mRNA-programmed translation pauses in the targeting of E. coli membrane proteins
Figures
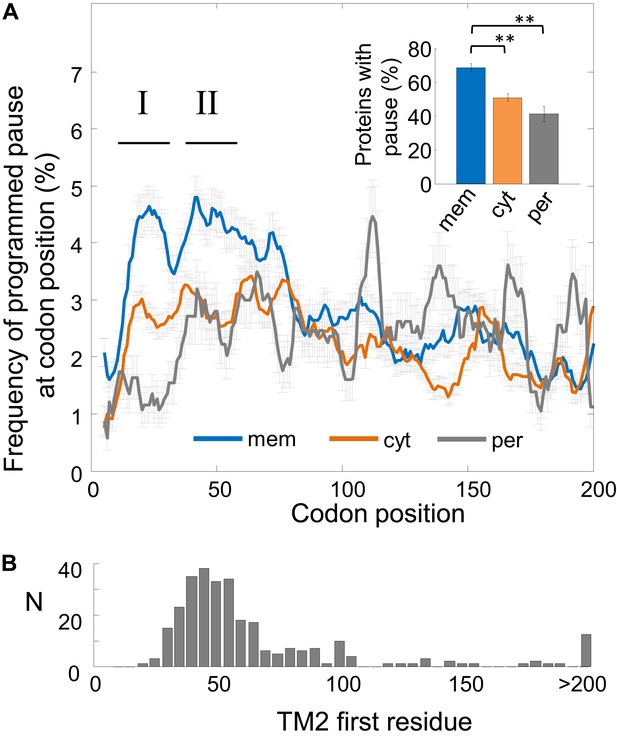
Frequency of programmed pauses across coding sequences.
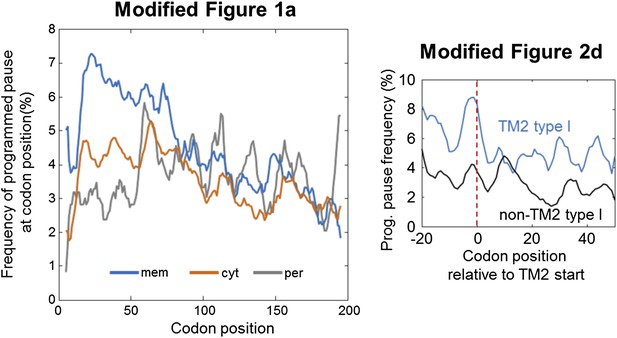
(A) Percentage of the programmed pauses in every codon across the coding sequences in membrane, cytoplasmic, and periplasmic proteins. Regions of elevated pause in membrane proteins are marked by I and II. Pause codon positions refer to codons in the A-site of the ribosome. Inset: percentage of proteins from different classes having at least one pause in the codon range 16–60. ** indicates p < 10−6. Error bars in figure and inset indicate s.e. for proportion. (B) Histogram of the distribution of the positions of the first codon of TM2 in E. coli membrane proteins that were analyzed in (A). N, number of proteins in which TM2 starts at the indicated position.
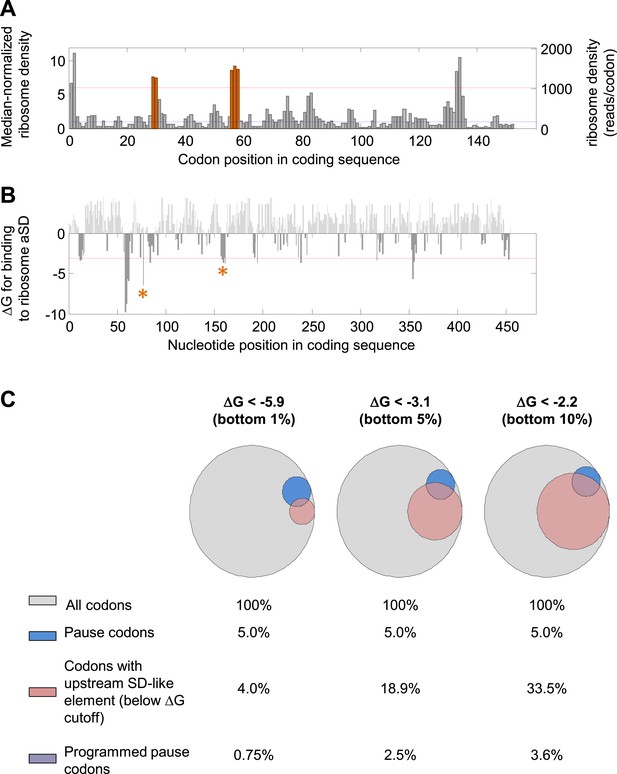
Detection of mRNA-programmed pauses.
