Bidirectional helical motility of cytoplasmic dynein around microtubules
Figures
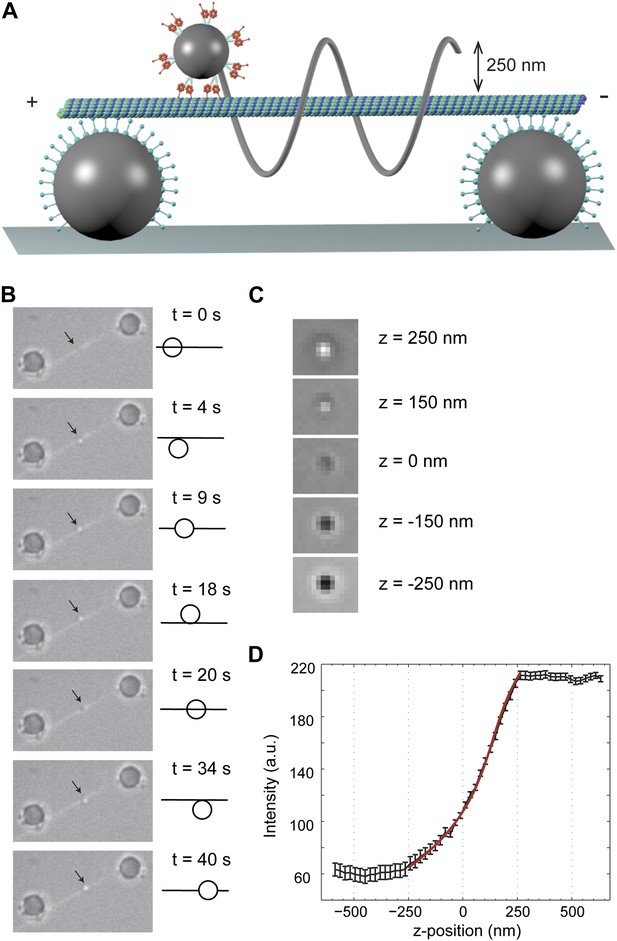
3D tracking of dynein-driven transport along MT.
(A) Schematic representation of the experimental geometry (not to scale). The MT suspended bridge is formed by attaching an MT (green) to the surface-immobilized beads (gray) that are coated with a chimeric protein containing the dynein MTBD. A 500 nm diameter cargo bead is coated with multiple dynein motors and trapped by a focused laser beam (not shown) for placement of the bead on the bridge. The bead center is expected to be separated by ∼250 nm from the MT. (B) Movement of a GST-Dyn331kD-coated bead along an MT bridge (left). The fluorescent image of the MTs has been superimposed onto the bright-field images. The bead moves in a left-handed helical manner along the MT. The schematic on the right represents the side view for the orientation of the bead relative to the MT. (C) Bright-field image of the cargo-bead in different z positions shows that the z position of the bead relative to MT can be determined by its brightness. Images are taken at z = −250 nm, −150 nm, 0 nm, +150 nm, +250 nm. (D) The averaged intensity of a 500 nm diameter bead under a brightfield illumination at variable z positions. The averaged intensity from 20 beads increases as the bead is moved from −250 nm to +250 nm in the z direction relative to the image plane. The red curve represents a fit to a third order polynomial (R2 = 0.998). The z position of a motor-coated bead was calculated from the calibration curve. Error bars represent SEM.
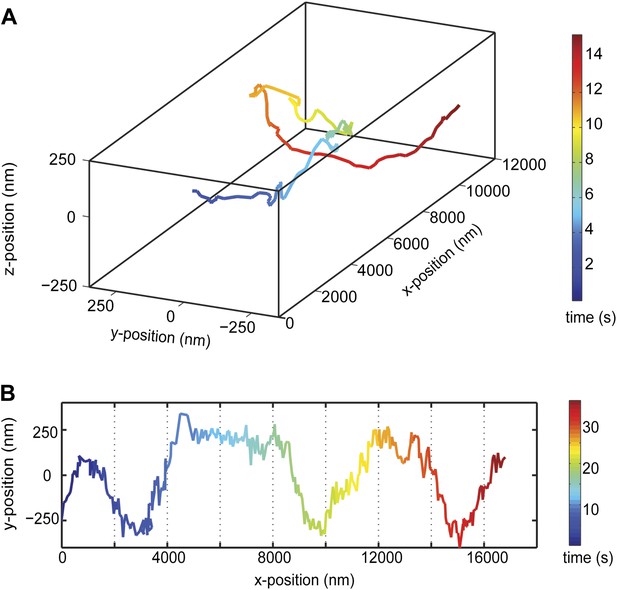
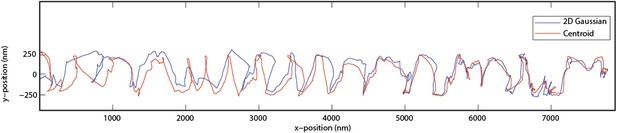
Kinesin-1 follows a single protofilament on a MT track.
