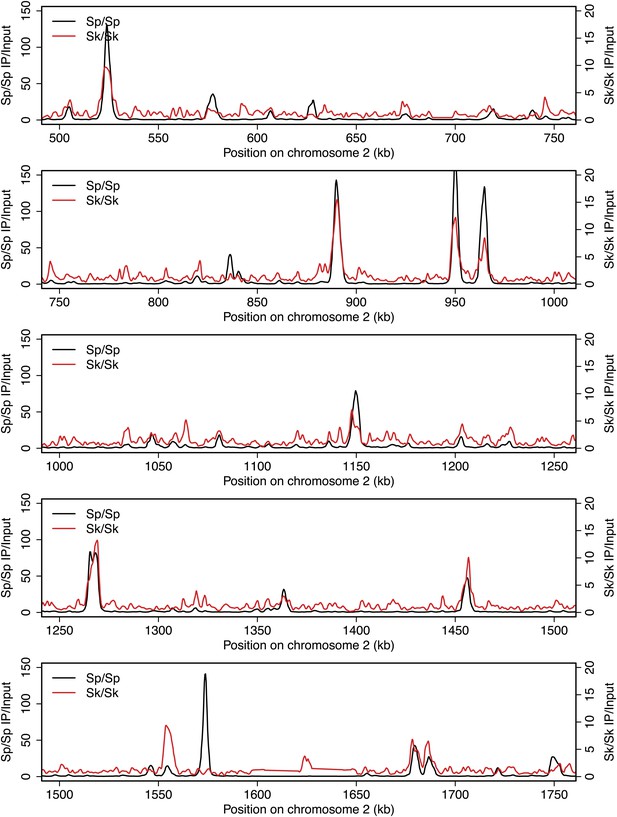
DSB hotspots in Sk and Sp.
We used ChIP-chip of Rec12-FLAG from rad50S Sk meiotic cultures to assay DSB hotspots and compared the Sk profile (red) to the published DSB hotspot maps of Sp (black) (Fowler et al., 2013). Traces are shown in Figure 2—figure supplements 1–11.
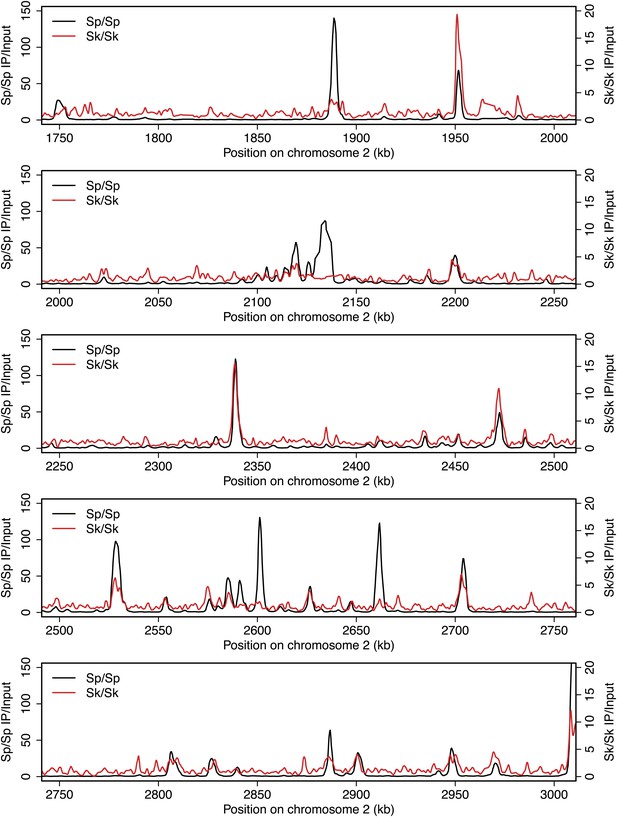
DSB hotspots in Sk and Sp.
We used ChIP-chip of Rec12-FLAG from rad50S Sk meiotic cultures to assay DSB hotspots and compared the Sk profile (red) to the published DSB hotspot maps of Sp (black) (Fowler et al., 2013). Traces are shown in Figure 2—figure supplements 1–11.
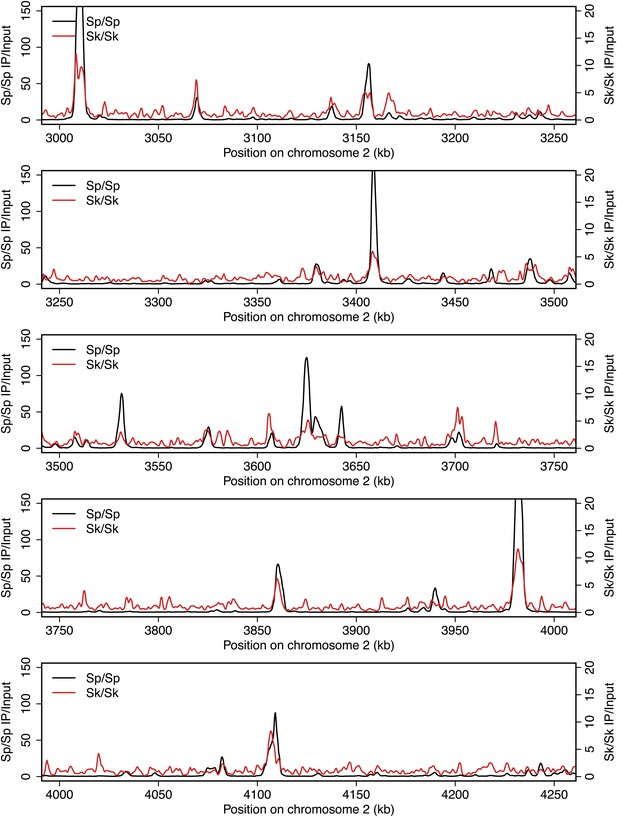
DSB hotspots in Sk and Sp.
