Chromosome mis-segregation and cytokinesis failure in trisomic human cells
Figures
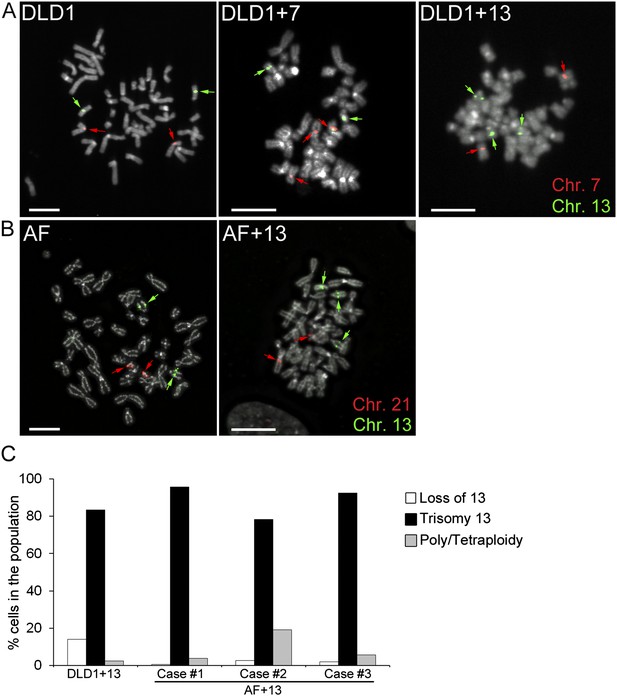
Trisomy 7 and 13 in DLD1 and AF cells.
(A–B) Chromosome-specific FISH staining for chromosomes 7 (red),13 (green), and 21 (red) in metaphase spreads confirms trisomy 7 in DLD1+7 and trisomy 13 in DLD1+13 and AF+13 cells. DNA is shown in grey. Arrows point to FISH signals. Scale bars, 5 μm. (C) Characterization of cells with trisomy 13 used in this study. The DLD1+13 cells were subcloned and the clone used for this study was analyzed by chromosome painting of metaphase spreads with chromosome 13-specific probes. Cells were classified as having lost chromosome 13, trisomic for chromosome 13, or poly/tetraploid when they carried six copies of chromosome 13 and ∼88 other chromosomes. The AF+13 cells were analyzed by interphase FISH at passage 1–2 with probes specific to chromosomes 13 and 21. Cells were classified has having lost/gained a copy of chromosome 13 (two or four nuclear signals for chr. 13, two signals for chr. 21), displaying trisomy 13 (three signals for chr. 13 and 2 signals for chr. 21), or displaying poly/tetraploidy (six signals for chr. 13 and four signals for chr. 21). A small fraction (1.9%) of the AF+13 Case #3 displayed loss/gain of chr. 21.
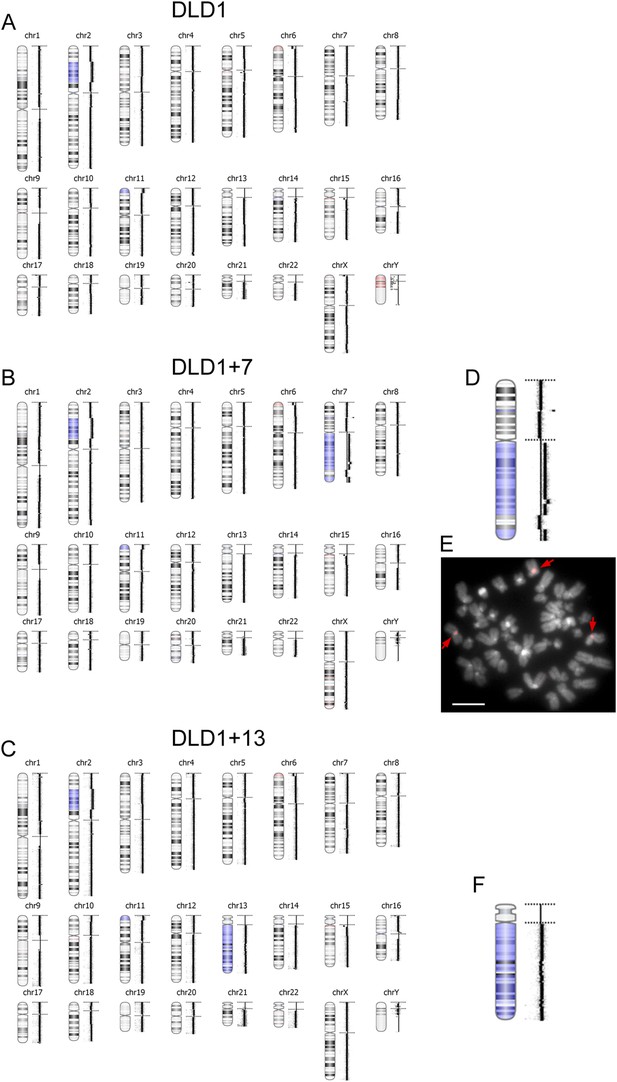
aCGH in DLD1, DLD1+7, and DLD1+13 cells.
