Cyclin D activates the Rb tumor suppressor by mono-phosphorylation
Figures
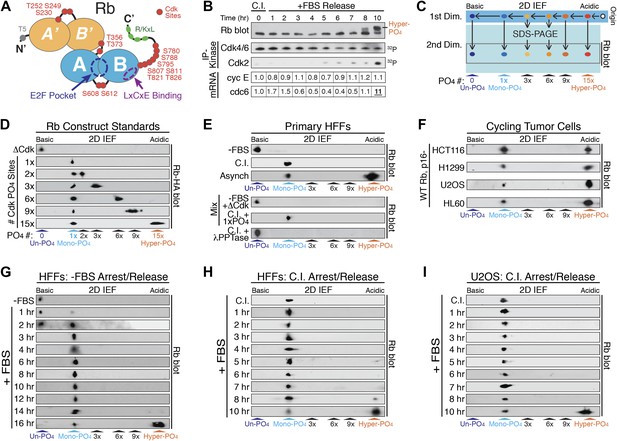
Rb is exclusively mono-phosphorylated in early G1 phase.
(A) Schematic diagram of human Rb Cdk phosphorylation sites, A'/B' and A/B pocket domains. (B) Kinetic analysis of contact inhibited early G1 phase arrested (+FBS) and released primary Human Fibroblasts (HFFs) by 1D SDS-PAGE Rb immunoblot, anti-Cdk4/6, and anti-Cdk2 immunoprecipitation-kinase assay, and qRT-PCR of cyclin E and cdc6 mRNA normalized to B-2-microglobulin levels. (C) Schematic diagram of two-dimensional isoelectric focusing (2D IEF). Immunoprecipitated Rb is loaded at origin on acidic end of IEF strip and separated first by pI. IEF strip is then soaked in SDS, run in second dimension into SDS-PAGE and immunoblotted for Rb. (D) 2D IEF Rb-HA immunoblot of Rb construct standards expressed in cycling cells and containing 0 (ΔCdk), 1x, 2x, 3x, 6x, 9x or 15x Cdk phosphorylation sites. (E) Top panels: 2D IEF Rb immunoblot of primary HFFs serum deprived G0 arrested (−FBS), contact inhibited early G1 phase arrested (+FBS), or asynchronously cycling. Bottom panels: 2D IEF Rb immunoblot of serum deprived G0 arrested (−FBS) HFFs mixed with ΔCdk Rb standard, contact inhibited early G1 phase arrested (+FBS) HFFs mixed with single Cdk site Rb standard and contact inhibited treated with λ phosphatase. (F) 2D IEF Rb immunoblot from cycling human tumor cell lines expressing wild-type Rb and deregulated cyclin D:Cdk4/6 due to p16 deletion, HCT116 colon carcinoma, H1299 lung adenocarcinoma, U2OS osteosarcoma, HL60 promyelocytic leukemia. (G) 2D IEF Rb immunoblot from serum deprived G0 arrested (−FBS) and released (+FBS) primary HFFs from 0 to 16 hr (H) 2D IEF Rb immunoblot from contact inhibited early G1 phase arrested (+FBS) and released HFFs from 0 to 10 hr (I) 2D IEF Rb immunoblot from contact inhibited early G1 phase arrested (+FBS) and released U2OS from 0 to 10 hr.
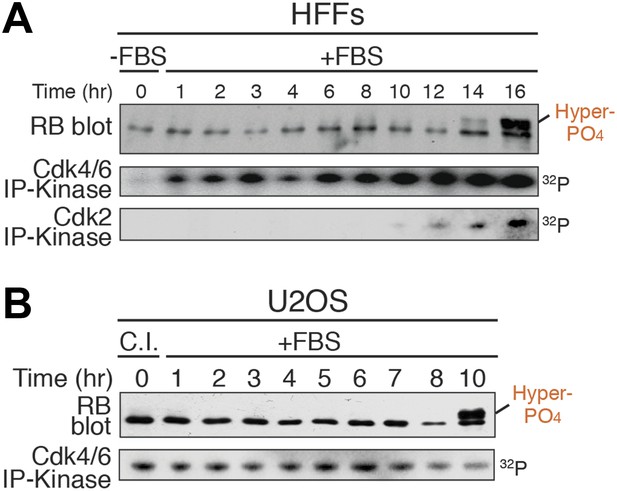
Rb is exclusively mono-phosphorylated in early G1 phase.
