Coordinated control of senescence by lncRNA and a novel T-box3 co-repressor complex
Figures
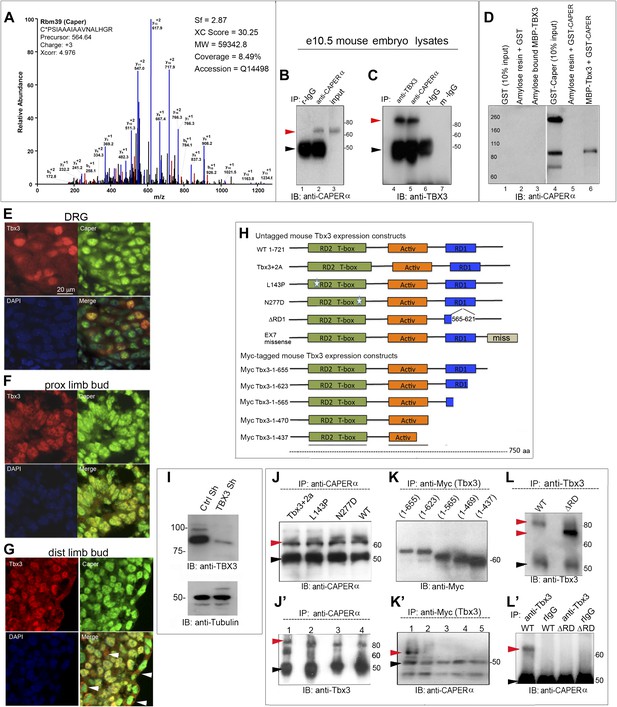
CAPERα and TBX3 directly interact via the TBX3 repressor domain.
(A) Representative spectrum for CAPERα identified in anti-TBX3 co-IP of HEK293 cell lysates. Mass spec analysis identified six specific CAPERα peptides, providing 8.5% sequence coverage of the protein. This spectrum shows fragmentation of one of these peptides, C*PSIAAAIAAVNALHGR, with diagnostic b- and y-series ions shown in red and blue, respectively. * indicates carbamidomethylation. (B) Anti-CAPERα immunoblot (IB) analysis of anti-CAPERα immunoprecipitated (IP'd, lane 2) e10.5 mouse embryo lysates. Black arrowheads indicate IgG heavy chain and red indicate protein of interest (CAPERα or TBX3). (C) Anti-Tbx3 IB of anti-Tbx3 (lane 4) and anti-Caperα (lane 5) IP'd mouse embryo lysates. Rabbit (r)-IgG (lanes1, 6) and mouse (m)-IgG (lane 7) are negative controls. (D) In vitro MBP pull down assay: MBP and MBP-Tbx3 bound amylose affinity columns were incubated with GST or GST-CAPERα. Bound proteins were eluted, subjected to SDS-PAGE followed by IB with anti-CAPERα antibody. (E–G) Colocalization of Tbx3 and Caperα in vivo shown by immunohistochemical analysis of sectioned e10.5 mouse embryo: embryonic dorsal root ganglion (DRG, E), proximal (F), and distal (G) limb bud with anti-Tbx3 (red) and anti-Caperα (green) antibodies and DAPI (blue). White arrowheads in G label representative ectodermal and mesenchymal cells with cytoplasmic Tbx3 and nuclear Caperα. (H) Schematic representation of mouse Tbx3 overexpression constructs.Tbx3 DNA binding domain (DBD) point, ΔRD and exon7 missense proteins are untagged and the C-terminal deletion mutants are Myc-tagged. (I) Anti-TBX3 IB of HEK293 cell lysates transfected with control or anti-TBX3 shRNA. (J) Anti-CAPERα IB of anti-CAPERα IP'd samples from HEK293 cells transfected with anti-TBX3 shRNA and expressing mouse Tbx3 proteins listed at top. Production and IP of endogenous CAPERα is not affected by production of mutant Tbx3 proteins. (J′) Anti-Tbx3 IB of anti-CAPERα IP'd samples from HEK293 cells transfected with anti-TBX3 shRNA and expressing Tbx3 proteins as in J. The DBD point mutant proteins (lanes 2, 3) interact with CAPERα as efficiently as wild type Tbx3 (lanes 1, 4). (K) Anti-Myc IB of anti-Myc IP'd samples from HEK293 cell lysates expressing Myc-tagged mouse Tbx3 C-terminal deletion mutants. The mutant proteins are expressed and efficiently IP'd. These cells were not treated with anti-TBX3 shRNA because the expression constructs produce a Myc- tagged mutants that can be IP'd independently of endogenous TBX3. (K′) anti-CAPERα IB of anti-Myc IP'd samples from HEK293 cell lysates expressing Myc-tagged mouse Tbx3 C-terminal deletion mutants. These cells were not treated with anti-TBX3 shRNA because the expression constructs produce a Myc- tagged mutants that can be IP'd independently of endogenous TBX3. (L) Anti-Tbx3 IB of anti-Tbx3 IP'd samples from HEK293 cells transfected with anti-TBX3 shRNA and expressing wt or repressor domain deletion mutant (ΔRD) mouseTbx3. The shRNA does not prevent production of the overexpression proteins. (L′) Anti-CAPERα IB of HEK293 cells transfected with anti-TBX3 shRNA and expressing mouse wt or ΔRD Tbx3 proteins and IP'd with anti-Tbx3 or IgG. Loss of the repressor domain prevents interaction with CAPERα. Black arrowheads indicate IgG heavy chain and red indicate protein of interest (CAPERα or TBX3). TBX3, CAPERα = human; Tbx3, Caperα = mouse.
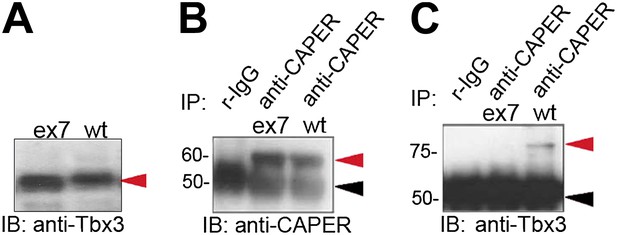
Missense mutation of the C-terminus of Tbx3 disrupts interaction with CAPERα.
(A) Anti-Tbx3 IB of exon 7 missense (ex7) and wt proteins expressed in HEK293 cells also transfected with anti-TBX3 shRNA. The overexpressed proteins are produced (red arrowhead). (B) anti-CAPERα and anti-TBX3 (C) IB of anti-CAPERα and negative control IP'd samples from HEK293 cells transfected with anti-TBX3 shRNA and overexpressing ex7 missense or wt Tbx3. Production and IP of endogenous CAPERα is not affected by production of mutant Tbx3 proteins. (C) Anti-Tbx3 IB of anti-CAPERα and negative control IP'd samples from HEK293 cells transfected with anti-TBX3 shRNA and overexpressing ex7 missense or wt Tbx3. The missense mutation disrupts interaction between Tbx3 and CAPERα. Black arrowheads indicate IgG heavy chain and red indicate protein of interest (CAPERα or TBX3). TBX3, CAPERα = human; Tbx3, Caperα = mouse.