We used ChIP-chip of Rec12-FLAG from rad50S Sk meiotic cultures to assay DSB hotspots and compared the Sk profile (red) to the published DSB hotspot maps of Sp (black) (Fowler et al., 2013). Traces are shown in Figure 2—figure supplements 1–11.
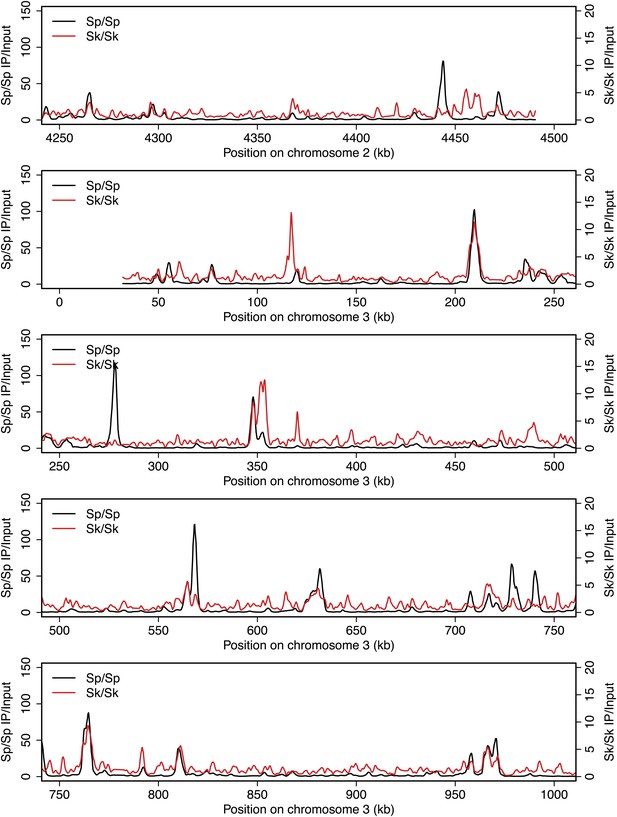
DSB hotspots in Sk and Sp.
We used ChIP-chip of Rec12-FLAG from rad50S Sk meiotic cultures to assay DSB hotspots and compared the Sk profile (red) to the published DSB hotspot maps of Sp (black) (Fowler et al., 2013). Traces are shown in Figure 2—figure supplements 1–11.
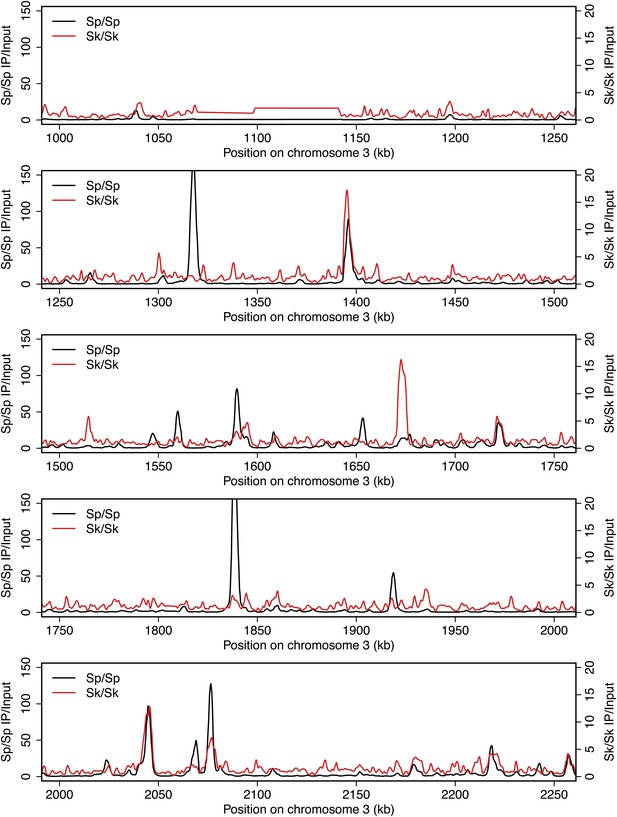
DSB hotspots in Sk and Sp.
We used ChIP-chip of Rec12-FLAG from rad50S Sk meiotic cultures to assay DSB hotspots and compared the Sk profile (red) to the published DSB hotspot maps of Sp (black) (Fowler et al., 2013). Traces are shown in Figure 2—figure supplements 1–11.
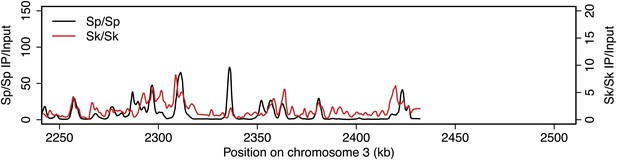
DSB hotspots in Sk and Sp.
We used ChIP-chip of Rec12-FLAG from rad50S Sk meiotic cultures to assay DSB hotspots and compared the Sk profile (red) to the published DSB hotspot maps of Sp (black) (Fowler et al., 2013). Traces are shown in Figure 2—figure supplements 1–11.