(A) Primary HFFs were synchronized in G0 by serum deprivation (−FBS) and released (+FBS), and analyzed by Rb immunoblot, and Cdk4/6 and Cdk2 immunoprecipitation-kinase assays. (B) p16-deleted U2OS tumor cells were arrested in early G1 phase by contact inhibition (+FBS) and released (+FBS), and analyzed by Rb immunoblot, and Cdk4/6 immunoprecipitation-kinase assay.
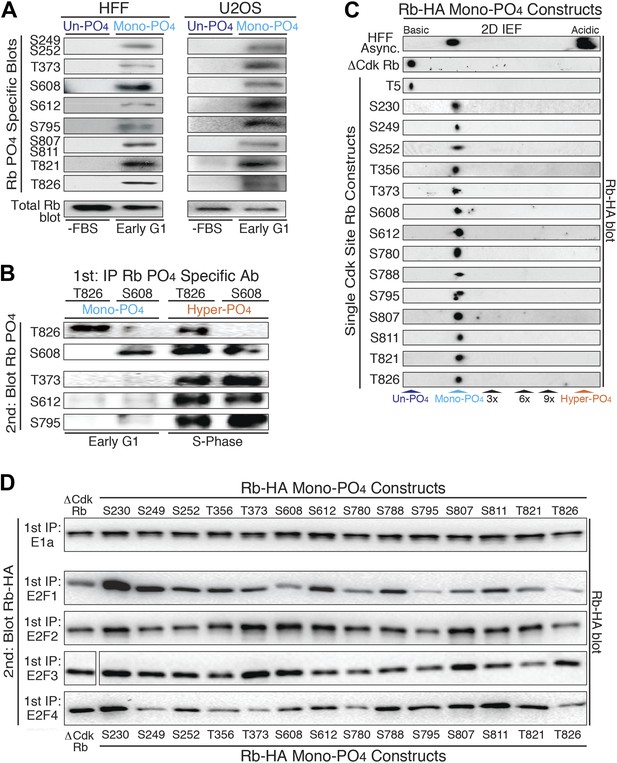
Mono-phosphorylated Rb exists as fourteen individual isoforms.
(A) Phospho-specific Rb immunoblot of un-phosphorylated Rb (G0, −FBS) and mono-phosphorylated Rb (contact inhibited early G1, +FBS) from HFF and U2OS cells. (B) T826 and S608 phospho-specific Rb immunoprecipitation of mono-phosphorylated Rb (contact inhibited early G1, +FBS) and hyper-phosphorylated Rb (S phase) from HFF cells, followed by phospho-specific Rb immunoblot analysis, as indicated. Note the absence of other phosphates at these locations on mono-phosphorylated Rb, but present on hyper-phosphorylated Rb. (C) 2D IEF Rb-HA immunoblot of single Cdk site Rb-HA constructs expressed in cycling cells. Numbering indicates single Cdk site location on Rb. (D) Immunoblot of Rb-HA single Cdk site constructs and control ΔCdk Rb construct from co-immunoprecipitated and co-expressed E1a, E2F1, E2F2, E2F3 or E2F4 (Myc tagged) as indicated. Numbering indicates single Cdk site location on Rb. Note that all 14 avidly bind to at least two E2F family members and there are no completely inactive mono-phosphorylated Rb isoforms.
