Divergent mechanisms regulate conserved cardiopharyngeal development and gene expression in distantly related ascidians
Figures
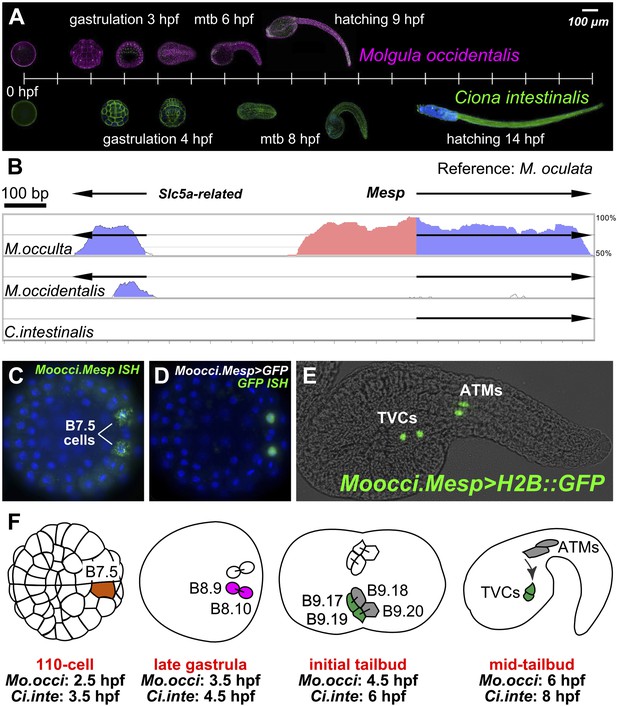
The B7.5 lineage in M. occidentalis.
(A) Diagram comparing M. occidentalis (top) and C. intestinalis (bottom) embryogenesis at 24°C. Embryos were stained with Alexa Fluor dye-conjugated phalloidin to visualize cell outlines and DAPI to visualize cell nuclei. (B) Diagram of mVISTA (Frazer et al., 2004; genome.lbl.gov/vista/) alignment of M. oculata Mesp (Moocul.Mesp) locus to orthologs in M. occulta, M. occidentalis, and C. intestinalis. Shaded peaks indicate sequence conservation above 70% over 100-bp windows (blue = protein-coding, pink = non-coding). Arrows indicate direction of transcription of protein-coding genes. Non-coding sequences upstream of Mesp are only conserved between M. oculata and M. occulta. M. occidentalis and C. intestinalis show considerable divergence even in protein-coding sequences. Note that microsynteny with SLC5A-related gene supports the orthology of these sequences among the Molgulids. (C) In situ hybridization (ISH) for Moocci.Mesp in 110-cell stage embryo (vegetal view), showing mRNA detection (green) in B7.5 blastomeres. Nuclei were counterstained with DAPI (blue). Staging is given by hours post-fertilization (hpf). (D) Vegetal view of a 110-cell stage embryo electoporated with Moocci.Mesp>GFP reporter construct. Reporter gene expression was detected by ISH for GFP transcripts (green). Nuclei were stained with DAPI (blue). (E) Lateral view of a mid-tailbud stage embryo electroporated with Moocci.Mesp>Histone2B::GFP reporter construct. GFP fluorescence reveals B7.5 descendants on left side of embryo: two trunk ventral cells (TVCs) and two anterior tail muscles (ATMs). (F) Diagram of B7.5 lineage divisions from 110-cell stage to mid-tailbud stage, inferred from previous C. intestinalis studies. Cells are named according to Conklin's method (Conklin, 1905). The lineage is bilaterally symmetric, but only cells on the left side are indicated and named. Relative staging given for M. occidentalis (Mo.occi) and C. intestinalis (Ci.inte). 110-cell and late gastrula: vegetal view. Initial tailbud: dorsal view. Mid-tailbud: lateral view. Anterior pole is on the left in all images and illustrations.
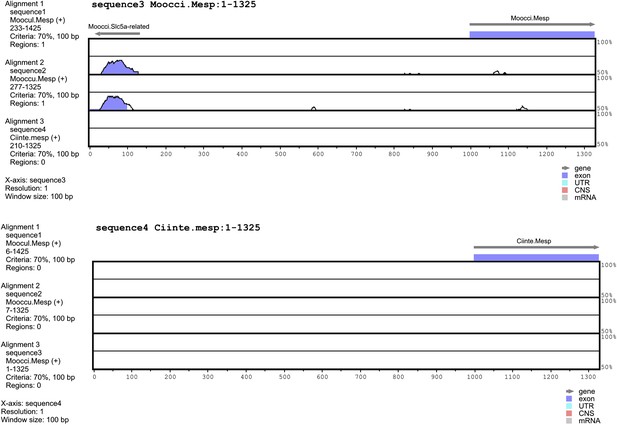
Alignment of 5′ flanking sequences from Mesp orthologs.
Top: diagram of mVISTA (Frazer et al., 2004; genome.lbl.gov/vista/) alignment of M. occidentalis Mesp (Moocci.Mesp) locus to orthologs in (from top to bottom) M. oculata, M. occulta, and C. intestinalis. Bottom: diagram of mVISTA alignment of C. intestinalis Mesp (Moocci.Mesp) locus to orthologs in (from top to bottom) M. oculata, M. occulta, and M. occidentalis. Blue-shaded peaks indicate protein-coding sequence conservation above 70% in 100-bp windows. Arrows indicate direction of transcription of protein-coding genes Slc5a-related and Mesp.
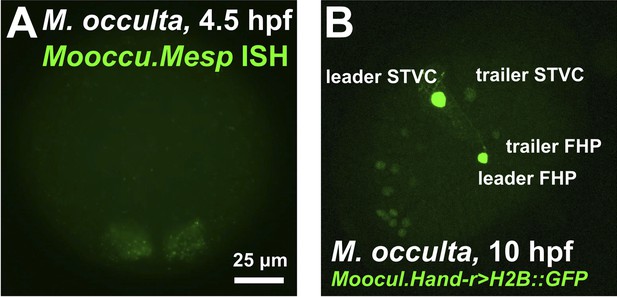
The B7.5 lineage of the tail-less species M. occulta.
