Semaphorin 5A inhibits synaptogenesis in early postnatal- and adult-born hippocampal dentate granule cells
Figures
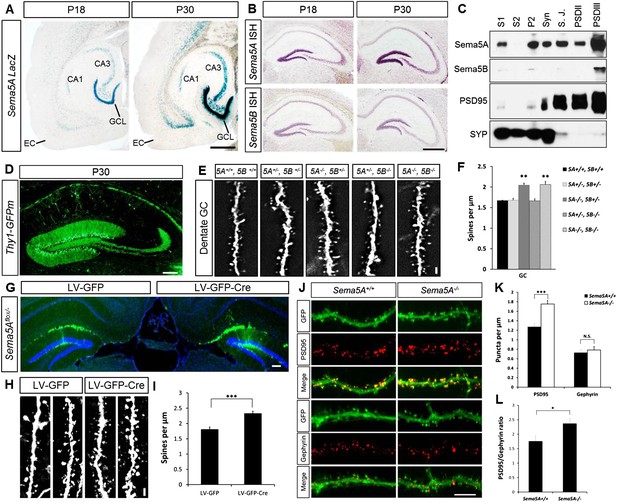
Sema5A, but not Sema5B, negatively regulates dendritic spine density of hippocampal GCs in vivo.
(A) Horizontal sections through the hippocampus of Sema5ALacZ/+ brains show robust β-gal activity in the GCL at P18 and P30. Weaker labeling is observed in the pyramidal cell layer of CA3 and a small segment of CA1. In the EC, labeling is confined to deep cortical layers. (B) In situ hybridization of P18 and P30 coronal sections with probes specific for Sema5A and Sema5B. (C) Western blot analysis of synaptic density fractions prepared from P18 mouse hippocampus. S1, homogenate; S2, cytosolic fraction; P2, membrane fraction; Syn, synaptosomal fraction; S.J., synaptic junction, and PSD purified postsynaptic fractions. Anti-PSD95 and anti-Synaptophysin (SYP) are shown as post- and pre-synaptic markers. (D) Coronal section of the P33 Thy1-GFPm hippocampus showing labeling of GCs. (E) Representative images of GFP-positive dendrites of GCs of WT (5A+/+, 5B+/+), Sema5A+/−, Sema5B+/− (5A+/−, 5B+/−); Sema5A−/−, Sema5B+/− (5A−/−, 5B+/−); Sema5A+/−, Sema5B−/− (5A+/−, 5B−/−) and double mutant (5A−/−, 5B−/−) mice. (F) Quantification of dendritic spine density of GCs shown in E. Values are represented as mean ± SEM from 3 to 4 mice per genotype (for details on spine quantification see Table 1). **indicates p < 0.01, two-tailed unpaired Student's t test. (G) Representative image of the DG following stereotaxic injection of LV-GFP (left side) or LV-syn-GFP-IRES-Cre (right side) viral vector into Sema5Aflox/− mice. (H) High magnification images of LV transduced GC dendrites. (I) Quantification of spine density of GFP+ GC dendrites. Values are represented as mean ± SEM from three independent mice per condition. (J) Cultured mouse hippocampal neurons at DIV21 obtained from Sema5A+/+ and Sema5A−/− pups. Cultures were transfected at DIV4 with a GFP expression construct and fixed at DIV21. GCs were identified by anti-Prox1 labeling (data not shown). Cultures were stained with anti-PSD95 or anti-gephyrin to identify excitatory synapses confined to dendritic spines, and inhibitory synapses confined to the dendritic shaft of GCs. (K) Quantification of PSD95 positive puncta reveals a significant increase in Sema5A−/− GCs. No significant (N.S.) difference in gephyrin positive puncta was observed between Sema5A+/+ and Sema5A−/− GCs. (L) The ratio of excitatory/inhibitory synapses is significantly increased in Sema5A−/− GCs. Number of neurons quantified: n = 32–34 neurons per condition from three mice per genotype. Values are represented as mean ± SEM. ***p < 0.001; *p < 0.05 two-tailed unpaired Student's t test. Scale bars, A and B = 500 µm, D and G = 200 µm, J = 10 µm, E and H = 1 µm.
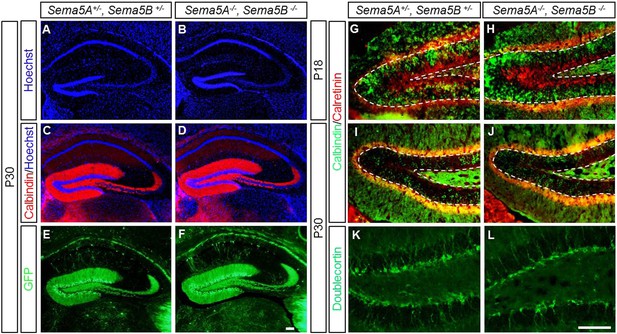
Normal maturation and patterning of dentate GCs in Sema5A−/− ; Sema5B−/− double mutants.
(A and B) Coronal sections of age-matched (P30) double-heterozygous and double-mutant Sema5A, Sema5B brains at the level of the rostral hippocampus. Tissue sections were stained with the nuclear dye Hoechst 33,342 to assess gross morphology of the dentate gyrus and the CA1 and CA3 pyramidal cell layers. (C and D) At P30, anti-calbindin immunolabeling revealed no obvious differences in GC axonal and dendritic processes between Sema5A+/−; Sema5B+/− and Sema5A−/−; Sema5B−/− mice. n = 3 mice per genotype. (E–F) Sema5 mice were intercrossed with the Thy1-GFPm reporter mouse. Consistent with anti-calbindin labeling, no obvious defects in GC dendritic protrusions, axonal targeting or pruning to the CA3 subfield was observed. n = 3–4 mice per genotype. (G–J) Double immunofluorescence labeling of calbindin (green), a marker for mature GCs, and calretinin (red), a marker for immature GCs. (G and H) Coronal sections of P18 DG and (I and J) P30 DG showed no differences in labeling between Sema5A+/−; Sema5B+/− and Sema5A−/−; Sema5B−/− mice. Dashed lines demarcate the inner and outer borders of the dentate granule cell layer. (K and L) P30 DG coronal section of Sema5A+/−; Sema5B+/− and Sema5A−/−; Sema5B−/− mice immunostained for doublecortin (DCX), a microtubule-associated protein expressed in immature GCs during migration. No differences in abundance or distribution of DCX positive cells were observed between the different genotypes. n = 3 mice per genotype. Scale bars = 100 µm (A–F) and for (G–L).