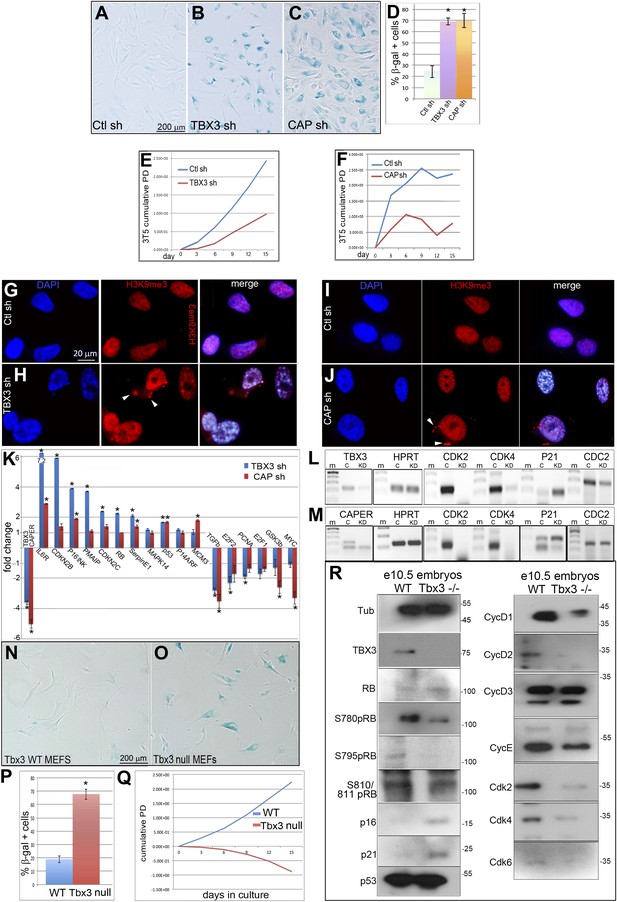
Knockdown of endogenous CAPERα and TBX3 in primary human fibroblasts and mouse embryos induces premature senescence and disrupts expression of cell cycle and senescence regulators.
(A–C) Representative bright field images of senescence associated β-galactosidase (SA-βG) assays of HFFs transduced with control, TBX3 shRNA A or CAPERα shRNA A. Only occasional cells in the control transduction have detectable lacZ staining (blue) whereas knockdown of either TBX3 or CAPERα results in marked changes in cell morphology and increased lacZ staining. (D) Bar graph quantitating % beta-galactosidase positive cells from four replicate plates of SA-βgal assays. * indicates p<0.001 compared to control. (E and F) 3T5 cell proliferation assay (Lessnick et al., 2002) of cumulative population doublings in HFFs transduced at passage 30 with control, TBX3 or CAPERα shRNAs. These are representative curves of duplicate experiments; each point on the curve is a measurement of cell count from a single plating followed over the course of the experiment as described in methods. (G–J) Immunohistochemical analysis of H3K9me3 immunoreactivity (red) and DAPI (blue) in HFFs after knockdown with control (G and I), TBX3 (H), or CAPERα (J) shRNAs. Individual channels are shown and the merged image is on the right. Note increased nuclear punctate staining consistent with Senescence-associated heterochromatin foci (SAHFs) in both channels and evidence of nuclear disruption (white arrowheads in red channel) after loss of either TBX3 or CAPERα. (K–M) Analysis of cell cycle and senescence marker transcript levels in HFFs transduced with control, TBX3, or CAPERα shRNAs. (K) Relative transcript levels assessed by quantitative real time-PCR (qPCR) of cDNA. Values reflect fold change in knockdown HFFs relative to control after normalization to HPRT levels. Note general pattern of expression changes are similar in TBX3 (blue) and CAPERα (red) knockdowns. Data are plotted as fold change mean ± standard deviation. * indicates p<0.05 relative to control. (L and M) Agarose gel of PCR amplicons of cDNAs reverse transcribed from TBX3 (L) or CAPERα (M) shRNA knockdown HFF RNA reveals similar decreases in cell cycle promoting genes CDK2 and 4 in TBX3 and CAPERα knockdowns and increased p21 levels. (N and O) SA-βgal assay of wild type and Tbx3 null MEFS reveals that Tbx3 is required to prevent premature senescence of primary murine embryonic fibroblasts (MEFs). (P) Quantitation of % beta-galactosidase positive cells from five replicate experiments exemplified in O, P. * indicates p<0.01. (Q) 3T5 cell proliferation assay of cumulative population doublings in wild-type and Tbx3 null MEFs. These are representative curves from duplicate experiments; each point on the curve is a measurement of cell count from a single plating followed over the course of the experiment as described in 'Materials and methods'. (R) IBs to assay levels of cell cycle and senescence proteins in wild type and Tbx3 null embryo lysates. Tubulin loading control is at top left (Tub). The changes at the protein level correlate with those observed at the RNA level (K–M) and RB is hypophosphorylated on multiple serine residues consistent with increased p16 and decreased CDK activity. TBX3, CAPERα = human; Tbx3, Caperα = mouse.
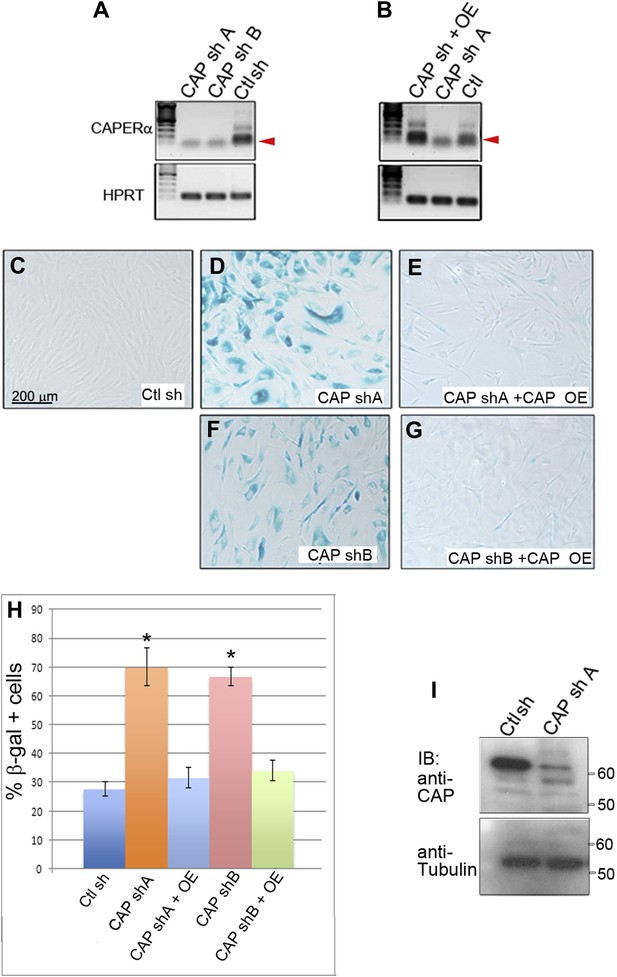
Effective knockdown of endogenous CAPERα in primary human foreskin fibroblasts using viral shRNA transduction.
(A) RT-PCR analysis of CAPERα and HPRT transcript levels in HFFs transduced with two different retroviruses producing anti-CAPERα shRNAs (CAP sh A and B) and control shRNA virus (Ctl sh). Red arrowhead indicates CAPERα-specific amplicon. (B) RT-PCR analysis of CAPERα and HPRT transcript levels in HFFs transduced with retroviruses producing anti-CAPERα shRNA A and a CAPERα overexpression virus (CAP OE). Note rescue of CAPERα expression by overexpression virus. (C–G) SA-βGal assays of HFFs transduced with control or CAPERα shRNAs A or B and rescue by CAPERα overexpression. (H) Quantitation of SA-βGal assays in C–G. * indicates p<0.01 compared to control shRNA. (I) Western blot showing depletion of endogenous CAPERα protein by CAP shRNA A. Anti-tubulin IB is loading control. This CAPERα shRNA ‘A’ was used for all subsequent CAPERα shRNA knockdown experiments. TBX3, CAPERα = human; Tbx3, Caperα = mouse.