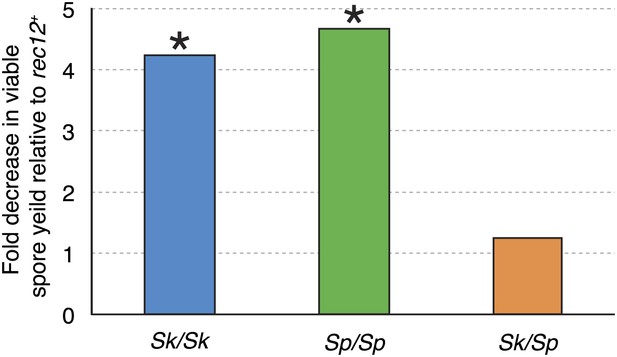
Recombination does not alter Sk/Sp hybrid fertility.
The average rec12+ viable spore yield of each diploid was divided by that of the corresponding rec12Δ mutant. For the pure species diploids, the viable spore yield was significantly lower in the absence of Rec12 (n ≥ 5 experiments for each genotype; * t test p<0.05). The viable spore yield of Sk/Sp hybrids, on the other hand, was not significantly different between rec12+ and rec12Δ Sk/Sp hybrids (p=0.42). This indicates that recombination likely hurts fertility just as much as it promotes fertility in Sk/Sp hybrid meiosis. These data are shown in a different format in Figure 5A.
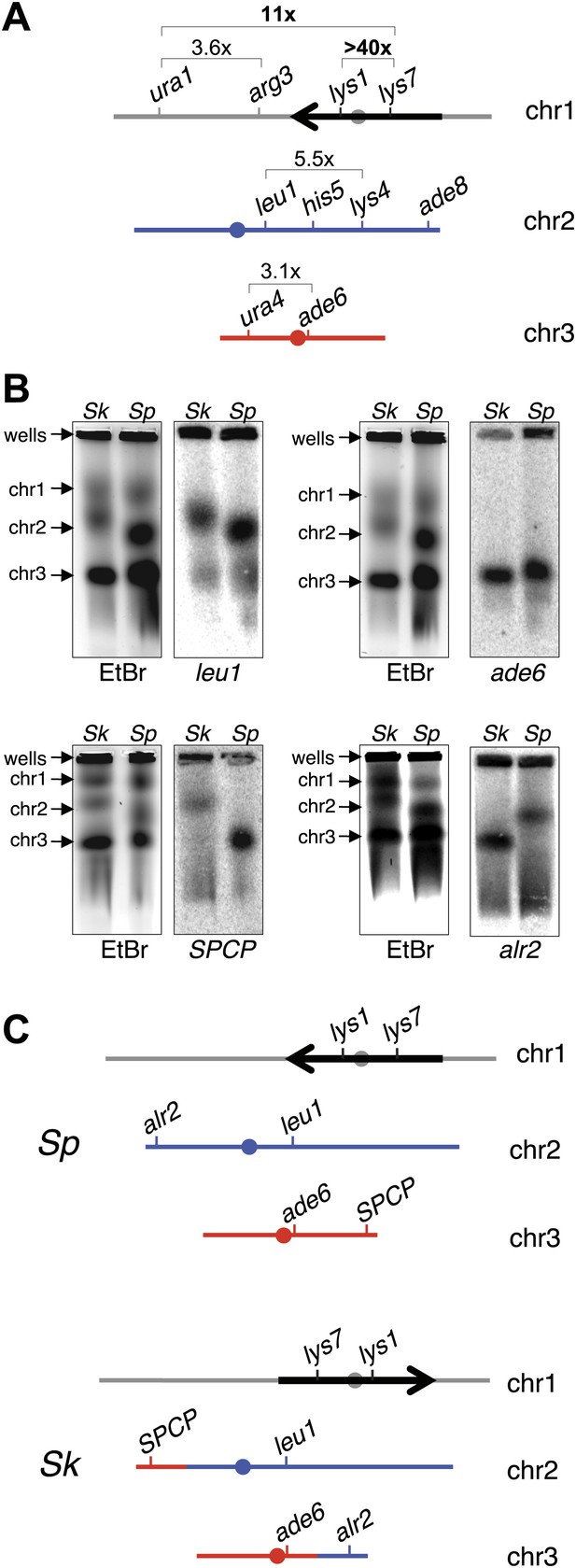
Chromosome rearrangements limit the recovery of recombinant progeny in Sk/Sp hybrids.
(A) A cartoon illustrating the fold decrease in recombinant frequencies in Sk/Sp spores compared to that in Sp/Sp spores. The detailed recombination data are in Figure 4—figure supplements 2 and 3. The Sp karyotype is depicted with a grey/black chromosome 1, blue chromosome 2, and red chromosome 3. The backwards black arrow indicates an inversion in Sp relative to the ancestral karyotype. (B) Pulsed-field gels separating Sp and Sk chromosomes and Southern blots of the gels probed with DNA from the indicated loci revealed a reciprocal translocation that includes several essential genes including alr2 and SPCP1E11.08 (abbreviated SPCP). The EtBr-stained gels are on the left and the Southern blots are on the right in each pair. (C) A cartoon summary of the karyotype differences between Sp and Sk. The arrow indicates the location of the inversion in Sp. A few landmark loci are shown.
Recombination frequencies in Sk.
We crossed Sk strains to measure recombination frequencies between the indicated loci. Sk cM were calculated from the data shown using Haldane’s formula and the Sp cM were calculated using the genome average of 0.16 cM/kilobase (Smith, 2009; Young et al., 2002) because crossover frequencies are relatively uniform in Sp and two of the intervals are too large to be measured directly.
