Microtubules provide directional information for core PCP function
Figures
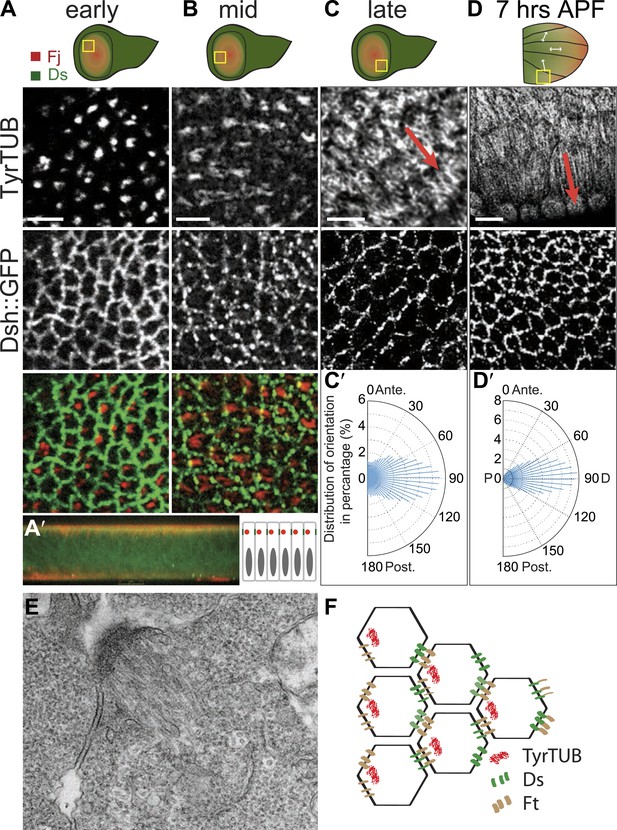
Apical junction-anchored MTs appear in early third-instar wing discs.
(A–C) Tubulin staining in progressively older third-instar wing pouch. Note the asymmetric accumulation of tubulin on the proximal side of the cell in early third-instar wing pouch and growing MTs observed in mid-late third-instar wing pouch. White arrows in the first wing disc cartoon show the proximal distal axis, with distal in the center of the wing pouch, and proximal tissue forming a ring around the future wing blade. (D) MTs in 7 hAPF pupal wing. Scale bar: 5 μm. (A′) A vertical cross-section of the early third-instar wing disc. Apical is at the top. (C′ and D′) Orientations of MTs, in the regions shown, calculated using OrientationJ, aligned with the P-D axis at each stage and location (arrows). (E) TEM micrograph of MTs in early third-instar wing pouch. Note anchored MTs at intercellular junction. (F) Schematic of organization of MTs, Ft and Ds in early third instar wing cells.
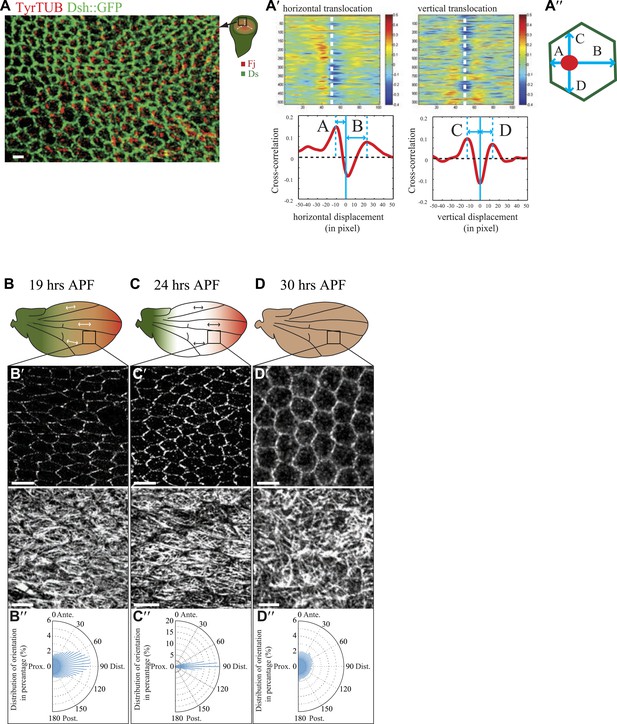
Organization of the apical MT cytoskeleton in early third-instar wing discs.