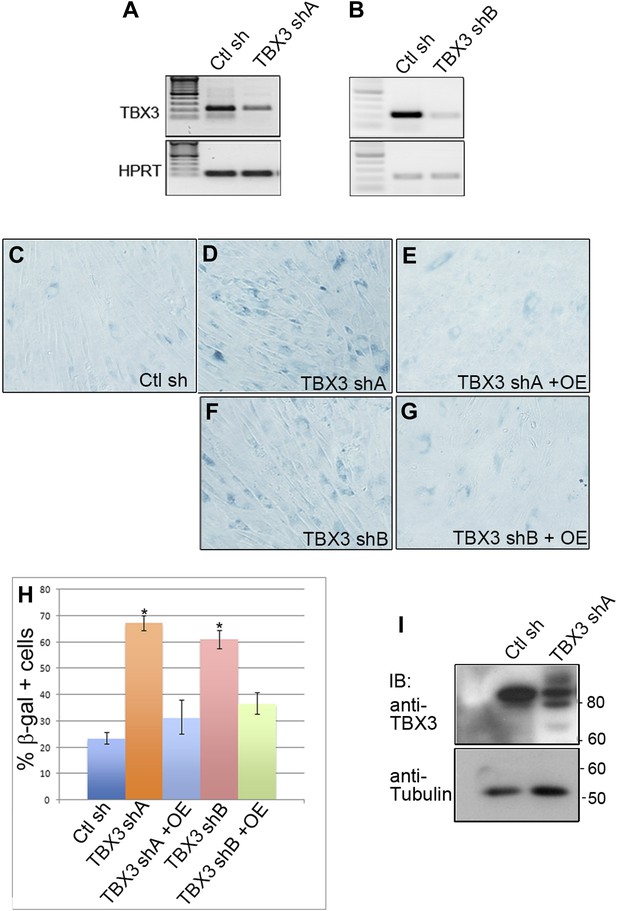
Effective knockdown of endogenous TBX3 in primary human foreskin fibroblasts using viral shRNA transduction.
(A) RT-PCR analysis of TBX3 and HPRT transcript levels in primary human foreskin fibroblasts (HFFs) transduced with control (Ctl sh) or TBX3 (TBX3 shA) shRNA retrovirus. (B) RT-PCR analysis of TBX3 and HPRT transcript levels in primary human foreskin fibroblasts (HFFs) transduced with control (Ctl sh) or TBX3 (TBX3 shB) shRNA retrovirus. (C–G) SA-βGal assays of HFFs transduced with control or TBX3 shRNAs A or B and rescue by Tbx3 overexpression. (H) Quantitation of SA-βGal assays in C–G. * indicates p<0.01 compared to control shRNA. (I) Western blot showing depletion of endogenous TBX3 protein by TBX3 shRNA A. Anti-tubulin IB is loading control. This TBX3 shRNA ‘A’ was used for all subsequent TBX3 shRNA knockdown experiments. TBX3, CAPERα = human; Tbx3, Caperα = mouse.
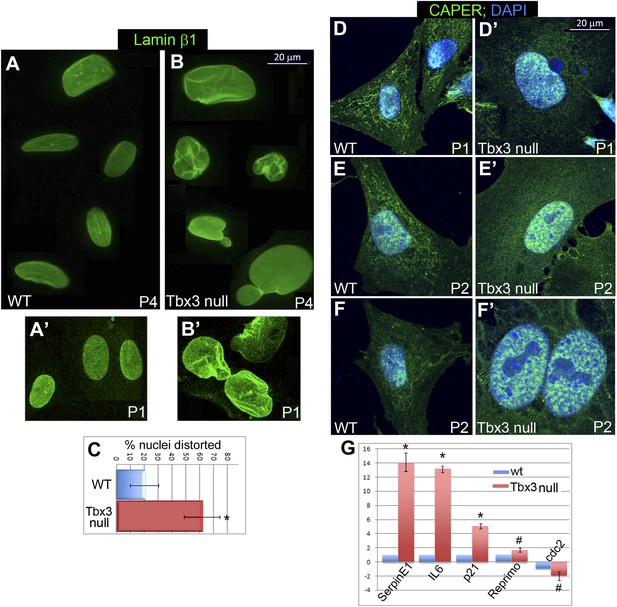
Tbx3 null murine embryonic fibroblasts (MEFS) have altered lamin β1 localization, nuclear disruption and mislocalized Caperα.
. (A–B′) Representative WT and Tbx3 null MEFs cells stained for laminβ1 at passage (P) 4 (A and B) and P1 (A′ and B′); note nuclear distortion and rupture in senescing Tbx3 null MEFs as early as P1. (C) Quantitation of % distorted nuclei in WT vs Tbx3 null MEFs. * indicates p<0.05. (D–F′) Immunohistochemistry for Caperα (green) and DNA (DAPI, blue) in control and Tbx3 null MEFs at P1 (D and D′) and P2 (E–F′). In mutant cells, Caperα signal shifts to nucleus from cytoplasm at P1, and large intranuclear Caperα+ foci are present by P2. (G) qPCR quantitation of senescence marker genes in WT vs Tbx3 null MEFs. Data are displayed as mean fold change ± standard deviation relative to WT after normalization to HPRT levels. * indicates p<0.01. # indicates p<0.05. TBX3, CAPERα = human; Tbx3, Caperα = mouse.
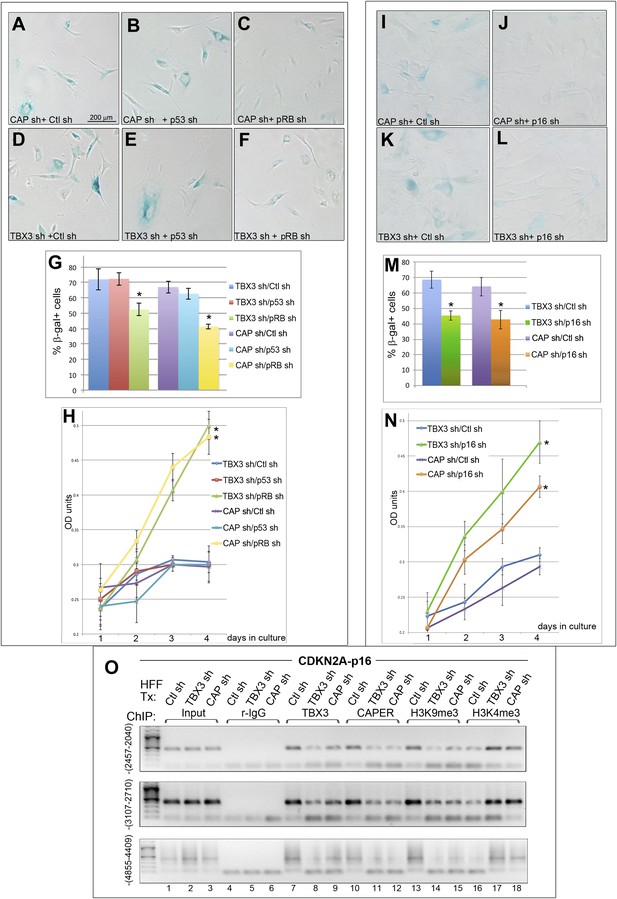
RB and p16 mediate senescence after CAPERα/TBX3 loss of function and CAPERα/TBX3 regulates chromatin structure of CDKN2A-p16.
(A–F) SA-βgal assays of HFFs stably transduced with control (Ctl) or p53 (Masutomi et al., 2003) or RB (Boehm et al., 2005) shRNAs subsequently transduced with CAPERα or TBX3 shRNAs. (G) % Quantitation of A–F from three replicate experiments. * indicates p<0.05 relative to Control or p53 shRNAs. (H) Cell proliferation assayed by crystal violet incorporation (OD units) in HFFs treated as in A–F. * indicates p<0.001 relative to Ctl or p53 shRNAs. (I–L) SA-βgal assays of HFFs stably transduced with control or p16 (Haga et al., 2007) shRNAs subsequently transduced with CAPERα or TBX3 shRNAs. (M) % Quantitation of I-L from three replicate experiments. * indicates p<0.05 relative to Ctl shRNA. (N) Cell proliferation assayed by crystal violet incorporation (OD units) in HFFs treated as in I–L. * indicates p<0.01 relative to Ctl shRNA. (O) ChIP-PCR with antibodies listed at top on three regions upstream of the CDKN2A-p16 transcriptional start site (TSS); position relative to (TSS) is indicated in parentheses at left of panels. PCR of input material used for the ChIP is shown under ‘Input’. The shRNA transduced is listed above each lane (HFF Tx). TBX3 knockdown decreases binding of TBX3 (lanes 8) and CAPERα (lanes 11) to all three regions. CAPERα knockdown has minimal effect on TBX3 binding (lanes 9). Knockdown of either TBX3 or CAPERα decreases the repressive chromatin mark H3K9me3 (lanes14, 15) and increases the activating chromatin mark H3K4me3 (lanes 17, 18). TBX3, CAPERα = human; Tbx3, Caperα = mouse.
