Recurrent loss of CenH3 is associated with independent transitions to holocentricity in insects
Figures
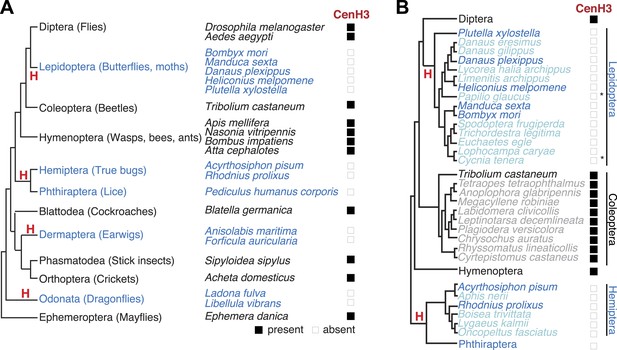
Holocentric insects lack CenH3.
(A) Phylogeny of insect orders (species) examined in this study. Holocentric insect orders are indicated in blue, and inferred multiple transitions to holocentricity in insects are labeled with ‘H’. Using protein homology searches of genomes or assembled transcriptomes, we inferred either the presence (black box) or absence (empty box) of CenH3. (B) CenH3 loss is widespread in Lepidoptera and Hemiptera, but not Coleoptera. Phylogenetic relationship of holocentric insects used for transcriptome assemblies (light blue), holocentric insects with sequenced genomes (blue), monocentric insects used for transcriptome assemblies (gray), and monocentric insects with sequenced genomes (black). The presence of contaminating microsporidian CenH3 transcripts is indicated with an asterisk.
-
Figure 1—source data 1
List of species with sequenced genomes and information to their corresponding analyzed proteomes.
- https://doi.org/10.7554/eLife.03676.004
-
Figure 1—source data 2
Statistics of mRNA-Seq assemblies.
Order, species name, centromere type (holocentromere [H] or monocentromere [M]), number of reads for the assembly, obtained transcripts longer than 250 base-pairs (only one transcript isoform) and number of significant tblastn alignments (E value 10–10) out of 16,644 annotated T. castaneum and 30,305 annotated D. melanogaster proteins.
- https://doi.org/10.7554/eLife.03676.005
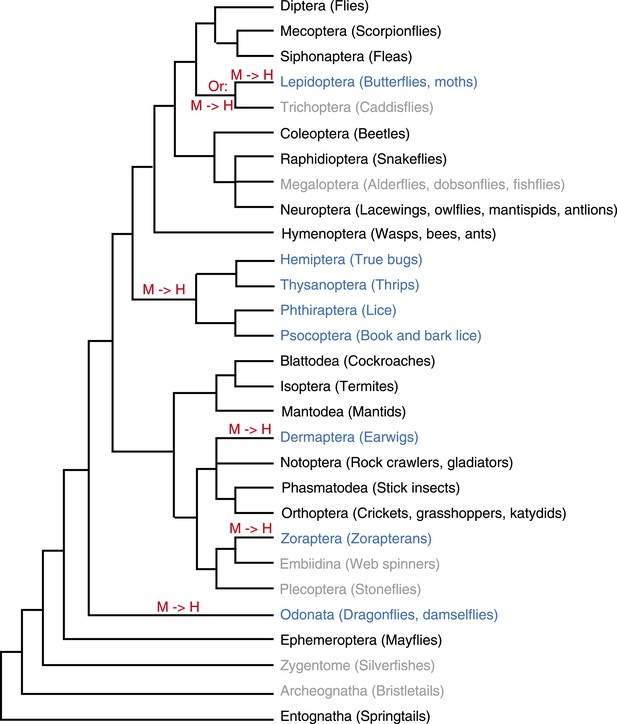
Evolution of holocentricity in insects.