(A–C) The images show the results of aCGH in the three cell lines, with regions shaded in blue corresponding to gains and regions shaded in red corresponding to losses. The data show that the DLD1+7 and DLD1+13 cell lines share CNVs, such as duplication of a portion of the short arm of chromosome 2, gain of a short portion of the p arm of chromosome 11, and loss of a short region of the p arm of chromosome 6, with the parental cell line. Moreover, the data highlight the gain of most of the q arm of chromosome 7 in the DLD1+7 cell line and gain of chromosome 13 in the DLD1+13 cell line. (D) 2× enlargement of the aCGH image of chr. 7 from the DLD1+7 cell line. (E) FISH staining of DLD1+7 metaphase spread using a probe (red) specific to the centromeric DNA of chromosome 7. The image shows that the trisomic chromosome 7q maintains a centromere. DNA is shown in grey. Arrows point to FISH signals. (F) 2× enlargement of the aCGH image of chr. 13 from the DLD1+13 cell line.
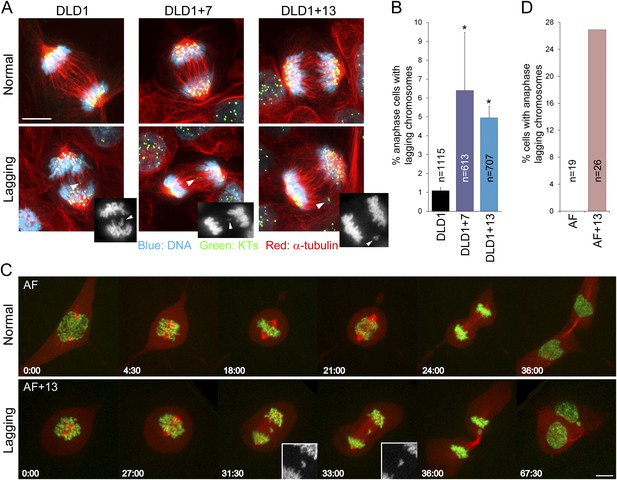
Increased rates of anaphase lagging chromosomes in cells with trisomy 7 or 13.
(A) Examples of normal anaphases (top row) and anaphase cells with lagging chromosomes (bottom row). Cells were immunostained for microtubules (red) and kinetochores (green). DNA is shown in blue. Images represent maximum intensity projections of Z-stacks. Arrowheads point at anaphase lagging chromosomes. Grey scale images at the bottom right corners of the images in the bottom row are single focal planes of DAPI-stained chromosomes shown for easier visualization of the lagging chromosomes. (B) Frequencies of anaphase lagging chromosomes were significantly higher (*χ2 test, p < 0.0001) in both DLD1+7 and DLD1+13 compared to DLD1 cells. Data are reported as mean ± S.E.M and represent the average of three independent experiments in which a total of 613–1115 anaphases were analyzed. (C) Time-lapse microscopy of AF and AF+13 cells undergoing mitosis. An example of AF undergoing normal mitosis is shown in the top row and an example of AF+13 displaying an anaphase lagging chromosome is shown in the bottom row. DNA is shown in green (H2B-GFP) and microtubules in red (RFP-tubulin). Images are maximum intensity projections of Z-stacks. Insets in the bottom row display enlarged views of the DNA alone (in grey scale) in the region around the lagging chromosome. (D) None of the 19 AF cells (from cases #1 and #2) imaged displayed anaphase lagging chromosomes, whereas 7 out of 26 AF+13 cells (5 out of 18 in case #1 and 3 out of 8 in case #2) displayed anaphase lagging chromosomes. Time stamps indicate elapsed time in min:sec. Scale bars, 5 μm.