(A) Example trace of a kinesin-1 coated bead, traveling along GMP-CPP MT filaments with a left handed spiraling motion. (B) Representative two dimensional trace. The average pitch of helical motion is 6500 ± 400 nm (mean ±SEM, N = 6).
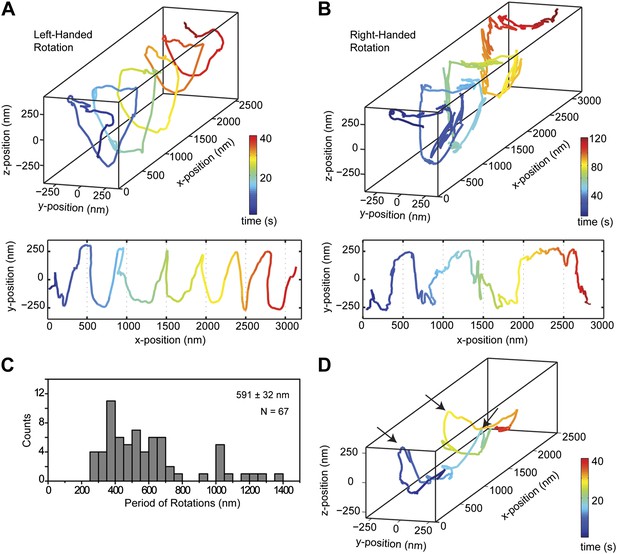
Dynein moves in both left- and right-handed helical paths along MT bridges.
(A and B) (top) Representative three-dimensional trace of a cargo bead-driven by GST-Dyn331kD motors shows left- (A) and right-handed (B) helical motion. (bottom) Two-dimensional projections of the traces shown at top. (C) Histogram of observed pitches per complete rotation. The average pitch is 591 ± 32 nm (mean ± SEM). The average pitch of the left-handed movement (546 ± 42 nm, SEM, N = 32) was shorter (t-test, p = 0.01) than that of the right-handed movement (749 ± 81 nm, SEM, N = 10). (D) Change in handedness of rotation during the transport of a cargo bead. An example trace shows that a cargo bead initially moves along GMP-CPP MTs with a right-handed helical motion. At around t = 10 s, the bead reverses its helical motion for half of the period. At t = 20 s, the bead switches back to right-handed rotation and takes another half turn around the MT. Finally, at t = 25 s, the bead resumes left-handed helical motion until it disassociates from the MT. Arrows show the transitions from one type of helical motion to the other.

Movement of a GST-Dyn331kD coated bead along an MT bridge.
(A) First 20 s of the 3D trace shown in Figure 2A is plotted in x, y, and z directions as a function of time. (B) The brightfield image of the bead shows the changes in the position and intensity of the bead as a function of time.
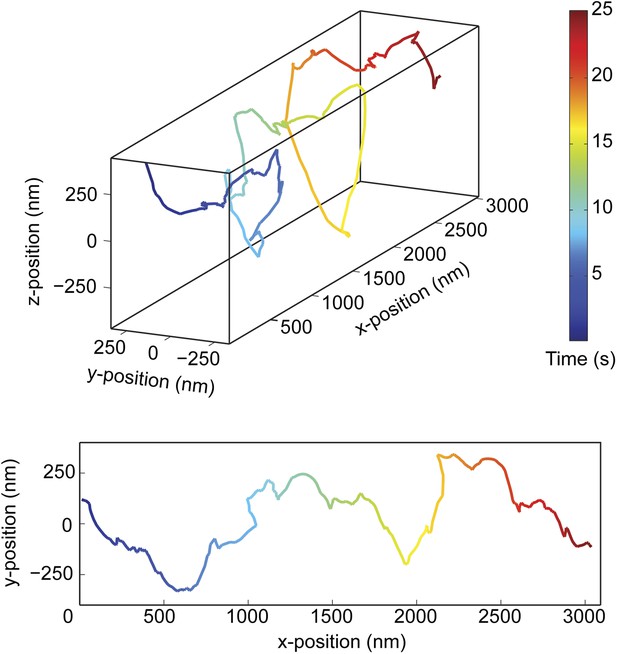
Additional example of the right-handed helical movement of a GST-Dyn331kD-coated bead along an MT bridge.