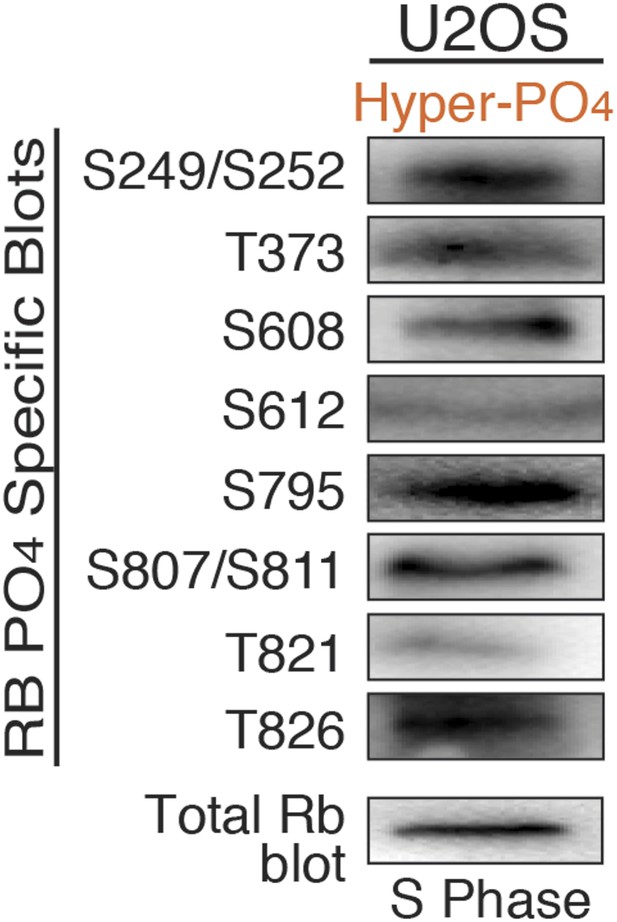
Mono-Phosphorylated Rb exists as fourteen individual isoforms.
U2OS S phase arrested cells (hydroxyurea) were immunoblotted with indicated phospho-specific Rb antibodies or pan Rb antibody.
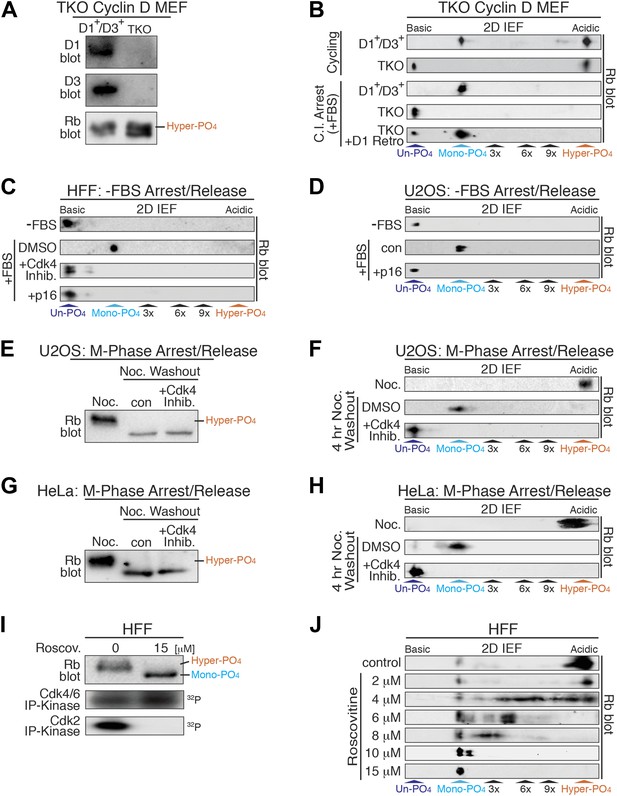
Cyclin D:Cdk/6 is the Rb mono-phosphorylation kinase.
(A) Immunoblot of cyclin D1, D3, Rb in cycling parental D1+/D3+ MEFs and triple knockout (TKO) cyclin D− MEFs. (B) 2D IEF Rb immunoblot from cycling parental D1+/D3+ MEFs and TKO cyclin D− MEFs, and contact inhibited early G1 phase arrested (+FBS) parental D1+/D3+ MEFs and TKO cyclin D− MEFs plus/minus retroviral cyclin D1 expression. (C) 2D IEF Rb immunoblot from serum deprived G0 arrested (−FBS) and released (+FBS) HFFs plus control DMSO, Cdk4 inhibitor (PD0332991) or retroviral p16 expression. (D) 2D IEF Rb immunoblot from serum deprived G0 arrested (−FBS) and released (+FBS) p16-deleted U2OS tumor cells or released (+FBS) plus TET-induced p16 expression. (E, F) 1D SDS-PAGE (E) and 2D IEF (F) Rb immunoblot of G2/M phase nocodazole arrested (Noc.) and released U2OS cells plus DMSO (con) or Cdk4 inhibitor (PD0332991). (G, H) 1D SDS-PAGE (G) and 2D IEF (H) Rb immunoblot of G2/M phase nocodazole arrested (Noc.) and released HeLa cells plus DMSO (con) or Cdk4 inhibitor (PD0332991). (I) Late G1 phase primary HFFs were treated with Cdk2 inhibitor Roscovitine [15 μM] or control (DMSO) and analyzed by 1D SDS-PAGE Rb immunoblot, anti-Cdk4/6 and anti-Cdk2 immunoprecipitation-kinase assays. (J) 2D IEF Rb immunoblot from late G1 phase HFFs treated with dose curve of Cdk2 inhibitor Roscovitine or control (DMSO).