(A) In situ hybridization for M. occulta Mesp (Mooccu.Mesp) in 110-cell st/early gastrula embryo, showing expression (green) in the B7.5 blastomeres. Image is taken from a vegetal view, anterior to the top. (B) Lateral view of a transiently-transfected M. occulta mid-tailbud-equivalent stage embryo electroporated with a Moocul.Hand-r reporter construct (Moocul.Hand-r>H2B::GFP). The reporter proteins (Histone2B::GFP, green) label the cell nuclei descendant of the trunk ventral cells (TVCs), namely the secondary TVCs (STVCs) and the first heart precursors (FHPs). Due to leader/trailer mosaic retention of the electroporated plasmid (Stolfi and Christiaen, 2012), the daughters of the leader cell (‘leader STVC’ and ‘leader HFP’) are more strongly labeled than the daughters of the trailer cell. Some staining is also seen in what may be anterior endoderm or A7.6 mesoderm, other known territories of Hand-r expression in M. occulta and other ascidian species. Anterior is to the bottom left.
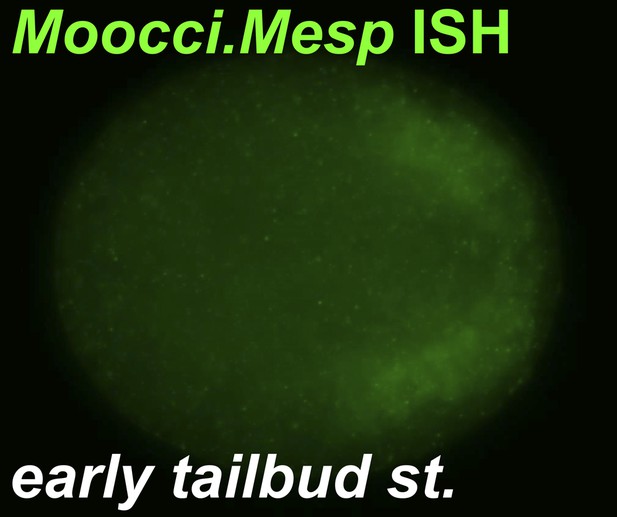
Moocci.Mesp in situ hybridization at tailbud stage.
In situ hybridization for Moocci.Mesp in initial/early tailbud embryo, showing no expression at this stage. Image is a coronal view with anterior to the left.
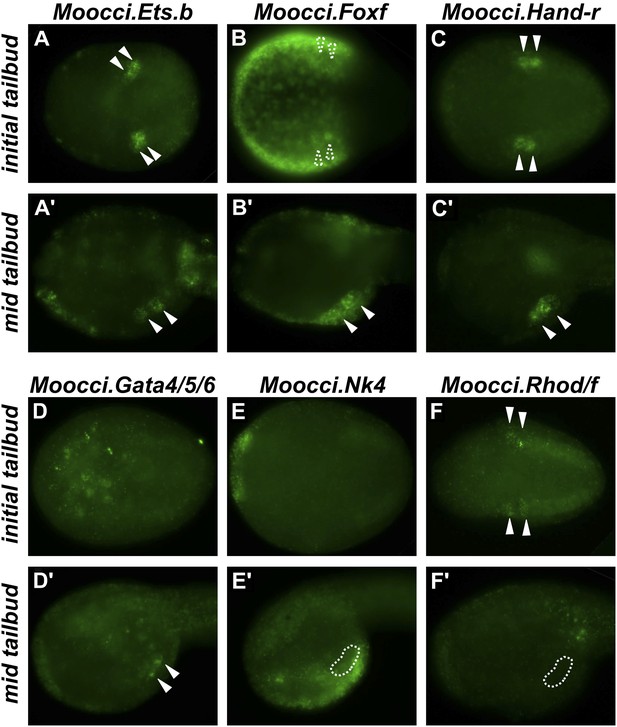
Expression of conserved TVC/heart markers in M. occidentalis embryos.
In situ hybridization (ISH) in M. occidentalis embryos for (A and A′) Moocci.Ets.b, (B and B′) Moocci.Foxf, (C and C′) Moocci.Hand-related (Moocci.Hand-r), (D and D′) Moocci.Gata4/5/6, (E and E′) Moocci.Nk4, (F and F′) Moocci.Rhod/f. ISH was performed on initial tailbud (A–F) and mid tailbud (A′–F′) stage embryos. Solid arrowheads indicate definitive expression in TVCs. Dotted arrowheads indicate potential expression of Moocci.Foxf in initial tailbud, obscured by strong epidermal expression. Dotted outline indicates probable position of TVCs, not visible due to lack of mRNA hybridization signal. Initial tailbud embryos were imaged ventrally or dorsally, while tailbud embryos were imaged laterally.
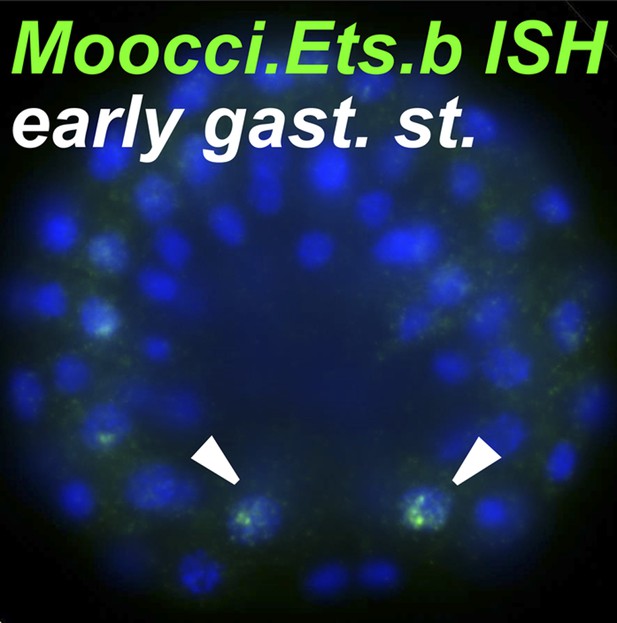
Ets.b expression in B7.5 of M. occidentalis.
In situ hybridization (ISH) for Moocci.Ets.b in 110-cell st/early gastrula embryo, showing expression (green) in the B7.5 blastomeres. Nuclei are counterstained with DAPI (blue). Embryo was imaged from a vegetal view, with anterior to the top.
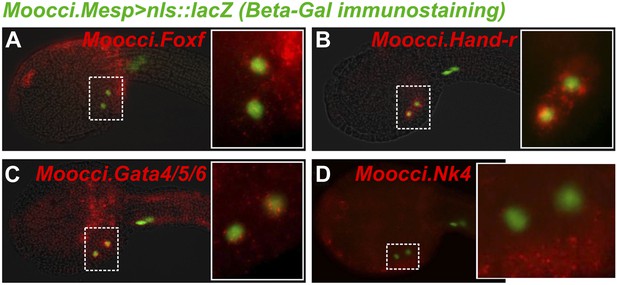
In situ hybridizations reveal TVC gene expression in M. occidentalis.
