A molecular tweezer antagonizes seminal amyloids and HIV infection
Figures
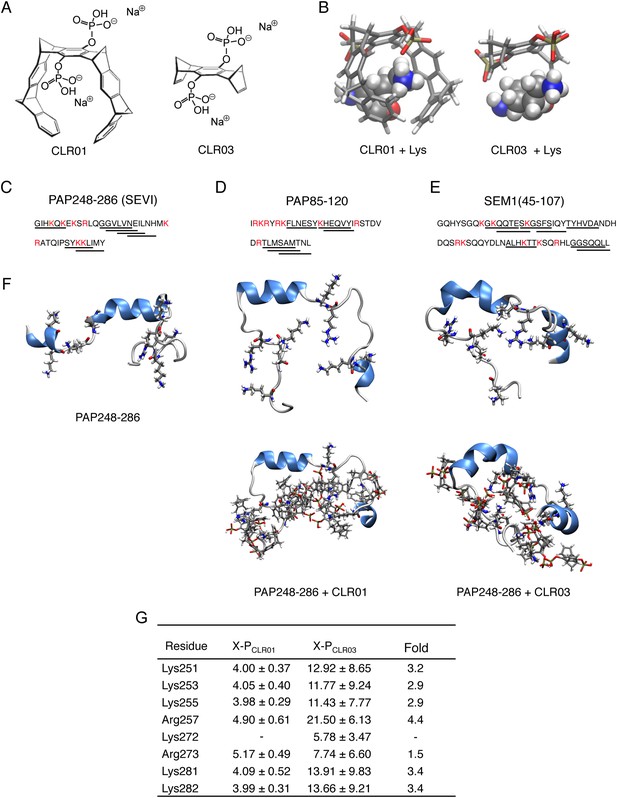
CLR01 binds to lysine and arginine residues.
(A) Chemical structures of CLR01 and CLR03. (B) Stick representation of the structures of CLR01 and CLR03 and their engagement of lysine side chains. (C–E) The primary sequences of PAP248-286 (C), PAP85-120 (D), and SEM1(45-107) (E) are provided. Lysine and arginine residues are in red and hexapeptides predicted to form steric zippers (Goldschmidt et al., 2010; Castellano and Shorter, 2012) are underlined. (F) The average structures of the most populated clusters derived from the REMD simulations of PAP248-286 (left), PAP248-286 with 7 CLR01 molecules (middle), and PAP248-286 with 8 CLR03 molecules (right) are shown in the upper row, CLR01 and CLR03 molecules are not shown for clarity. The lower row shows, for each case, a representative structure of the most populated cluster including CLR01 and CLR03. (G) CLR03 establishes only labile interactions with PAP248-286 as shown by the large X-P distances (Å) between one P atom of CLR03 and the nitrogen atom of the lysine side chain (or carbon atom of the guanidinium moiety of arginine). Contrarily, the complexes between CLR01 and Lys or Arg were conserved during all the REMD simulations.
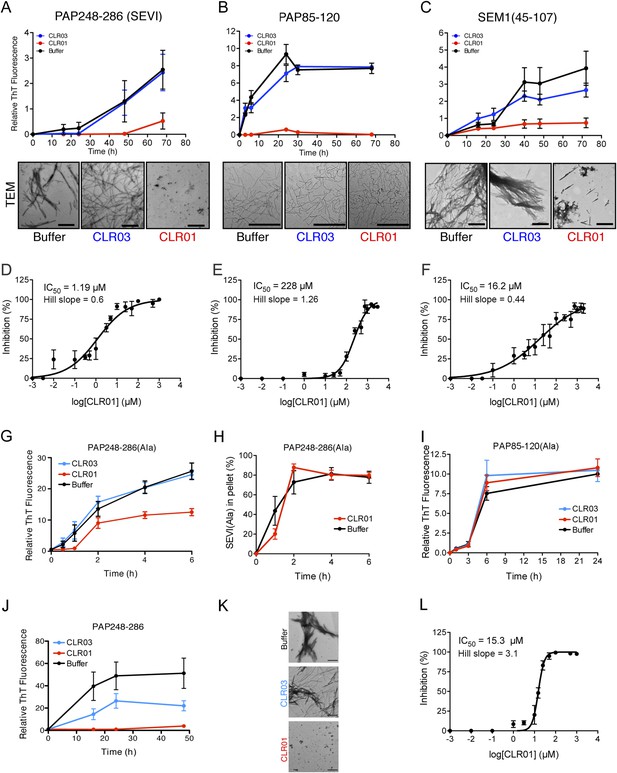
CLR01 inhibits formation of seminal amyloid fibrils.