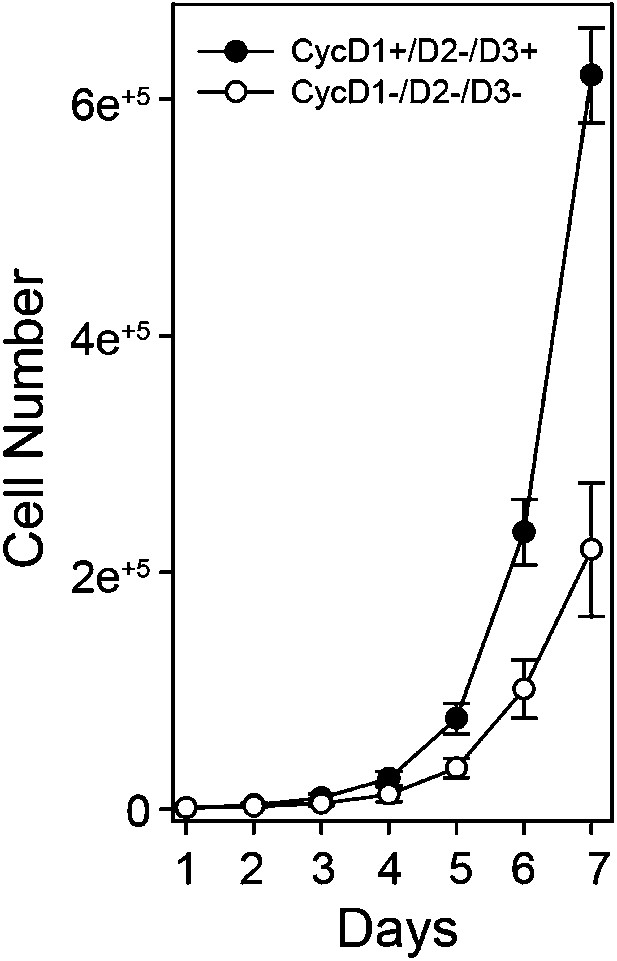
Cyclin D:Cdk/6 is the Rb mono-phosphorylation kinase.
Growth curve analysis of parental D1+/D3+ MEFs and triple knockout (TKO) cyclin D− MEFs plated at low density (10,000) in 10% FBS and counted each day over 7 days.
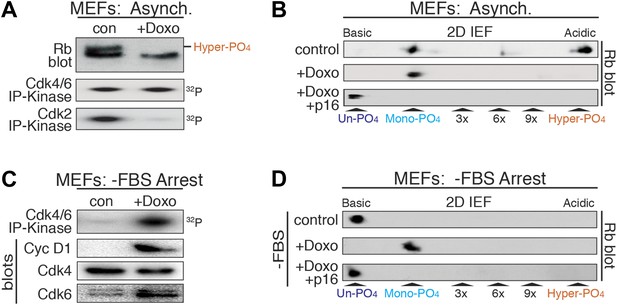
DNA damage induces cyclin D:Cdk4/6 activity and mono-phosphorylated Rb.
(A) Asynchronously cycling MEFs (+FBS) treated with Doxorubicin (+Doxo, 100 ng/ml) were analyzed by 1D SDS-PAGE Rb immunoblot, anti-Cdk4/6 and anti-Cdk2 immunoprecipitation-kinase assays. (B) 2D IEF Rb immunoblot from asynchronously cycling MEFs (+FBS) treated with control or Doxorubicin (+Doxo) or Doxorubicin plus retroviral p16 (+Doxo/+p16) expression. (C) Serum-deprived G0 arrested MEFs (−FBS) treated with Doxorubicin (+Doxo/−FBS) were immunoblot analyzed for cyclin D1, Cdk4, Cdk6, and Cdk4/6 immunoprecipitation-kinase activity. Note activation of cyclin D:Cdk4/6 in absence of serum growth factors. (D) 2D IEF Rb immunoblot from serum-deprived G0 arrested MEFs (−FBS) treated with control or Doxorubicin (+Doxo) or Doxorubicin plus retroviral p16 (+Doxo/+p16) expression.