Mid-tailbud M. occidentalis embryos electroporated with Moocci.Mesp>nls::lacZ, assayed with β-galactosidase immunodetection (IHC, green) couple to in situ hybridization (ISH, red) for (A) Moocci.Foxf, (B) Moocci.Hand-r, (C) Moocci.Gata4/5/6, and (D) Moocci.Nk4. Expression of these genes was detected in the TVCs, with the possible exception of Nk4. All embryos are oriented with anterior to the left.
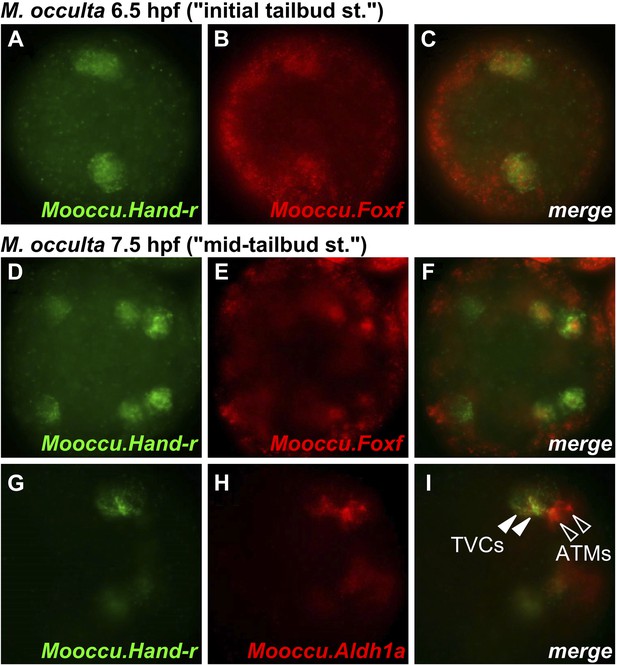
Hand-r and FoxF co-expression reveals TVCs of M. occulta.
Coronal view of M. occulta embryos at (A–C) 6.5 hpf (equivalent to the initial tailbud stage of tailed M. oculata) and (D–I) 7.5 hpf (equivalent to the mid-tailbud stage). Embryos were assayed for expression of Mooccu.Hand-r (green—A, D, G) and Mooccu.Foxf (red—B, E) or Mooccu.Aldh1a (red—H) by double in situ hybridization. Merged green and red channels show co-expression in TVCs (C, F, I). Mooccu.Foxf expression in epidermis seen in F suggests position of TVCs in ventro-lateral regions of the trunk similar to what is observed in M. occidentalis and C. intestinalis. Mooccu.Aldh1a-expressing cells immediately posterior to TVCs are likely the anterior tail muscles. Anterior is to the left in all panels.
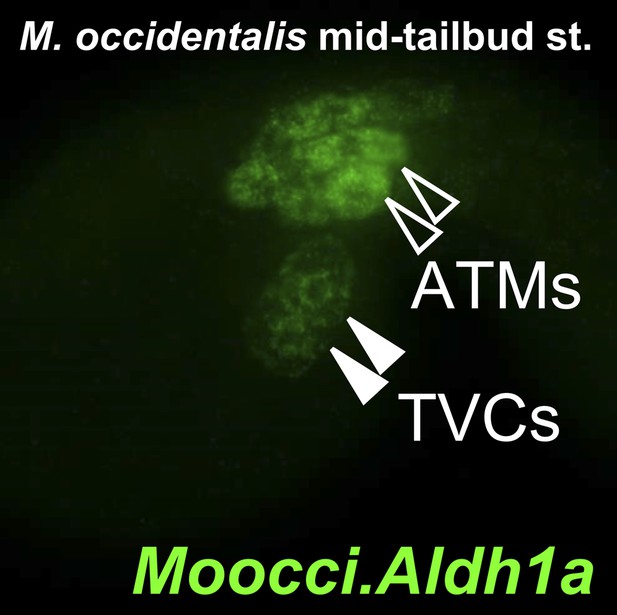

Aldh1a expression in M. occidentalis embryo.
In situ hybridization for Moocci.Aldh1a showing expression in TVCs and ATMs in a mid-tailbud embryo.
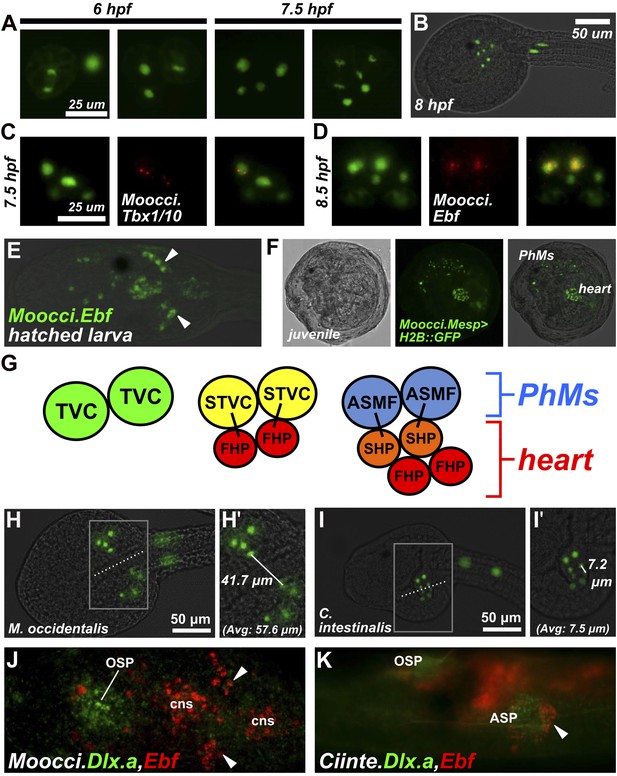
Specification of heart vs atrial siphon muscle precursors in M. occidentalis.