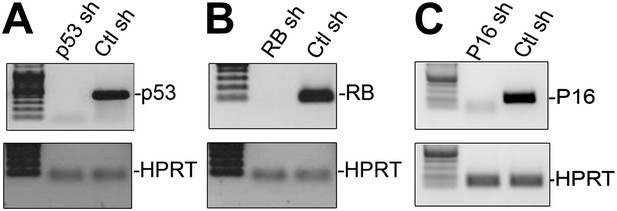
Effective knockdown of p53, RB and p16 in HFFs.
(A–C) RT-PCR analysis of p53 (A), RB (B) and p16 (C) transcript levels relative to HPRT after shRNA-mediated KD in HFFs. The shRNAs employed for these knockdowns were obtained from Addgene and have been previously employed by numerous investigators (Masutomi et al., 2003; Boehm et al., 2005; Haga et al., 2007; Hong et al., 2009; Elzi et al., 2012). TBX3, CAPERα = human; Tbx3, Caperα = mouse.
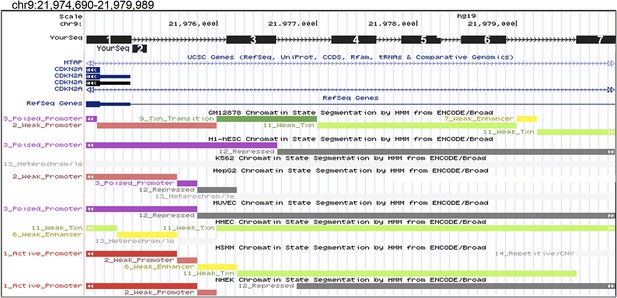
UCSC Genome Browser view of the CDKN2A locus and 5′ regions screened for binding by CAPERα and TBX3.
Seven regions tested upstream of CDKN2A-p16 promoter by ChIP with anti-TBX3 and anti-CAPERα antibodies. Amplicons are numbered black boxes 1–7 ‘Your Seq’ at top superimposed on window from UCSC genome browser. Chromatin states in various cell types based are noted by colored bars below. Of these 7 regions, 3 were bound by both TBX3 and CAPERα: regions 3, 4 and 5 (data are presented in Figure 3O). TBX3, CAPERα = human; Tbx3, Caperα = mouse.
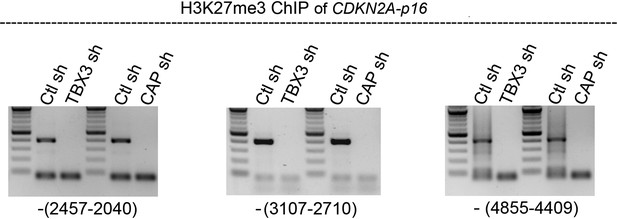
CDKN2a-p16 H3K27 trimethylation markedly decreases in HFFS after knockdown of CAPERα or TBX3 consistent with activation of CDKN2a-p16 expression.
ChIP-PCR of CDKN2A-p16 regulatory elements with anti-H3K27me3 in control, TBX3 or CAPERα shRNA-transduced HFFs. Locations of amplicons relative to transcription start site are noted in parentheses below each panel and correspond to regions 3, 4 and 5 in Figure 3—figure supplement 2. TBX3, CAPERα = human; Tbx3, Caperα = mouse.
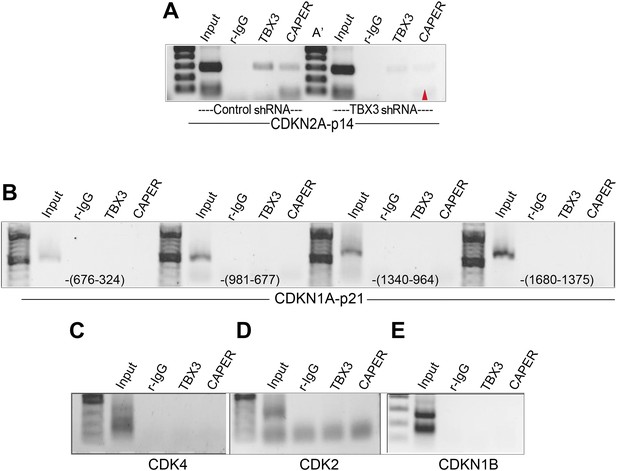
Testing CAPERα and TBX3 binding to p14, p21, CDK2, CDK4, and CDKN1B regulatory elements.
(A) ChIP-PCR of CDKN2A-p14 promoter with antibodies listed at top in control (C) and TBX3 siRNA (C′) transduced HFFs. Red arrowhead indicates loss of CAPERα binding after TBX3 knockdown. (B–E) ChIP/PCR of HFF chromatin showing lack of TBX3 and CAPERα binding to known regulatory elements (Baksh et al., 2002; Wang et al., 2005; Louie et al., 2010) of: (B) CDKN1A-p21 (location relative to transcription start site is noted in parentheses at the bottom of the panels (C) CDK4 (D) CDK2 (E) CDKN1B. TBX3, CAPERα = human; Tbx3, Caperα = mouse.
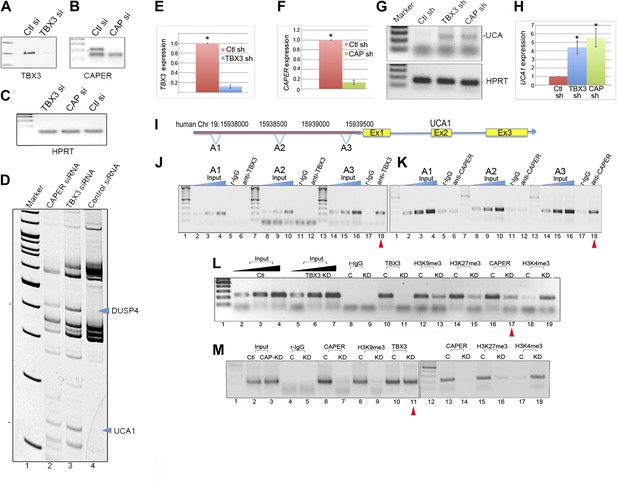
CAPERα/TBX3 directly represses expression of the long noncoding RNA UCA1.
(A–C) Gel showing RT-PCR analysis of TBX3, CAPERα, and HPRT expression in control, TBX3 and CAPERα siRNA-transfected HEK293 cells. The siRNAs effectively decreased transcript levels of their targets. (D) Differential display: representative PAGE gel of cDNAs derived from random primed, RT-PCR'd mRNAs from CAPERα, TBX3 and control siRNA transfected HEK293 cells. Blue arrowheads denote upregulated transcripts subsequently identified by sequencing as DUSP4 and UCA1. (E and F) qPCR analysis of TBX3 and CAPERα transcript levels in control and TBX3 or CAPERα shRNA transduced HFFs (repeat of experiment shown in Figure 2—figure supplements 1A and 2A). (G) RT-PCR analysis of UCA1 and HPRT gene expression in control, TBX3 or CAPERα shRNA-transduced HFFs. (H) qPCR analysis of UCA1 transcript levels in control, TBX3 or CAPERα shRNA transduced HFFs. Results confirm differential display result that KD of TBX3 or CAPERα results in increase in UCA1 transcript levels. (I) Schematic representation of the UCA1 locus with primer sets employed for ChIP-PCR amplification of denoted regions 5′ of gene (A1, A2, A3). (J) Anti-TBX3 ChIP-PCR of regions of the UCA1 promoter in HFFs; only A3 is ChIP'd by TBX3 (lane 18, red arrowhead). (K) Anti-CAPERα ChIP-PCR of regions of the UCA1 promoter in HFFs; only A3 chromatin is ChIP'd (lane 18, red arrowhead). (L) ChIP-PCR analysis of UCA1/A3 chromatin from in HFFs transduced with control (C) or TBX3 (KD) shRNA; ChIP antibodies are listed at top. Note decreased CAPERα binding after TBX3 KD (lane 17, red arrowhead), gain of activating mark H3K4me3 and loss of repressive marks H3K9me3 and H3K27me3. (M) ChIP-PCR analysis of UCA1/A3 with antibodies listed at top of panel in HFFs transduced with control (C) or CAPERα shRNAs. Note continued TBX3 binding despite CAPERα KD (lane 11, red arrowhead) and changes in chromatin marks parallel those seen in with TBX3 KD in panel L. TBX3, CAPERα = human; Tbx3, Caperα = mouse.