All insect orders and their phylogenetic relationship (Whiting, 2002; Grimaldi and Engel, 2005; Savard et al., 2006) are displayed. Holocentric orders are highlighted in blue, monocentric orders are in black, and orders with no diagnostic karyotype data are in gray. Lineages with transitions to holocentricity are labeled with ‘M → H’ in red. Although cytological analyses of the two trichopteran insects proposed the possibility of holocentric chromosomes (Suomalainen, 1966), we did not find the data diagnostic for this conclusion. Therefore, we depicted the trichopteran order in gray and indicate that the holocentric transition could have either occurred in the common ancestor of Trichoptera and Lepidoptera or on the lepidopteran lineage (‘Materials and methods’). The phylogenetic position of the holocentric zorapteran order is unresolved. Analyses of morphological data proposed a monophyletic relationship with Dermaptera (Jarvis et al., 2005; Terry and Whiting, 2005), which likely places the transition to holocentricity in the common ancestor of both orders and reduces the number of independent transitions to holocentricity from 5 to 4.

Insect CenH3 comparison.
Protein alignment including CenH3s from four representative species from Hymenoptera (Apis mellifera, Bombus impatiens, Atta cephalotes, and Nasonia vitripennis), one species from Coleoptera (Tribolium castaneum), two species from Diptera (Drosophila melanogaster and Aedes aegypti), cricket (Acheta domesticus), stick insect (Sipyloidea sipylus), cockroach (Blatella germanica), and mayfly (Ephemera danica) CenH3s compared to canonical histone H3 (blue) that is invariant among these species. CenH3-characteristic amino acid changes (red) and loop1 (gray box) are highlighted. Although the CenH3 histone fold domain is relatively well conserved, the N-terminal tail is not and cannot even be considered homologous between the different insect orders.
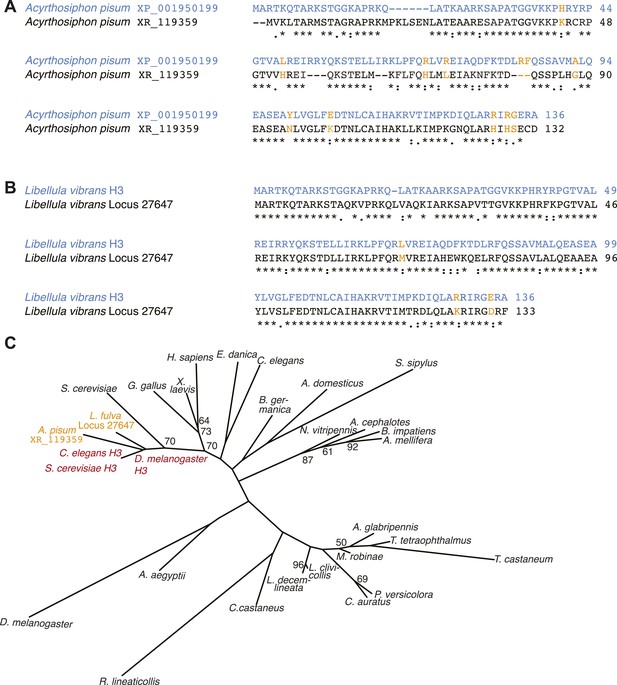
Additional H3-like variants in holocentric insects.
(A) Acyrthosiphon pisum alignment of H3 (blue) and an H3-derived variant that is missing many canonical H3 residues in the histone fold domain. Changes and deficiencies of completely conserved amino acids of H3 homologs (Malik and Henikoff, 2003) are highlighted in yellow. (B) Libellula vibrans alignment of H3 (blue) and a recently derived H3-like variant. The two proteins are 80.2% identical. (C) Phylogeny of various animal and fungal H3 (red), H3-like (orange), and CenH3 proteins (black). Maximum likelihood tree of histone fold domains including the two identified H3-derived variants in the A. pisum and L. vibrans genomes. Bootstrap percentages above 50 are indicated.