CLR01 inhibits fibril formation by PAP248-286 (1 mM) (A), PAP85-120 (1 mM) (B), and SEM1(45-107) (500 µM) (C). Peptides were incubated with equimolar CLR01, CLR03, or buffer and agitated at 1400 rpm at 37°C. Aliquots were removed at various time points and fibrillization was assessed using the amyloid-binding dye, Thioflavin-T (ThT). Values represent means ±SEM (n = 4 for PAP248-286; n = 3 for PAP85-120; n = 9 for SEM1(45-107)). Transmission electron microscopy (TEM) images of PAP248-286 (A), PAP85-120 (B), and SEM1(45-107) (C) agitated in the presence of CLR01, CLR03, or buffer. PAP248-286 and SEM1(45-107) samples were visualized after 72 hr of agitation, PAP85-120 after 24 hr. Scale bar: 500 nm. (D) Dose-response curve for CLR01 inhibition of PAP248-286 (1 mM) fibrillization after 72 hr of agitation. The IC50 value and Hill slope are indicated. Values represent means ±SEM (n = 3–12). (E) Dose-response curve for CLR01 inhibition of PAP85-120 (1 mM) fibrillization after 24 hr of agitation. The IC50 value and Hill slope are indicated. Values represent means ±SEM (n = 3–7). (F) Dose–response curve for CLR01 inhibition of SEM1(45-107) (500 µM) fibrillization after 72 hr of agitation. The IC50 value and Hill slope are indicated. Values represent means ±SEM (n = 3–12). (G, H) CLR01 is unable to block PAP248-286(Ala) fibrillization. Lyophilized PAP248-286(Ala) was dissolved in PBS (100 µM), incubated with CLR01 (100 µM), CLR03 (100 µM), or buffer and agitated at 1400 rpm at 37°C. Aliquots were removed at various time points and fibril assembly was assessed using ThT fluorescence (G) or sedimentation analysis (H). Values represent means ±SEM (n = 7–9) (G) or means ±SEM (n = 3) (H). (I) CLR01 is unable to block PAP85-120(Ala) fibrillization. Lyophilized PAP85-120(Ala) was dissolved in PBS (150 µM), incubated with CLR01 (150 µM), CLR03 (150 µM), or buffer and agitated at 1400 rpm at 37°C. At the indicated times, fibril assembly was assessed using ThT fluorescence. Values represent means ±SEM (n = 3). (J) Inhibition of seeded PAP248-286 fibrillization by CLR01. Lyophilized PAP248-286 peptide was reconstituted at 1 mM in PBS and incubated with 1 mM CLR01, 1 mM CLR03, or buffer. Preformed SEVI fibrils (2% wt/wt) were added to each condition and agitated at 1400 rpm at 37°C. Aliquots were removed at various time points and fibrillization was assessed using ThT fluorescence. Values represent means ±SEM (n = 6). (K) Electron microscopy visualization of CLR01 inhibition of seeded PAP248-286 fibrillization after 48 hr of agitation. Scale bar: 500 nm. (L) Dose-response curve for CLR01 inhibition of PAP248-286 fibrillization seeded by preformed SEVI fibrils (2% wt/wt) after 48 hr of agitation. The IC50 value and Hill slope are indicated. Values represent means ±SEM (n = 3–7).