(A and B) dskA gene as an example. (A) Median-normalized ribosome density profile for the E. coli gene dskA based on data from Li et al. (2012). Y axis scales to the left and right indicate the median-normalized and un-normalized values, respectively. Blue dashed line indicates median density of the gene. Red dashed line indicates the threshold for pause calling (density > ∼sixfold over the median). Orange indicates codons with programmed pauses (i.e., having upstream SD-like elements). (B) Calculated affinity of sequences in the coding sequences for the ribosome anti-Shine–Dalgarno sequence (aSD). Red dashed line indicates the energetic threshold for SD-like element calling (ΔG < −3.1). ΔG values above 0 are shown in light gray. Asterisk marks SD-like elements that explain the programmed pauses indicated in orange in (A). (C) Venn diagrams representing the effect of modifying the energetic threshold for SD calling. A total of 868,507 codons from well-expressed genes were analyzed. Blue, fraction of slowly translated codons (density > 6.06); pink, fraction of codons with at least one upstream sequence element predicted to bind the ribosome aSD with the indicated ΔG (‘Materials and methods’). The intersection of blue and pink makes up the ‘programmed pause’ codons in each energetic threshold. Energetic thresholds analyzed represent the bottom 1%, 5%, or 10% genome-wide.
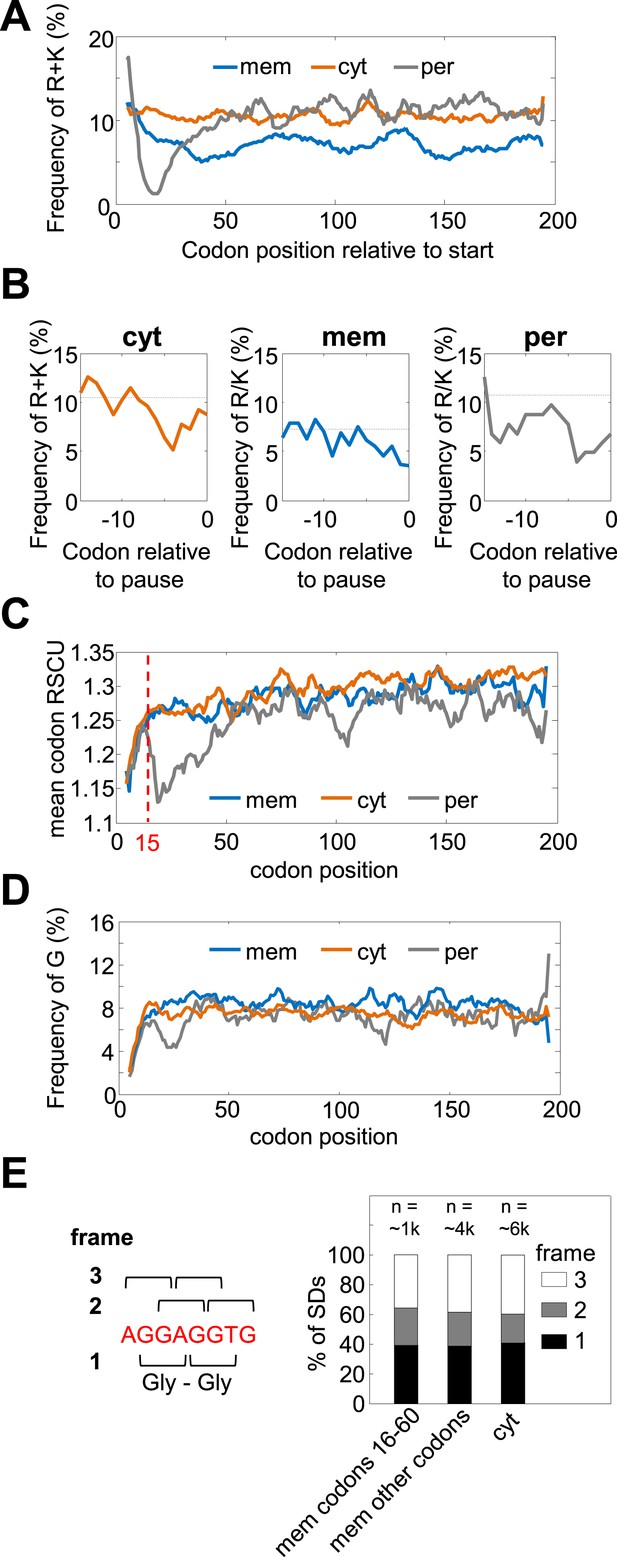
Amino acid and codon biases are not mediating pauses in membrane proteins' codons 16–60.