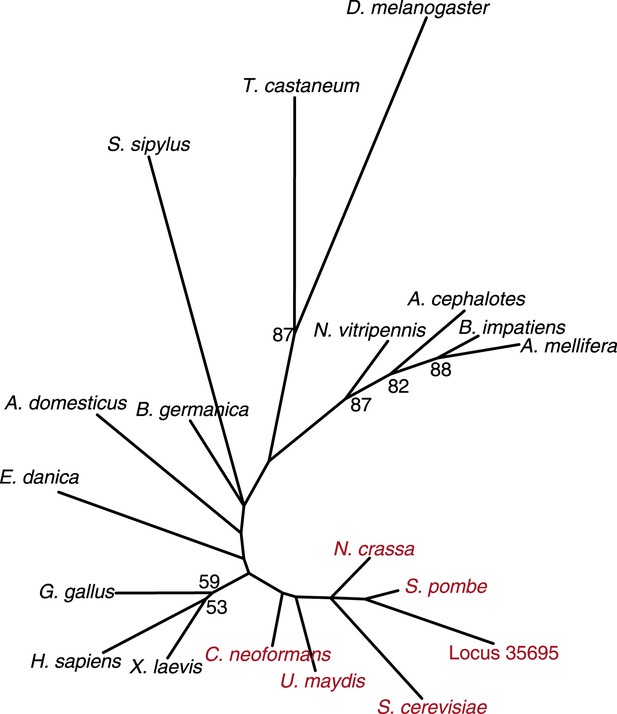
Fungal CenH3 transcript contaminants in the cockroach assembly.
Maximum likelihood derived CenH3 phylogeny of various animal (black) and fungal (red) CenH3 proteins based on their histone fold domains. Locus 35,695 encoding CenH3 (orange) found in the cockroach B. germanica assembly clearly groups with fungal homologs. Bootstrap percentages above 50 are indicated.
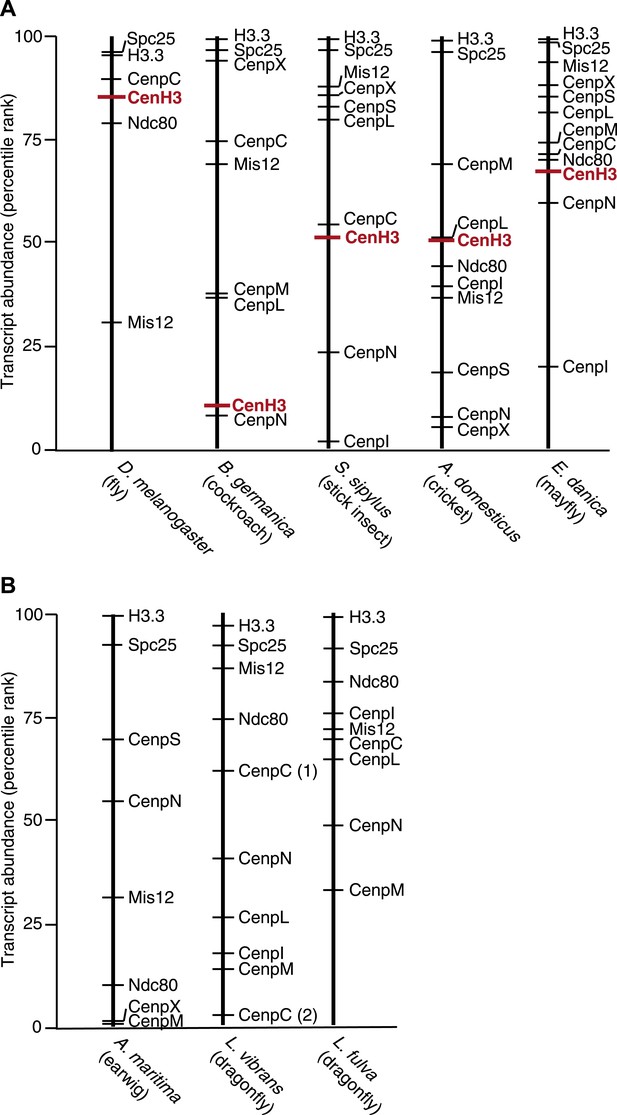
CenH3 transcript abundance in mRNA-Seq assemblies.