(A) Tubulin staining in third-instar wing pouch. Note the asymmetric accumulation of tubulin on the proximal side of the cell in early third-instar wing pouch. Scale bar, 2.5 Δm. White arrows show the proximal distal axis in the wing disc, with distal in the center of the wing pouch, and proximal tissue forming a ring around the future wing blade. (A′) Correlation analysis of image (A) for tubulin and Dsh. Note the increasing overlap of signal when the image of tubulin is shifted leftward (proximal) relative to the Dsh image, resulting in a peak at distance A. A rightward shift (distal) initially shows a decrease in correlation of tubulin and Dsh signals, with a peak at larger distance B, indicating that, on average, the tubulin signals are closer to the proximal than to the distal sides of cells. Similar dorsal or ventral translocations show that tubulin signals are, on average, equidistant between dorsal and ventral sides of the cell. (B–D) Orientations of apical MTs correlate with core PCP protein polarization and Fj/Ds gradients throughout wing development. Spatiotemporal correlation of core PCP protein polarity (Dsh::GFP) with MT alignment (TyrTUB) at time points from 19 to 30 hAPF. Rose plots show distributions of MT orientation (P-D axis, corresponding to the Fj and Ds gradients, is plotted as horizontal (90°); Plots, derived from OrientationJ, are composed of 36 bins of 5° each). Note that orientation remains polarized until 30 hAPF, at which time orientations become randomized. Scale bars: 5 μm.
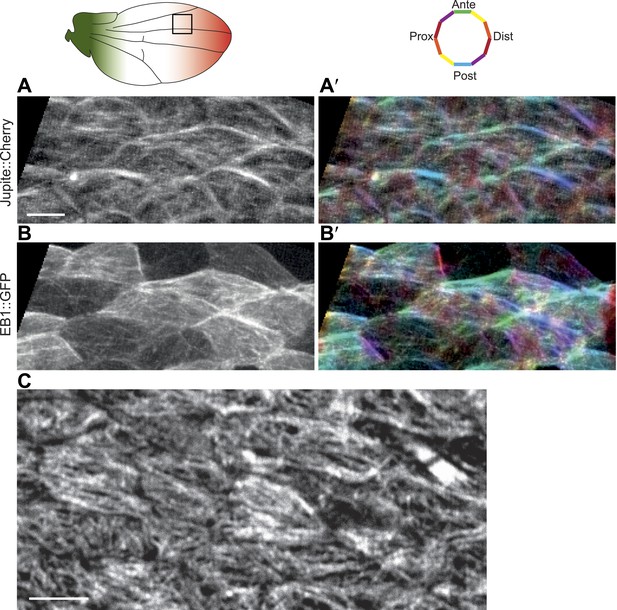
Correlation between EB1 comets and MT orientation in Drosophila wing epithelium.
(A and B) Overlays of 84 frames from time-lapse videos (Video 1) in 24 hAPF pupal wing expressing EB1::GFP and Jupiter::Cherry (Jupiter is a MT associated protein). (A′ and B′) Orientations of MTs and EB1 comets (panels on the left) are color-coded using OrientationJ (18). Scale bar: 5 μm. (C) MT staining with anti-α-tubulin antibody is showing identical pattern as staining with anti-Tyr-tubulin antibody. Scale bar: 5 μm.
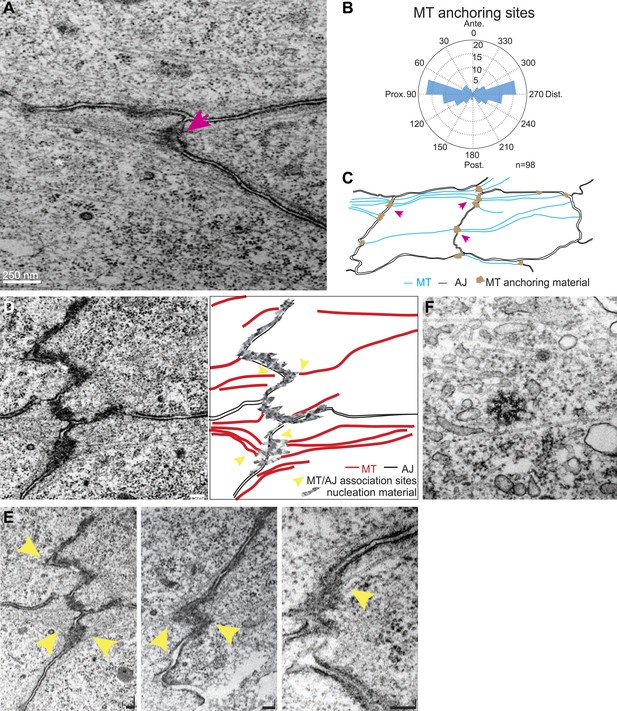
TEM micrographs showing apical MT organization in 24 hAPF pupal wings.