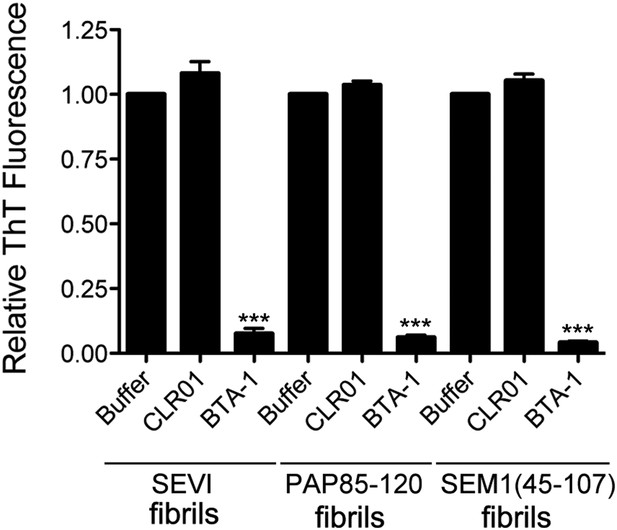
CLR01 does not displace ThT from SEVI, PAP85-120, or SEM1(45-107) fibrils.
Preformed SEVI, PAP85-120, or SEM1(45-107) fibrils (5 µM monomer) were preincubated with ThT (25 µM) for 30 min at room temperature. Buffer (PBS), CLR01 (250 µM), or a known competitor of ThT binding, BTA-1 (250 µM) were then added and incubated for 10 min at room temperature. ThT displacement was then assessed by fluorescence measurements. Values represent means ±SEM (n = 3). A one-way ANOVA with the post hoc Dunnett's multiple comparisons test was used to compare the buffer control to the CLR01 or BTA-1 condition (*** denotes p < 0.0001).
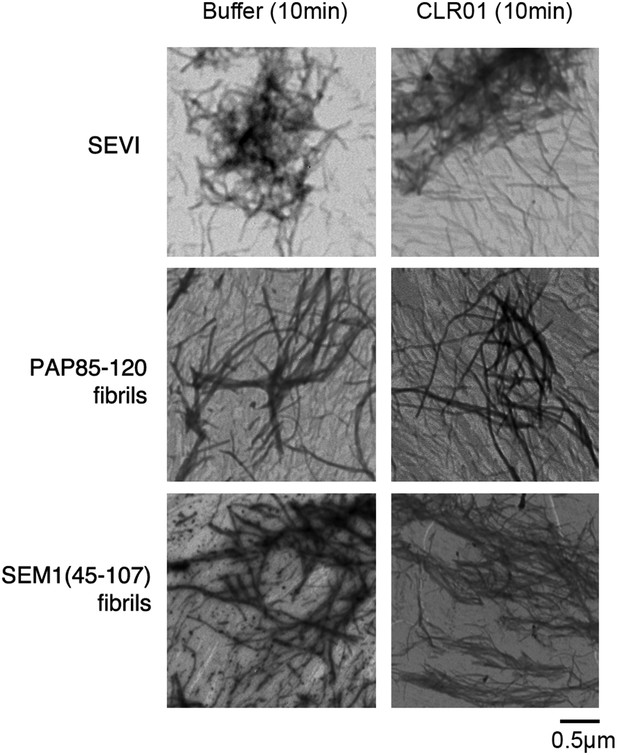
CLR01 does not impede adsorption of SEVI, PAP85-120, or SEM1(45-107) fibrils to the EM grid.
Preformed SEVI, PAP85-120 fibrils, or SEM1(45-107) fibrils (20 µM) were treated with a 10-fold excess of CLR01 or buffer for 10 min and processed for TEM. Note the presence of abundant fibrils under each condition indicating that CLR01 does not impede fibril adsorption to the EM grid.
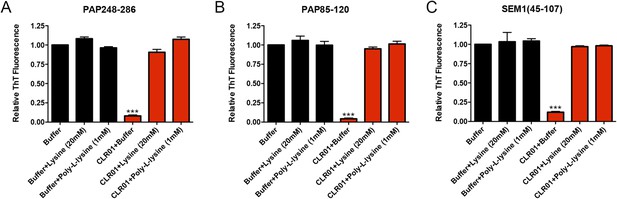
Lysine and poly-L-lysine antagonize the ability of CLR01 to inhibit spontaneous formation of seminal amyloid fibrils.