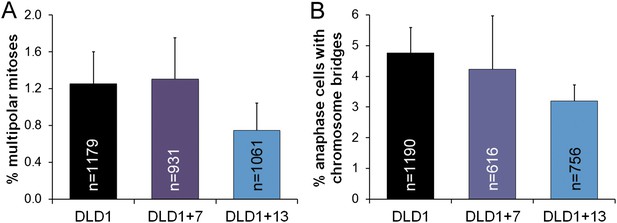
Similar frequencies of multipolar mitoses (A) and anaphase chromosome bridges (B) in diploid vs aneuploid DLD1 cells.
The data are reported as mean ± S.E.M. and represent the average of three experiments in which a total of 931–1179 mitotic cells (A) or 616–1190 anaphase cells (B) were analyzed.
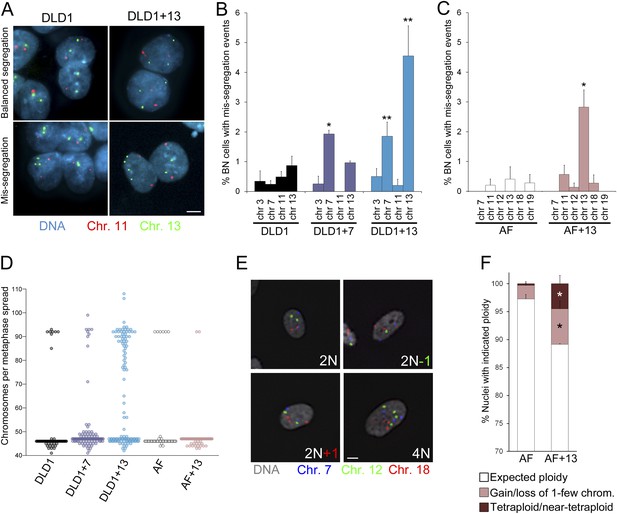
Increased chromosome mis-segregation rates and karyotypic heterogeneity in cells with trisomy 7 or 13.
(A–C) Combination of cytokinesis-block assay and FISH staining with chromosome-specific probes shows higher chromosome mis-segregation rates in DLD1+7 and DLD1+13 compared to DLD1 cells and AF+13 compared to AF cells. (A) Examples of FISH-stained binucleate (BN) DLD1 and DLD1+13 cells. Scale bar, 5 μm. (B–C) Frequencies of BN cells displaying mis-segregation events. *Two-tailed χ2 test, p < 0.005; **two-tailed χ2 test, p < 0.0001, when compared to mis-segregation of the same chromosome in the euploid cell line. Data are presented as mean ± S.E.M. and represent the average of at least three independent experiments/samples in which a total of 460–1229 BN cells were analyzed. (D) Beeswarm plot displaying data from chromosome counts in metaphase spreads from the five cell lines. DLD1+7, DLD1+13, and AF+13 (modal chromosome number 47, shown by the high concentration of sampled points) displayed increased karyotypic heterogeneity compared to DLD1 and AF cells (modal chromosome number 46, shown by the high concentration of sampled points), respectively. In addition, DLD1+13 cells displayed a large sub-population of near-tetraploid cells (modal chromosome number 92). Chromosome counts were performed on 89–303 metaphase spreads. (E–F) FISH staining with chromosome-specific probes in interphase nuclei reveals higher rates of aneuploidy and tetraploidy in AF+13 vs AF cells. (E) FISH staining in interphase AF and AF+13 cells with probes specific for chromosomes 7 (blue), 12 (green), and 18 (red). Scale bar, 5 μm. (F) Quantification of interphase FISH data shown in (E). Cells were classified as having gained or lost 1-few chromosomes or as tetraploid (4 signals for each of the three chromosomes analyzed). The data show a larger fraction of cells with gain/loss of 1-few chromosomes in the AF+13 population compared to AF cells (*χ2 test, p < 0.0001). Additionally, AF+13 cells displayed a larger sub-population of tetraploid cells (*χ2 test, p < 0.0001). Data are presented as mean + S.E.M. and represent the average of three different samples in which a total of 2,117–2,410 cells were analyzed.