(A and B) Frequencies of positively charged residues (R+K) that were shown to slow-down translation (Charneski and Hurst, 2013) in regions of programmed pauses. (A) Frequencies of R+K along the coding sequences in membrane (mem), cytosolic (cyt), and periplasmic (per) proteins. Membrane proteins are depleted of K and R in codons 16–60 (and also throughout the sequence) compared to cytosolic proteins. (B) Frequency of R+K in regions immediately preceding programmed pauses that occur in codons 16–60. Programmed pause events that occur in codons 16–60 were analyzed for each protein class separately, and the peptide sequences directly preceding pauses were aligned such that position 0 constitutes the pause codon position. The frequency of K+R in every position is shown as solid line. Dashed lines indicate the average frequencies of K+R in the entire protein sequences. No significant excess of positively charged amino acids preceding pauses is observed. (C) Codon rarity along coding sequence position, as indicated by mean relative synonymous codon usage (RSCU) (Sharp et al., 1986). Most of the bias for rare codons occurs before codon 15 (dashed line), while the pause-enrichment in membrane proteins occurs only later. (D) Frequencies of G (Gly) residues, whose G-nucleotide-rich codons may mediate pause (Li et al., 2012), along the coding sequences. Membrane proteins show only a slight elevation of Gly occurrence, which is not specific to the region of codons 16–60. (E) Analysis of coding reading frames of SD-like sequences preceding pauses indicates that most such motifs do not occur in frames that would encode glycines. Left: the canonical SD sequence (red) and definition of frames. Frame 1 was defined as the frame in which the canonical SD encodes Gly–Gly dipeptide. Right: distribution of pause-preceding SD motifs in each frame. The analysis was done separately for membrane proteins in the codon range 16–60, for the other codons in membrane proteins, or for cytosolic proteins.
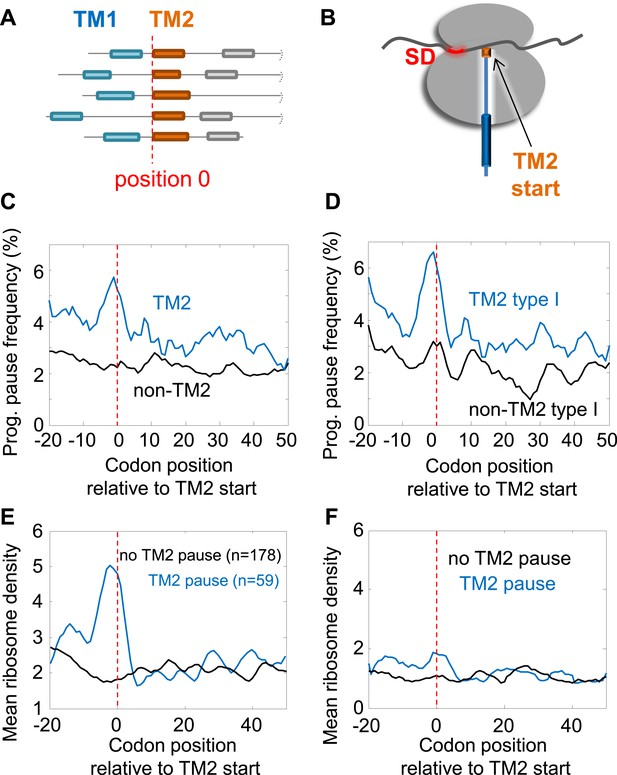
Pause before translation of TM2.
(A) Membrane proteins were aligned such that the first residue of TM2 is aligned to position 0 (red line). (B) Visual scheme of the stage at which pause occurs during translation. Red marks SD-like element in the mRNA. Blue line and cylinders depict nascent polypeptide and possible TM location in the tunnel, respectively. (C and D) The frequency of codons having programmed pauses in every position was analyzed, either in all membrane proteins (C) or only in proteins having intracellular N-termini (D). Profiles of proteins aligned to TM2 are in blue. Profiles of proteins aligned to all other TMs are in black. (E) Average ribosome density profiles of proteins with (blue) or without (black) identifiable programmed pauses in codons −5 to +1 relative to TM2 start. (F) Same as (E) but only in mRNA nucleotide positions lacking upstream SD-like elements.