(A) PAP248-286 (1 mM) was incubated with buffer or CLR01 (100 µM) in the presence or absence of lysine (20 mM) or poly-L-lysine (1 mM) and agitated at 1400 rpm at 37°C for 68 hr. Fibrillization was assessed via ThT fluorescence. Values represent means ±SEM (n = 3). A one-way ANOVA with the post hoc Dunnett's multiple comparisons test was used to compare the buffer alone control to the other conditions (*** denotes p < 0.0001). (B) PAP248-286 (100 µM) was incubated with CLR01 (100 µM) or buffer in the presence or absence of lysine (20 mM) or poly-L-lysine (1 mM) and agitated at 1400 rpm at 37°C for 68 hr. Fibrillization was assessed via ThT fluorescence. Values represent means ±SEM (n = 3). A one-way ANOVA with the post hoc Dunnett's multiple comparisons test was used to compare the buffer alone control to the other conditions (*** denotes p < 0.0001). (C) SEM1(45-107) (0.5 mM) was incubated with CLR01 (100 µM) or buffer in the presence or absence of lysine (20 mM) or poly-L-lysine (1 mM) and agitated at 1400 rpm at 37°C for 68 hr. Fibrillization was assessed via ThT fluorescence. Values represent means ±SEM (n = 3). A one-way ANOVA with the post hoc Dunnett's multiple comparisons test was used to compare the buffer alone control to the other conditions (*** denotes p < 0.0001).
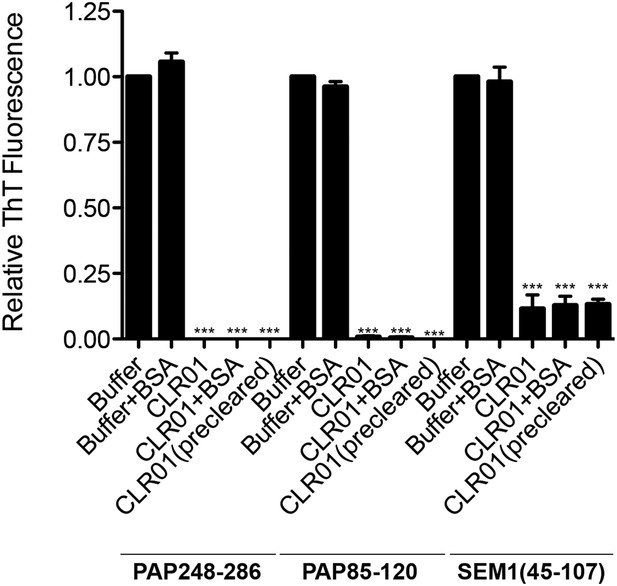
BSA or preclearing CLR01 has no effect on the ability of CLR01 to inhibit formation of seminal amyloid fibrils.
PAP248-286 (1 mM), PAP85-120 (1 mM), or SEM1(45-107) (500 µM) was incubated with CLR01 (1 mM) or buffer in the presence or absence of BSA (10 mg/ml) and agitated at 1400 rpm at 37°C for 68 hr. In some reactions, CLR01 was first cleared by centrifugation at 16,100×g for 20 min at 25°C (precleared). Fibrillization was assessed via ThT fluorescence. Values represent means ±SEM (n = 3). A one-way ANOVA with the post hoc Dunnett's multiple comparisons test was used to compare the buffer alone control to the other conditions (*** denotes p < 0.0001).