(A) TEM micrograph of MTs in 24 hAPF pupal wing. Note MTs anchored at both sides of intercellular junctions (arrow). (B) Graph depicting the localization of MT anchoring sites in the cell relative to the cell centroid. Plot is composed of 20 bins of 18° each. (C) Organization of the apical MT cytoskeleton traced from a single micrograph showing MTs spanning the cell preferentially in the P-D orientation. (D) TEM revealing planar MTs with anchoring sites on adjacent P-D cell membranes juxtaposed between two neighboring cells, with a sketch of the junctions and MTs. (E) Three additional images. The first is a lighter exposure of the image in D, to better reveal the structure of the junctions. Yellow arrowheads show associations of planar MTs with apical intercellular junctions. Scale bars, 100 nm. (F) TEM micrograpf showing no association of centrosome with MTs.
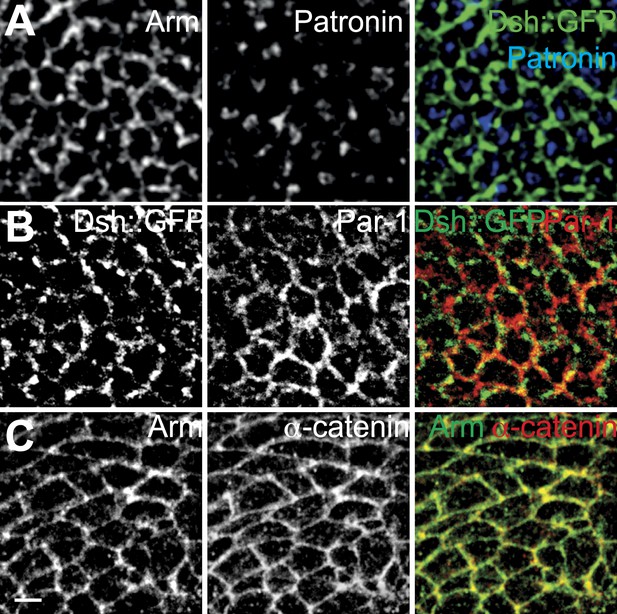
Distributions of potential MT interacting proteins.
(A) Patronin, (B) Par-1 and (C) α-catenin. Patronin is not membrane associated, and Par-1 and α-catenin are cortical but symmetric.
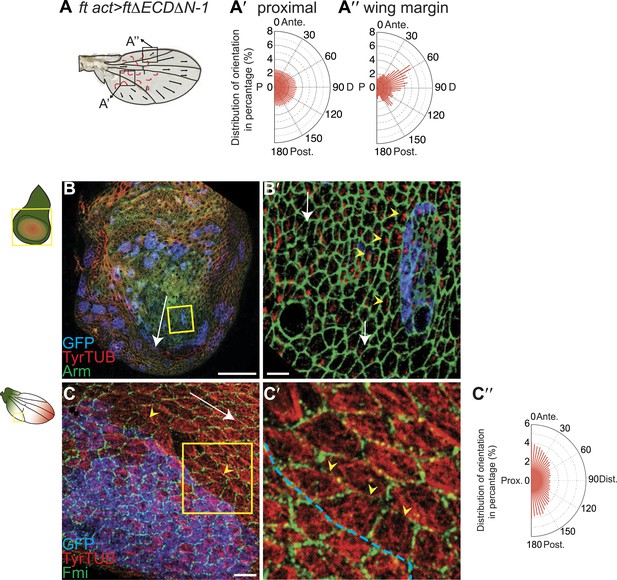
MTs are misaligned in ft and ds mutant wing.
(A–A″) Analysis of hair polarity and MT orientation in ftl(2) fd / ftGRV ActP-Gal4 UAS-FtΔECDΔN-1 mutant flies. (A′) MTs in the proximal and central part of the wing, where hair polarity is disturbed, are randomized. (A″) MTs in the distal/peripheral part of the wing are oriented with the hairs pointing toward the margin (all plots are composed of 36 bins of 5° each). (B and C) Ft-dependent instructive re-organization of the MT cytoskeleton. Wildtype cells around ft clones only see Ds in their mutant neighbors, and therefore preferentially accumulate Ft at their junction with the mutant cell. The orientation of MT dots is changed in wildtype cells around ft clones in third instar (B and B′) and 24 hr pupal wings (C and C′). White arrows show P-D axis and yellow arrowheads show MT spots initiating toward the clone rather than proximally (B′) or MTs organized perpendicular to the clone boundary (C′) rather than proximally-distally (right panels are high magnification images of left panel boxes). (C″) Orientations of MTs in the wildtype cells bordering ft clones calculated using OrientationJ. Scale bars: 50 μm (B and C) and 5 μm (B′–C′). The P-D axis is defined as a radius from the center of the wing disc to the circumference, as described in Figure 1.