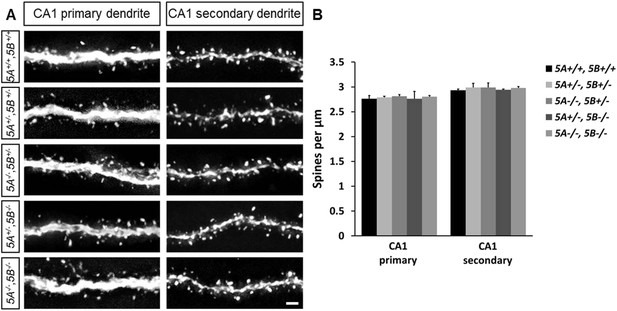
Sema5 mutant mice do not show altered spine density in CA1 pyramidal neurons.
(A) Representative images of GFP-labeled primary and secondary dendrites of CA1 pyramidal neurons of P30-33 WT (5A+/+, 5B+/+), Sema5A+/−, Sema5B +/− (5A+/−, 5B+/−); Sema5A−/−, Sema5B+/− (5A−/−, 5B+/−); Sema5A+/−, Sema5B−/−(5A+/−, 5B−/−) and double mutant (5A−/−, 5B−/−) mice. (B) Quantification of dendritic spine density for conditions shown in A. Values are represented as mean ± SEM from 3 to 4 mice per genotype (for details on spine quantification see Table 1). Two-tailed unpaired Student's t test. Scale bar = 1 µm.
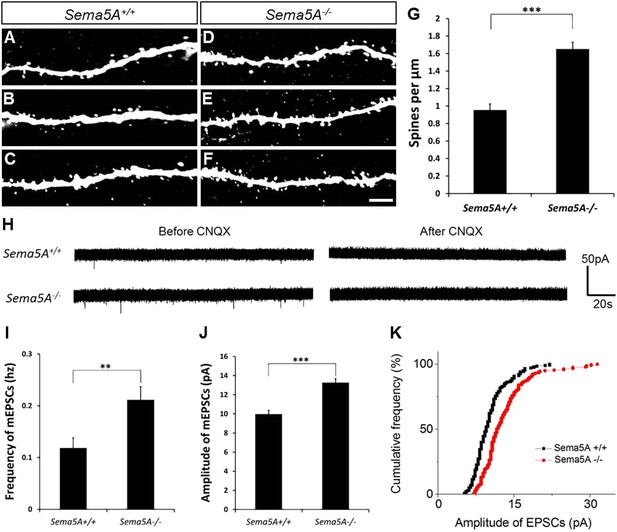
In adult-born GCs, Sema5A negatively regulates dendritic spine density and affects their electrophysiological properties.
(A–F) Representative images of dendritic segments of adult-born, 19- to 21-day-old retrovirally-labeled GCs in 3-month-old Sema5A+/+ and Sema5A−/− mice. (G) Quantification of dendritic spine density in adult-born GCs of Sema5A−/− (n = 34 neurons) and Sema5A+/+ (n = 35 neurons) mice. Error bars represent ± SEM, n = 3 independent animals per genotype. ***indicates p < 0.0001, two-tailed unpaired Student's t test. Scale bar = 5 µm. (H–K) Recordings of spontaneous mEPSCs from retrovirally-labeled adult-born GCs at 19–21 days postmitosis. (H) Shown are sample voltage-clamp whole-cell recording traces (Vm = −65 mV) from Sema5A+/+ (top) and Sema5A−/− (bottom) animals before and after bath application of CNQX (20 μM). (I) Quantification of AMPA-type mEPSC mean frequency and (J) mean amplitude. (K) Relative cumulative frequency distribution of mEPSC amplitudes in Sema5A+/+ (black) and Sema5A−/− (red) GCs. The distribution of mEPSC amplitudes in mutants is significantly shifted to the right, indicating an increase in the proportion of larger amplitude events. Recordings were performed in the presence of TTX (1 μM) and bicuculline (10 μM). Values are represented as mean ± SEM. Sema5A+/+, n = 13 cells/6 mice and Sema5A−/−, n = 13 cells/6 mice. **p < 0.01, ***p < 0.001, Student's t test.
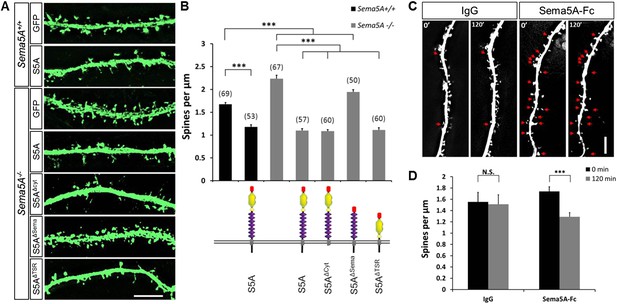
Sema5A inhibits dendritic spine density in primary dentate GCs.