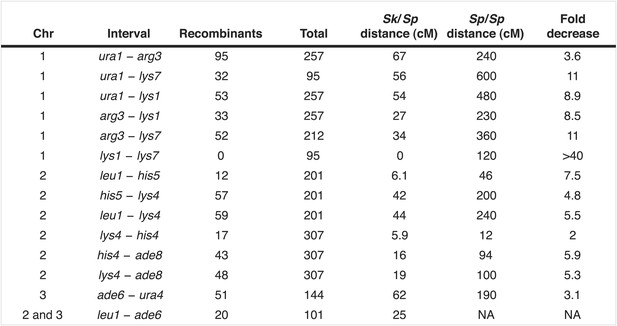
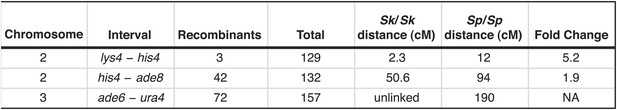
Recombination frequencies in Sk/Sp hybrids are low relative to Sp.
We crossed Sk and Sp strains to measure recombination frequencies between the indicated loci. Sk/Sp cM were calculated from the data shown using Haldane's formula and the Sp cM were calculated using the genome average of 0.16 cM/ kilobase (Smith, 2009; Young et al., 2002) because crossover frequencies are fairly uniform in Sp and most of the intervals are too large to be measured directly. The numbers of each parental and recombinant type are listed in Figure 4—figure supplement 3. NA, not applicable.
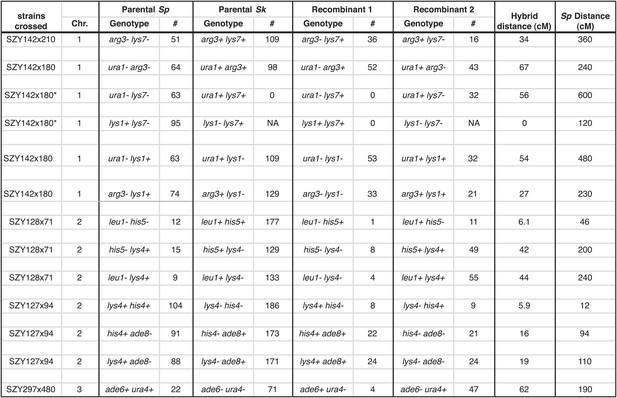
Sk alleles are underrepresented in the progeny of Sk/Sp hybrids.
This shows the numbers of each parental and recombinant class that were used to calculate recombination frequencies in Sk/Sp hybrids. These data are shown in simplified form in Figure 4—figure supplement 2. We crossed Sk and Sp strains to measure recombination frequencies between the indicated loci. Both parental and recombinant classes illustrate the effects of meiotic drive on the inheritance of each locus. In each cross, Sk-derived alleles are overrepresented. Sk/Sp cM were calculated from the data shown using Haldane’s formula and the Sp cM were calculated using the genome average of 0.16 cM/ kilobase (Smith, 2009; Young et al., 2002) because crossover frequencies are relatively uniform in Sp and most of the intervals are too large to be measured directly. *In the SZY142x180 cross, not all progeny were used to calculate recombination frequencies between lys7 and other markers. This was because the cross contained both lys7− and lys1− alleles. The lys7+ lys1− and lys7− lys1− genotypes could not be distinguished because they have the same phenotype. For that reason, we considered only Lys1+ (G418 sensitive) progeny so that we could use the Lys phenotype to genotype lys7. In all crosses, only spores containing one copy of the assayed chromosome were considered.
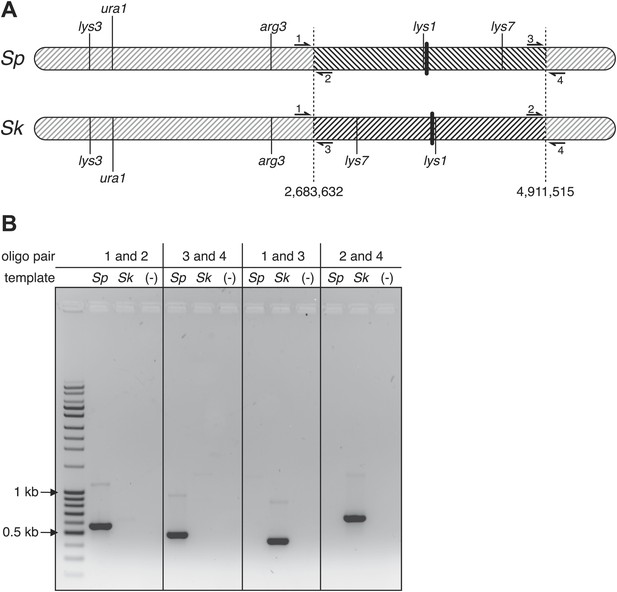
Sp has an inversion on chromosome 1 relative to Sk.
(A) Scaled schematic illustrating the breakpoints of the inversion shown as dashed vertical lines at positions 2,683,632 and 4,911,515 on the Sp chromosome. Several genetic markers, the centromeres (thick vertical lines), and the locations of PCR oligos (numbered 1–4) used to verify the inversion are shown. (B) The inversion was verified using PCR: ethidium bromide stained gel of PCR products using the indicated oligos on Sk and Sp DNA templates. Oligo pairs 1+2 and 3+4 produce a band in Sp, but not Sk. Oligo pairs 1+3 and 2+4 produce a band in Sk, but not Sp.