Abundance of transcripts encoding for CenH3 and other kinetochore components in mRNA-Seq assemblies of (A) monocentric insects and (B) holocentric insects. Transcripts were rank-ordered based on their normalized abundance. Ranked percentiles corresponding to transcripts encoding for CenH3 homologs (red lines) and kinetochore proteins (black lines) are indicated, where low percentiles correspond to low abundance, while high percentiles correspond to high abundance.
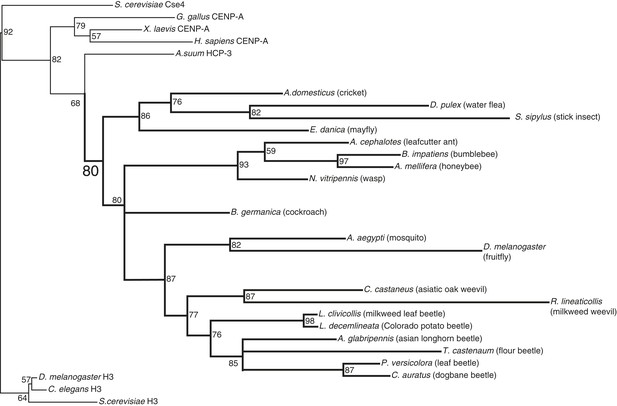
CenH3 phylogeny in insects.
Maximum likelihood phylogeny of histone fold domains of canonical H3 proteins as well as those from previously and newly (this study) identified CenH3 proteins. Branches with bootstrap percentages below 50 were collapsed. H3 and CenH3 homologs form distinct clades. Arthropod CenH3s constitute their own monophyletic clade (supported by significant bootstrap support) separate from nematode, vertebrate, and fungal CenH3s, although there are two topological inconsistencies with accepted arthropod phylogeny within the arthropod CenH3s. First, the position of the Daphnia pulex CenH3 is within the insect clade, whereas, based on the species phylogeny D. pulex CenH3 is expected to branch ancestrally to all insect CenH3s. Second, the position of the cockroach CenH3 between the hymenopteran and dipteran clades is not consistent with the closer phylogenetic relationship of cockroaches to crickets and stick insects. Nevertheless, our findings suggest that CenH3 homologs have been inherited via vertical transmission in monocentric insects and were independently lost in holocentric insects.
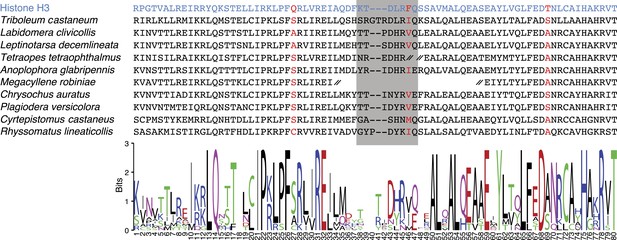
CenH3 homologs in Coleoptera (beetles).
Top: protein alignment is shown for canonical histone H3 (blue), T. castaneum CenH3, and other coleopteran CenH3s identified in transcriptome assemblies. CenH3-characteristic amino acid changes (red) and loop1 (grey box) are highlighted. CenH3 derived protein fragment ends are indicated by a double hash. Bottom: sequence logo for coleopteran CenH3 alignment is presented (http://weblogo.berkeley.edu/logo.cgi).