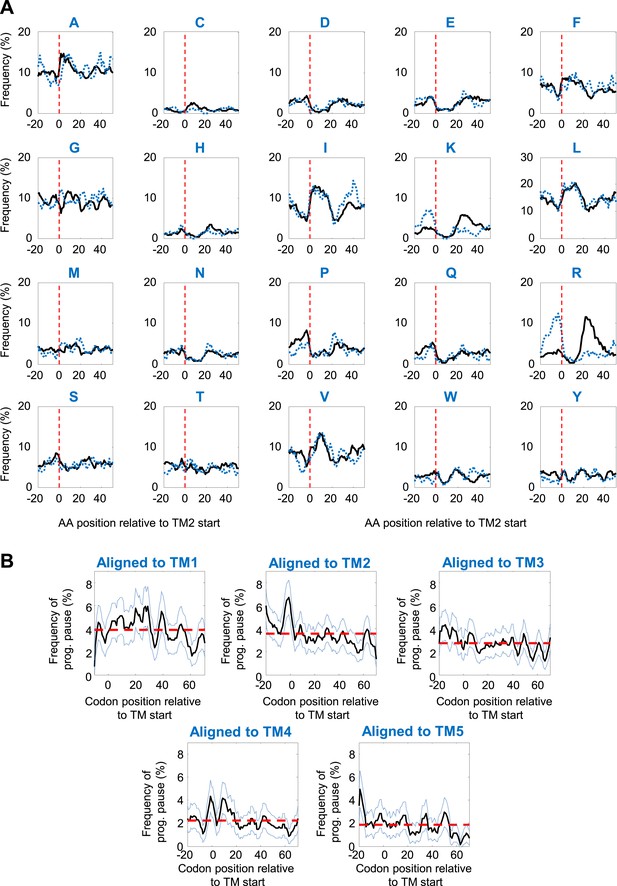
Specificity of programmed pause before TM2.
(A) Amino-acid frequencies in various positions relative to TM2 start. Black and blue dashed lines depict proteins with intracellular or extracellular N-temini, respectively. (B) Programmed pause profiles of coding sequences aligned to different TMs. Membrane proteins with cytosolic N-termini were aligned such that the first residue of TM-X (X = 1–5) is aligned to position 0. The frequency of codons having programmed pauses in every position was analyzed, similar to Figure 2D. Light blue dashed lines indicate 95% confidence intervals. Red dashed lines depict the average frequency over the range.
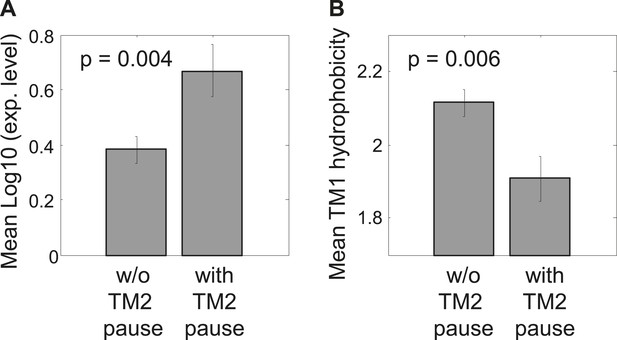
Comparison between proteins with and without identifiable pauses before TM2.
(A) Mean expression levels of genes in both groups. The gene expression was calculated for each gene as the logarithm of the median ribosome density in its coding sequence, reflecting the level of protein translation. Error bars indicate SEM. (B) Mean TM1 hydrophobicity in both protein groups. For each protein, TM1 and 5 flanking residues down- and up-stream were taken. Hydrophobicity was calculated by applying a sliding window of 18 residues and calculating the average Kyte–Doolitle score. The maximal score was taken to represent the TM hydrophobicity. Error bars indicate SEM.