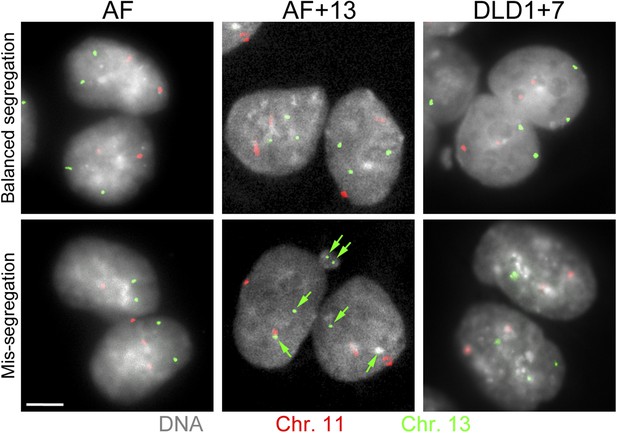
Combined cytokinesis-block assay and FISH staining with chromosome-specific probes.
The images show examples of binucleate AF, AF+13, and DLD1+7 cells hybridized with FISH probes specific to chromosomes 11 and 13. Green arrows in image of mis-segregation in AF+13 cell point at signals for chromosome 13. Scale bar, 5 μm.
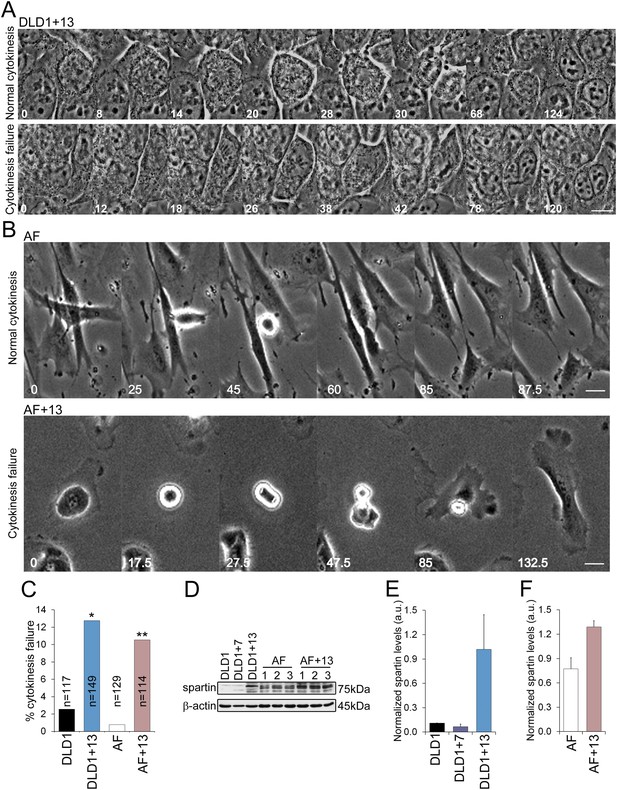
DLD1+13 and AF+13 cells overexpress SPG20 and fail cytokinesis at high rates.