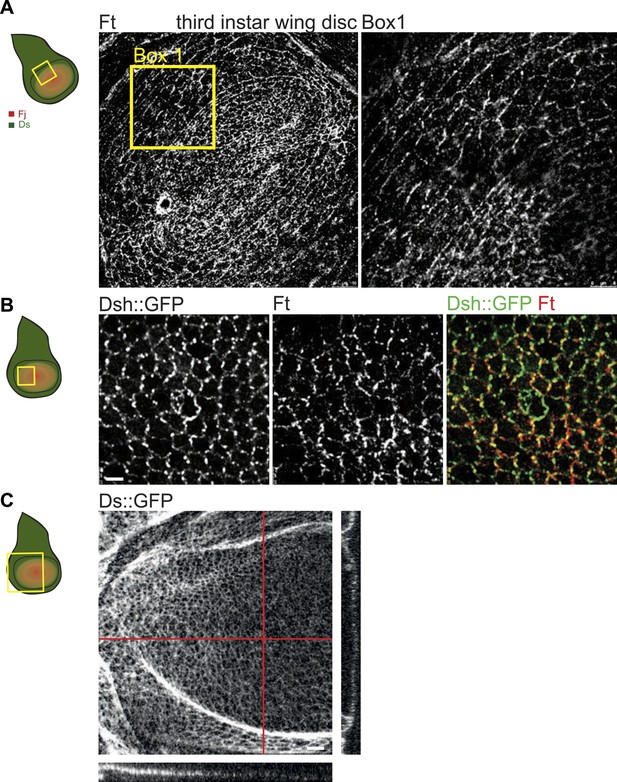
Orientations of Ft, Ds and core proteins correlate in wing disc.
(A) Ft staining in third-instar wing pouch. Right panel is a high magnification image of the box in the left panel. (B) Dsh::GFP and Ft localization in a mid third-instar wing disc. Note asymmetric localization of Dsh and Ft that overlap on the P/D boundaries. Scale bar: 2.5 μm. (C) Image of Ds::GFP in third-instar wing disc. On the right and below are vertical section of wing pounch (along red lines) showing the gradient of Ds::GFP along the P/D axis. Scale bar: 2.5 μm.
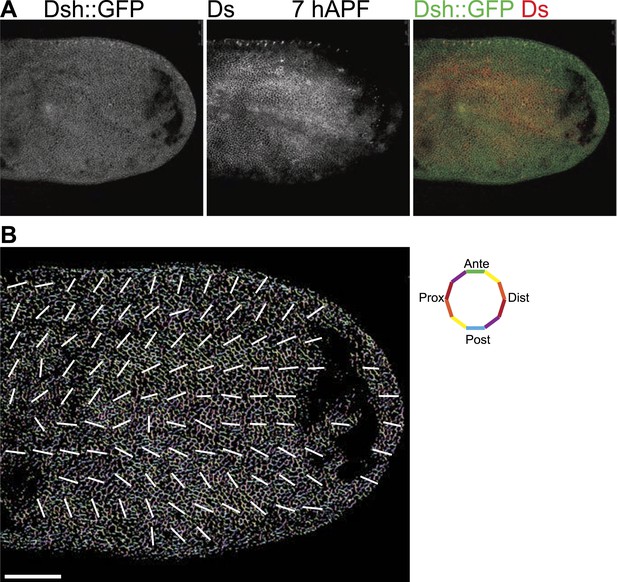
Orientations of Ds gradient and core proteins correlate in early pupal wing (7 hAPF).
(A) Images of 7 hAPF pupal wings showing Ds staining and Dsh::GFP. Note the Ds staining projecting into the central part of the wing. High Ds expression retracts fully into the hinge around or shortly after 24 hAPF (Figure 3—figure supplement 3A,B). (B) A low magnification image of Dsh::GFP at 7 hAPF with overall polarity derived from OrientationJ showing the radial (P/D axis from wing hinge toward wing margin) pattern. Scale bar: 50 μm.