(A) First and second asymmetric division of TVCs in M. occidentalis embryos. B7.5 lineage was visualized by electroporation of Moocci.Mesp>H2B::GFP (green). The first division occurs at ∼6 hpf at 24°C, and the second division occurs at ∼7.5 hpf at 24°C. (B) Result of two asymmetric divisions of TVCs on left side of embryo electroporated with Moocci.Mesp>H2B::GFP (green). At 8 hpf at 24°C, a cluster of 6 cells derived from the TVCs is located in the ventro-lateral region of the trunk. From top to bottom: 2 atrial siphon muscle founder cells (ASMFs), 2 second heart precursors (SHPs), and 2 first heart precursors (FHPs). (C) In situ hybridization (ISH, red) for Moocci.Tbx1/10 + β-galactosidase immunodetection (IHC, green) in embryos electoporated with Moocci.Mesp>nls::lacZ. Moocci.Tbx1/10 nascent transcripts are detected as two dots in the nuclei of Secondary TVCs (STVCs), between the first and second asymmetric divisions. (D) ISH + IHC for Moocci.Ebf (red) in embryos electoporated with Moocci.Mesp>nls::lacZ (green), revealing Moocci.EBF expression in ASMFs after the second asymmetric division. (E) ISH for Moocci.Ebf in a swimming larva, viewed dorsally, revealing Moocci.Ebf+ migrating atrial siphon muscle precursors (ASMPs, arrowheads). (F) Lateral view of a M. occidentalis juvenile (>100 hpf) electroporated with Moocci.Mesp>H2B::GFP. GFP + nuclei reveal contributions of B7.5 lineage to atrial siphon muscle-derived pharyngeal muscles (PhMs) and heart. (G) Diagram of the TVC divisions giving rise to the pharyngeal muscles and heart of the adult. (H) Ventral view of a M. occidentalis embryo electroporated with Moocci.Mesp>H2B::GFP, just after the first asymmetric division of the TVCs. (H′) Inset from (H) is focused on the distance between FHPs on either side of the embryo (mean = 57.6 μm, n = 9). (I) Ventrolateral view of a C. intestinalis embryo electroporated with Ciinte.Mesp>H2B::GFP, right after the first asymmetric division. (I′) The distance between FHPs from either side is very small (mean = 7.5 μm, n = 14) since they contact each other at the midline to form a single cluster of cells. (J) Double ISH for Moocci.Dlx.a (green) and Moocci.Ebf (red) in a M. occidentalis larva, viewed dorsally. Moocci.Dlx.a marks the siphon primordia, while Moocci.Ebf marks siphon muscle precursors. Arrowheads point to atrial siphon muscle precursors (ASMPs), which have migrated dorsally but do not encounter an atrial siphon primordium around which to encircle. In contrast, oral siphon muscle precursors form a ring around the oral siphon primordium (OSP). Other Moocci.Ebf+ cells seen are neurons or neuronal precursors in the central nervous system (cns). (K) Double ISH for Ciinte.Dlx.a (green) and Ciinte.Ebf (red) in a C. intestinalis larva viewed dorsolaterally. Arrowhead indicates Ciinte.Ebf+ ASMPs on one side of the embryo encircling one of two bilaterally paired atrial siphon primordia (ASP). The same configuration of Ciinte.Ebf+ muscle precursors is seen encircling the OSP.
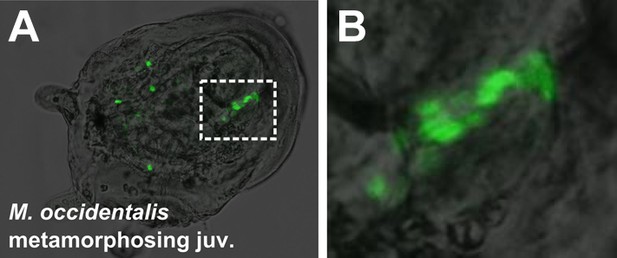
Mosaic transgene labeling of M. occidentalis suggests bilateral origin of juvenile heart.
(A) Dorso-lateral view of a metamorphosing juvenile (>100 hpf) developing from embryo electroporated with Moocci.Mesp>H2B::GFP. H2B::GFP labels nuclei of right side of and subset of atrial siphon/body-wall muscles. (B) Magnified view of boxed area in A. Mosaicism of H2B::GFP-labeling of juvenile tissues is interpreted as reflecting left/right mosaic uptake and/or retention of Moocci.Mesp>H2B::GFP plasmid during embryonic stages.
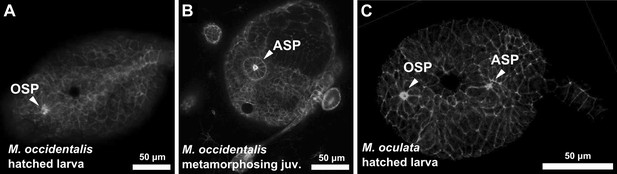
Delayed atrial siphon primordium specification in M. occidentalis.
(A) Dorsal view of hatched larva (13 hpf) stained with Alexa Fluor 546 phalloidin, showing oral siphon primordium (OSP) as a distinct structure, while the atrial siphon primordium (ASP) has not formed yet. (B) Dorsal view of metamorphosing juvenile (>72 hpf) stained with Alexa Fluor 546 phalloidin showing fully formed ASP as a rosette of apically constricted cells. OSP is out of the plane of view. (C) Dorsal view of M. oculata hatched larva stained with Alexa Fluor 633 phalloidin, showing formation of both OSP and ASP at this stage.
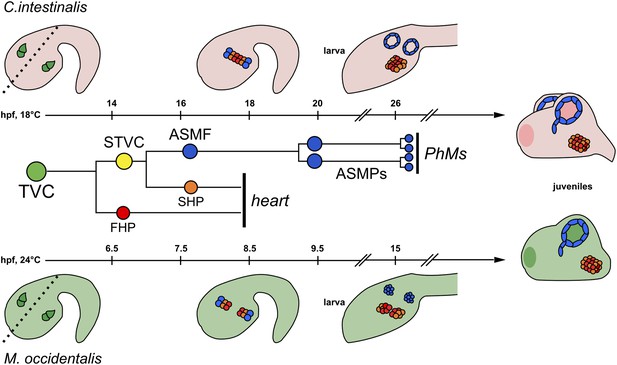
Cardiopharyngeal development in C. intestinalis vs. M. occidentalis.
Schematic diagram comparing cardiopharyngeal development in C. intestinalis (top) and M. occidentalis (bottom) embryos. The B7.5 cell lineage, which is identical in the two species, is shown in the middle, and each progenitor type is color-coded to match the embryo diagrams. Dashed line in left-most drawings indicates embryonic midline. Although both FHPs and SHPs contribute to the juvenile heart, the relative contributions of their descendants have yet to be elucidated, in either Molgula or Ciona. Refer to text for other details.