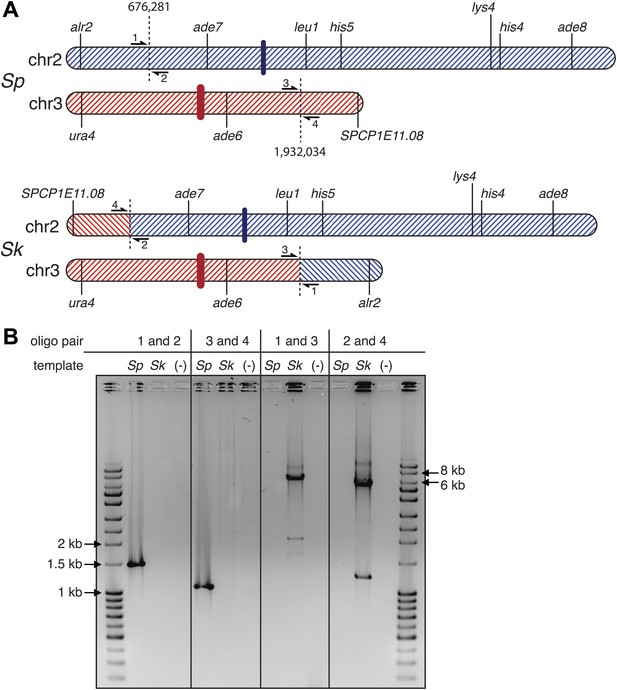
Sk has a reciprocal translocation between chromosomes 2 and 3.
The translocation likely occurred as homologous recombination between repetitive elements. (A) Scaled schematic illustrating the location of the two full-length transposons in Sk where the translocation occured, shown as dashed vertical lines at positions 676,281 and 1,932,034 on the Sp chromosomes. The position on chr2 marks the start of a single LTR also found in Sp. Several genetic markers, two (of many) essential genes within the translocated segments, the centromeres (thick colored vertical lines), and the locations of PCR oligos (numbered 1–4) used to verify the translocation are shown. (B) The translocation junction was verified using PCR: ethidium bromide stained gel of PCR products using the indicated oligos on Sk and Sp DNA templates. Oligo pairs 1+2 and 3+4 produce a band in Sp, but not Sk. Oligo pairs 1+3 and 2+4 produce a band in Sk, but not Sp. The larger size of the Sk bands corresponds to the presence of a transposon at each location in Sk, but not in Sp. We believe the smaller bands in the Sk lanes are artefacts of PCR amplification across transposons because we see them even when we PCR across known Sp transposons.
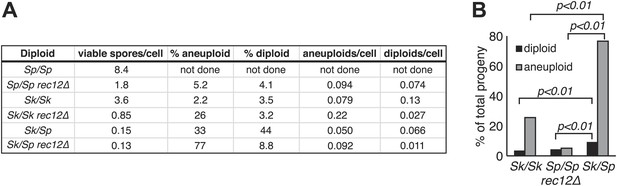
Increased aneuploidy amongst viable Sk/Sp gametes is recombination-independent.
(A) We calculated both viable spore yield (viable spores/cell) as well as the fraction of viable spores that are aneuploid or diploid (‘Materials and methods’). In the absence of Rec12, the relative frequencies of aneuploids and diploids are elevated in all cases. However, there is significantly more aneuploidy and diploidy of viable spores produced by rec12Δ Sk/Sp hybrids than by rec12Δ pure species controls. This shows the phenotype is not caused solely by recombination defects. In addition, Sk/Sp diploids do not generate more aneuploids or diploids relative to the number of cells induced to undergo meiosis compared to pure-species controls. Some of these data are presented in a different format in Figures 1H and Figure 3. (B) A bar graph illustrating the fraction of the viable spores produced by the indicated rec12Δ diploids that are aneuploid or diploid (G-test, n >300 for each).
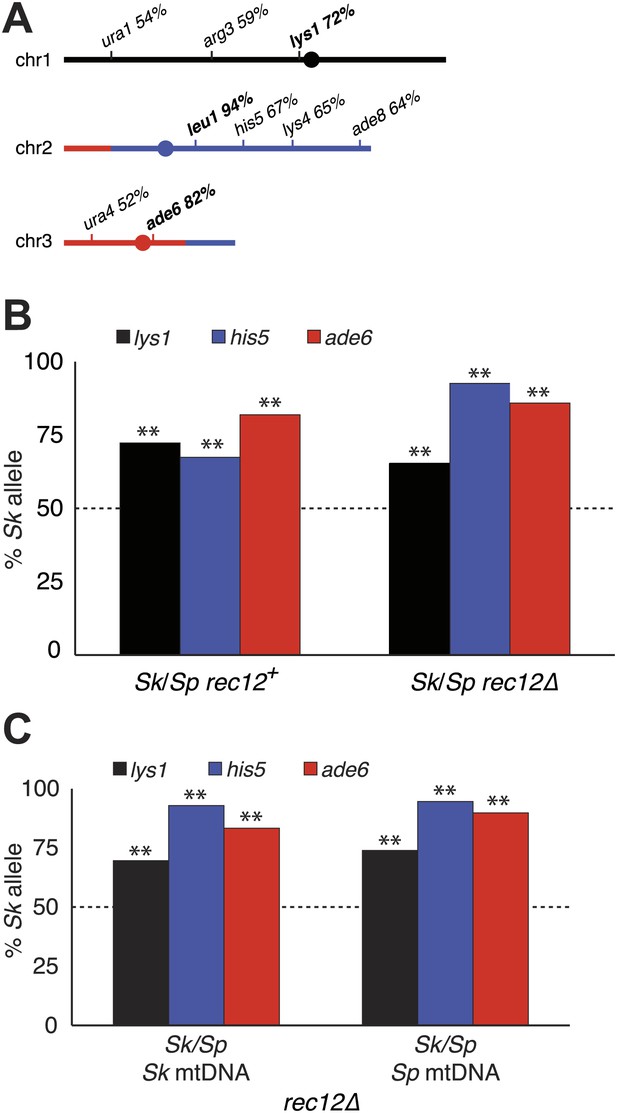
Alleles on all three Sk chromosomes show drive (independent of mitochondrial DNA type).