(A) High power deconvolution images of dendritic segments of GCs from WT (Sema5A+/+) and mutant (Sema5A−/−) hippocampal cultures transfected either with eGFP only, or eGFP plus full-length Sema5A (S5A), the Sema5A deletion constructs S5AΔcyt, S5AΔSema, or S5AΔTSR plasmid DNA. (B) Quantification of dendritic spine density of DIV21 prox1+ GCs. Numbers in brackets indicate the number of cells analyzed per condition. (C) Time-lapse study of Sema5A-Fc induced GC dendritic spine loss. DIV21 dentate cultures were treated for 120 min with Sema5A-Fc or control IgG. (D) Quantification of dendritic spine density in prox1+ GCs. Values are represented as mean ± SEM from 3 to 4 cultures, established from 3 to 4 different animals for each condition. ***p < 0.001, two-tailed unpaired Student's t test. Scale bars in A and C = 10 µm.
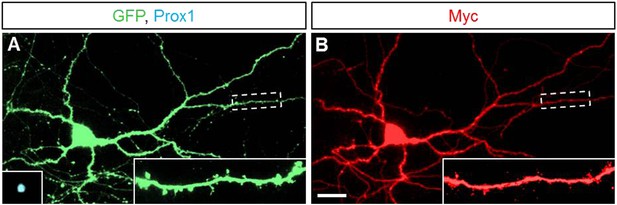
Sema5A is localized to neurites in dissociated hippocampal neurons.
(A and B) Representative images of primary hippocampal neurons transfected with GFP and 6x-myc-Sema5A plasmid DNA. At DIV21, cultures were labeled with anti-GFP (green), anti-prox1 (blue, small insert in A), and anti-myc (red).
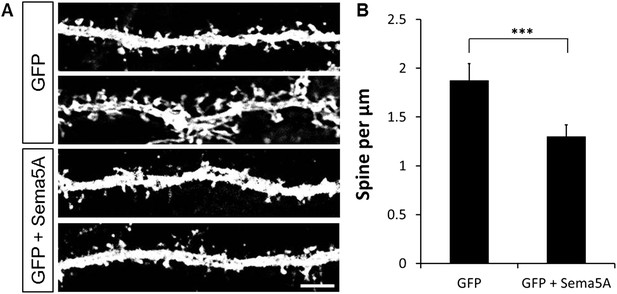
Ectopic expression of Sema5A in hippocampal pyramidal neurons decreases dendritic spine density.
(A) Representative images of dendritic segments from prox1-negative pyramidal neurons in WT hippocampal cultures at DIV21, transfected either with GFP only, or GFP plus full-length Sema5A (S5A) expression constructs. (B) Quantification of spine density revealed a significant reduction in pyramidal neurons expressing full-length Sema5A. Values are represented as mean ± SEM from three cultures, established from three different animals, for each condition. ***p < 0.001, two-tailed unpaired Student's t test. Scale bar = 10 µm.
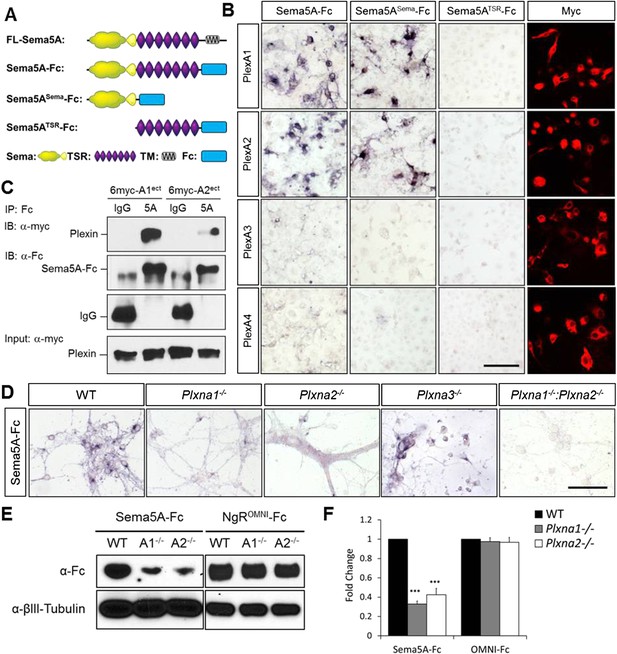
Direct binding of Sema5A to PlexA1 and PlexA2.
(A) Schematic diagrams of Sema5A domain deletion constructs used for binding studies. (B) Binding of different Sema5A-Fc fusion proteins to COS7 cells transiently expressing myc-tagged PlexA1, PlexA2, PlexA3, or PlexA4. Sema5A-Fc and Sema5ASema-Fc bind selectively to PlexA1 and PlexA2. PlexA expression was confirmed by anti-myc immunostaining. (C) Co-precipitation of purified ectodomain of Sema5A (Sema5A-Fc) with the ectodomains of PlexA1 (6x-myc-PlexA1ect) or PlexA2 (6x-myc-PlexA2ect). Immunoprecipitation (IP) with anti-Fc shows that Sema5A-Fc (5A), but not control IgG, forms a complex with the ectodomains of PlexA1 and PlexA2. The precipitated plexin fragments can be detected by anti-myc immunoblotting (top blot). Immunoblotting with an antibody specific for the Fc fragment detects Sema5A-Fc and control IgG in the precipitate (second blot from top). Note that the higher molecular weight band is Sema5A-Fc and the lower molecular weight band is intact (not fully reduced ∼150-kDa) IgG. Anti-IgG immunoblotting shows that control IgG at 50-kDa was successfully pulled down with protein A/G agarose-beads (third blot from top). To demonstrate that equal amounts of plexinA ectodomain were used for the IP experiment, an aliquot of the input was probed with anti-myc (blot at bottom). (D) Binding of Sema5A-Fc to primary hippocampal neurons established from WT, Plxna1, Plxna2 and Plxna3 single null mice as well as Plxna1; Plxna2 double null mice. (E) Hippocampal neurons from WT, Plxna1−/−, and Plxna2−/− mice were incubated with either Sema5A-Fc (50 nM) or NgROMNI-Fc (50 nM) for 1 hr. Cells were rinsed extensively, lysed, and subjected to WB analysis. Sema5A-Fc binding to Plxna1−/− and Plxna2−/− neurons is reduced compared to WT neurons. NgROMNI-Fc binding was strong to hippocampal neurons and independent of the Plxna genotype. Anti-βIII tubulin in lysates is shown as a loading control. (F) Quantification of Western blot signals from three independent experiments. Scale bars in B and D = 50 µm.