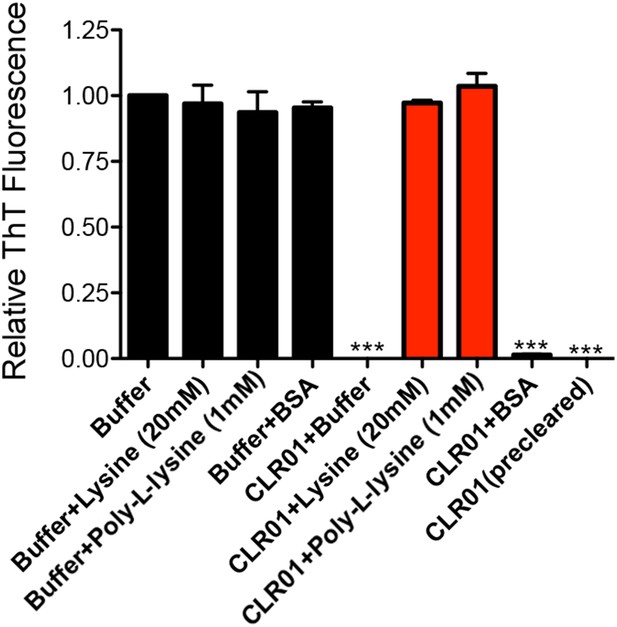
Lysine or poly-L-lysine, but not BSA or preclearing CLR01 solutions, antagonize the ability of CLR01 to inhibit seeded assembly of SEVI fibrils.
PAP248-286 (1 mM) was incubated with agitation at 37°C for 24 hr with preformed SEVI fibrils (2% wt/wt) plus CLR01 (100 µM) or buffer in the presence or absence of lysine (20 mM), poly-L-lysine (1 mM), or BSA (10 mg/ml). In some reactions, CLR01 was first cleared by centrifugation at 16,100×g for 20 min at 25°C (precleared). Fibrillization was assessed via ThT fluorescence. Values represent means ±SEM (n = 3). A one-way ANOVA with the post hoc Dunnett's multiple comparisons test was used to compare the buffer alone control to the other conditions (*** denotes p < 0.0001).
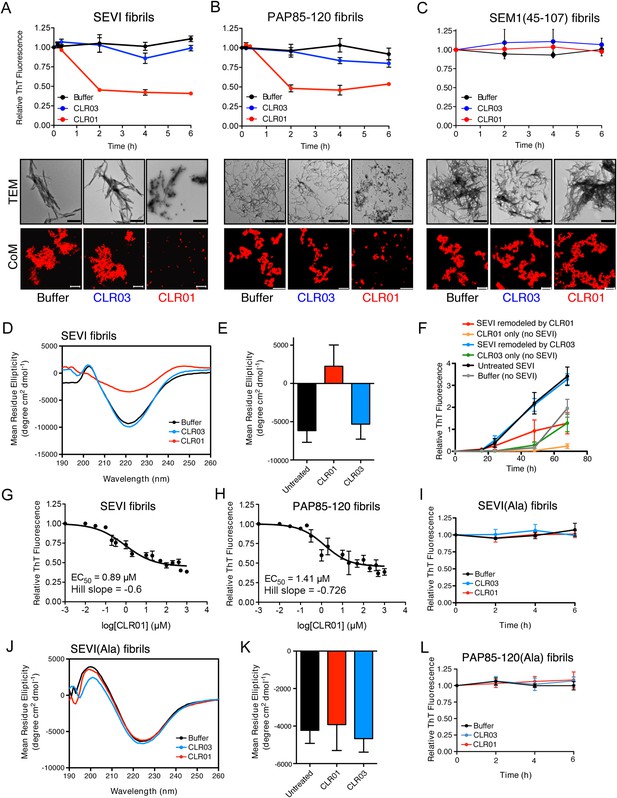
CLR01 rapidly remodels SEVI and PAP85-120 fibrils.