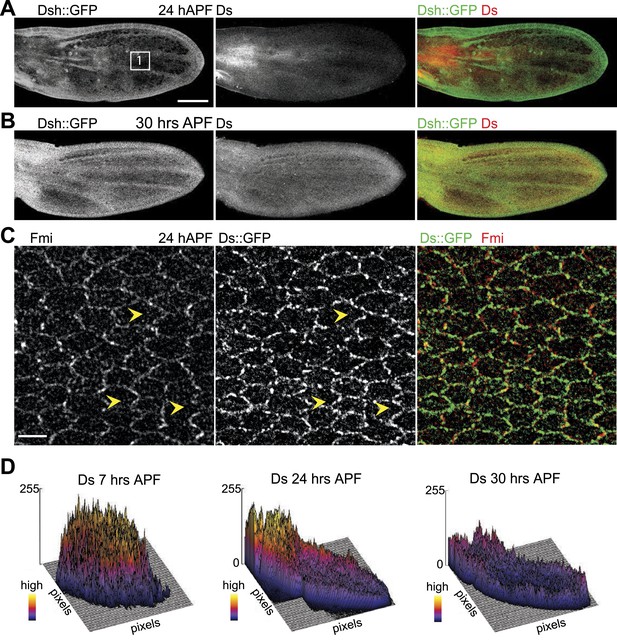
Orientations of Ds gradient and core proteins correlate in late pupal wing (24 hAPF).
(A and B) Images of 24 hAPF and 30 hAPF pupal wings showing Ds staining and Dsh::GFP. (C) Note the strong overlap of Fmi and Ds subcellular localization in the central part of the wing at 24 hAPF. (D) Surface plots of Ds staining at different pupal ages showing the shape of the overall gradient. Scale bars: 100 μm for top image and 5 μm for bottom.
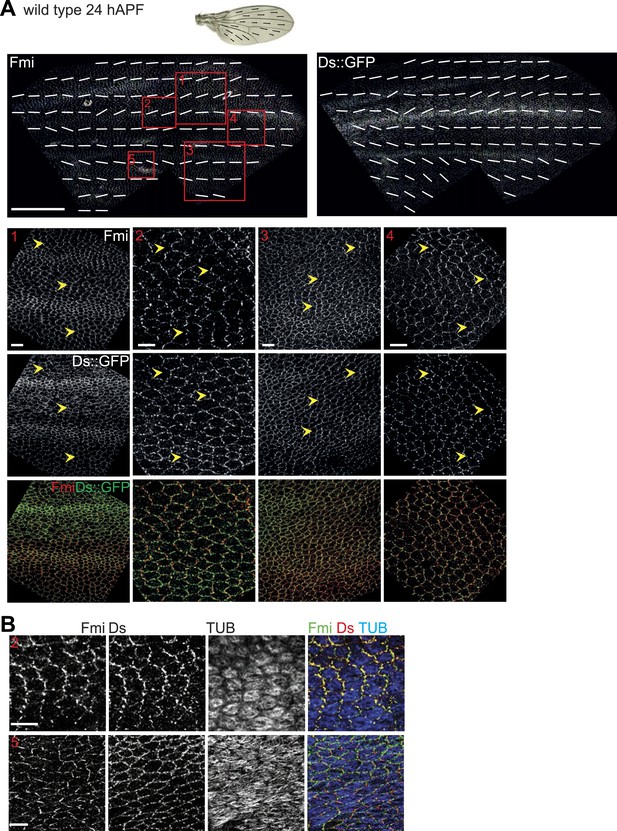
Orientations of Ds gradient, MTs and core proteins correlate in late pupal wing.
The mostly parallel pattern of Ds localization at or shortly after 24 hAPF generally correlates with both the MTs and core PCP polarity, although subtle divergence in the posterior suggests the possibility of other inputs that organize the MTs. (A) A low magnification image of Fmi staining and Ds::GFP at 26 hAPF with overall polarity derived from OrientationJ. Below are high magnification images from areas marked with red boxes (1–4) in top panels. Scale bars: 100 μm for top image and 7.5 μm for bottom. (B) Note the strong overlap of Fmi and Ds subcellular localization and the correspondence with MT orientation at 24 hAPF. The boxes in B correspond to the equivalent region in panel A—Box 2 and Box 5. Scale bar: 7.5 μm.