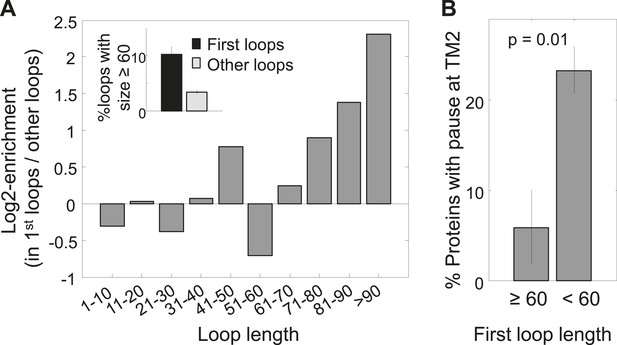
Enrichment of long first loops and its effect on pausing.
(A) Enrichment of various loop lengths in first loops compared to all other loops. Shown are Log2 values of ratios. Inset: occurrence of long loops of length ≥60 in first loops or other loops. Error bars indicate s.e. for proportion. (B) Occurrence of pause before TM2 in proteins with long or short first loops. Error bars indicate s.e. for proportion.
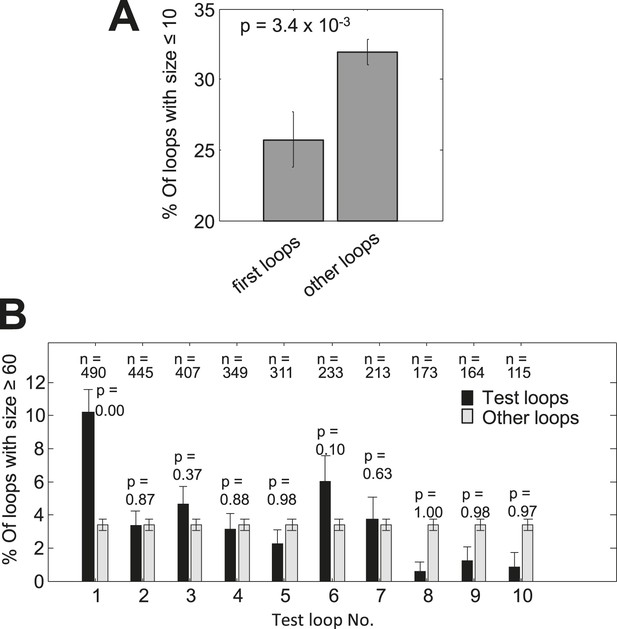
Analysis of loop lengths.
(A) Occurrence of short loops of length ≤10 in first loops or other loops. (B) Occurrence of loops with length ≥60 in increasing loop locations or all other loops. Similar to the inset in Figure 3A. The sample sizes of the test loops and the p-values (hypergeometric test) for enrichment are given above the bars. Error bars indicate s.e. for proportion.
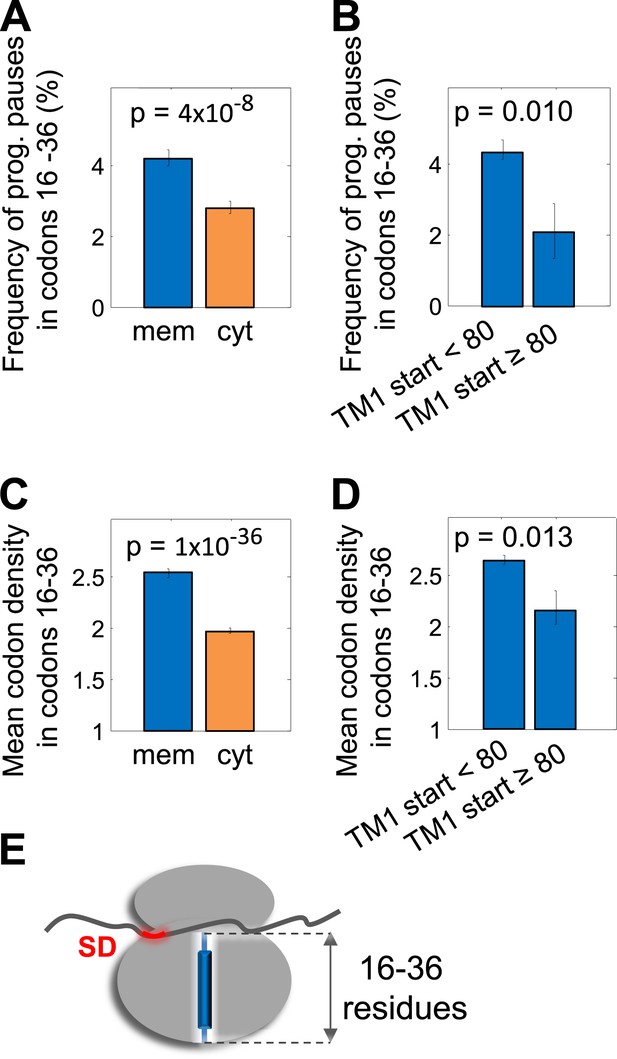
Pause in codon range 16–36.