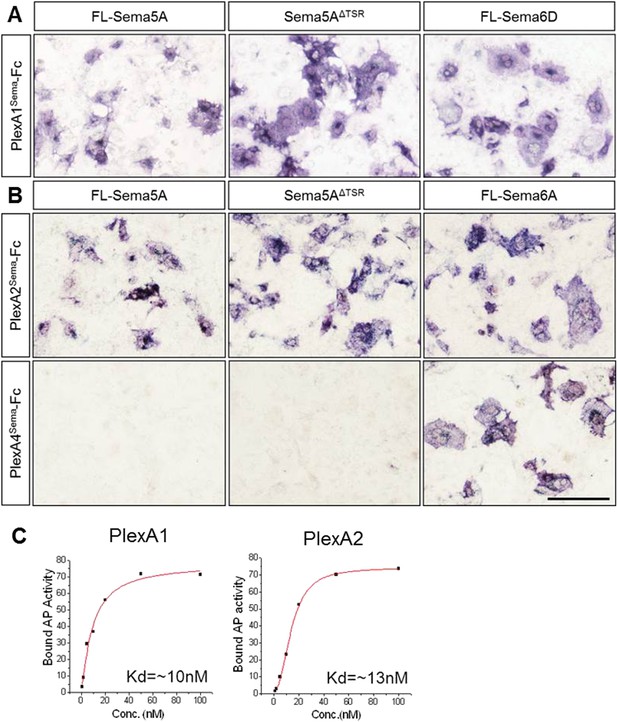
Molecular basis of PlexA1 and PlexA2 binding to Sema5A.
Full-length Sema5A (FL-Sema5A), a membrane bound Sema5A deletion construct that lacks the seven thrombospondin type-1 repeats (Sema5AΔTSR), or class 6 semaphorins, were expressed in transiently transfected COS7 cells and assayed for binding of soluble PlexASema-Fc fusion protein oligomerized with an anti-Fc antibody conjugated to human alkaline phosphatase. (A) The sema-domain of PlexA1 (PlexA1Sema-Fc) binds to membrane bound FL-Sema5A, Sema5AΔTSR, and FL-Sema6D. (B) PlexA2Sema-Fc, but not PlexA4Sema-Fc, binds to Sema5A and SemaS5AΔTSR expressed on the surface of COS7 cells. Sema6D is shown as a positive control for PlexA1Sema-Fc binding, and Sema6A is shown as a positive control for PlexA2Sema-Fc and PlexA4Sema-Fc binding. Collectively, these binding studies reveal that the sema-domain of Sema5A interacts with the soluble sema domains of PlexA1 and PlexA2, but not PlexA4. (C) Saturation binding curve of Sema5A-Fc binding to COS7 cells expressing PlexA1 or PlexA2. For quantification of binding, Sema5A-Fc was oligomerized with anti-Fc conjugated to human placental alkaline phosphate. The calculated dissociation constants (KD) for the Sema5A/PlexA1 and the Sema5A/PlexA2 interactions are 10 nM and 13 nM, respectively. Scale bar = 50 μm.
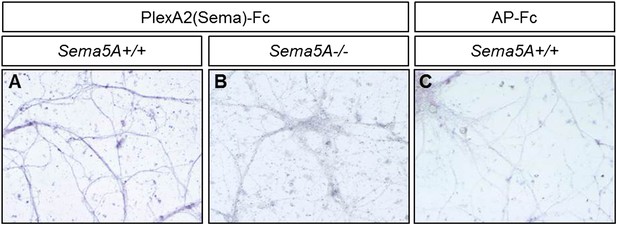
Sema5A can be detected on the neuronal surface and is localized to neurites.
(A–C) Visualization of endogenously expressed Sema5A on the surface of DIV14 dissociated dentate gyrus cultures. (A) PlexA2Sema-Fc strongly binds to Sema5A+/+ cultures (B) but not to Sema5A−/− cultures. (C) AP-Fc binding to Sema5A+/+ cultures was used as a control.
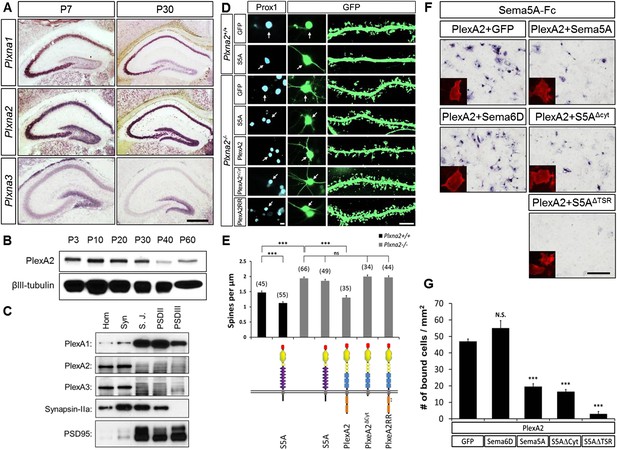
In dentate GCs, PlexA2 is necessary for Sema5A-mediated regulation of dendritic spine density.