(A–B) Time-lapse microscopy indicates that cells with trisomy 13 frequently fail cytokinesis. Time stamps indicate elapsed time in minutes. Scale bars, 10 μm. (A) Still images from time-lapse phase contrast videos of DLD1+13 cells undergoing mitosis and completing (top row) or failing (bottom row) cytokinesis. (B) Still images from time-lapse phase contrast videos of AF (top row) and AF+13 (bottom row) cells undergoing mitosis and completing (AF, top row) or failing (AF+13, bottom row) cytokinesis. (C) Quantification of cytokinesis failure from phase-contrast time-lapse videos showing that the rates of cytokinesis failure in DLD1+13 and AF+13 cells are significantly higher than those observed in their diploid counterparts, DLD1 and AF cells (*χ2 test, p < 0.01; ** χ2 test, p < 0.001). (D) Western blot analysis of Spartin across DLD1 cell lines, three samples of AF and three samples of AF+13 cells. β-actin was used as a loading control. (E) Quantification of spartin levels (normalized to β-actin) in DLD1, DLD1+7, and DLD1+13 cells. The data reported are the average of three independent experiments and are displayed as mean + S.E.M. (F) Quantification of spartin levels (normalized to β-actin) in AF and AF+13 cells. The data reported are the average of the three AF and AF+13 samples shown in (E) and are displayed as mean + S.E.M.
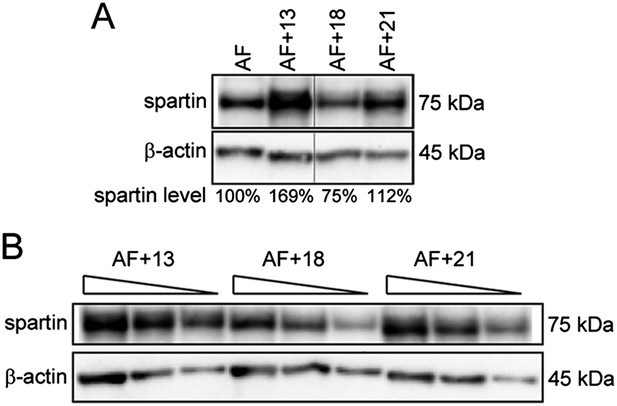
Trisomies other than 13 are not associated with spartin overexpression.
(A) Western blot analysis of Spartin levels in AF control and AF+13, AF+18, and AF+21 trisomic cells. Spartin levels were normalized to β-actin and are shown as percentage of the amount in the euploid control. (B) Western blot with decreasing amounts (30, 20, and 10 μg) of total extracts from the trisomies used in (A) performed for titration of the western blot conditions.
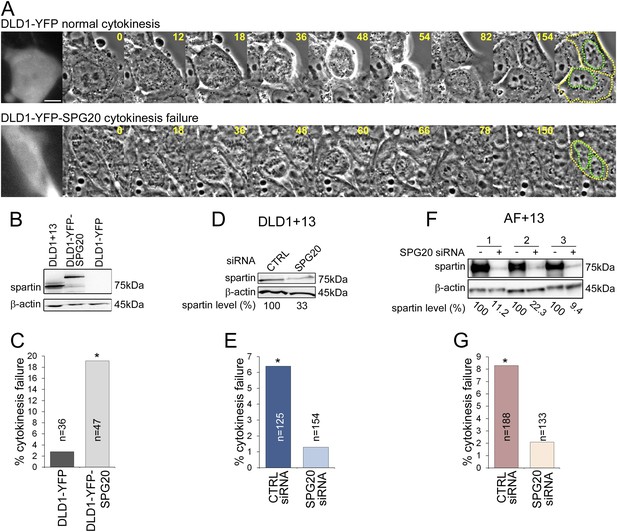
Spartin overexpression induces cytokinesis failure.