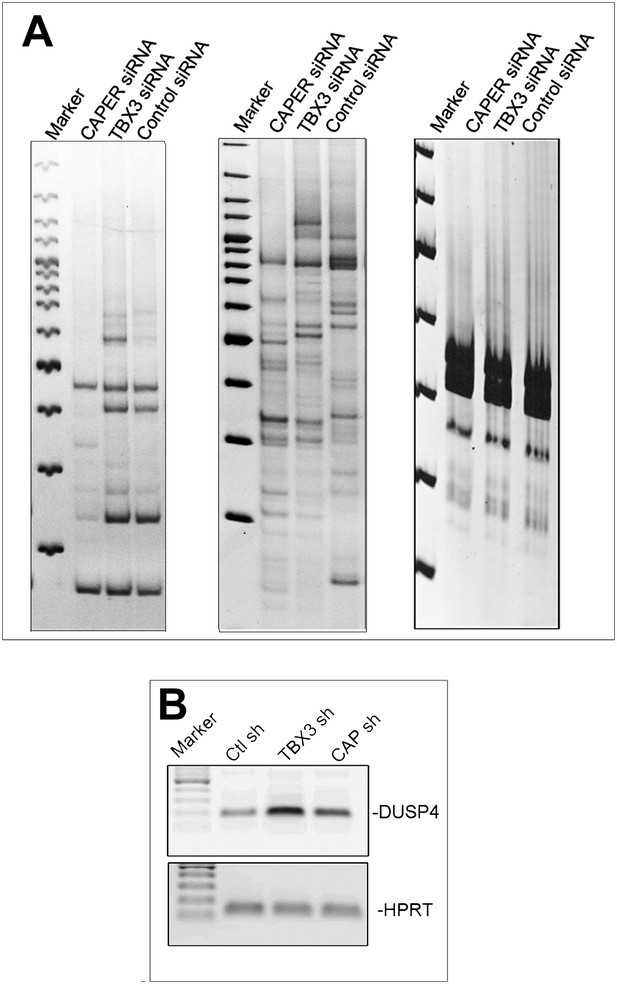
Validation of differential display findings.
(A) Additional representative differential display gels with transcripts unchanged or independently affected by knockdown of CAPERα or TBX3 in HEK293 cells. (B) RT-PCR validating differential display result of increased DUSP4 transcripts (Figure 4D) after CAPERα or TBX3 KD in HEK293 cells. TBX3, CAPERα = human; Tbx3, Caperα = mouse.
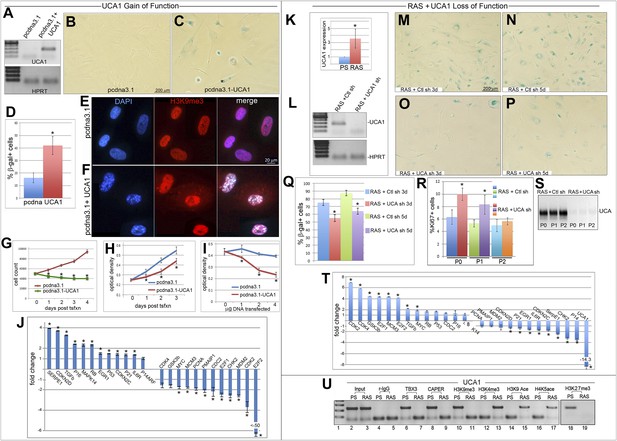
UCA1 expression is sufficient to induce senescence and required for normal execution of oncogene-induced senescence.
(A) UCA1 and HPRT transcripts assessed by RT-PCR in control and UCA1-overexpressing HFFs. (B and C) Representative bright field images of SA-βgal assay of cultured HFFs transfected with control and UCA1 overexpression plasmids. (D) % quantitation of SA-βgal cells from five replicates in control and UCA1 overexpressing HFFs. * indicates p<0.05. (E and F) Immunohistochemical analysis reveals co-localization of H3K9me3 and DAPI in SAHFs in HFFs transfected with UCA1 overexpression plasmid (F) but not control plasmid (E). (G) Cell count of control and UCA1 overexpressing HFFs 3 days post transfection. Mean ± SD of 3 plates is shown at each time point. * indicates p<0.005 relative to control. (H) Crystal violet assay of cell growth in control and UCA1 overexpressing HFFs transfected with 2 μg of expression or control vector and assayed daily for 3 days post- transfection. * indicates p<0.01 relative to control. (I) Crystal violet assay of HFFs cultured for 3 days after transfecting 0, 1, 2, or 4 μg of control or UCA1 overexpression plasmid. * indicates p<0.01 relative to control. (J) Transcript levels assessed by qPCR; values reflect fold change in UCA1-overexpressing HFFs relative to control after normalization to HPRT levels. * indicates p<0.05 relative to control. (K) qPCR analysis of UCA1 expression in untransduced, presenescent (PS) HFFs and HFFs transduced with constitutively active G12VRAS (RAS). * indicates p<0.05 relative to PS. (L) Efficient knockdown of UCA1 transcripts in RAS HFFs with UCA1 shRNA (quantitated in panel T). (M–P) SA-βgal assays of RAS HFFs transduced with either control or UCA1 shRNA at 3 (M and O) and 5 (N and P) days post transduction. (Q) % quantitation of SA-βgal cells from six replicate experiments as represented in panels M–P. * indicates p<0.001 relative to control. (R) % quantitation of Ki67 + cells from three replicates in control vs UCA1 shRNA transduced RAS HFFs. * indicates p<0.001 relative to control. (S) RT-PCR for UCA1 transcripts shows persistent knockdown of UCA1 in RAS shRNA cells with increasing passage (P0–P2). (T) qPCR analysis of fold changes in transcript levels of cell cycle and senescence genes after UCA1 shRNA knockdown in RAS HFFs. * indicates p<0.05 relative to control. (U) ChIP-PCR analysis of UCA1 region A3 with antibodies listed at top in PS and RAS HFFs. Note gain of activating (H3K4me3, H3K9ace, H4K5ace) and loss of repressive marks (H3K9me3, H3K27me3) at the UCA1 locus after oncogene-induced senescence by RAS. TBX3, CAPERα = human; Tbx3, Caperα = mouse.
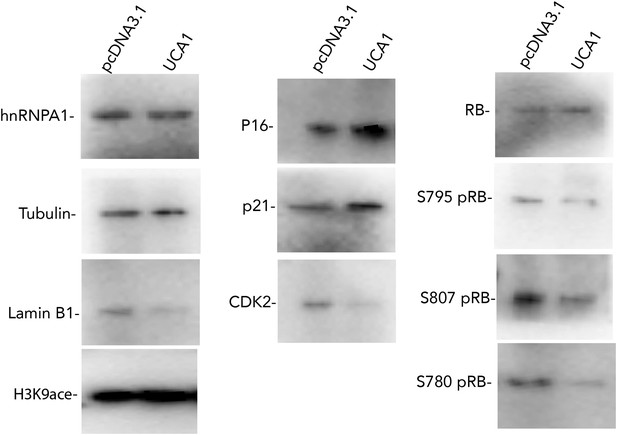
Western blots showing changes in protein levels in response to UCA1 overexpression in HFFs.
pcDNA3.1 are control transfected cells and UCA1 were transfected with UCA1 expression plasmid in pcDNA3.1 (as in Figure 5A). Note increased p16 and p21 levels and hypophosphorylation of RB. TBX3, CAPERα = human; Tbx3, Caperα = mouse.