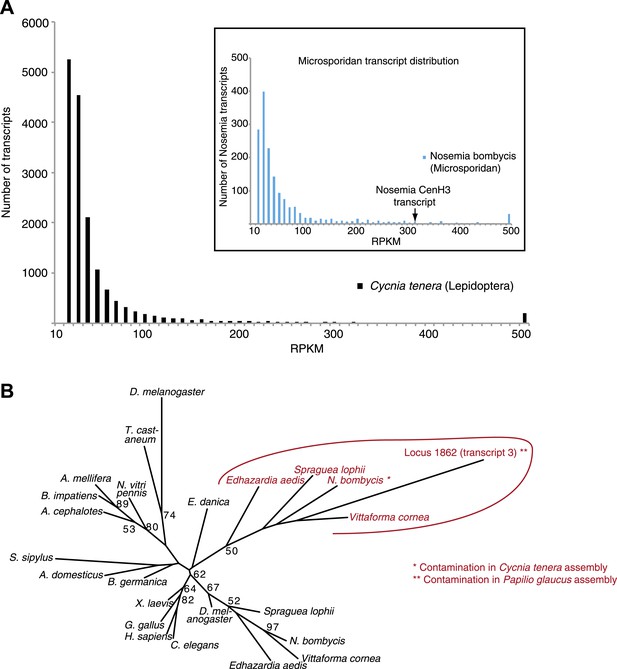
Contaminating microsporidian CenH3 transcripts in the two lepidopteran assemblies.
(A) Transcript abundance profiles in RPKM of the Cycnia tenera mRNA-Seq assembly. Only assembled transcripts longer than 250 bp were selected, and in cases of multiple different isoforms the longest isoform was chosen. To identify all the contaminating microsporidian Nosema bombycis derived transcripts, assembled transcripts were aligned to the N. bombycis CQ1 genome and significant hits (E value <10–5) were chosen. Main panel: a histogram of RPKM values of all transcripts (total 16,643). Inset: a histogram of RPKM values of N. bombycis transcripts (total 1646). (B) Phylogenetic tree of various animal and microsporidian CenH3 proteins. Maximum likelihood phylogeny of H3 and CenH3 histone fold domains (excluding loop 1) including the Locus 1862 (transcript 3) encoding CenH3 found in the Papilio glaucus assembly. Microsporidian CenH3 homologs and the putative homolog in the P. glaucus assembly are shown in red and orange, respectively. Bootstrap percentages above 50 are indicated.
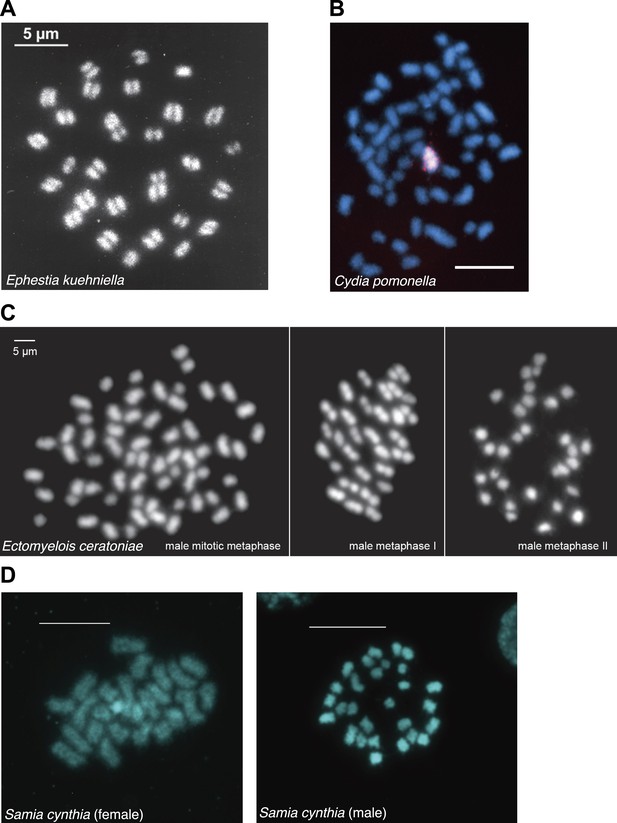
Chromosome spreads from various lepidopteran species generously provided by Frantisek Marec and Atsuo Yoshido.