(A) Time-lapse microscopy of DLD1 cells transiently transfected with a YFP-N1 vector (DLD1-YFP, top row) or a YFP-SPG20-N1 vector (DLD1-YFP-SPG20, bottom row). Representative still images of time-lapse videos show a DLD1-YFP cell undergoing mitosis and completing cytokinesis (top row) and a DLD1-YFP-SPG20 cell undergoing mitosis and failing cytokinesis (bottom row). YFP expression was verified by fluorescence imaging and it is shown in the first panel for each time-lapse series. A copy of the last frame was added at the end of the sequence to highlight cell (yellow) and nuclear (green) outlines. Scale bar, 10 μm. (B) Western blot analysis of Spartin levels in DLD1+13, DLD1-YFP, and DLD1-YFP-SPG20 cells shows that the levels of YFP-Spartin in DLD1-YFP-SPG20 cells (center lane) are much higher than the levels of Spartin in DLD1-YFP cells. WB of Spartin in DLD1+13 cells is shown for comparison. (C) Quantification of cytokinesis failure rates in DLD1-YFP and DLD1-YFP-SPG20 showing that overexpression of SPG20 leads to increased rates of cytokinesis failure (*χ2 test, p < 0.01 when comparing DLD1-YFP-SPG20 to DLD1-YFP cells). (D) Western blot analysis of Spartin levels in DLD1+13 cells treated with a SPG20 siRNA or with a control siRNA. (E) Reducing the levels of Spartin by SPG20 siRNA significantly reduces the rate of cytokinesis failure in DLD1+13 cells (*χ2 test, p < 0.02 when comparing cells treated with a control siRNA to cells treated with SPG20 siRNA). (F) Western blot analysis of Spartin levels in AF+13 cells treated with a SPG20 siRNA or with a control siRNA. (G) Reducing the levels of Spartin by SPG20 siRNA significantly reduces the rates of cytokinesis failure in AF+13 cells (*χ2 test, p < 0.02 when comparing cells treated with a control siRNA to cells treated with SPG20 siRNA). Average values from three independent experiments; variability between AF+13 samples was not significant.
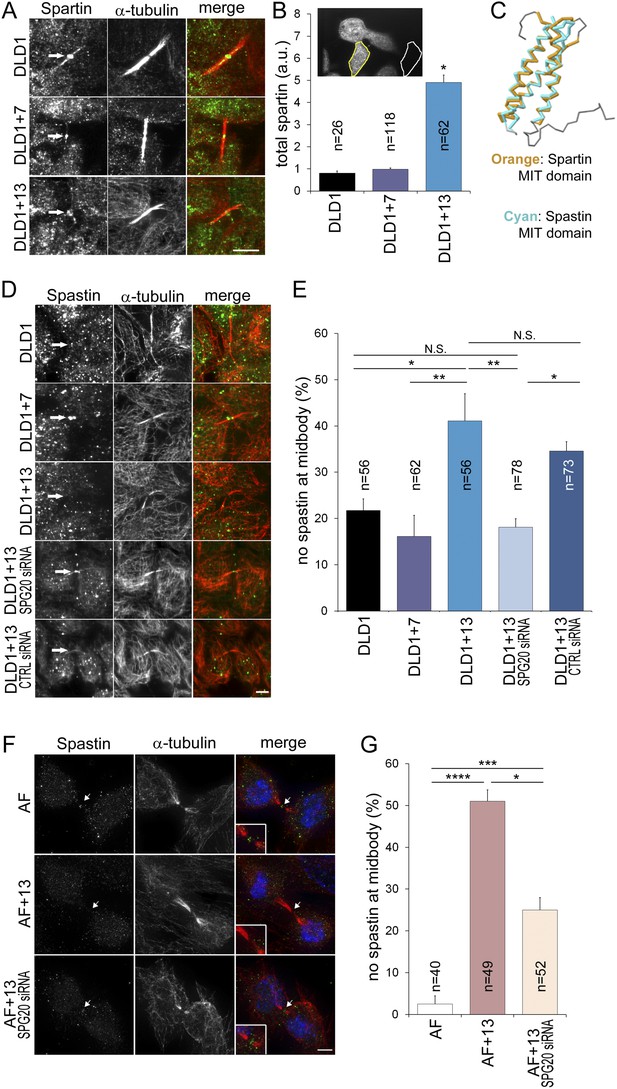
Spartin overexpression impairs Spastin localization to the midbody.