(A–C) Preformed SEVI fibrils (A), PAP85-120 fibrils (B), and SEM1(45-107) fibrils (C) (20 µM) were treated with a 10-fold excess of CLR01 or CLR03 or buffer for 0–6 hr at 37°C. Fibril integrity was assessed using ThT. Values represent means ±SEM (n = 3). TEM (middle panel) and confocal microscopy (bottom panel) of SEVI (A), PAP85-120 fibrils (B), and SEM1(45-107) fibrils (C) obtained after 2 hr treatment with CLR01, CLR03, or buffer. Scale bar for TEM images: 500 nm. For confocal microcopy (CoM), samples were stained with Proteostat dye. Scale bar: 20 µm. (D) CD spectrum of 50 µM SEVI fibrils incubated with 50 µM CLR01, 50 µM CLR03, or buffer. A representative spectrum is shown. (E) The mean residue ellipticity (MRE) at 218 nm was averaged from three independent experiments. Values represent means ±SEM. (F) Seeding with CLR01-remodeled SEVI products. SEVI fibrils (20 µM) were treated with 200 µM CLR01, 200 µM CLR03, or buffer for 3 hr and reaction products were used to seed PAP248-286 fibrillization (0.1% fibril seed, 1 mM peptide). Buffer conditions with no fibril seed present were also included. Fibrillization was monitored by ThT fluorescence. Values represent means ±SEM (n = 8). (G) Dose–response curve for CLR01 remodeling of SEVI fibrils (20 µM) after 2 hr of treatment. The EC50 value and Hill slope are indicated. Values represent means ±SEM (n = 3–15). (H) Dose-response curve for CLR01 remodeling of PAP85-120 fibrils (20 µM) after 2 hr of treatment. The EC50 value and Hill slope are indicated. Values represent means ±SEM (n = 3–14). (I) CLR01 is unable to remodel SEVI(Ala) fibrils. Preformed SEVI(Ala) fibrils (20 µM) were treated with 200 µM CLR01, 200 µM CLR03, or buffer for 0–6 hr. Fibril integrity was assessed using ThT fluorescence. Values represent means ±SEM (n = 3). (J) CD spectrum of 50 µM SEVI(Ala) fibrils incubated with either 50 µM CLR01, 50 µM CLR03, or buffer. One representative example is shown. (K) The mean residue ellipticity (MRE) at 218 nm was averaged from three independent experiments. Values represent means ±SEM (n = 3). (L) Preformed PAP85-120(Ala) fibrils (20 μM) were treated with CLR01 or CLR03 (200 μM) or buffer for 0–6 hr. Fibril integrity was assessed using ThT fluorescence. Values represent means ±SEM (n = 3).
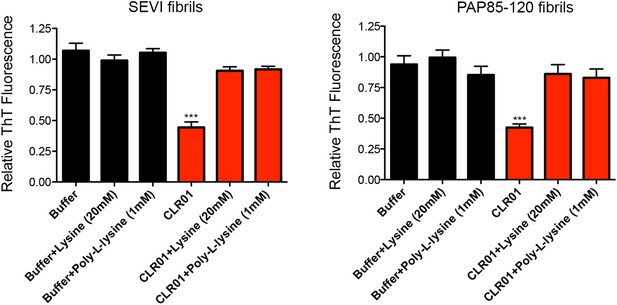
Lysine and poly-L-lysine antagonize the ability of CLR01 to remodel SEVI and PAP85-120 fibrils.
SEVI (left) or PAP85-120 fibrils (right, 20 µM monomer) were treated with buffer or CLR01 (100 µM) in the presence or absence of lysine (20 mM) or poly-L-lysine (1 mM) for 2 hr at 37°C. Fibril remodeling was then assessed by ThT fluorescence. Values represent means ±SEM (n = 3–8). A one-way ANOVA with the post hoc Dunnett's multiple comparisons test was used to compare the buffer alone control to the other conditions (*** denotes p < 0.0001).
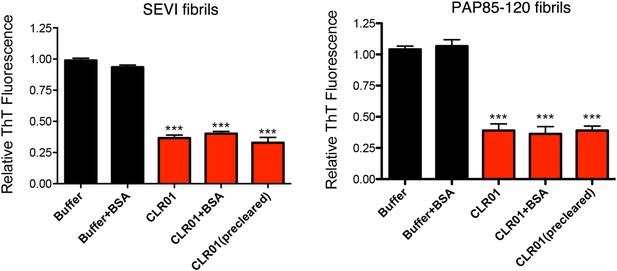
BSA or preclearing CLR01 has no effect on the ability of CLR01 to remodel SEVI and PAP85-120 fibrils.