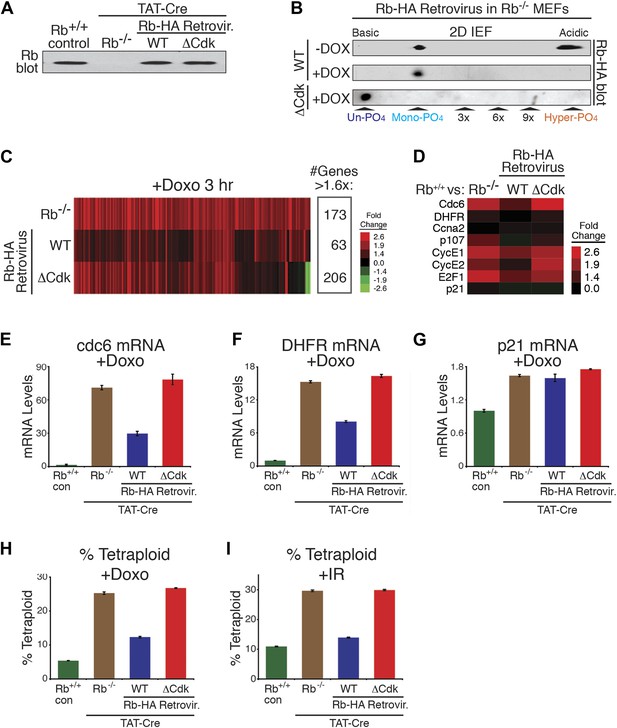
Mono-phosphorylated Rb is active during a DNA damage response.
(A) Rb immunoblot of contact inhibited parental MEFs, conditionally deleted Rb−/− MEFs, and retrovirally-expressed wild type (WT) Rb-HA or non-phosphorylatable ΔCdk Rb-HA in deleted Rb−/− MEFs. (B) 2D IEF Rb immunoblot of cycling and doxorubicin (+Dox) (100 ng/ml) treated WT Rb-HA MEFs, and ΔCdk Rb-HA MEFs in deleted Rb−/− background. (C) Microarray heat map of mRNA levels from early G1 phase Rb−/− MEFs, WT Rb-HA MEFs, and ΔCdk Rb-HA MEFs compared to parental MEFs treated with Doxorubicin (+Doxo) for 3 hr. Only genes increased >1.6-fold are shown. (D) Focused mRNA expression analysis of E2F target genes and p21 control gene from (C). (E–G) qRT-PCR mRNA analysis of endogenous E2F-dependent target genes, cdc6 (E) and DHFR (F), and a non-E2F control gene, p21 (G), from parental MEFs (con), Rb−/− MEFs, WT Rb-HA MEFs, and ΔCdk Rb-HA MEFs in deleted Rb−/− background treated with doxorubicin (+Doxo). Mean values were normalized to β2-microglobulin levels, reported as fold change from parental MEFs (con). Error bars indicate SEM from three independent experiments. (H and I) Quantification of percent tetraploid (>4n DNA) nuclei in parental MEFs (con), Rb−/− MEFs, WT Rb-HA MEFs, and ΔCdk Rb-HA MEFs in deleted Rb−/− background four days after treatment with Doxorubicin (+Doxo) (H) or 2 days post-treatment with ionizing radiation (20 grays) (I). Error bars indicate SEM from three independent experiments.