(A–B) Spartin localization at the midbody is not affected by the high levels of intracellular Spartin in DLD1+13. (A) Images showing Spartin localization (arrows) at the midbody of the three DLD1 cell lines. (B) The image shows interphase DLD1+13 cells immunostained for Spartin and the yellow and white outline indicate the regions of interest (ROI) selected for measurements of total intracellular fluorescence (yellow ROI) and background fluorescence (white ROI). The data in the graph report the total intracellular Spartin fluorescence intensity after background subtraction in randomly sampled interphase cells and are represented as mean ± S.E.M (*t-test, p < 0.0001 when comparing DLD1+13 to either DLD1 or DLD1+7). (C) Superimposition of the MIT domains of Spartin (orange) and Spastin (cyan) illustrates the considerable degree of structural homology between the two. PDB ID #2DL1 (Suetake et al., 2009) and #3EAB (Yang et al., 2008) for Spartin and Spastin, respectively. (D–E) Spastin localization at the midbody is impaired in DLD1+13 cells, and is rescued by SPG20 siRNA. (D) Images of midbodies of cells immunostained for Spastin (green) and microtubules (red). Arrows point at sites of Spastin localization or lack thereof. (E) Frequencies of cells lacking Spastin at the midbody. Higher frequencies of cells lacking Spastin at the midbody can be found in DLD1+13 cells compared to either DLD1 or DLD1+7 cells. Spastin localization could be rescued by knocking down Spartin levels in DLD1+13 cells via SPG20 siRNA. Statistical significance was calculated using a χ2 test (*p < 0.05; **p < 0.01; N.S. = not significant). Data are presented as mean ± S.E.M. and represent the average of three independent experiments. (F–G) Spastin localization at the midbody is impaired in AF+13 cells, and is partially rescued by SPG20 siRNA. (F) Images of midbodies of cells immunostained for Spastin (green) and microtubules (red). DNA is shown in blue in the merged images. Arrows point at sites of Spastin localization or lack thereof. The insets show 3× magnifications of the midbody region. (G) Frequencies of cells lacking Spastin at the midbody are higher in AF+13 compared to AF cells. Spastin localization was partially rescued by SPG20 siRNA-mediated Spartin knock down in AF+13 cells. Statistical significance was calculated using a χ2 test (*p < 0.05; ***p < 0.001; ****p < 0.0001). Data are presented as mean ± S.E.M. and represent the average of three independent experiments/samples. Scale bars, 5 μm.
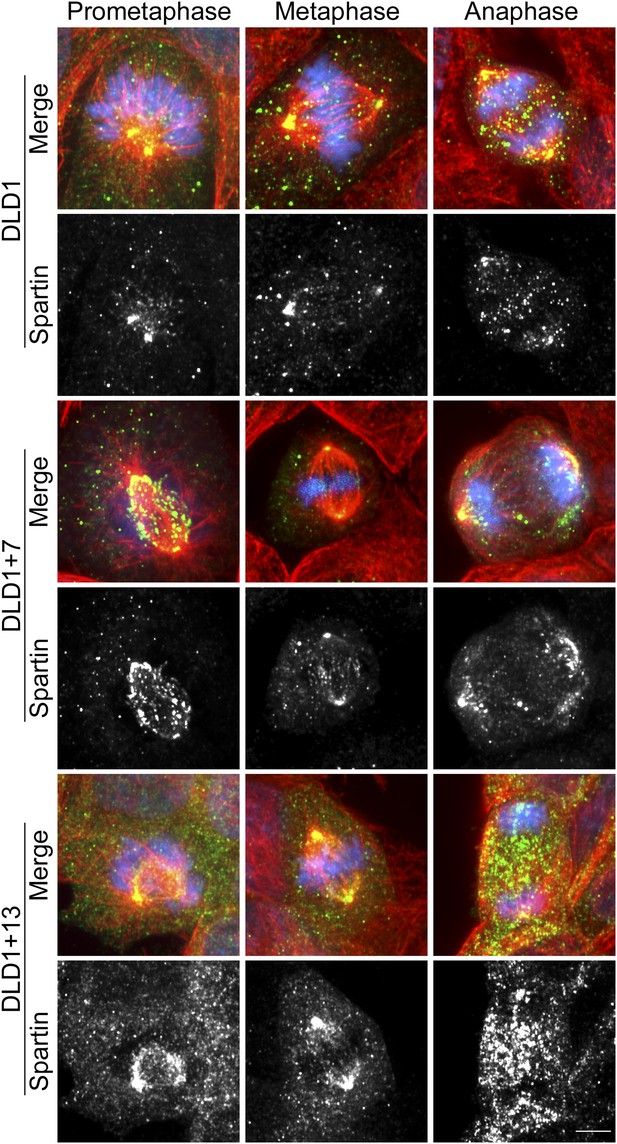
Spartin localization during mitosis.