SEVI (left) or PAP85-120 fibrils (right, 20 µM monomer) were treated with buffer or CLR01 (100 µM) in the presence or absence of BSA (10 mg/ml) for 2 hr at 37°C. In some reactions, CLR01 was first cleared by centrifugation at 16,100×g for 20 min at 25°C (precleared). Fibril remodeling was then assessed by ThT fluorescence. Values represent means ±SEM (n = 3). A one-way ANOVA with the post hoc Dunnett's multiple comparisons test was used to compare the buffer alone control to the other conditions (*** denotes p < 0.0001).
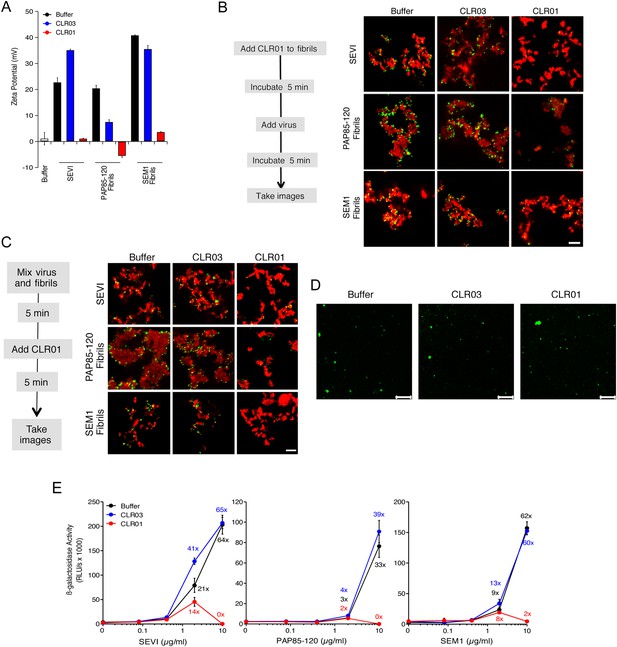
CLR01 neutralizes the positive surface charge of seminal amyloids and abrogates their ability to bind virions and enhance HIV infection.
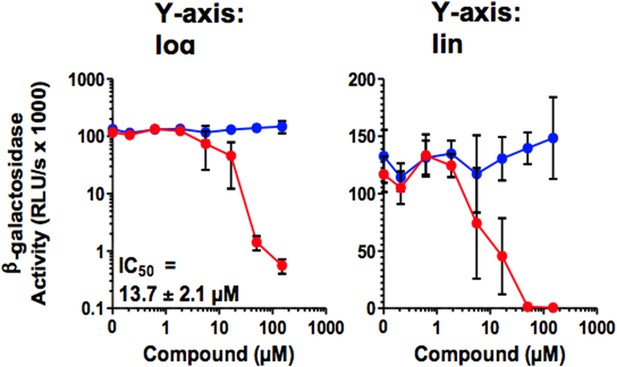
(A) Surface charge of seminal amyloids determined by zeta potential measurements. Fibrils were mixed with buffer, CLR01, or CLR03 and centrifuged for 10 min at 20,000×g. The pellets were resuspended in 1 mM KCl and zeta potential was measured. Values represent means ±SD (n = 3). (B) Fibrils (200 µg/ml) were pretreated with PBS, CLR01, or CLR03 in 20-fold excess for 5 min and stained with Proteostat dye. MLV-Gag-YFP particles (green) were added 1:2 and allowed to incubate with pretreated fibrils (red) for 5 min. Samples were analyzed using confocal microscopy. Scale bar: 5 µm. (C) CLR01 displaces virions from fibrils. MLV-Gag-YFP particles (green) were incubated with Proteostat-stained seminal amyloids (red) for 5 min. PBS, CLR01, or CLR03 were added at 20-fold excess, and after an additional 5 min of incubation, samples were analyzed by confocal microscopy. Scale bar: 5 µm. (D) CLR01 does not quench the fluorescence signal of MLV-Gag-YFP particles. MLV-Gag-YFP particles were treated with 300 µM CLR01 or CLR03 for 30 min at 37°C. Thereafter, virions were recovered in the pellet fraction via high-speed centrifugation and aliquots analyzed by confocal microscopy. Scale bar = 20 µm. (E) CLR01 antagonizes the HIV-1 enhancing activity of seminal amyloids. SEVI, PAP85-120, and SEM1(49-107) fibrils were mixed with a 20-fold excess of CLR01 or CLR03. CCR5-tropic HIV-1 was added and samples were used to inoculate TZM-bl cells. Infection rates were determined 3 days post infection. Values represent mean β-galactosidase activities derived from triplicate infections ±SD (RLU/s: relative light units per second). Numbers above symbols indicate the n-fold enhancement of infection.