(A) Sk alleles were inherited by significantly more than 50% of the viable spores produced by Sk/Sp hybrids, except ura1 and ura4 (G-test p<0.01; n >100 for each). The markers nearest to the meiotic drive loci (i.e., those showing the greatest bias towards Sk inheritance) are shown in boldface. The color scheme is the same as that in Figure 4. The data underlying these numbers are shown in Figure 4—figure supplement 3, and Figure 6—figure supplement 1. (B) The Sk alleles of lys1, his5 and ade6 show significant drive both in the presence and absence of recombination (**p<0.01, n >300 for lys1 and his5, n >80 for ade6). The amount of his5 drive is greater in the absence to Rec12 due to enhanced linkage with the driving locus. The data underlying this graph are shown in Figure 6—figure supplement 1. (C) Incompatibilities between the Sk mitochondrial DNA and Sp nuclear genes are not responsible for the drive phenotype because we observed the same drive in rec12Δ Sk/Sp hybrids with either Sk or Sp-derived mitochondrial DNA (**p<0.01, n >200 for lys1 and his5, n >50 for ade6). The data underlying this graph are shown in Figure 6—figure supplement 4.
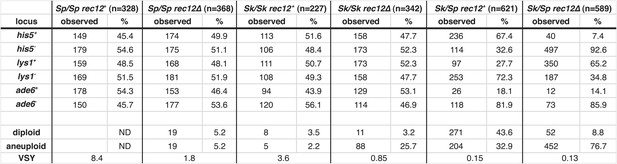
Summary of Sk/Sp hybrid and pure species diploid meiotic phenotypes and distribution of alleles in their progeny.
We induced Sk/Sp hybrids as well as Sk/Sk and Sp/Sp control diploids to undergo meiosis and measured their fertility using a viable spore yield assay (VSY). All diploids were heterozygous for genetic markers on chromosome 1 (lys1), chromsome 2 (his5) and chromsome 3 (ade6). We genotyped the viable spores from each diploid using these markers. In the rec12+ Sk/Sp hybrids, the Sk parent contributed the his5+, lys1−, and ade6− alleles. In cases where the diploids had co-dominant markers on chromosomes 2 and 3, we were also able to determine the number of chromosome 3 aneuploids and diploids amongst the viable spores. Parts of these data are also presented in Figure 1H, Figure 3, Figure 5, Figure 6, Figure 8, and Figure 8—figure supplement 2.
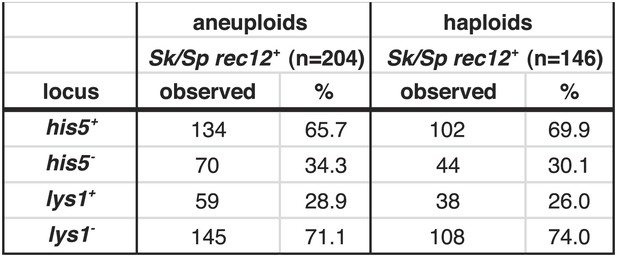
Biased transmission favoring Sk alleles on chromosomes 1 and 2 is observed in aneuploid and haploid spores.
The Sk/Sp rec12+ data presented in Figure 6B and Figure 6—figure supplement 1 were sorted to display aneuploid and haploid spores seperately. The diploid was heterozygous for genetic markers on chromosome 1 (lys1), chromsome 2 (his5) and chromsome 3 (ade6). We genotyped the viable spores of each diploid using these markers. The Sk parent contributed the his5+ and lys1− alleles.
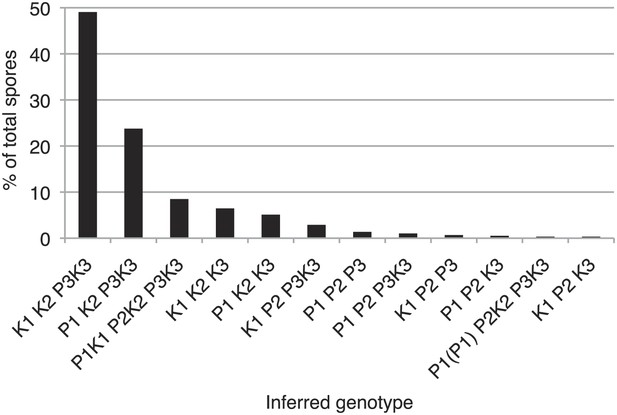
Distribution of progeny from rec12Δ Sk/Sp hybrid meiosis.
rec12Δ Sk/Sp hybrid diploids were sporulated and we genotyped the viable spores using genetic markers that distinguished each chromosome (n = 589). Chromosomes 2 and 3 contained codominant markers, which allowed us to definitively determine which strains had two copies of these chromosomes. We assumed that all strains that inherited two copies of chromosome 2 and chromosme 3 were diploids, because chromosome 1 aneuploids are inviable in Sp (Niwa et al., 2006). We assume the same is likely true in Sk and Sk/Sp hybrids because we very rarely recover marker combinations consistent with aneuploidy for chromosomes 1 or 2 from rec12Δ crosses. We assume that the few such progeny we do recover are rare recombinants (Davis and Smith, 2003). The plot shows the distribution of the viable spores that inherited each chromosome. Sp chromosomes are designated P1-P3, Sk chromosomes are designated K1-K3. This analysis is limited in that it assumes no recombination. However, recombination does occur at a low frequency (Davis and Smith, 2003). For example, all strains tested that have P2 K3 or K2 P3 (incompatible combinations) were recombinant.