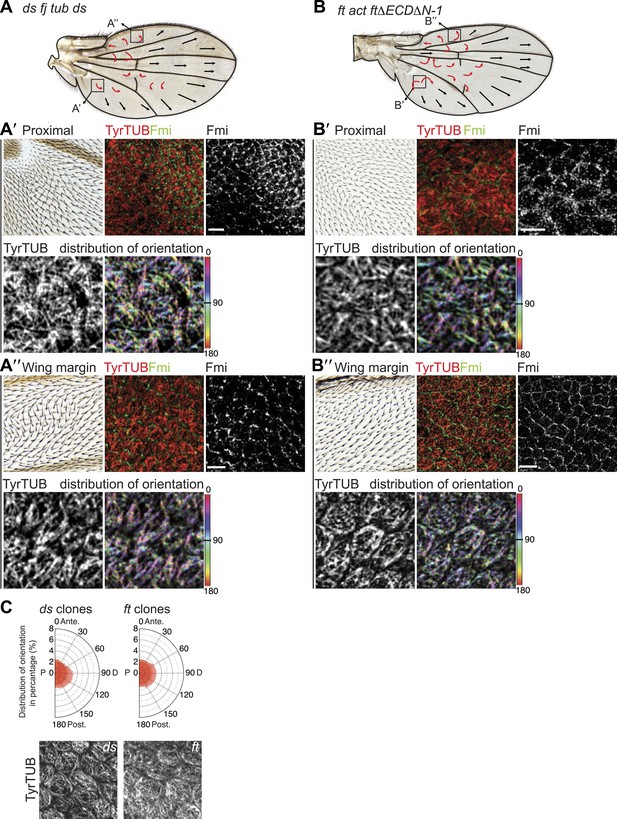
MTs in ft mutant wings are randomized.
(A) Hair polarity in ftl(2) fd/ftGRV ActP-Gal4 UAS-FtΔECDΔN-1 adult wing and higher magnification images from the boxes showing hair polarity, Fmi staining pattern and tubulin orientation in 28 hAPF pupal wing. Note that MT orientation is randomized in the central proximal part of the wing (A′), but is more ordered and oriented toward the margin closer to the periphery (A″). (C) MTs in ds and ft clones are randomized. (B) Hair polarity in ds38k fjN7/dsUA071 fjd1; UAS-Ds/TubP-Gal4 adult wing and higher magnification images from the boxes showing hair polarity, Fmi staining pattern and tubulin orientation in 28 hAPF pupal wing. Note that MT orientation is randomized in the central proximal part of the wing (B′), but is more ordered and oriented toward the margin closer to the periphery (B″). (C) MTs in ds and ft clones are randomized.
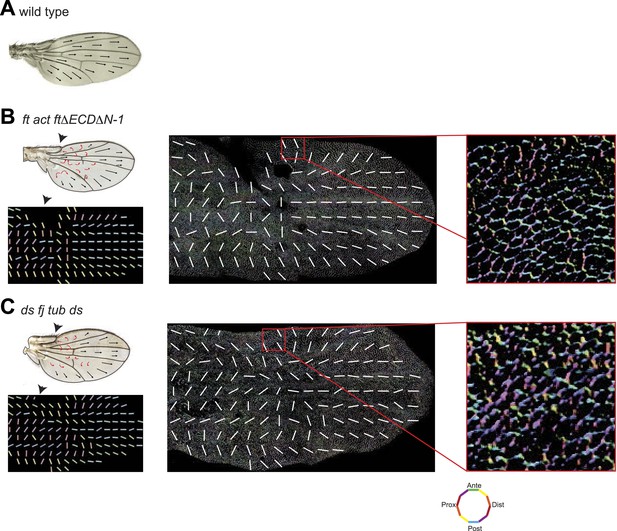

Wing hair polarity in ft mutant wings reflects core PCP orientation in these wings.
(A) Hair polarity in wildtype wing. (B) Hair polarity in ftl(2) fd/ftGRV ActP-Gal4 UAS-FtΔECDΔN-1 adult wing (left top) and core PCP polarity in 28 hAPF pupal wing (middle) in same genotype. Right panel is high magnification image of middle panel box. (C) Hair polarity in ds38k fjN7/dsUA071 fjd1; UAS-Ds/TubP-Gal4 adult wing (left top) and core PCP polarity in 28 hAPF pupal wing (middle) in same genotype. Right panel is high magnification image of middle panel box. Orientations of cell boundaries are color-coded using OrientationJ (18).

Overexpression of Wnt4 affects MT polarity.
(A) Wing hair and MT polarity in wildtype wing. (B) Wing hair and MT polarity in en-Gal4 UAS-Wnt4 are disturbed in wildtype cells outside the expressing area of Wnt4 between L3 and L4 veins. Note that wing hairs are pointing toward posterior region where Wnt4 is expressed and that MTs are oriented along the Wnt4 gradient generated outside the expression area of Wnt4. (C) Orientations of MTs in the wildtype cells between L3 and L4 veins calculated using OrientationJ.