(A) Coronal sections of P7 and P30 mouse hippocampus labeled with riboprobes specific for Plxna1, Plxna2, and Plxna3 transcripts. In the DG, strong expression of Plxna2 is observed, while Plxna1 is virtually absent. Plxna3 mRNA expression is largely confined to more immature GCs located near the inner surface of the GCL. (B) Developmental time course of PlexA2 protein expression in the mouse hippocampus between P3 and P60, normalized to βIII-tubulin. (C) Distribution of PlexA1, PlexA2, and PlexA3 in synaptic density fractions prepared from P23 mouse hippocampus. The three PlexA family members are present in the synaptosomal (syn) fraction. PlexA1, but not PlexA2 or PlexA3, is enriched in the synaptic junction (S.J.) and postsynaptic density (PSD) fractions. As controls, markers enriched in postsynaptic fractions (anti-PSD95) and presynaptic fractions (anti-Synapsin-IIa) are shown. n = 4 independent experiments. (D) Representative images of DIV21 GCs from Plxna2+/+ and Plxna2−/− mice transfected with GFP alone or GFP plus myc-tagged FL-Sema5A (S5A), FL-PlexA2 (PlexA2), PlexA2Δcyt, or PlexA2RR expression constructs. Cultures were fixed and stained with anti-Prox1 (blue) and anti-GFP (green). The cell bodies of GFP-expressing GCs are labeled with an arrow and a dendritic segment of the same neuron is shown. (E) Quantification of spine density of GCs shown in panel D. WT neurons (black bars) and Plxna2−/− mutant neurons (grey bars). Numbers in brackets indicate GCs analyzed per condition. Values are represented as mean ± SEM from three animals for each condition. (F) Cis interaction between Sema5A and PlexinA2. Membrane-bound Sema5A presented in cis, and soluble Sema5A-Fc presented in trans, compete for binding to PlexinA2. Sema5A-Fc binding to COS7 cells transiently transfected to co-express PlexA2 and GFP, PlexA2 and Sema6D, PlexA2 and Sema5A, PlexA2 and Sema5A lacking the cytoplasmic domain (S5AΔcyt), or PlexA2 and Sema5A lacking the seven thrombospondin type-1 domains (S5AΔTSR). Prior to binding, Sema5A-Fc was super-clustered with anti-Fc conjugated to human placental alkaline phosphatase (Robak et al., 2009). Surface expression of recombinant PlexA2 was confirmed in a parallel experiment by immunofluorescence labeling of its N-terminal myc epitope (small inserts). (G) Quantification of Sema5A-Fc bound to COS7 cells co-expressing PlexA2 together with GFP, Sema6D, Sema5A, S5AΔcyt, or S5AΔTSR. Cis expression of Sema5A constructs that harbor the sema domain significantly attenuates the binding of bath applied Sema5A-Fc to PlexA2. Error bars represent SEM from three independent experiments. ***indicates p < 0.0001. Two-tailed unpaired Student's t test. Scale bar in A = 50 µm.
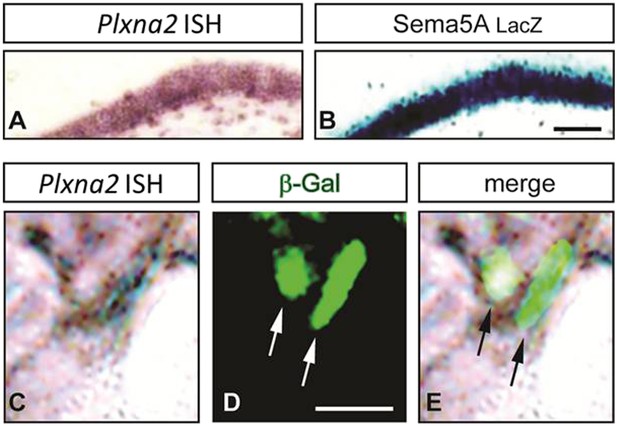
Sema5A and PlexA2 are co-expressed in GCs and interact in cis and in trans.
(A and B) Coronal sections of adult Sema5AlacZ/lacZ mice were subjected to staining for Plxna2 mRNA (purple) or nuclear β-galactosidase (blue) by X-gal. (C–E) High magnification view of the dorsal DG. Overlapping labeling for Plxna2 mRNA and nuclear β-galactosidase was observed in GCs. Scale bars in A and B = 500 μm.
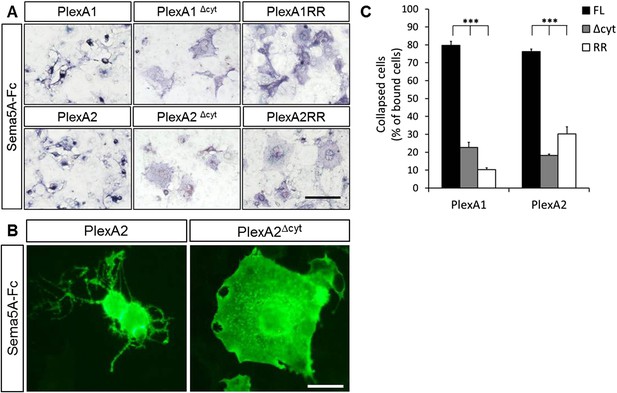
Sema5A-Fc–induced collapse of PlexA1- and PlexA2- expressing COS7 cells requires PlexA rasGAP activity.