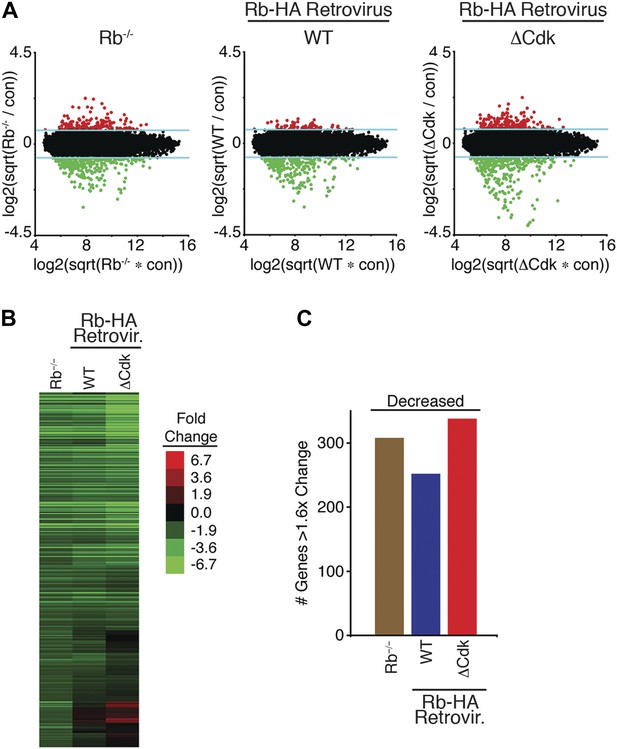
Mono-phosphorylated Rb is active during a DNA damage response.
(A) Total RNA isolated from contact inhibited and released Rb−/− MEFs, WT Rb-HA MEFs and ΔCdk Rb-HA MEFs in deleted Rb−/− background compared to the control parental MEFs treated with 100 ng/ml doxorubicin for 3 hr was used to probe whole-genome microarrays. Blue line indicates 1.6-fold increase/decrease filter. Red and green dots indicate genes with increased or decreased expression >1.6-fold, respectively. (B) Microarray heat map of mRNA levels from early G1 phase Rb−/− MEFs, WT Rb-HA MEFs and ΔCdk Rb-HA MEFs in deleted Rb−/− background compared to parental MEFs treated with Doxorubicin (+Doxo) for 3 hr restricted to 308 genes that were decreased >1.6-fold. (C) Total number of genes altered from (B).
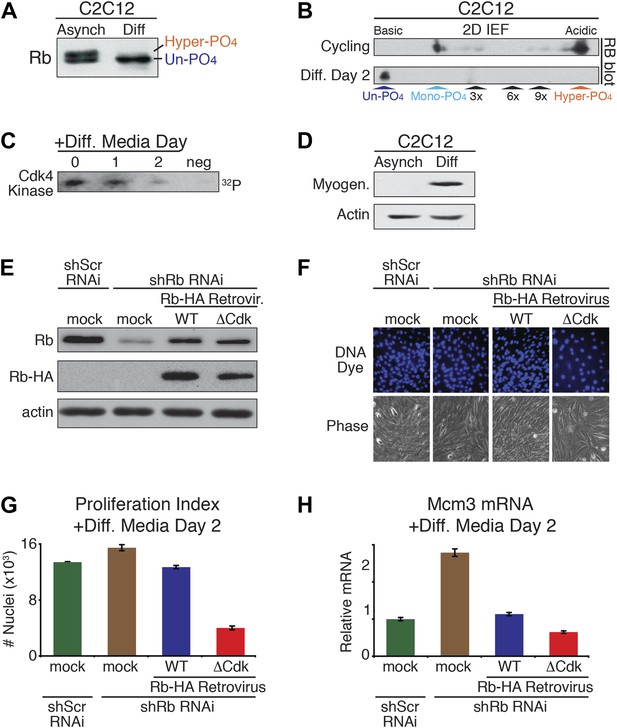
Un-phosphorylated Rb promotes cell cycle exit and differentiation.