(top) Representative three-dimensional trace of a cargo bead-driven by GST-Dyn331kD motors shows right-handed helical motion. (bottom) Two-dimensional projection of the trace shown at top.
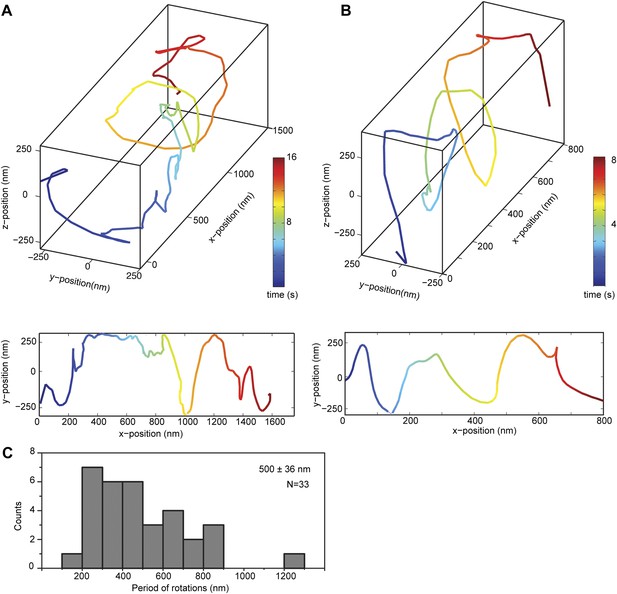
Bidirectional helical motility of cargo beads driven by full-length dynein along MT bridges.
(A and B) (top) Representative three-dimensional trace of a cargo bead driven by full-length dynein motors shows left- (A) and right-handed (B) helical motion. (bottom) Two-dimensional projections of the traces shown at top. The average pitch of the left-handed movement (576 ± 66 nm, SEM, N = 14) was longer (t-test, p <0.01) than that of the right-handed movement (399 ± 94 nm, SEM, N = 19). (C) Histogram of observed pitches per complete rotation. The average pitch is 500 ± 36 nm (mean ± SEM).
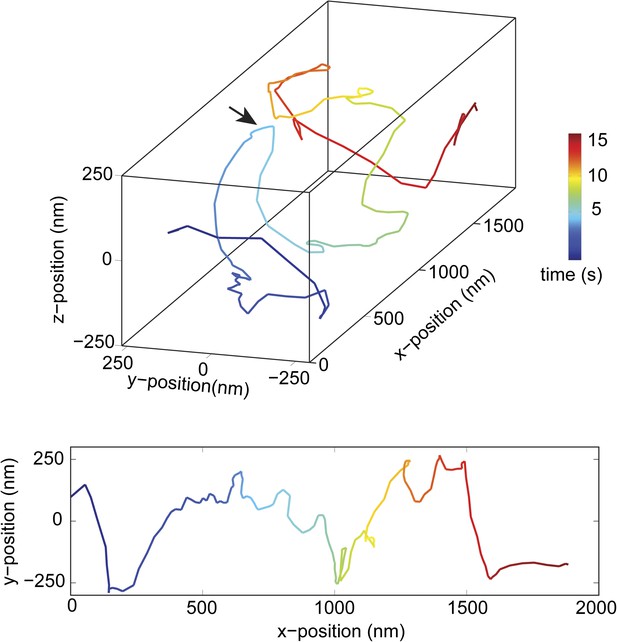
Change in handedness of rotation during the transport of a cargo bead driven by full-length dynein motors.
An example trace shows that a cargo bead initially moves along a GMP-CPP MT bridge with a right-handed helical motion. At around t = 5 s, the bead reverses its helical directionality (arrow) and moves in a left-handed helical path.