(A) COS7 cells were transiently transfected to express full-length (FL) PlexA1, PlexA2, deletion mutants lacking their cytoplasmic domains (PlexA1Δcyt, PlexA2Δcyt), or point mutants selectively deficient for the rasGAP activity (PlexA1RR and PlexA2RR) that resides in the PlexA cytoplasmic portion. Cells were then incubated with Sema5A-Fc oligomerized with anti-Fc conjugated to placental alkaline phosphatase. Sema5A-Fc induces collapse of PlexA1- and PlexA2-expressing COS7 cells. Deletion of the PlexA1 or PlexA2 cytoplasmic domain, or mutation of the rasGAP activity in either plexin, does not impair binding of Sema5A-Fc but attenuates COS7 cell collapse. (B) All PlexA constructs contain an N-terminal myc-tag. High magnification anti-myc labeling of PlexA2 and PlexA2Δcyt transfected cells treated with Sema5A-Fc. Immunolabeling was carried out under non-permeable conditions. (C) Quantification of the percentile of Sema5A-Fc binding cells that show a collapsed morphology. Values are represented as mean ± SEM from three independent experiments. p value ***p < 0.0001, two-tailed unpaired Student's t test. Scale bar = 50 μm.
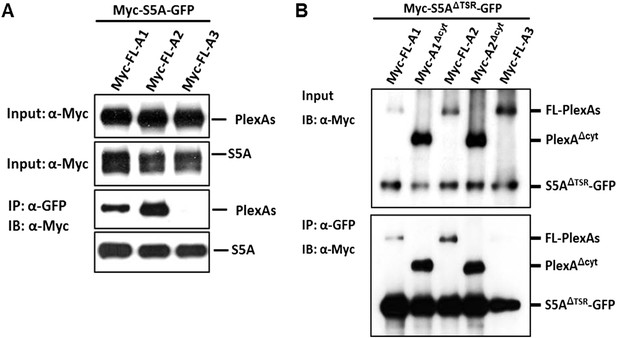
Molecular basis of Sema5A cis interaction with PlexA1 and PlexA2.
(A) Cis interaction between PlexA1 or PlexA2 with Sema5A. Co-expression of Sema5A-eGFP (Myc-S5A-GFP) with myc-tagged full-length (FL) expression constructs of PlexA family members (Myc-FL-A1, Myc-FL-A2, or Myc-FL-A3) in HEK293T cells allows for co-immunoprecipitation (co-IP) of Sema5A with PlexA1 and PlexA2, but not PlexA3. (B) Myc-tagged Sema5A lacking the seven thrombospondin type1 repeats was C-terminally fused to GFP (Myc-S5AΔTSR-GFP) and co-transfected into HEK293T with FL PlexA1, PlexA2, PlexA3, or PlexA1 and PlexA2 deletion constructs lacking the cytoplasmic domain (Myc-FL-A1Δcyt or Myc-FL-A2Δcyt). Immunoprecipitation with anti-GFP followed by anti-myc immunoblotting (IB) demonstrates an interaction between the Sema5A sema domain and the PlexA1 and PlexA2 ectodomains, but not the PlexA3 ectodomain.
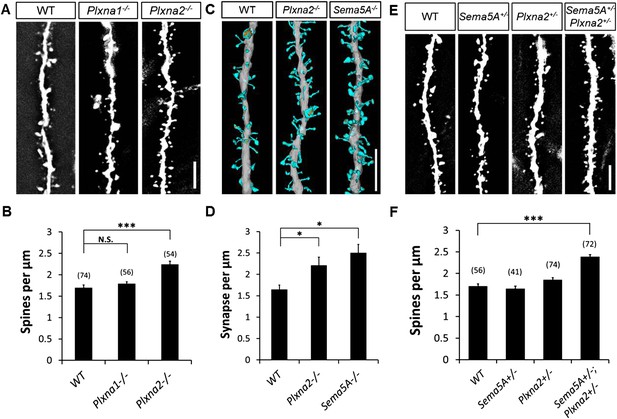
Sema5A and Plxna2 genetically interact, and Plxna2−/− and Sema5A−/− mice show increased synapse density in dentate GCs.
(A) Representative images of outer dendritic segments of GCs from P30 Thy1-eGFPm WT, Plxna1−/− and Plxna2−/− hippocampal sections. (B) Quantification of GC dendritic spine density reveals a significant increase in Plxna2−/− mice compared to Plxna1−/− and WT controls. Numbers in brackets indicate the dendritic segments analyzed per condition. Values are represented as mean ± SEM from three to four animals of each genotype. ***p < 0.001, two-tailed unpaired Student's t test. (C) Three-dimensional reconstruction of GC dendritic segments from P32 WT, Plxna2−/− and Sema5A−/− mice. The dendritic shaft is colored in gray, dendritic spines in blue and the PSDs in orange. A total of 110, 208, and 165 spines were reconstructed and analyzed for WT, Plxna2−/− and Sema5A−/− mice, respectively (n = 1 animal per genotype). (D) Quantification of spine density. Values are represented as mean ± SEM from three to four dendritic segments (each 20 to 22 µm in length) per genotype, *p < 0.05, two-tailed unpaired Student's t test. (E) Representative images of GC dendritic segments of P30 Thy1-eGFPm WT, Sema5A+/−, Plxna2+/−, and compound heterozygous Sema5A+/−; Plxna2+/− mice. (F) Quantification of GC spine densities of genotypes in E reveals significantly more spines in compound heterozygotes than WT and single heterozygotes. Numbers in brackets indicate the number of dendrites analyzed per condition. Values are represented as mean ± SEM from four animals per genotype. ***p < 0.0001, two-tailed unpaired Student's t test. Scale bars in A, C and D = 5 µm.