(A and C) Comparison of the frequency of programmed pauses (A) or mean ribosome density (C) in codon range 16 to 36 between cytoplasmic and membrane proteins. Error bars indicate s.e. for proportion (A and B) or SEM (C and D). (B and D) Same comparisons between membrane proteins in which the first residue of TM1 occurs after or before position 80 of the polypeptide. (E) Visual scheme of the stage at which pause occurs during translation, similar to Figure 2B.
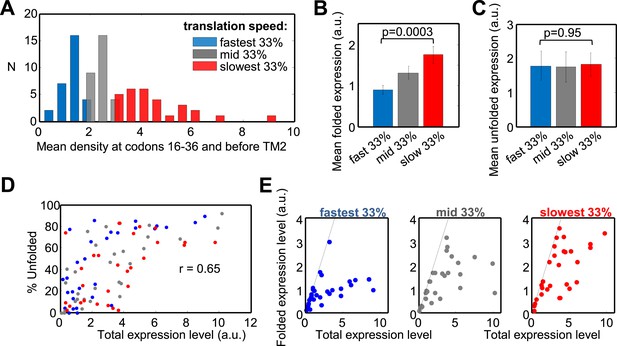
Effect of translation rate at codons 16–36 and before TM2 on membrane protein overexpression.
(A) Histogram of translation speeds. For each gene, the ribosome density at codons 16–36 and before TM2 was averaged. The distribution of values in the different genes is shown (N, number of genes with indicated density). Colors indicate groups corresponding to different translation speeds at these codon ranges and accompany the rest of the figure. (B and C) Mean (±SEM) expression values of the folded (B) and unfolded (C) forms of proteins from the three different groups. (D) Effect of the total expression level on the percentage of unfolding in the different protein groups. (E) Folded vs total levels of expression in the three protein groups. Each dot represents a different protein. Dashed line depicts a 1:1 ratio wherein all of the expressed protein is detected in its folded form. The experiment was repeated three times.
-
Figure 5—source data 1
Quantification of protein overexpression.
- https://doi.org/10.7554/eLife.03440.013
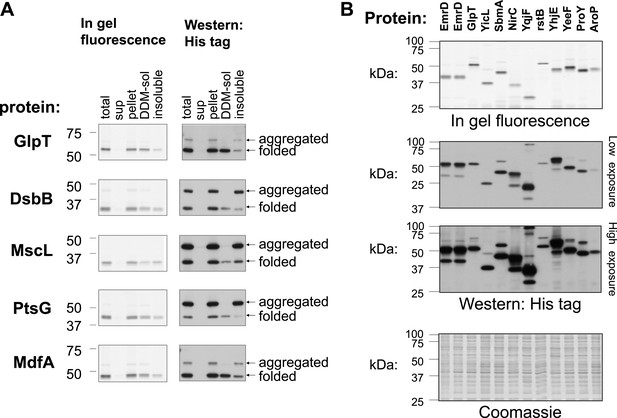
Detection of folded and unfolded membrane protein–GFP-His8 hybrids.
SDS-PAGE analysis of various membrane proteins overexpressed in E. coli. In-gel fluorescence detects only folded GFP. Western blotting with detection of all His-tagged proteins reveals both the folded and unfolded forms. The Western and fluorescence measurements were done on the same gels. (A) Fractionation and solubility of selected membrane proteins overexpressed in E. coli. Cells expressing the indicated proteins were disrupted by sonication (total) and ultracentrifuged to separate the cytosolic fraction (sup) from the membranes and aggregates (pellet). The pellet fraction was then solubilized with the mild detergent DDM which only solubilizes non-aggregated membrane proteins. Detergent-solubility was determined by ultracentrifugation (DDM-sol and insoluble). Only the lower, folded band is soluble with DDM in all cases. (B) SDS-PAGE analysis of various membrane proteins overexpressed in E. coli. Upper panel: in-gel fluorescence. Middle panels: low and high exposure of Western blotting. Lower panel: Coomassie stain of all cellular proteins, as a control for protein amounts. Two first lanes in all gels were from independent cell cultures expressing EmrD and served as standard.