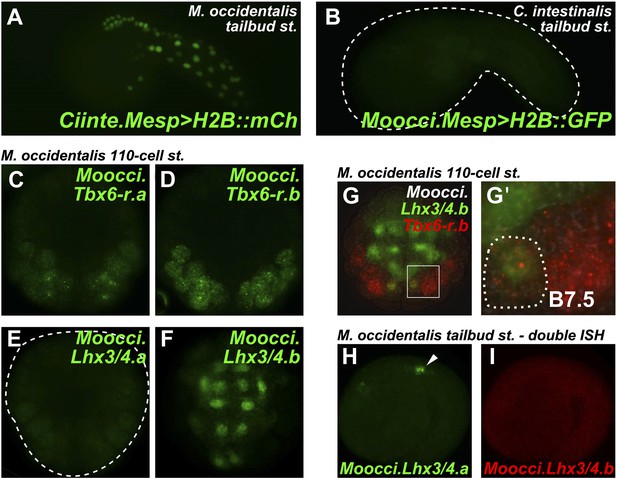
Developmental system drift of Mesp regulation between C. intestinalis and M. occidentalis.
(A) M. occidentalis tailbud embryo electroporated with a Ciinte.Mesp>H2B::mCherry reporter construct. Weak reporter gene expression was observed in the B7.5 lineage and occasionally in other territories including B-line mesenchyme and tail muscle cells, and A-line neural plate derivatives. (B) C. intestinalis tailbud embryo electroporated with Moocci.Mesp>H2B::GFP reporter. No fluorescence was seen in any cells, indicating complete lack of activity of Moocci.Mesp enhancer in wild-type C. intestinalis embryos. (C) In situ hybridization (ISH) for Moocci.Tbx6-r.a, (D) Moocci.Tbx6-r.b, (E) Moocci.Lhx3/4.a, and (F) Moocci.Lhx3/4.b in 110-cell stage embryos. (G) Double ISH in 110-cell stage embryo reveals co-expression of Moocci.Lhx3/4b (green) and Moocci.Tbx6-r.b (red) exactly in the B7.5 cells of M. occidentalis. (G′) Magnified view of inset in (G). (H) Double ISH for Moocci.Lhx3/4.a (green) and (I) Moocci.Lhx3/4.b (red) in a mid-tailbud embryo. Moocci.Lhx3/4.a but not Moocci.Lhx3/4.b is expressed in motor ganglion neurons (arrowhead).
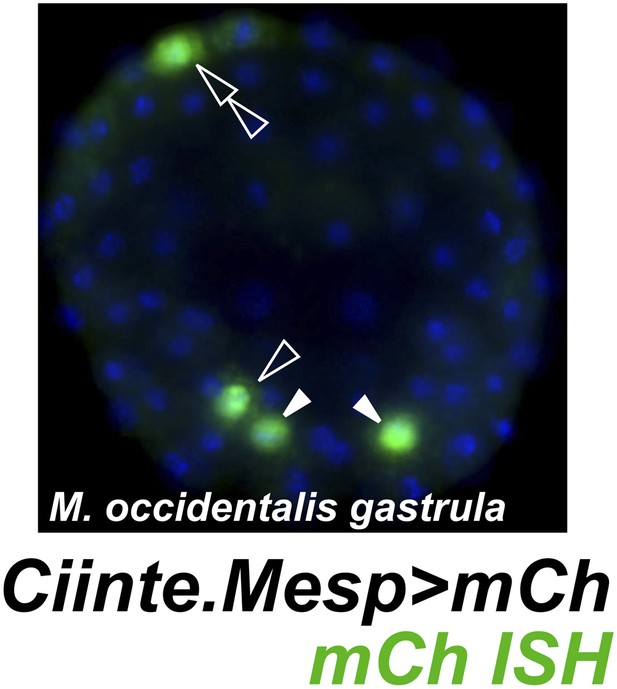
Weak and leaky expression of Ciinte.Mesp reporter in M. occidentalis embryos.
In situ hybridization for mCherry mRNA (green) in a M. occidentalis early gastrula stage embryo electroporated with Ciinte.Mesp>mCherry. Nuclei counterstained with DAPI (blue). Embryo is viewed vegetally, anterior to the top. Expression in B7.5 blastomeres (solid arrowheads) was observed in 42% of embryos. Ectopic expression in other B-line cells (hollow arrowhead) was seen in 24% of embryos. Ectopic expression in A-line neural precursors (hollow double arrowhead) was seen in 8% of embryos. In contrast, in situ hybridization revealed that Moocci.Mesp reporter construct is expressed in the TVCs in 60% of electroporated M. occidentalis embryos, with 0% embryos showing any ectopic reporter gene expression (data not shown, see Figure 1D for example). n = 50 embryos for each construct.
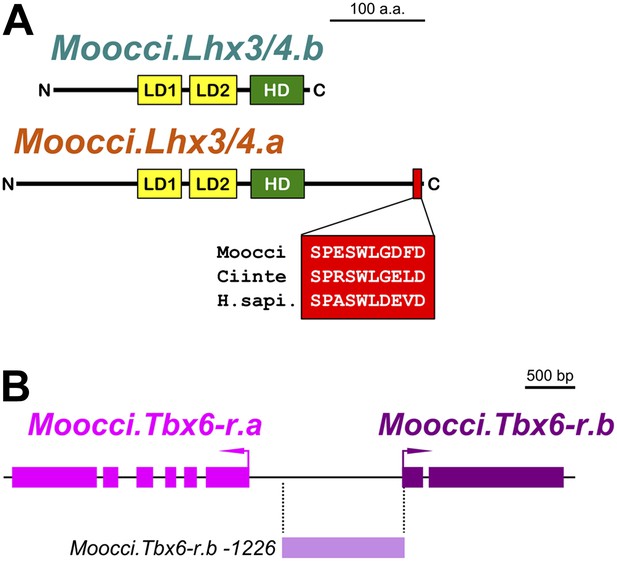
Configuration of Lhx3/4 protein domains and Tbx6-r locus in M. occidentalis.
(A) Schematic of Moocci.Lhx3/4.b and Moocci.Lhx3/4.a proteins. LIM domains (LD1 and LD2) are in yellow, and the homeodomain (HD) is in green. Moocci.Lhx3/4.a retains a more extensive C-terminus including a highly conserved motif (highlighted in red) of unknown function. Alignment to Lhx3/4 orthologs from C. intestinalis (Ciinte) and humans (H.sapi.) is shown in inset. (B) Schematic representing the Tbx6-related locus in M. occidentalis, showing head-to-head configuration of Tbx6-r.a and Tbx6-r.b. Exons are represented by thick blocks. Tbx6-r.a is encoded by 6 exons, while Tbx6-r.b is encoded by only 2 exons. The region corresponding to the Moocci.Tbx6-r.b driver used in this study is shown in periwinkle.