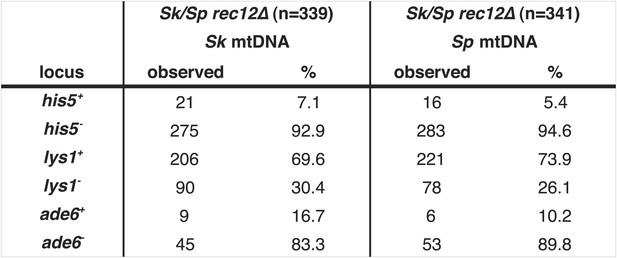
Meiotic drive in Sk/Sp hybrids is independent of mitochondrial DNA.
We induced Sk/Sp hybrids heterozygous for genetic markers on chromosome 1 (lys1), chromosome 2 (his5) and chromosome 3 (ade6) containing either Sk or Sp mitochondrial (mt) DNA to undergo meiosis. We genotyped the viable spores using these markers and indicated the number that inherited each allele. The Sk parent contributed the his5−, lys1+, and ade6− alleles.
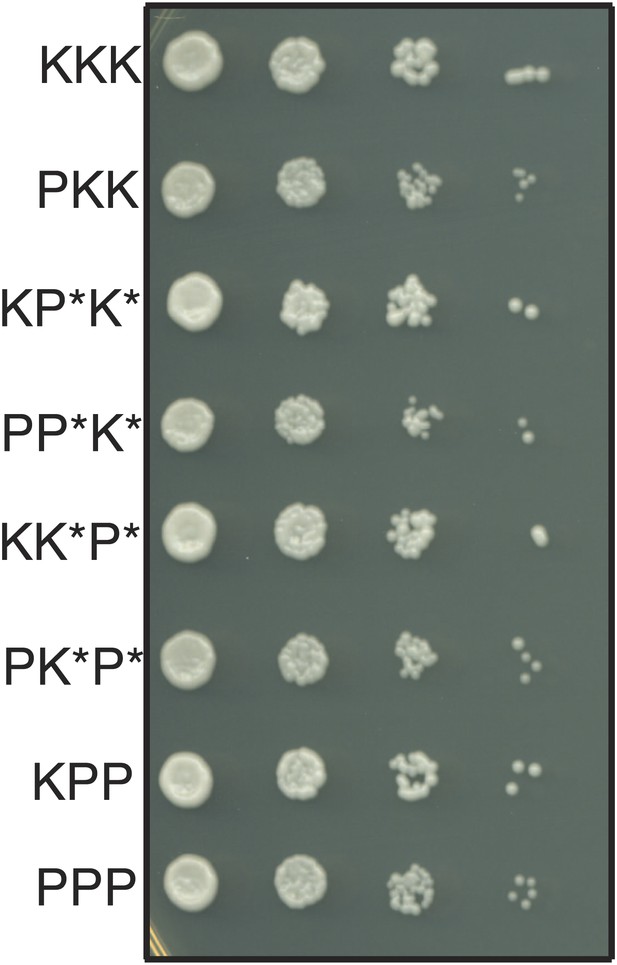
The haploid progeny of Sk/Sp hybrids have similar growth rates.
The progeny of rec12Δ Sk/Sp hybrids with the indicated chromosomes were diluted and grown on rich YEA medium. KKK indicates the Sk parental genotype, whereas PPP indicates the Sp parental genotype. The strains were genotyped using lys1, his5 and ade6 alleles on chromosomes 1, 2, and 3, respectively. Strains that inherit intact chromosomes 2 and 3 from different species are non-viable because they lack many essential genes. However, we do rarely recover viable recombinant strains that have alleles from Sk chromosome 2 and Sp chromosome 3 (and vice versa). Potentially recombinant chromosomes are denoted with an *. All haploid strains recovered have growth rates similar to that of the parental species, suggesting mitotic growth defects do not underlie differential recovery of the genotypes.
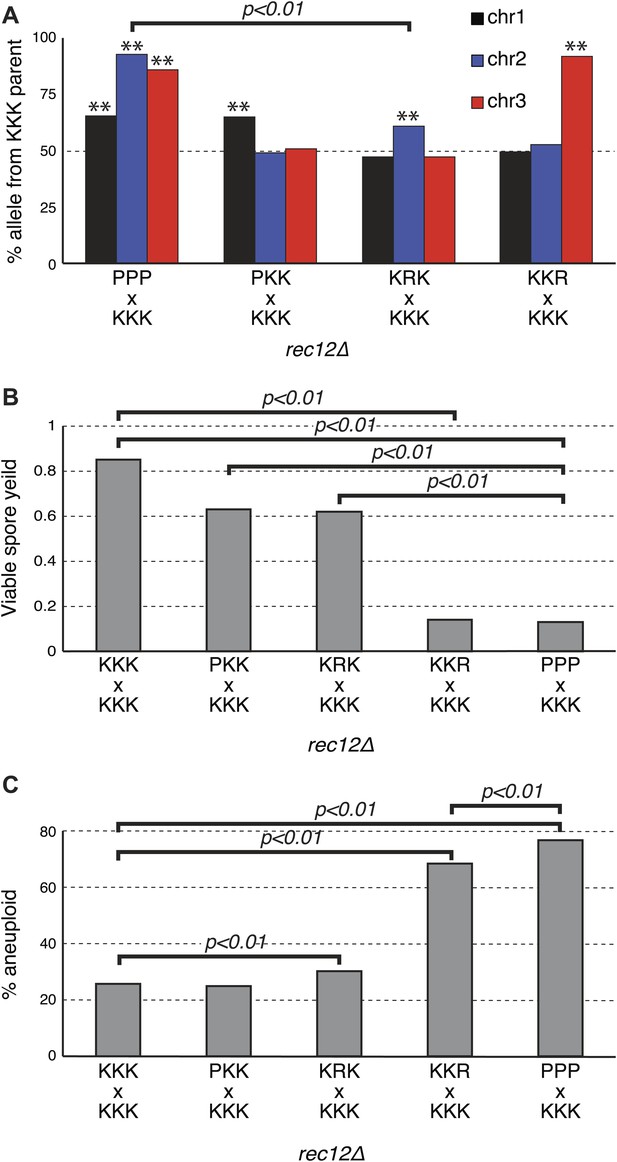
Sk drive alleles are autonomous and contribute to hybrid infertility.