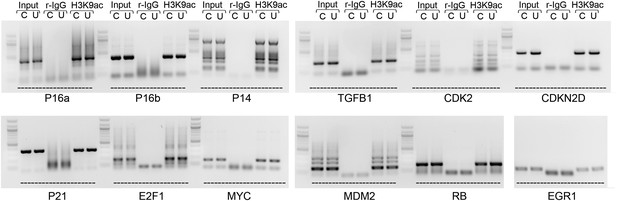
ChIP-PCR assay for H3K9 acetylation of known regulatory elements of prosenescence and cell cycle genes whose expression is dyregulated after UCA1 overexpression.
Input, rabbit IgG negative control ChIP, and H3K9acetylation ChIP in control “C” or UCA1 “U” transfected HFFs for gene regulatory regions as labeled at bottom (primer sequences listed in ChIP primers section of methods). P16 a and b refer to amplicons –(2457–2040) and –(3107–2710), respectively. No changes in H3K9ace levels were detected in response to UCA1 overexpression, suggesting that altered chromatin structure and subsequent increased transcription are not the cause of observed changes in transcript levels detected with UCA1 overexpression and shown in Figure 5J. TBX3, CAPERα = human; Tbx3, Caperα = mouse.
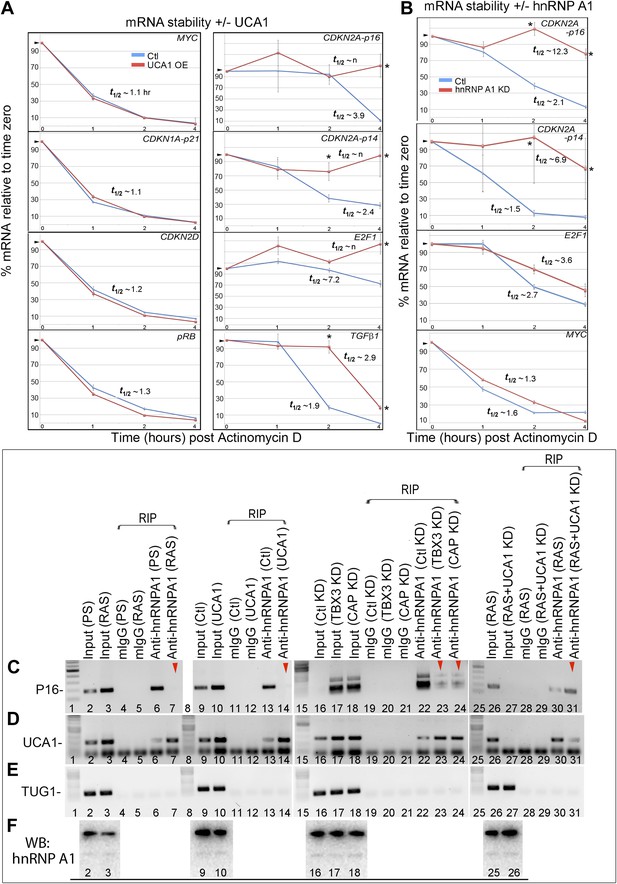
UCA1 stabilizes CDKN2A-p16 mRNA levels during senescence by sequestering hnRNP A1.
(A) Graphs of transcript levels assayed by RT-qPCR in HFFs transfected with control (blue) or UCA1 (red) expression plasmids and treated with Actinomycin (D). Y axis shows % mRNA level relative to time zero and X axis shows time in hours assayed post treatment. The estimated half-lives (t1/2) were obtained using linear regression; the best fit lines, their equations and R values are shown in Figure 6—figure supplement 1. * indicates p<0.04 for p16INK and p<0.01 for all others. (B) Assay of mRNA levels in HFFs transfected with control or hnRNPA1 siRNA and treated with Actinomycin D. Axes and t1/2 calculations are as in panel A. * indicates p<0.05. (C–E) Agarose gels of RT-PCR products assessing levels of CDKN2A-p16 (p16, panel C), UCA1 (panel D), and negative control lncRNA TUG1 (panel E) transcripts in PS and RAS HFFs treated as labeled at top and subjected to RIP with anti-hnRNPA1 antibody. mIgG lanes are negative controls for RIP assays. Gels from left to right show: PS vs RAS; control vs UCA1 overexpression; control vs TBX3 or CAPERα knockdown; RAS vs RAS/UCA1 knockdown. (C) Lane 7 (red arrowhead) shows loss of p16INK /hnRNP A1 interaction in RAS. Lane 14 (red arrowhead) shows loss of p16INK /hnRNP A1 interaction with UCA1 overexpression. Lanes 23 and 24 show loss of p16INK /hnRNP A1 interaction after TBX3 or CAPERα knockdown. Lane 27 shows that UCA1 knockdown decreases the total amount of p16INK mRNA in RAS cells. Lane 31 shows that UCA1 knockdown increases p16INK mRNA/hnRNP A1 binding (red arrowhead) in RAS cells, even though there is less total p16INK (lane 27). (F) Panels show immunoblots to detect hnRNP A1 protein in input samples assayed in panels C–E. Lanes are numbered to correspond with panels above. TBX3, CAPERα = human; Tbx3, Caperα = mouse.
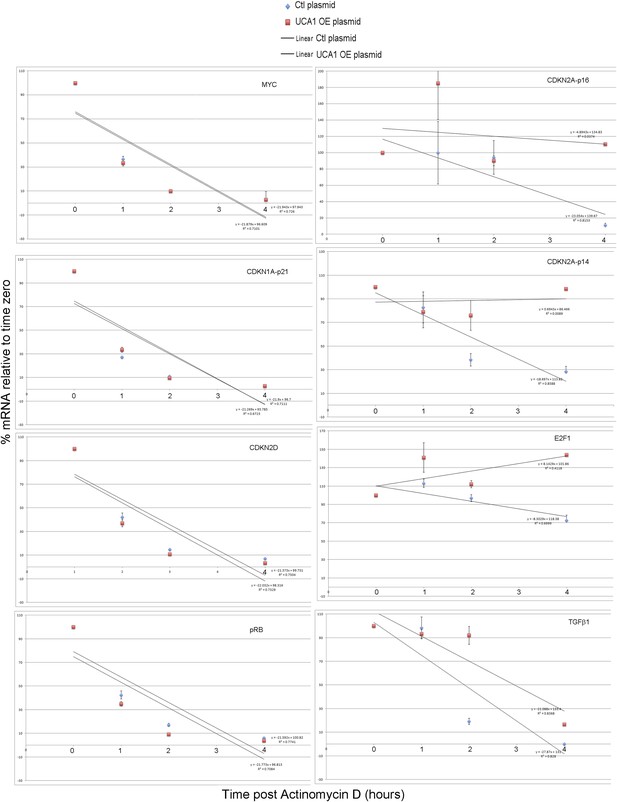
Graphs showing best fit lines, their equations, and R values used to calculate estimated mRNA half-life values shown in Figure 6A.
TBX3, CAPERα = human; Tbx3, Caperα = mouse.
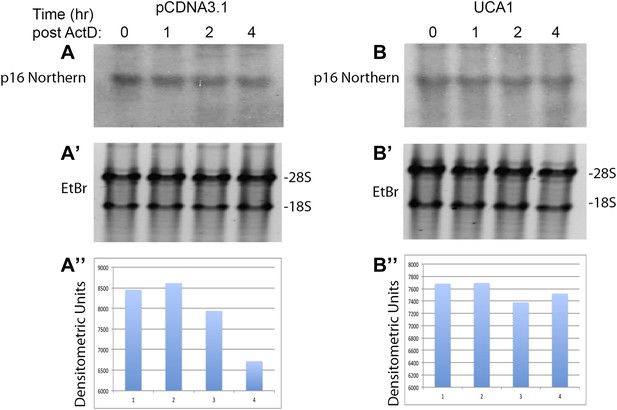
Northern blot assay of p16INK mRNA levels in the absence and presence of UCA1.