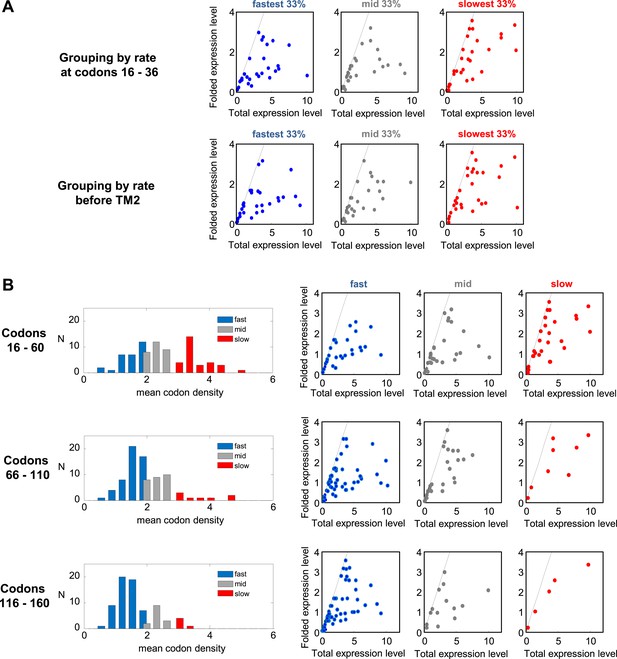
Effect of translation rates at various positions on protein overexpression.
(A) Membrane proteins were divided to 3 groups, similar to Figure 5A, according to the rate of translation at codon range 16–36 (upper panel) or before TM2 (codons [−5]–[+1] relative to TM2 start) (lower panel). The folded vs total levels of expression in the three protein groups are plotted, similar to Figure 5E. (B) Translation rates in different codon ranges and how they affect expression. Left panels: distributions of average translation rates in different codon ranges, as estimated from mean codon densities in different proteins (similar to Figure 5A). In increasing downstream codon regions, the number of slow proteins decreases. Right panels: folded vs total levels of expression in the three protein groups (similar to Figure 5E). Slowly translated proteins appear to benefit from better folding, regardless of where the pauses occur.
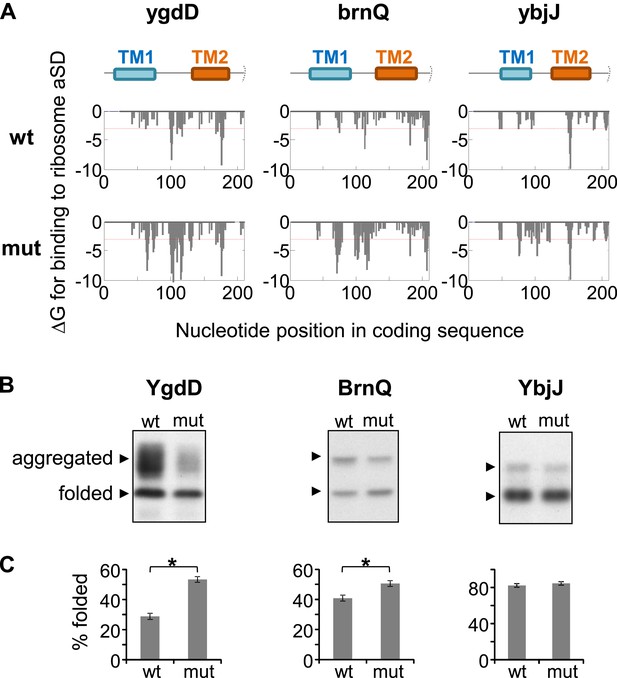
Effect of silent mutations introducing SD-like motifs on protein folding and aggregation.
(A) Effect of mutations on affinity for ribosomal anti-Shine–Dalgarno sequence. Lower panels show calculated binding energies to anti-Shine–Dalgarno sequence, similar to Figure 1—figure supplement 1B. wt, wild-type; mut, mutant. Upper panel indicates the nucleotide positions in the coding sequence that code for TM1 and TM2. Note that all proteins and coding sequences are longer than the range of 70 amino-acids and 210 nucleotides shown. (B) Western blot analysis of protein–GFP-His8 hybrids, similar to Figure 5—figure supplement 1. (C) Densitomentry-based quantification of the percentage of the folded state from the total protein. Error bars represent SEM from three independent experiments. * indicates p < 0.01 (one tailed, paired t test).