Aneuploidy is largely caused by heterozygosity of Sk and Sp DNA on chromosome 3. (A) Comparison of meiotic drive phenotypes between rec12Δ diploids generated by mating Sk to Sp or to haploid strains obtained from Sk/Sp hybrids. ‘R’ indicates a recombinant chromosome (Figure 8—figure supplement 1), which is compatible with all Sk chromosomes but does not contain a meiotic drive allele. All Sk chromosomes can drive autonomously (** indicates drive; G-test p<0.01). However, the drive of Sk chr2 is lower in the KRK/KKK diploid than in pure PPP/KKK hybrids (G-test; n >500 for chromosomes 1 and 2, n >80 for chromosome 3 in each cross). The PPP/KKK data are also shown in Figure 6B. (B) Fertility defects of hybrids parallel the amount of drive observed amongst the viable spores (see A, p-values obtained from t-tests, averages of at least five experiments are shown). This is consistent with drive causing spore death. (C) The high aneuploidy amongst the viable progeny of Sk/Sp hybrids is largely due to heterozygosity of one or more loci on chromosome 3 (G-test, n >500 for each cross). The PPP/KKK and KKK/KKK viable spore yield and aneuploid data are also shown in Figure 5. The data underlying these graphs are summarized in Figure 8—figure supplement 2.
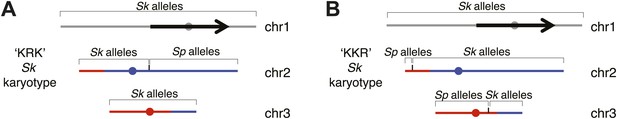
Genotype of recombinant strains used in Figure 8.
We sequenced these two genomes to determine which parent (Sk or Sp) contributed which components of the genome using single nucleotide polymorphisms (SNPs) to distinguish them. Both strains have the Sk karyotype. The black/grey indicates DNA that is located on chromosome 1 in Sp, blue indicates DNA that is located on chromosome 2 in Sp and red indicates DNA that is located on chromosome 3 in Sp. (A) Strain KRK has only Sk alleles on chromosomes 1 and 3. Chromosome 2 in this strain was generated by a crossover between Sk and Sp between SNPs 2,113,023 and 2,115,195 (which flank the mat1 locus). (B) Strain KKR has only Sk alleles on chromosome 1. On chromosome 2 only the left-most tip has Sp alleles. This chromosome was generated by a recombination event between Sk chromosome 2 and Sp chromosome 3. The crossover occurred between SNPs at positions 2,290,808 and 2,292,393 on Sp chromosome 3. Chromosome 3 in this strain was generated by a recombination event between Sk chromosome 3 and Sp chromosome 3 that occurred between SNPs located at positions 1,558,635 and 1,559,674 on Sp chromosome 3.
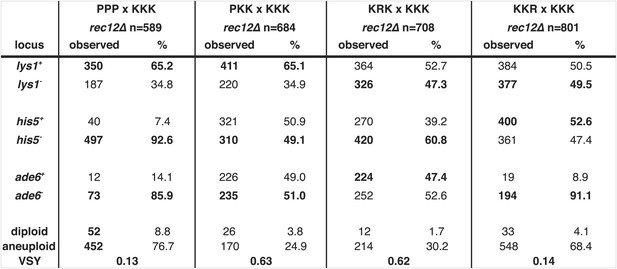
Summary of meiotic phenotypes for Sk/Sp hybrids and diploids with one heterozygous chromosome and the distribution of alleles in their viable progeny.
Detailed data underlying the plots in Figure 8. We induced Sk/Sp hybrids as well as diploids with only one heterozygous chromosome to undergo meiosis and measured their fertility using a viable spore yeild assay (VSY). All diploids were heterozygous for genetic markers on chromosome 1 (lys1), chromosome 2 (his5) and chromosome 3 (ade6). Sk chromosomes in the crosses to generate the diploids (top) are shown as ‘K’s whereas Sp chromosomes are shown as ‘P’s. The exact genotypes of the strains containing ‘R’ chromosomes can be found in Figure 8—figure supplement 1. We genotyped the viable spore progeny of each diploid. The alleles shown in bold were contributed by the pure Sk (KKK) parent. We used co-dominant markers on chromosomes 2 and 3 to determine the number of chromosome 3 aneuploids and diploids amongst the viable spore progeny. Some of the PPPxKKK data are also shown in Figure 5, Figure 6B, and Figure 6—figure supplement 1.
Additional files
-
Supplementary file 1
Yeast strain description.
- https://doi.org/10.7554/eLife.02630.033
-
Supplementary file 2
Oligo/primer description.
- https://doi.org/10.7554/eLife.02630.034