Cells were immunostained for Spartin (green) and microtubules (red) and counterstained with DAPI for DNA (blue). For each cell line, the merged images (top) and Spartin alone (bottom) are shown. Images are maximum intensity projections of Z-stacks. Spartin localizes to the spindle poles during prometaphase, metaphase, and anaphase in DLD1, DLD1+7 and DLD1+13. An eccess of cytoplasmic spartin (away from the mitotic spindle) can be noticed in DLD1+13 cells, but not in the other two cell lines. Scale bar, 5 μm.
Videos
Representative video showing normal mitosis in an AF cell.
Images were acquired by spinning-disk confocal microscopy at 1 min intervals and they are played back at 5 frames per second. DNA is shown in green (H2B-GFP) and microtubules in red (RFP-tubulin).
Representative video showing anaphase lagging chromosome during mitosis in an AF+13 cell.
Images were acquired by spinning-disk confocal microscopy at 1 min intervals and they are played back at 5 frames per second. DNA is shown green (H2B-GFP) and microtubules in red (RFP-tubulin).
Representative video showing normal cytokinesis in a DLD1+13 cell.
Images were acquired by phase contrast microscopy at 2 min intervals, and they are played back at 7 frames per second. Scale bar, 5 μm.
Representative video showing cytokinesis failure in a DLD1+13 cell.
Images were acquired by phase contrast microscopy at 2 min intervals, and they are played back at 7 frames per second. Scale bar, 5 μm.
Representative video showing normal cytokinesis in an AF cell.
Images were acquired by phase contrast microscopy at 2.5 min intervals, and they are played back at 5 frames per second. Scale bar, 5 μm.
Representative video showing cytokinesis failure in an AF+13 cell.
Images were acquired by phase contrast microscopy at 2.5 min intervals, and they are played back at 5 frames per second. Scale bar, 5 μm.
Representative video showing normal cytokinesis in a DLD1 cell transiently transfected with a YFP vector (control).
Near-simultaneous phase contrast and epifluorescence images were acquired at 4 min intervals at a single focal plane using the Nikon perfect focus function. For clarity, the fluorescent image is displayed only in the first frame, whereas the rest of the video shows only phase contrast images played back at 7 frames per second. Scale bar, 10 μm.
Representative video showing cytokinesis failure in a DLD1 cell transiently transfected with a YFP-SPG20 vector (SPG20 overexpression).
Near-simultaneous phase contrast and epifluorescence images were acquired at 4 min intervals at a single focal plane using the Nikon perfect focus function. For clarity, the fluorescent image is displayed only in the first frame, whereas the rest of the video shows only phase contrast images played back at 7 frames per second. Scale bar, 10 μm.
Tables
Euploid and trisomic amniocytes used in this study
| Name | Karyotype | Gestational age (week) |
|---|---|---|
| AF 1 | 46,XX | 17 |
| AF 2 | 46,XY | 20 |
| AF 3 | 46,XX | 21 |
| AF+13 1 | 47,XX,+13 | 16 |
| AF+13 2 | 47,XX,+13 | 21 |
| AF+13 3 | 47,XX,+13 | 21 |
| AF+18 | 47,XX,+18 | 17 |
| AF+21 | 47,XX,+21 | 17 |