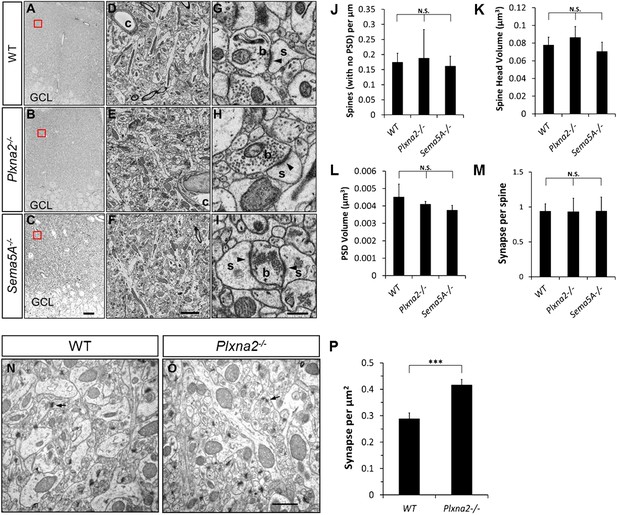
Ultra-structural analysis of the Plxna2−/− and Sema5A−/− dentate gyrus molecular layer (ML).
The rostral DG of P32 mice was subjected to serial block-face scanning electron microscopy (sEM). (A, D and G) Coronal sections show part of the granule cell layer (GCL) and the entire width of the ML of the hippocampus of WT (A), Plxna2−/− (D) and Sema5A−/− mice (G). The area boxed in red is shown at higher magnification in panels B and C for WT mice, panels E and F for Plxna2−/− mice, and panels H and I for Sema5A−/− mice. (B, C, E, F, H and I) Examples of higher magnification images of dentate ML, c = capillary. Examples of spine synapses composed of a dendritic spine (s) with a large head, a post-synaptic density (PSD, arrowhead) and a presynaptic bouton (b) with neurotransmitter vesicles. For 3D reconstruction, dendritic segments (each 20–22 µm in length) and their spines were traced in individual sEM images. (J) The number of dendritic protrusions that do not show a PSD is very similar among the three genotypes: WT (0.17 ± 0.03 spines/µm), Plxna2−/− (0.19 ± 0.09 spines/µm) and Sema5A−/− (0.16 ± 0.03 spines/µm) mice. (K) Quantification of spine head volume of WT (0.08 ± 0.01 µm3), Plxna2−/− (0.09 ± 0.01 µm3), and Sema5A−/− (0.07 ± 0.01 µm3) GCs did not reveal a significant difference. (L) Quantification of the surface area of the PSD revealed no significant differences between WT (0.06 ± 0.009 µm2), Plxna2−/− (0.06 ± 0.003 µm2), and Sema5A−/− (0.05 ± 0.003 µm2) mice. (M) The number of presynaptic boutons that contact a dendritic spine (i.e., synapses formed per spine) is not significantly different between WT, Plxna2−/− and Sema5A−/− GCs. (N and O) Two-dimensional EM images of P30 littermate Plxna2+/+ (WT, n = 2) and Plxna2−/− (n = 2) mice (arrow points to PSD). (P) Quantification of synapse density in 2D EM images. Scale bars in A, D, G = 20 µm; in B, E, H = 2 µm, in C, F, I = 0.2 µm, in N, O = 0.8 µm. For quantification of 3D EM images, n = 110, 208, and 165 spines were analyzed for WT (n = 1), Plxna2−/− (n = 1) and Sema5A−/− (n = 1) mice, respectively. Values are represented as mean ± SEM from three to four dendritic segments for each genotype. ***p < 0.001, N.S. = not significant (p > 0.05) two-tailed unpaired Student's t test.
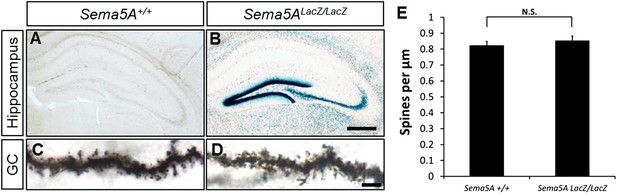
Sema5ALacZ/LacZ mice have normal GC dendritic spine density.
(A and B) Coronal brain sections of P30 WT control and littermate Sema5ALacZ/LacZ mice. Expression of Sema5A was visualized by β-galactosidase activity. (C and D) To visualize GCs dendritic spines, brain sections were stained by Golgi-Cox labeling. Representative images of dendrites taken from P30 mice. Scale bar in A and B = 200 μm, in C and D = 5 μm. (E) Quantification of GC dendritic spine density of Sema5A+/+ (0.83 ± 0.03 spines/μm) and Sema5ALacZ/LacZ (0.85 ± 0.03 spines/μm) mice. n = 15 neurons for Sema5A+/+ and n = 20 neurons for Sema5ALacZ/LacZ mice. Error bars represent SEM. N.S. p =0.43, two-tailed unpaired Student's t test.
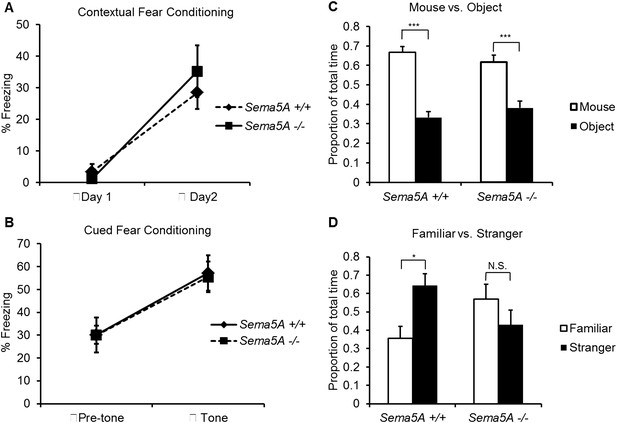
Sema5A−/− mice show abnormal social interaction.