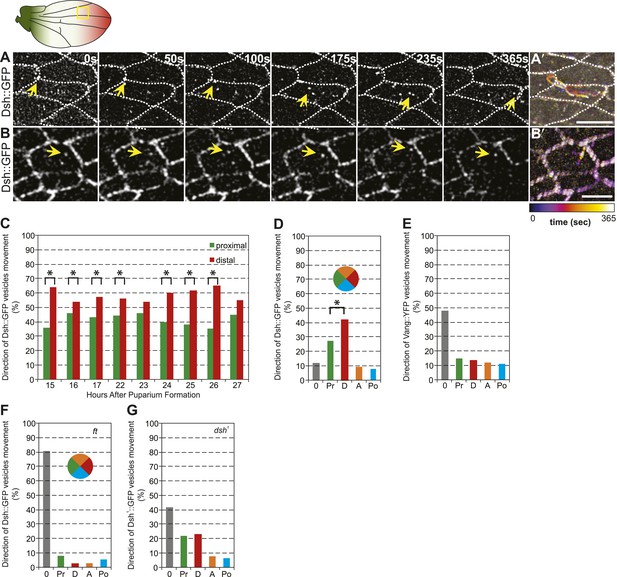
Dsh::GFP vesicles move preferentially toward the distal side of the cell.
(A and B) Timelapse images of Dsh::GFP vesicles in 24 hAPF pupal wing. The majority of Dsh::GFP vesicles displayed directed, fast transcytotic movement with a distal bias (A). A minority of vesicles moved slowly along irregular paths (B) (derived from Video 2 and Video 3). (A′ and B′) Overlays of 73 frames from timelapse videos in A and B. Proximal is to the left and anterior is at the top. (C) The ratios of transcytosing Dsh::GFP vesicles at different pupal ages (n ≥ 1192). (D) Ratio of Dsh::GFP vesicles moving toward proximal, distal, anterior or posterior, or not moving (n = 50). (E) Similar plot of Vang::YFP vesicles showing absence of directed trafficking (n = 81). (F) Dsh::GFP vesicle movement inside ft clones showing little net movement and no significant bias (n = 42). (G) Dsh1::GFP vesicles in dsh1 wings showing a bias to P-D vesicle movement among vesicles showing net movement (n = 120). Numbers were too small to test significance of a possible difference between P and D. Scale bars, 5 μm. Significant differences between proximal and distal at p ≤ 0.05 using the binomial test are marked with *.
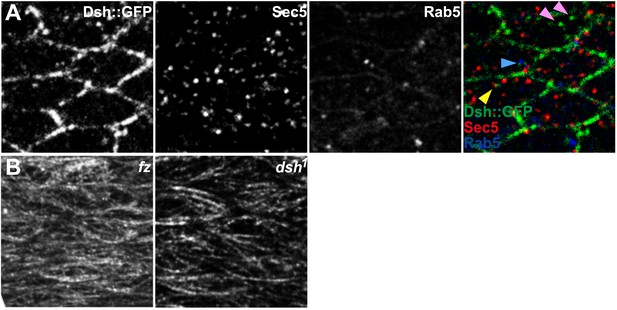
Vesicle identification and tubulin staining.
(A) 24 hAPF Dsh::GFP wing co-labeled for Sec5 and Rab5. Many Dsh::GFP vesicles do not label with Sec5 or Rab5 (pink arrowheads), while some stain for Sec5 (yellow arrowhead) or Rab5 (blue arrowhead). (B) Tubulin staining in fz and dsh1 mutant flies. Compare to Figure 1—figure supplement 1C.
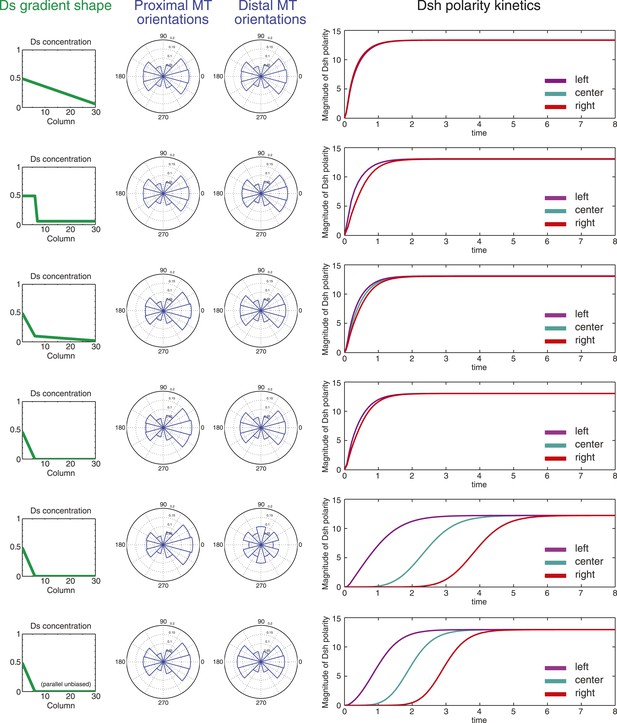
Simulations of polarization kinetics with different shapes of Ds gradients or imposed MT structures, using a mathematical model incorporating a simple representation of the Ft/Ds/Fj system to polarize MTs.