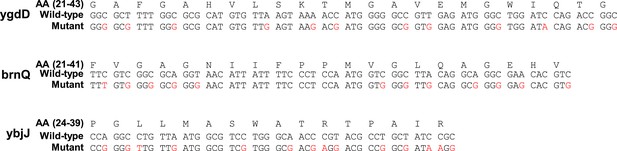
Silent mutations introducing SD-like motifs.
Amino-acid (AA) and coding sequences of test genes and mutants. Only the range of amino-acids whose codons were mutated is shown. Amino-acid range is indicated in parenthesis. Mutated nucleotides are indicated in red.
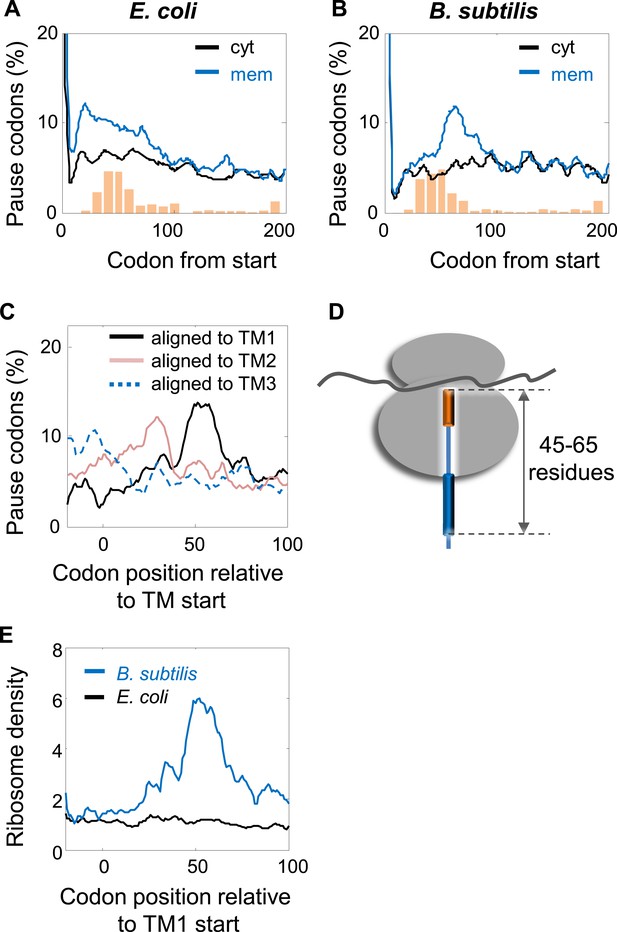
Comparison of pausing events in B. subtilis and E. coli.
(A and B) Percentage of pauses in codons across coding sequences in membrane (mem) and cytoplasmic (cyt) proteins in E. coli (A) and B. subtilis (B). Orange histograms at the bottom indicate the distribution of the position of the first codon of TM2 (similar to Figure 1B). (C) B. subtilis membrane proteins were aligned such that the first residues of TM1 or TM2 or TM3 are aligned to the position 0. The percentage of codons having pauses in every position was analyzed. (D) Visual scheme of the stage at which pause occurs during translation in B. subtilis, similar to Figure 2B. (E) Effect of position relative to TM1 on ribosome densities in positions lacking upstream SD-like elements.
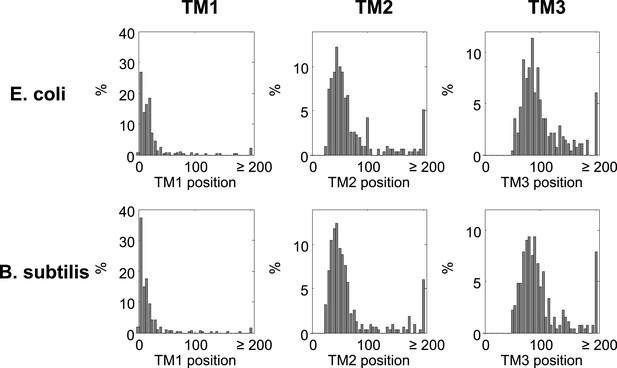
Histograms of TM locations in B. subtilis and E. coli.
The first TM residue was taken as the TM position, similar to Figure 1B. N, number of proteins in which the TM starts at the indicated position.