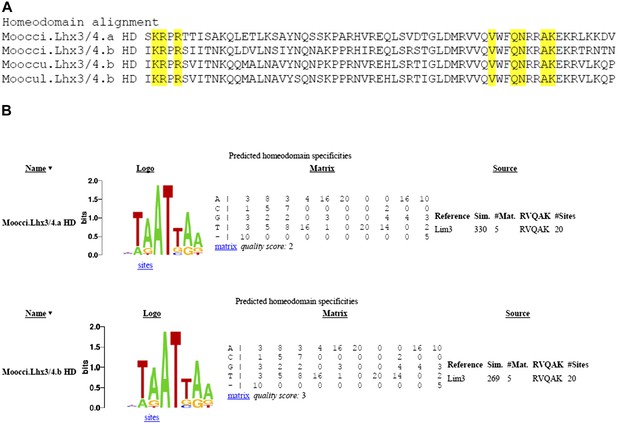
Divergent Molgula Lhx3/4.b homeodomains are not predicted to have altered DNA binding specificities.
(A) Alignment of homeodomains (HDs) from a set of Molgula Lhx3/4 family proteins, with HD recognition positions highlighted in yellow. HD recognition positions are invariant while intervening sequence is highly diverged between Lhx3/4.a and Lhx3/4.b. (B) Logos and matrices for predicted homeodmain specificities for Moocci.Lhx3/4.a (top) and Moocci.Lhx3/4.b (bottom), generated by the Homeodomain Specificity Prediction web page (http://stormo.wustl.edu/cgi-bin/flyhd/hd_pred.cgi; Noyes et al., 2008). The two predicted binding specificities are identical to one another, due to perfect conservation of the HD recognition positions.
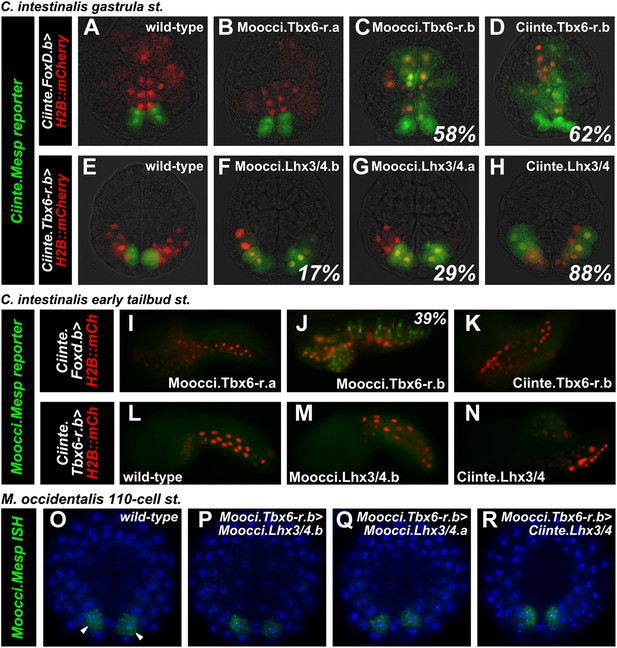
Divergence of Mesp regulation due to changes in cis and trans.
(A) Wild-type C. intestinalis gastrula embryo showing expression of Foxd.b reporter (red, Shi and Levine, 2008) in vegetal pole cells and Ciinte.Mesp reporter (green) in the B7.5 cells. (B) Electroporation of Ciinte.Foxd.b>Moocci.Tbx6-r.a has no effect on Ciinte.Mesp reporter expression. (C) Electroporation of Ciinte.Foxd.b>Moocci.Tbx6-r.b results in ectopic Ciinte.Mesp reporter expression in the vegetal pole in 58% of embryos (n = 100). (D) This is indistinguishable from the effect of Ciinte.Foxd.b>Ciinte.Tbx6-r.b, which results in ectopic Ciinte.Mesp reporter activation in 62% of embryos (n = 100). (E) Wild-type C. intestinalis gastrula embryo showing expression of Tbx6-r.b reporter (Christiaen et al., 2009a, red) in all B-line cells and Ciinte.Mesp reporter (green) in the B7.5 cells. (F) Electroporation of Ciinte.Tbx6-r.b>Moocci.Lhx3/4.b results in ectopic Ciinte.Mesp reporter expression in other B-line cells in 17% of embryos (n = 100) (G) Ciinte.Tbx6-r.b>Moocci.Lhx3/4.a results in ectopic Ciinte.Mesp reporter expression in 29% of embryos (n = 100). (H) Ciinte.Tbx6-r.b>Ciinte.Lhx3/4 induces ectopic Ciinte.Mesp reporter expression in 88% of embryos (n = 100). (I) C. intestinalis tailbud embryo showing no expression of Moocci.Mesp reporter, as expected. (J) Moocci.Mesp reporter expression is not rescued by electroporation of Ciinte.Foxd.b>Moocci.Tbx6-r.a but (K) electroporation of Ciinte.Foxd.b>Moocci.Tbx6-r.b is sufficient to activate Moocci.Mesp reporter in 39% of embryos (n = 100). (K) Moocci.Mesp reporter is not transactivated upon electroporation of Foxd.b>Ciinte.Tbx6-r.b. (L) Wild-type C. intestinalis tailbud embryo, showing no expression of electroporated Moocci.Mesp reporter. (M) Electroporation of Ciinte.Tbx6-r.b>Moocci.Lhx3/4.b or (N) Ciinte.Tbx6-r.b>Ciinte.Lhx3/4 is not sufficient to activate co-electroporated Moocci.Mesp reporter expression. (O–R) ISH showing Moocci.Mesp expression (green, arrowheads) in 110-cell stage M. occidentalis embryos. Ectopic Moocci.Mesp was not observed upon overexpression of any Lhx3/4 orthologs in the posterior embryo using the Moocci.Tbx6-r.b driver (P–R). All gastrula stage embryos are viewed vegetally, with anterior to the top of the image. All tailbud embryos are viewed laterally, anterior to the left. M. occidentalis embryo nuclei were visualized by staining with DAPI (blue).
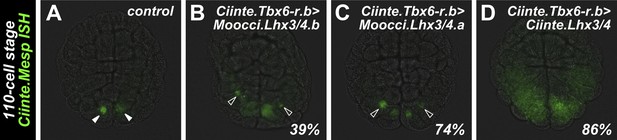
Lhx3/4 proteins from M. occidentalis are sufficient to activate ectopic expression of endogenous Mesp in C. intestinalis.