(A and B) Fear conditioning. Sema5A+/+ and Sema5A−/− mice show no difference in freezing behavior under contextual fear conditioning (A) and cued fear conditioning (B) paradigms. Number of animals tested: n = 6 Sema5A+/+ mice and n = 8 Sema5A−/− mice. Error bars represent ± SEM. (C and D) Three-chambered social interaction test: (C) given the choice, both Sema5A+/+ and Sema5A−/− mice spent significantly more time engaged in nose-to-nose interaction with a mouse than sniffing an inanimate object. (D) Sema5A+/+ mice spent significantly more time engaged in nose-to-nose interaction with a stranger mouse compared to a familiar mouse, whereas Sema5A−/− mice showed no preference. Number of animals tested: n = 8 Sema5A+/+ mice and n = 8 Sema5A−/− mice for mouse vs inanimate object, and n = 7 Sema5A+/+ mice and n = 7 Sema5A−/− mice for familiar vs stranger mouse. Error bars represent ± SEM. n.s. not significant, ***p < 0.001 and *p < 0.05 by two-tailed Student's t test.
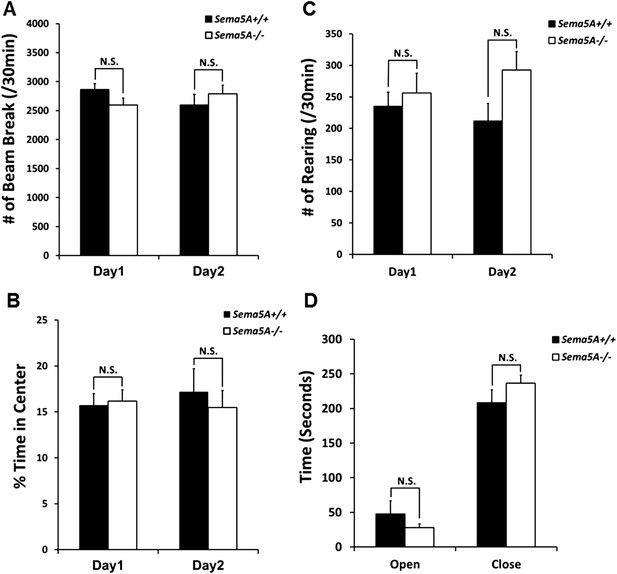
Sema5A−/− mice show normal anxiety levels and locomotor activity.
(A–C) Adult congenic C57b/6 Sema5A+/+ and Sema5A−/− mice subjected to the open field test show similar locomotion, as measured by total beam breaks (A), and also normal anxiety levels, as measured by percentage of time spent in the center of the field (B) and number of rearing events (C). Number of animals tested: n = 7 Sema5A+/+ mice and n = 9 Sema5A−/− mice. Data represent ± SEM. (D) Elevated plus maze. Sema5A−/− mice show no abnormality in anxiety level, as measured by time spent in the open arm vs the closed arm. Number of animals tested: n = 7 Sema5A+/+ mice and n = 9 Sema5A−/− mice. Data represent mean ± SEM. N.S. p > 0.05. Two-tailed unpaired Student's t test.
Tables
Quantification of dendritic spine density in the hippocampus of Thy1-eGFP WT, Sema5 and Plxna mice
| Genotype | Age | Dentate GC dendrite | CA1 primary dendrite | CA1 secondary dendrite | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| N | p Value (vs wt) | Spine density (mean ± SEM) | Distance from soma | n | p Value (vs wt) | Spine density (mean ± SEM) | Distance from soma | n | p Value (vs wt) | Spine density (mean ± SEM) | Distance from soma | ||
| WT | P30–33 | 4 animals (74 neurons) | 1.67 ± 0.01 | 50–100 μm | 3 animals (26 neurons) | 2.76 ± 0.07 | 50–100 μm | 3 animals (15 neurons) | 2.93 ± 0.02 | 50–100 μm | |||
| Sema5A+/−; Sema5B+/− | P30–33 | 3 animals (41 neurons) | 0.8795 | 1.68 ± 0.05 | 50–100 μm | 3 animals (24 neurons) | 0.6549 | 2.79 ± 0.019 | 50–100 μm | 3 animals (24 neurons) | 0.6333 | 2.99 ± 0.09 | 50–100 μm |
| Sema5A−/−; Sema5B+/− | P30–33 | 4 animals (65 neurons) | 0.0012 | 2.05 ± 0.05 | 50–100 μm | 3 animals (27 neurons) | 0.5122 | 2.81 ± 0.03 | 50–100 μm | 3 animals (21 neurons) | 0.618 | 2.99 ± 0.09 | 50–100 μm |
| Sema5A+/−; Sema5B−/− | P30–33 | 3 animals (46 neurons) | 0.8844 | 1.66 ± 0.05 | 50–100 μm | 3 animals (26 neurons) | 0.9797 | 2.76 ± 0.15 | 50–100 μm | 3 animals (21 neurons) | 0.9661 | 2.94 ± 0.02 | 50–100 μm |
| Sema5A−/−; Sema5B−/− | P30–33 | 3 animals (30 neurons) | 0.0066 | 2.06 ± 0.08 | 50–100 μm | 3 animals (27 neurons) | 0.5389 | 2.81 ± 0.02 | 50–100 μm | 3 animals (23 neurons) | 0.4006 | 2.98 ± 0.03 | 50–100 μm |
| Plxna1−/− | P30–33 | 3 animals (56 neurons) | 0.154 | 1.79 ± 0.05 | 50–100 μm | ||||||||
| Plxna2−/− | P30–33 | 3 animals (54 neurons) | <0.0001 | 2.24 ± 0.07 | 50–100 μm | ||||||||
| Plxna2+/− | P30–33 | 4 animals (74 neurons) | 0.0359 | 1.86 ± 0.05 | 50–100 μm | ||||||||
| Sema5A+/−; Plxna2+/− | P30–33 | 4 animals (72 neurons) | <0.0001 | 2.39 ± 0.05 | 50–100 μm | ||||||||