(A) Top panel shows Northern blot of HFF cells transfected with control plasmid pcDNA3.1 and treated with Actinomycin D for the times (hr) indicated at top. (A′) The ethidum bromide stained gel prior to transfer is shown for loading control and RNA quality. (A″) The signals obtained by probing for p16INK mRNA in A were subjected to densitometric quantitation. Note decrease in signal at 2 and 4 hr consistent with the decay/t1/2 obtained in Figure 6A. (B) Top panel shows Northern blot of HFF cells transfected with UCA1 expression plasmid and treated with Actinomycin D for the times (hr) indicated at top. (B′) The ethidum bromide stained gel prior to transfer is shown for loading control and RNA quality. (B″) The signals obtained by probing for p16INK mRNA in B were subjected to densitometric quantitation. Note that UCA1 expression results in minimal decrease in signal at 2 and 4 hr, consistent with UCA1-mediated mRNA stabilization observed in Figure 6A. TBX3, CAPERα = human; Tbx3, Caperα = mouse.
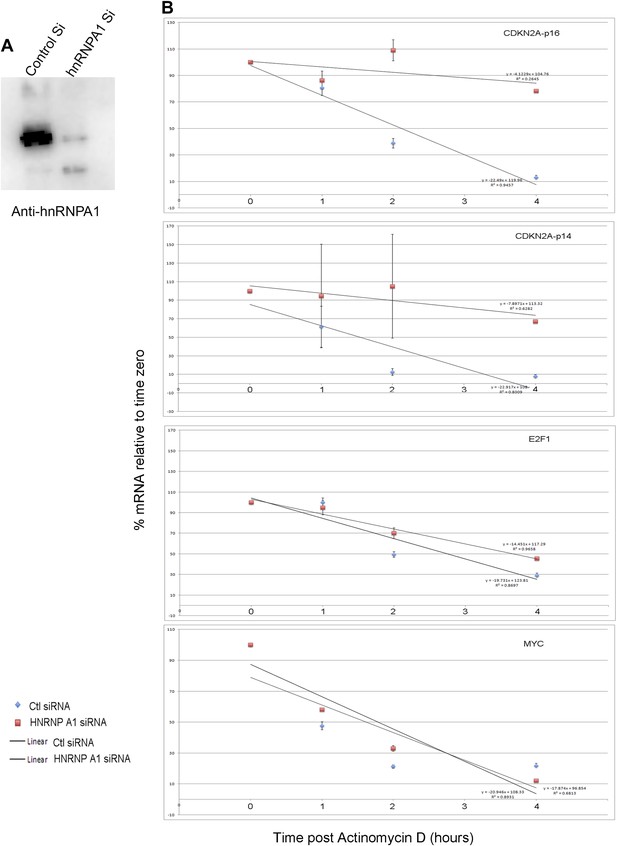
Graphs showing best fit lines, their equations and R values used to calculate estimated half-life values after hnRNP A1 siRNA knockdown shown in Figure 6B.
(A) Western blot assaying hnRNP A1 protein levels in HFFs after transfection of control or anti-hnRNP A1 siRNA. (B) Graphs of best fit lines, equations and R values for half-lives shown in Figure 6B. TBX3, CAPERα = human; Tbx3, Caperα = mouse.
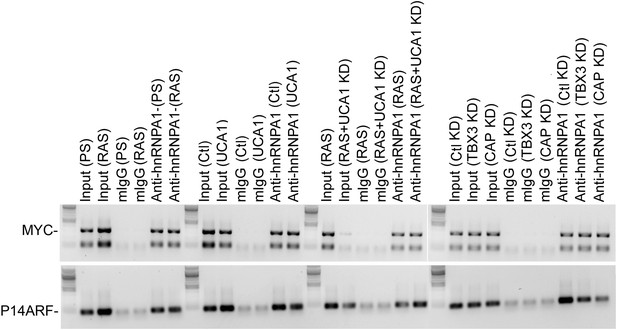
RNA Immunoprecipitation analysis of hnRNP A1 interactions with Myc and p14ARF mRNAs.
RIP-PCR of MYC and CDKN2A-p14 mRNAs shows they are bound by hnRNP A1 but these interactions are unaffected by OIS/RAS, UCA1 overexpression, or knockdown of TBX3 or CAPERα. TBX3, CAPERα = human; Tbx3, Caperα = mouse.
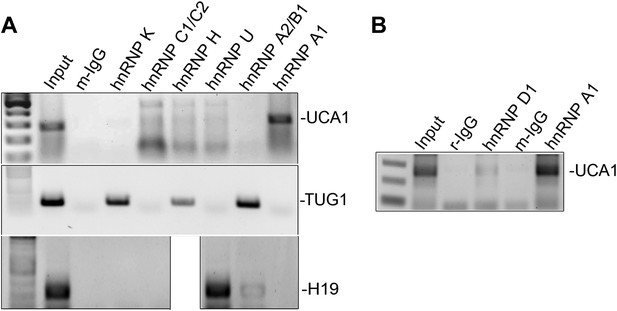
RIP-PCR of HFF lysates using antibodies listed at top.
Only hnRNP A1 (A) and hnRNP D (B) bind UCA1 lncRNA, while TUG1 and H19 lncRNAs are bound by other hnRNPs. TBX3, CAPERα = human; Tbx3, Caperα = mouse.
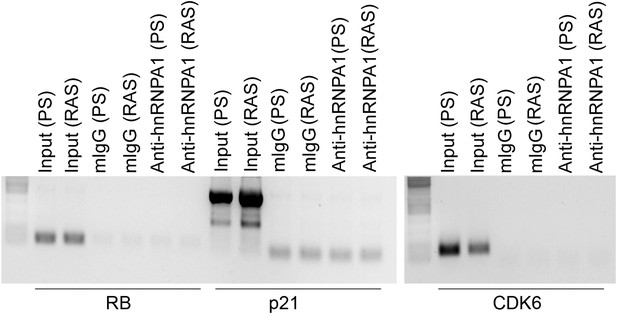
RIP-PCR indicates that RB, p21, and CDK6 mRNAs do not interact with hnRNP A1 in PS or RAS HFFs.
TBX3, CAPERα = human; Tbx3, Caperα = mouse.
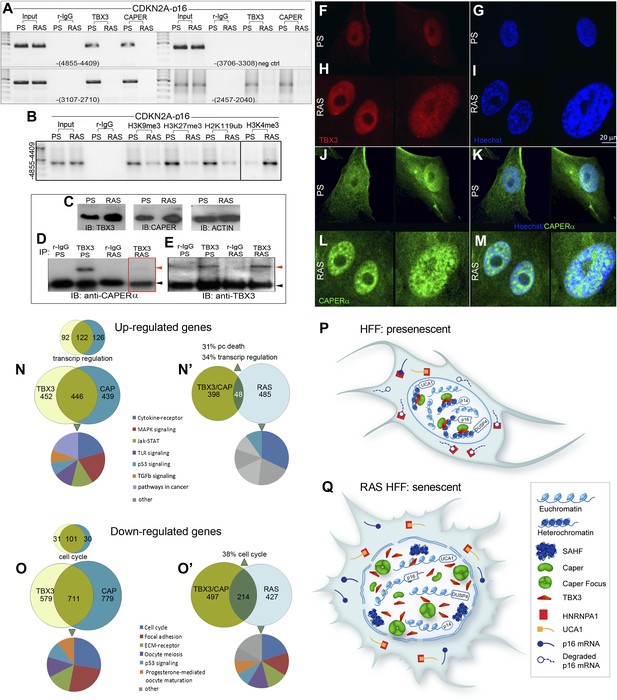
Disruption of the CAPERα/TBX3 repressor by OIS activates CDKN2A-p16 and UCA1 to trigger a senescence transcriptional response.