In situ hybridization for Ciinte.Mesp in 110-cell stage C. intestinalis embryos. (A) Expression of Ciinte.Mesp is restricted to the B7.5 blastomeres (solid arrowheads) in control embryos. (B) Ectopic Ciinte.Mesp expression is seen in a few cells (hollow arrowheads) in 39% (n = 100) of embryos electroporated with Ciinte.Tbx6-r.b>Moocci.Lhx3/4.b. (C) Ectopic Ciinte.Mesp expression is seen in a few cells (hollow arrowheads) in 74% (n = 100) of embryos electroporated with Ciinte.Tbx6-r.b>Moocci.Lhx3/4.a. (D) Ectopic Ciinte.Mesp expression is seen in a broad posterior swath of cells in 86% (n = 50) of embryos electroporated with Ciinte.Tbx6-r.b>Ciinte.Lhx3/4. All embryos viewed vegetally with the anterior to the top.
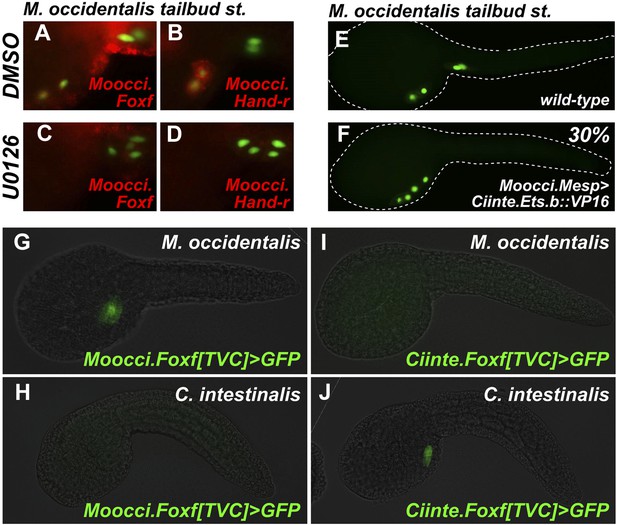
Mutual unintelligibility of orthologous Foxf enhancers in spite of shared requirement for the MAPK pathway in TVC fate specification.
In situ hybridization (ISH, red) + β-galactosidase immunodetection (IHC, green) in embryos electoporated with Moocci.Mesp>nls::lacZ and treated with DMSO (control treatment), showing normal TVC induction in 85% of embryos (n = 49) and TVC-localized expression of (A) Moocci.Foxf and (B) Moocci.Hand-r in 96% (n = 25) and 100% (n = 23) of embryos, respectively. Treatment with MEK inhibitor U0126 inhibited TVC induction and migration in 66% of embryos (n = 128). Treatment with U0126 also abolished TVC-specific (C) Moocci.Foxf expression in 91% of embryos (n = 64) and (D) Moocci.Hand-r expression in 91% of embryos (n = 64). (E) Wild-type M. occidentalis tailbud embryo with B7.5 lineage labeled by electroporation of Moocci.Mesp>H2B::GFP. Note two TVCs and two ATMs. (F) TVC induction and migration was enhanced upon electroporation with Moocci.Mesp>Ciinte.Ets.b::VP16, which converted ATMs into TVC-like cells in 30% of electroporated embryos (n = 200). (G) A fragment from −1517 to −425 upstream of the Moocci.Foxf start codon (Moocci.Foxf[TVC]), fused to the minimal promoter (−230/+21) from Moocci.Foxf and GFP (bpMoFF>GFP), recapitulates expression in the TVCs of M. occidentalis. (H) This same exact construct is silent in C. intestinalis embryos. (I) Conversely, a published TVC enhancer from the Ciinte.Foxf gene (Beh et al., 2007) fused to bpMoFF>GFP is silent in M. occidentalis embryos. (J) This same construct is strongly expressed in the TVCs of C. intestinalis. All panels represent lateral views.
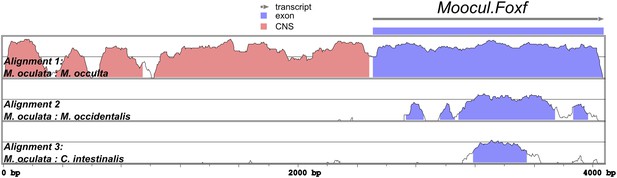
Alignment of Foxf coding and non-coding sequences.
Diagram of mVISTA alignment of M. oculata Foxf (Moocul.Foxf) locus aligned to orthologs in (from top to bottom) M. occulta, M. occidentalis, and C. intestinalis. Blue-shaded peaks indicate protein-coding sequence conservation above 70% in 100-bp windows. Red-shaded peaks indicate non-coding sequence conservation above 70% in 100-bp windows. Sequence conservation between the Roscovite Molgulids (M. oculata and M. occulta) and M. occidentalis is limited to protein-coding portions, and conservation between Molgula and Ciona is limited to a narrow portion of the sequence coding for the Foxf DNA-binding domain. Only the first exon of Foxf is shown.
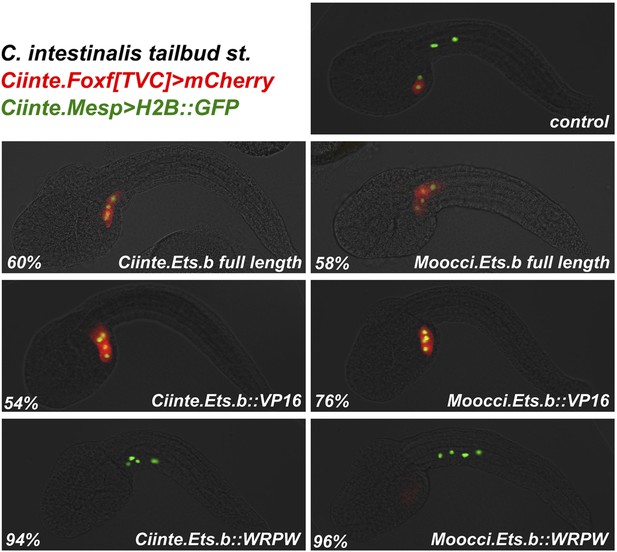

Moocci.Ets.b and Ciinte.Ets.b are functionally very similar.
C. intestinalis embryos co-electroporated with Ciinte.Mesp>H2B::GFP (green), Ciinte.Foxf>mCherry (red), and various Ciinte.Mesp-driven perturbation constructs. Percentages represent the frequency of the phenotype seen in the representative image displayed for each condition. Overexpression of full-length Ets.b proteins from both C. intestinalis and M. occidentalis results ectopic Foxf+ TVCs at the expense of ATMs. The same conversion of ATMs to TVCs is seen for overexpression of Ets.b::VP16 fusions. Conversely, Ets.b::WRPW fusions repress TVC induction and Ciinte.Foxf reporter expression. As expected from the lack of sequence divergence predicted to affect DNA-binding specificities, Ets.b proteins from both species appear to be equally suited to trans-activating Ciinte.Foxf in C. intestinalis embryos. n = 100 for all conditions.