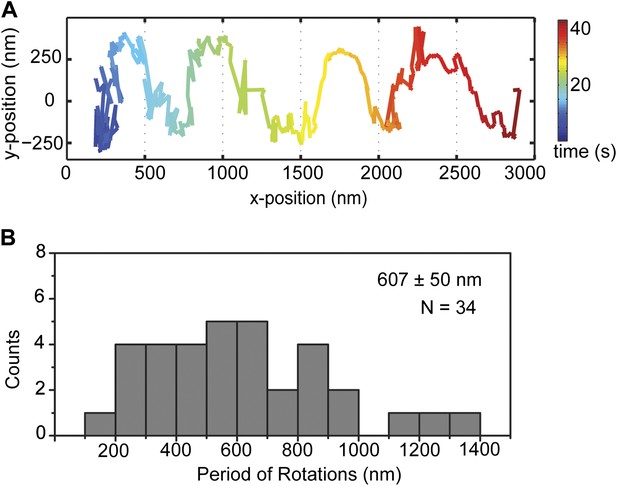
GST-Dyn331kD on taxol stabilized MTs.
(A) Representative two-dimensional trace of cargo beads driven by GST-Dyn331kD on taxol stabilized MTs which contains the mixture of 12, 13, 14 protofilaments with different rotational pitches. (B) Histogram of observed pitches per complete rotation. The results are similar to the cargo beads driven on GMP-CPP MTs which have 14 protofilaments with ∼6400 nm rotational pitch. The average pitch is 607 ± 50 nm (mean ± S.E.M.).
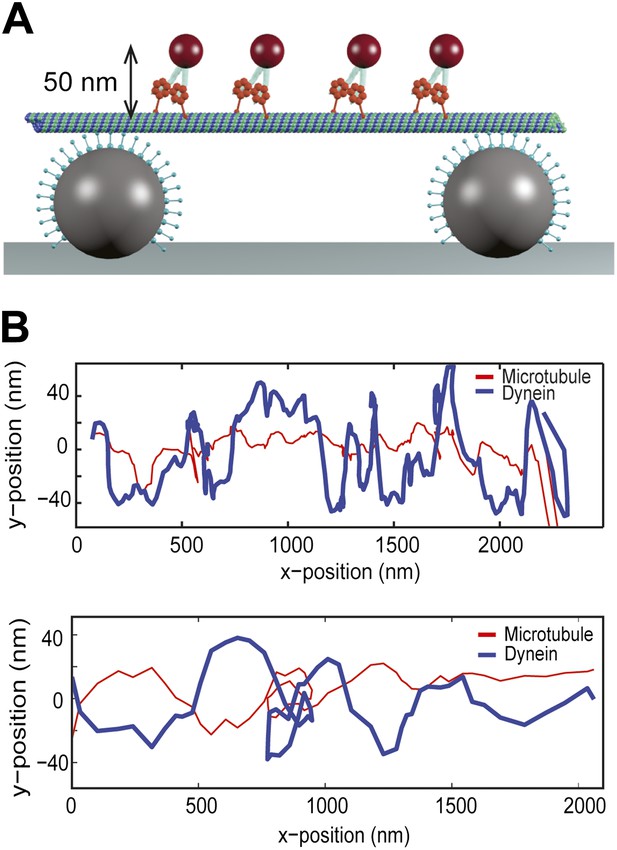
Single dynein motors frequently switch the direction of their sideways movement.
(A) Schematic representation of quantum-dot labeled single dynein motors on the MT bridges (not to scale). Expected amplitude of rotations is ∼50 nm. (B) Two example traces show 2D projection of dynein motors along the MT, using fluorescent tracking. MT filaments remain nearly straight between the bridges (persistence length is 5.2 mm) and oscillate due to the thermal fluctuation. The red trace represents the fluctuation of the MT bridge in the perpendicular axis, determined by the position of a quantum dot stably bound to a MT. The red trace was subtracted from the traces of quantum dots attached to single dynein motors (blue trace). Single motors do not show signs of regular helical movement.
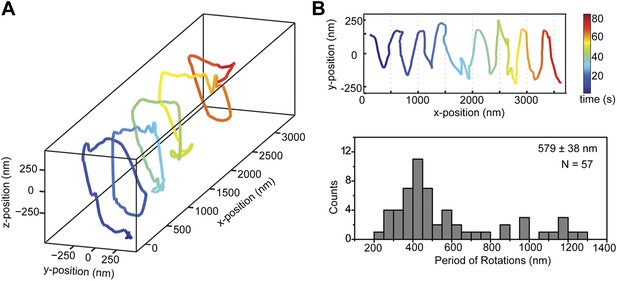
Dynein monomers prefer to move in a right-handed helix.