(A) ChIP-PCR of regions upstream of the CDKN2A-p16 transcriptional start site (position relative to TSS in parentheses) in PS and RAS HFFs; the −-3706–3308 amplicon is a negative control. OIS disrupts binding of p16 regulatory elements (initially identified in Figure 3O) by TBX3 and CAPERα. (B) ChIP-PCR of p16 -4855 element shown in A. Decreased TBX3 and CAPERα binding in RAS correlates with loss of repressive chromatin marks and gain of activating marks. Evaluation of chromatin marks on the other CDKN2A-p16 CAPERα/TBX3- responsive regulatory elements is shown in Figure 7—figure supplement 1A. (C) IBs for TBX3, CAPERα, and actin loading control show increased amount of both proteins in RAS compared to PS HFFs. (D) Anti-TBX3 and anti-CAPERα IBs of IP'd proteins from PS and RAS HFFs. (F–M) Immunocytochemical staining of PS (F, G, J, K) and RAS (H, I, L, M) HFFS for TBX3 (F and H), Hoechst (DNA; G and I), CAPERα (J and L). Panels K and M are merged Hoechst/CAPERα. Scale bar for all panels is sown at lower right of panel I. (N–O′) Functional analyses of genome wide transcriptional profiles of TBX3 KD, CAPERα KD, and control HFFs. All comparisons were statistically significant with p values <<<<0.0001; see Figure 7—source data 3 for hypergeometric test, as implemented in the R statistical language, used to test significance of the number of genes found to be co-regulated between samples. (N) Venn diagrams show highly significant number of CAPERα/TBX3 co-upregulated transcripts (446 total), especially in the GO biologic process (BP) category of transcriptional regulation (122 transcripts) as assayed with DAVID. Pie chart shows KEGG pathway analysis of co-regulated genes. (N′) Venn diagram showing 48 CAPERα/TBX3 co-upregulated transcripts also upregulated by RAS/OIS (Loayza-Puch et al., 2013), especially in BP categories of transcriptional regulation and programmed cell (pc) death. qPCR validation of coregulated genes is in S. Figure 6A. Pie chart shows KEGG pathway analysis of OIS dataset. (O and O′) As in N and N′ but for downregulated genes. Pie chart in O′ shows KEGG pathway analysis of OIS data set; note most pathways are the same as in TBX3/CAPERα. (P and Q) Models of CAPERα/TBX3 repressor and UCA1 function in proliferating (PS) HFFs vs RAS HFFs. In PS cell nuclei, CAPERα/TBX3 represses UCA1, p16, p14, and DUSP4 promoters in heterochromatin which permits ongoing cell proliferation. RAS disrupts the CAPERα/TBX3 complex and CAPERα relocates to dense intranuclear foci. Pro-senescence genes including UCA1 and p16 are converted to euchromatin and their expression/products induce senescence. In the cytoplasm of PS cells, hnRNP A1 binds and destabilizes p16 mRNA, but activation of UCA1 expression in OIS allows UCA1 to sequester hnRNP A1 and stabilize p16 mRNA. TBX3, CAPERα = human; Tbx3, Caperα = mouse.
-
Figure 7—source data 1
Differentially expressed genes after knockdown of CAPERα in HFFs detected by RNA-Seq.
- https://doi.org/10.7554/eLife.02805.027
-
Figure 7—source data 2
Differentially expressed genes after knockdown of TBX3 in HFFs detected by RNA-Seq.
- https://doi.org/10.7554/eLife.02805.028
-
Figure 7—source data 3
Determining the statistical significance of shared differentially expressed genes using the hypergeometric test, as implemented in the R statistical language (phyper).
- https://doi.org/10.7554/eLife.02805.029
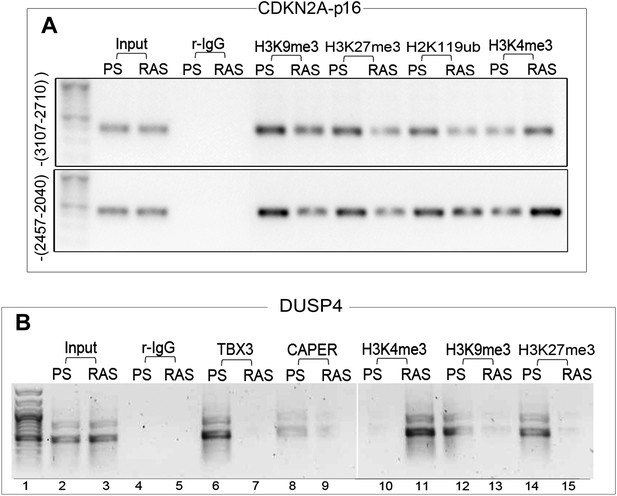
Repression of CDKN2A-p16 and DUSP4 by CAPERα /TBX3 correlates with chromatin architecture and is relieved during oncogene induced senescence.
(A) ChIP-PCR to assess chromatin marks on CDKN2A-p16 regulatory elements in PS and RAS HFFs; antibodies are listed at top. (B) ChIP-PCR of DUSP4 promoter in PS and RAS HFFs; antibodies are listed at top. TBX3 and CAPERα bind the DUSP4 promoter in PS (lanes 6, 8) but not RAS HFFs (lanes 7, 9), and their occupancy correlates with altered chromatin marks consistent with de-repression in OIS/RAS cells (lanes 10–15). TBX3, CAPERα = human; Tbx3, Caperα = mouse.
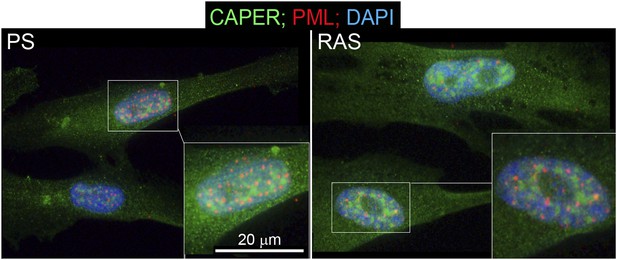
CAPERα relocalization due to oncogene-induced senescence is independent of PML bodies.
Immunocytochemical assay for endogenous CAPERα (green), PML (red), and DNA (DAPI, blue) in PS and RAS HFFs. TBX3, CAPERα = human; Tbx3, Caperα = mouse.
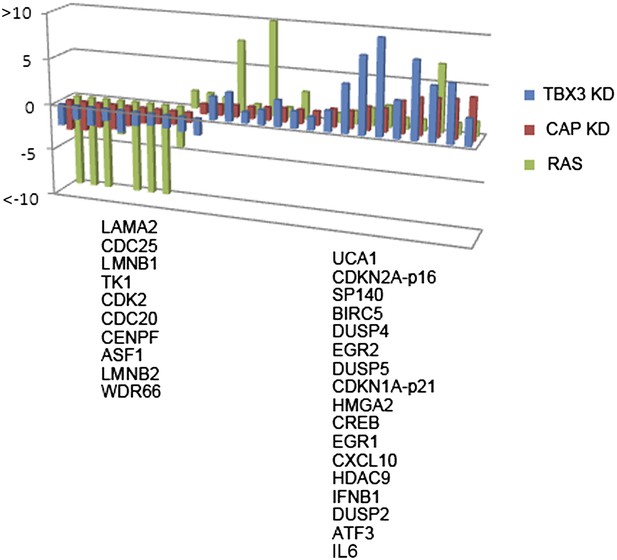
Validation of RNA-Seq identified expression changes induced by CAPERα and TBX3 KD.
qPCR validation of a subset of transcripts with altered expression detected by genome wide RNA-Seq on cDNA prepared from CAPERα (red) and TBX3 (blue) KD, and RAS HFFs (green). Downregulated transcripts are listed at left, upregulated at right. TBX3, CAPERα = human; Tbx3, Caperα = mouse.
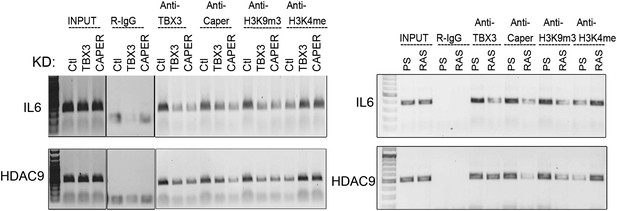
IL6 and HDAC9 are direct targets of CAPERα/TBX3.
ChIP-PCR with antibodies listed at top showing CAPERα/TBX3 directly binds IL6 (and HDAC9) control elements. Effects of TBX3 or CAPER KD on chromatin marks are shown compared with control KD. ChIP-PCR examining CAPERα/TBX3 binding to IL6 and HDAC9 control elements in PS and RAS HFFs; loss of binding correlates with altered chromatin marks. TBX3, CAPERα = human; Tbx3, Caperα = mouse.