(A) Rb immunoblot of cycling C2C12 myoblasts (Asynch), and after 2 days in differentiation medium (Diff). (B) 2D IEF Rb immunoblot from cycling C2C12 myoblasts (Asynch) and after 2 days in differentiation medium (Diff). (C) Cdk4/6 immunoprecipitation-kinase assay of cycling C2C12 myoblasts (0) and at 1 and 2 days post-addition of differentiation medium (Diff). Negative (neg) control, irrelevant antibody. (D) Myogenin immunoblot of cycling C2C12 myoblasts (Asynch), and after 2 days in differentiation medium (Diff). (E) Immunoblot of endogenous Rb, and retroviral expressed wild type (WT) Rb-HA or ΔCdk Rb-HA in C2C12 myoblasts co-infected with short hairpin (sh) RNAi retroviruses targeting endogenous Rb (3' UTR) or scrambled (Scr) control. (F and G) Proliferation analysis (number of nuclei) of C2C12 myoblasts co-infected with short hairpin (sh) RNAi retroviruses targeting endogenous Rb (3' UTR) or scrambled (Scr) control, and wild type (WT) Rb-HA or ΔCdk Rb-HA retroviruses. Cells were stained with Hoechst 33342 DNA dye, visualized by microscopy (F) and quantified by flow cytometry (G) (number of nuclei x 103) at 2 days post-addition of differentiation medium. Error bars indicate SEM. (H) qRT-PCR mRNA analysis of Mcm3 DNA replication factor in C2C12 myoblasts treated as above (G) for 2 days in differentiation media. Mean values were normalized to β2-microglobulin levels and reported as fold change from C2C12 myotubes expressing endogenous Rb (ScrshRNA). Error bars indicate SEM from three independent experiments.
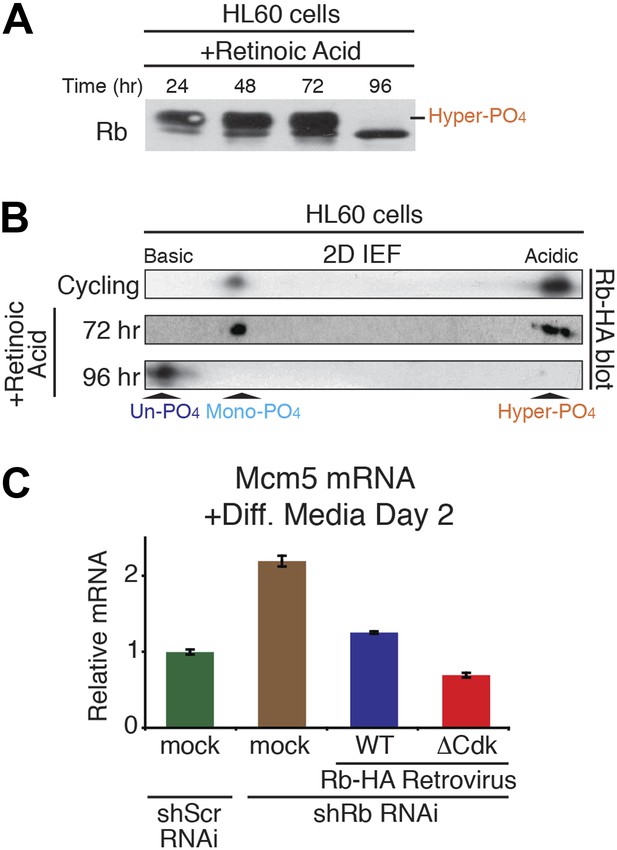
Un-phosphorylated Rb promotes cell cycle exit and differentiation.
(A and B) 1D SDS-PAGE (A) and 2D IEF (B) Rb immunoblot of human HL60 promyelocytic cells treated with retinoic acid to undergo cell cycle exit and differentiation. (C) qRT-PCR mRNA analysis of Mcm5 DNA replication factor in C2C12 myoblasts treated for 2 days in differentiation media. Mean values were normalized to β2-microglobulin levels and reported as fold change from C2C12 myotubes expressing endogenous Rb (ScrshRNA). Error bars indicate SEM from three independent experiments.
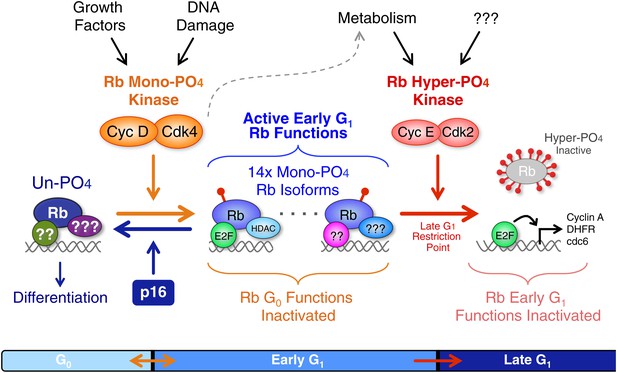
Revised working model of G1 cell cycle progression.