Input Ds gradients are shown on the left. Resulting (or imposed, on the right side of the last example) MT organization on the proximal or distal portion of the gradient are plotted (center). Kinetics of polarization of cells in column 8 (proximal = left), column 15 (center) or column 23 (distal = right; see Figure 5—figure supplement 1).
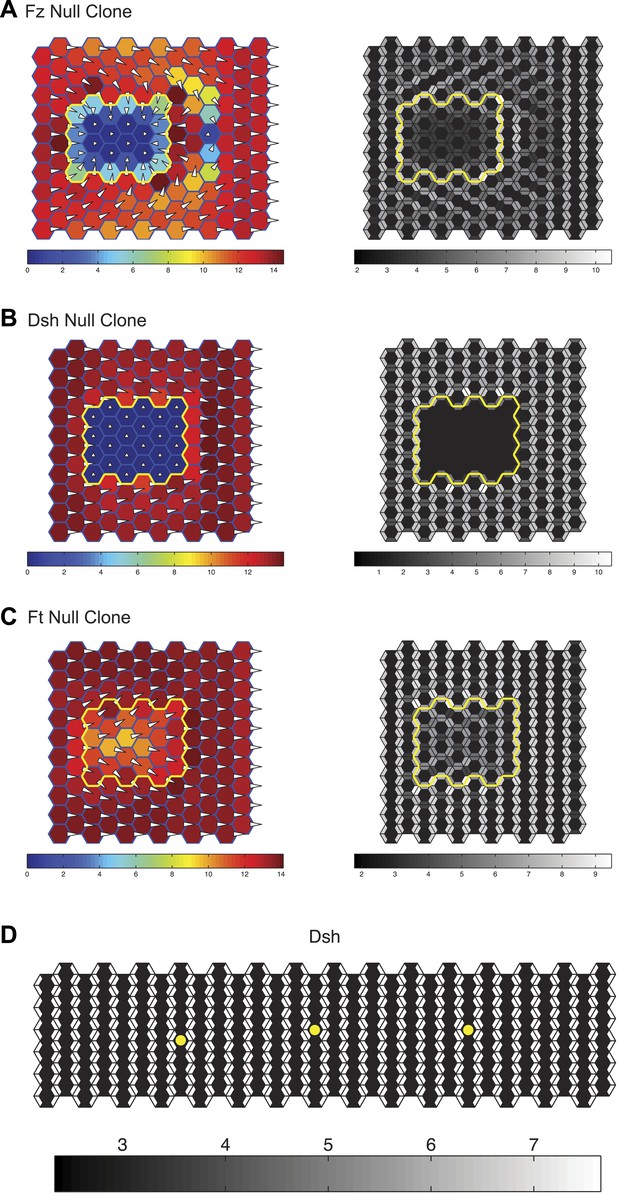
Simulation results.
Simulations of clones of fz (A), dsh (B) and ft (C) null clones validating that the mathematical model captures the domineering non-autonomy phenotypes as well as the ability of the core system to propagate through ft mutant tissue. The color scale represents the magnitude of the vector sum of Dsh in each cell. The length of the hair and its distance from the center of the cell is also plotted proportionally to the vector sum of Dsh in each cell. (D) A 6 × 30 cell grid used to simulate polarization kinetics. Cells with kinetics plotted in Figure 5 are marked with asterisks. Greyscales represent the combined quantity of Dsh on either side of each shared cell boundary, as would be seen with light microscopy.
Videos
Live in vivo imaging of EB1::GFP and Cherry::Jupiter.
Live in vivo imaging of EB1::GFP comets and Jupiter::Cherry labeled MTs in 24 hAPF pupal wing. Refers to Figure 1—figure supplement 2.
Live in vivo imaging of Dsh::GFP.
Live in vivo imaging of Dsh::GFP vesicles in 24 hAPF pupal wing. Refers to Figure 4A.
Live in vivo imaging of Dsh::GFP.
Live in vivo imaging of Dsh::GFP vesicles in 24 hAPF pupal wing. Refers to Figure 4B.
Live in vivo imaging of Dsh1::GFP in dsh1 mutant flies.
Live in vivo imaging of Dsh1::GFP vesicles in 24 hAPF dsh1 pupal wing. Refers to Figure 4G.
Additional files
-
Supplementary file 1
Description of modelling
- https://doi.org/10.7554/eLife.02893.024