Ets.b proteins are not sufficient to transactivate Moocci.Foxf reporter construct activation in C. intestinalis embryos.
C. intestinalis embryos co-electroporated with Ciinte.Mesp>H2B::GFP (green), Moocci.Foxf>mCherry (red), and various Ciinte.Mesp-driven perturbation constructs. Activation of Moocci.Foxf reporter construct was impervious in C. intestinalis TVCs even upon overexpression of Ets.b full-length or VP16 fusions from either C. intestinalis or M. occidentalis. n = 100 for all conditions.
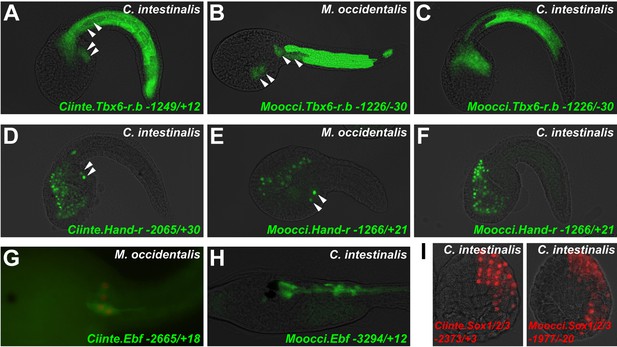
Cross-species reporter construct assays reveal multiple cases of cis-regulatory unintelligibility.
(A) C. intestinalis embryo electroporated with Ciinte.Tbx6-r.b>GFP reporter construct (Christiaen et al., 2009a), which drives GFP expression in tail muscles and the B7.5 lineage cells (arrowheads), recapitulating endogenous Ciinte.Tbx6-r.b expression. (B) M. occidentalis embryo electroporated with Moocci.Tbx6-r.b>GFP reporter construct, which recapitulates expression in tail muscle cells including B7.5 lineage cells (arrowheads). (C) C. intestinalis embryo electroporated with Moocci.Tbx6-r.b>GFP, which drives expression in B-line tail muscle and mesenchyme cells but is excluded from the B7.5 lineage. (D) C. intestinalis embryo electroporated with Ciinte.Hand-r>H2B::GFP reporter (Davidson and Levine, 2003), which drives H2B::GFP expression in anterior endoderm, A7.6 lineage, and TVCs (arrowheads), recapitulating endogenous Ciinte.Hand-r expression. (E) M. occidentalis embryo electroporated with Moccci.Hand-r>H2B::GFP construct, which recapitulates the same expression pattern. (F) C. intestinalis embryo electroporated with Moocci.Hand-r>H2B::GFP, which drives expression in endoderm and A7.6 lineage, but not in B7.5. (G) M. occidentalis embryo electroporated with Ciinte.Ebf neuron-specific YFP (green) and H2B::mCherry (red) reporter constructs electroporated (Stolfi and Levine, 2011), which drive very weak expression in a limited subset of motor ganglion neurons (green and red). (H) C. intestinalis embryo electroporated with a Moocci.Ebf>YFP reporter, which drives robust reporter gene expression in several brain, motor ganglion, and nerve cord neurons. (I) C. intestinalis embryos electroporated with Ciinte.Sox1/2/3 (left) and Moocci.Sox1/2/3 (right) H2B::mCherry reporter constructs, both of which recapitulate Sox1/2/3 expression in ectoderm. Panels A–F are lateral views of tailbud embryos, panels G is a dorsal view of a tailbud embryo, panel H is a dorso-lateral view of a hatched larva, and panel I shows vegetal views of mid-gastrula stage embryos. Anterior is to the right, except in Panel I, in which anterior is to the top.
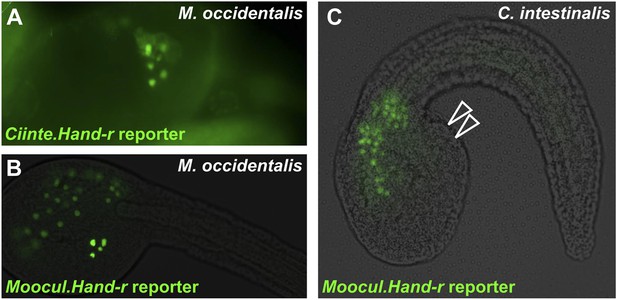
Asymmetric intelligibility between Molgula and Ciona Hand-r TVC enhancers.
(A) β-galactosidase immunodetection on M. occidentalis late tailbud embryo electroporated with Ciinte.Hand-r>nls::lacZ., showing reporter gene expression in TVC descendents (heart precursors and atrial siphon muscle founder cells) on one side of the embryo. (B) M. occidentalis embryo electroporated with Moocul.Hand-r>H2B::GFP, showing recapitulation of endogenous Moocci.Hand-r expression in endoderm, A7.6 lineage, and TVCs. (C) C. intestinalis embryo electroporated with Moocul.Hand-r>H2B::GFP showing slight expression in A7.6 lineage and B-line mesenchyme cells, but no expression in TVCs (hollow arrowheads).
Tables
Genome assembly statistics
| Species | N50 | Mean contig length | Total | Total number of base pairs | CEGMA C1 | CEGMA P2 |
|---|---|---|---|---|---|---|
| M. occidentalis | 26,298 | 5072 | 51,761 | 262,547,660 | 81.45 | 96.77 |
| M. occulta | 13,011 | 3233 | 58,489 | 189,110,562 | 77.42 | 98.79 |
| M. oculata | 34,042 | 6270 | 25,497 | 159,886,716 | 89.92 | 99.19 |
-
The contig N50 length, mean contig length, total number of contigs, total number of base pairs and CEGMA scores were collected for each draft assembly. The CEGMA scores is a metric of completeness measured against highly Conserved eukaryotic genes. Alignments of 70% or greater of the protein length are called complete (C1) and all other statistically significant alignments are called partial (P2).
Additional files
-
Supplementary file 1
DNA sequences of probes, enhancers, promoters, protein-coding cDNAs, in situ hybridization probe templates, primers, etc used in this study.
- https://doi.org/10.7554/eLife.03728.029
-
Supplementary file 2
Table of newly proposed tunicate gene names and symbols (Stolfi et al., 2014), and their aliases and synonyms.
- https://doi.org/10.7554/eLife.03728.030