(A) Representative three-dimensional trace of a cargo bead driven by monomers shows right-handed helical motion. (B) (top) Representative two-dimensional trace for monomeric Dyn331kD. (bottom) Histogram of the periods of rotations shows that the average pitch is 579 ± 38 nm (mean ±SEM). The average pitch of the left-handed movement (658 ± 92 nm, SEM, N = 17) was longer (t-test, p=0.05) than that of the right-handed movement (490 ± 40 nm, SEM, N = 24).
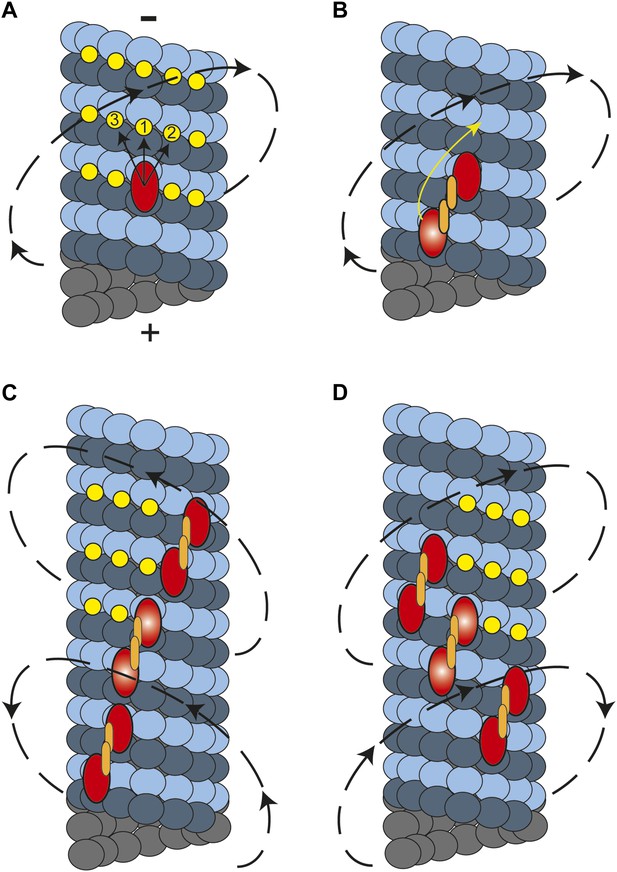
A model for the helical movement of cytoplasmic dynein.
(A) Top view of a monomeric dynein (red oval) stepping toward the MT-minus end (arrows). The yellow circles represent the putative binding sites for the highlighted dyneins. The closest available binding sites are numbered from 1 to 3. The nearest (8 nm) binding site is along the same protofilament (1). The binding site on the right (2) has a shorter distance (9.3 nm) than the one in left (3, 10.8 nm), resulting in a net preference to step rightward. (B) A dynein dimer prefers to orient on an MT with the leading head positioned on the right of the trailing head. When the trailing head (bright red oval) moves forward, it prefers to step rightwards to be positioned on the right hand side of its partner. (C) When multiple dimers carry a cargo bead, helical directionality may be affected by the number and orientation of the motors associated with an MT track. In this orientation, tubulin binding sites to the right for the motor in the middle (bright red ovals) may be obstructed for the motor in the lead. This results in a tendency to move in a left-handed helical pattern. (D) Due to the finite run length of dynein motors, MT-associated motors dissociate and new ones attach to the track. Changes in the orientation of MT-bound motors, switch the directionality of helical movement.
Videos
Example recording of left-handed rotations of a kinesin-1 coated bead at 100 ms time resolution via bright-field microscopy. Fluorescent image of MT has been superimposed onto the bright-field image for illustration purposes.
https://doi.org/10.7554/eLife.03205.005Example recording of left-handed rotations of GST-Dyn331kD-coated bead at 100 ms time resolution via bright-field microscopy.
https://doi.org/10.7554/eLife.03205.012Example recording of right-handed rotations of GST-Dyn331kD-coated bead at 100 ms time resolution via bright-field microscopy.
https://doi.org/10.7554/eLife.03205.013Example recording of right- and left-handed rotations of full-length dynein-coated bead at 100 ms time resolution via bright-field microscopy.
https://doi.org/10.7554/eLife.03205.014Example recording of right-handed rotations of monomeric Dyn331kD-coated bead. Video recorded via bright-field microscopy with time resolution of 100 ms.
https://doi.org/10.7554/eLife.03205.017Additional files
-
Supplementary file 1
Matlab code used in data analysis.
- https://doi.org/10.7554/eLife.03205.019