(A) Metaphase 1 oocytes of Ephestia kuehniella chromosomes (unpublished). (B) Cydia pomonella mitotic chromosomes (Fukova et al., 2007). (C) Male mitosis and meiosis of Ectomyelois ceratoniae chromosomes (Mediouni et al., 2004). (D) Female (left) and male (right) mitotic chromosomes of Samia cynthia (unpublished).
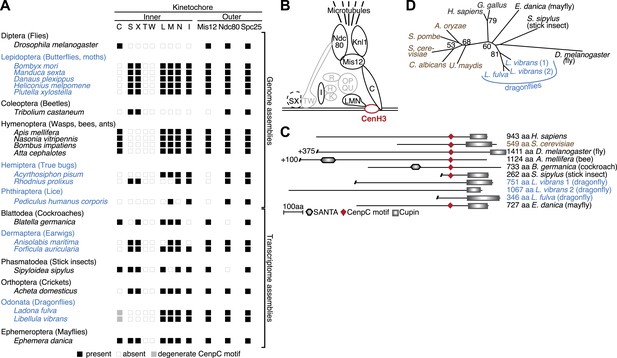
Evolution of the kinetochore composition in insects.
(A) Kinetochore proteins in insects. Using protein homology searches of the genome or assembled transcriptomes of monocentric (indicated in black) and holocentric insects (indicated in blue), we inferred either the presence (black box) or absence (empty box) of inner and outer kinetochore protein components. The presence of a putative CenpC homolog without a recognizable CenpC motif in Odonata is indicated by a filled gray box. We also found a weak match to a putative Mis12 homolog in T. castaneum (TC001997); however its weak homology relative to other insect Mis12 proteins leads us to assign this to be only a tentative match. (B) Schematic structure of a kinetochore largely based on its characterization in vertebrates and fungi (Hori et al., 2008; Westermann and Schleiffer, 2013). Components found in insects are highlighted in black, whereas components that are absent in all insects examined are in gray. The dashed line around the CenpS/X complex indicates that its kinetochore localization is unlikely in insects. (C) Putative CenpC proteins in insects. Schematics of structural domains and the CenpC motif of human, yeast, and insect CenpC proteins are shown. (D) Maximum likelihood tree of cupin domains of dragonfly (blue), fungal (brown), and other animal (black) CenpC proteins. Bootstrap percentages above 50 are indicated.
-
Figure 2—source data 1
Accession numbers or sequences for all insect kinetochore proteins analyzed or described in this study.
- https://doi.org/10.7554/eLife.03676.016

CenpC-Cupin domain alignment.
Top: protein alignment of various fungal (brown), animal (black), and the three dragonfly (blue) cupin domains. Bottom: sequence logo (http://weblogo.berkeley.edu/logo.cgi) for alignment showing conservation of structurally important residues (Dunwell et al., 2001).
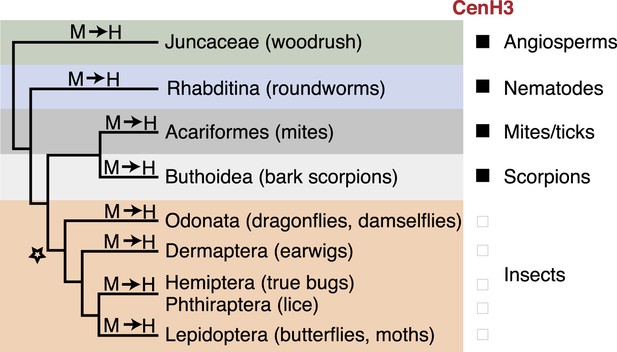

CenH3 losses associated with holocentric transitions are unique to insects.
Phylogenetic relationship of holocentric lineages are schematized. Transitions to holocentricity (M → H) and the absence (black box) or presence (empty box) of CenH3 are indicated. Inferred evolutionary origin of a first ‘potentiating event’ (indicated with a star) together with subsequent recurrent transitions to holocentricity allowing the loss of CenH3 in four insect lineages.