Un-phosphorylated Rb regulates G0 cell cycle exit and differentiation. Growth factor signaling and DNA damage stimulate activation of cyclin D:Cdk4/6 complexes that diversify Rb into 14 mono-phosphorylated isoforms that independently bind specific cellular factors to regulate early G1 phase functions and the DNA damage response. Cyclin D:Cdk4/6 mono-phosphorylation of Rb inactivates un-phosphorylated Rb G0 functions and thereby prevents cells from exiting the cell cycle. Activation of cyclin E:Cdk2 complexes inactivates all 14 mono-phosphorylated Rb isoforms by hyper-phosphorylation (>12x phosphates) at the late G1 Restriction Point. Cyclin A:Cdk2 and cyclin B:Cdk1 maintain Rb in an inactive hyper-phosphorylated state during S, G2 and M phases. As cells complete cytokinesis, hyper-phosphorylated Rb is de-phosphorylated by phosphatases and rapidly mono-phosphorylated by cyclin D:Cdk4/6 complexes. We speculate that an unknown metabolic sensor is upstream of cyclin E:Cdk2 activation.
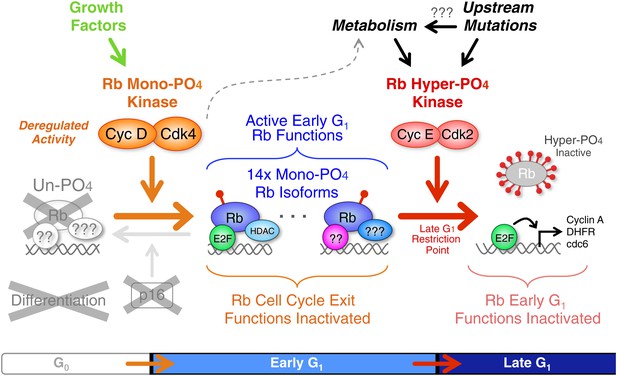
Deregulated cyclin D:Cdk4/6 in cancer mono-phosphorylates Rb to prevent cell cycle exit.
Deregulation of cyclin D:Cdk4/6 activity in cells occurs by a variety of mechanisms, including: p16 deletion, cyclin D1, D2 and D3 amplification or overexpression, and mutation or overexpression of Cdk4 or Cdk6. Cyclin D:Cdk4/6 mono-phosphorylation of Rb simultaneously inactivates Rb's G0 functions and activates Rb's early G1 phase functions thereby driving cells from a low metabolism G0 quiescence into a high metabolism early G1 arrested state that also prevents subsequent cell cycle exit or differentiation. Similar to p53 mutations and Bcl2 overexpression, deregulated cyclin D:Cdk4/6 activity is a well tolerated priming oncogenic mutation that avoids activation of oncogene-induced apoptosis. The net effect is a subtle, but irreversible, oncogenic step forward. We predict that additional oncogenic and metabolic pathways ultimately converge on and activate cyclin E:Cdk2 complexes to inactivate Rb by hyper-phosphorylation at the Restriction Point and drive cells into late G1 phase.
Tables
Mono-phosphorylated Rb is active during a DNA damage response
| Induced | Repressed | ||
| DNA replication | 24% | Localization | 18% |
| Cell cycle | 20% | Post-translational protein modification | 10% |
| Regulation of transcription | 18% | Biosynthetic process | 9% |
| Organelle organization and biogenesis | 12% | Organelle organization and biogenesis | 8% |
| Response to stress | 12% | Intracellular signaling cascade | 8% |
| DNA repair | 8% | Proteolysis | 6% |
| DNA packaging | 7% | Catabolic process | 6% |
| Macromolecular complex assembly | 6% | Cell cycle | 6% |
| Cellular component assembly | 6% | Nervous system development | 6% |
| Chromatin assembly | 6% | Response to external stimulus | 4% |
| Response to wounding | 3% | ||
| Dephosphorylation | 2% |
-
Gene ontology of mRNA differences showing a >1.6-fold increase/decrease level by microarray analysis between parental MEFs and Rb−/− MEFs treated with 100 ng/ml doxorubicin for 3 hr after release from contact inhibition.