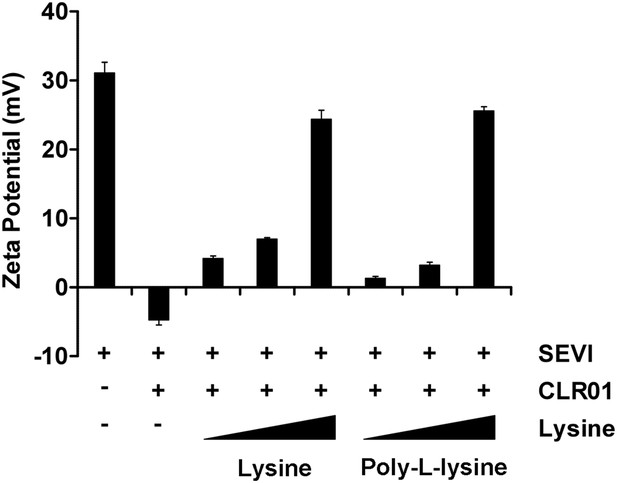
Lysine and poly-L-lysine dose-dependently antagonize CLR01 binding to SEVI.
SEVI fibrils were mixed with CLR01 and increasing amounts of lysine (1, 10 and 100-fold excess over CLR01) or poly-L-lysine (0.02, 0.25 and 2.5-fold excess of lysine monomer equivalents over CLR01) and centrifuged for 10 min at 20,000×g. The pellets were resuspended in 1 mM KCl and zeta potential was measured. Values represent means ±SD (n = 3).
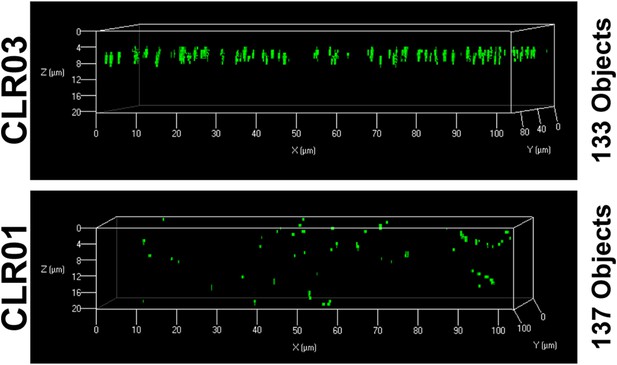
CLR01 does not quench the fluorescence signal of MLV-Gag-YFP particles.
MLV-Gag-YFP particles were treated with 300 µM CLR01 or CLR03 for 30 min at 37°C. Thereafter, virions were recovered in the pellet fraction via high-speed centrifugation and aliquots analyzed by confocal microscopy. Z-stack images of the samples revealed a similar amount of fluorescent particles after treatment with CLR01 or CLR03 but altered distribution in the slides.
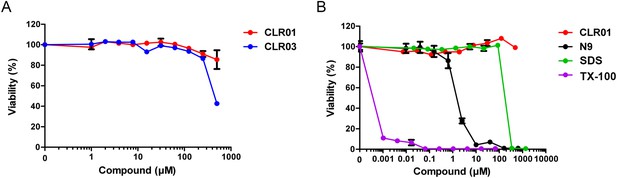
CLR01 is not cytotoxic.
TZM-bl cells were incubated for 3 days with the indicated concentrations of (A) CLR01 and CLR03 and (B) CLR01, nonoxynol 9 (N9), sodium dodecyl sulfate (SDS) and Triton X-100 (TX-100). Cell viability was measured in an MTT assay. Values represent means ±SD (n = 3).
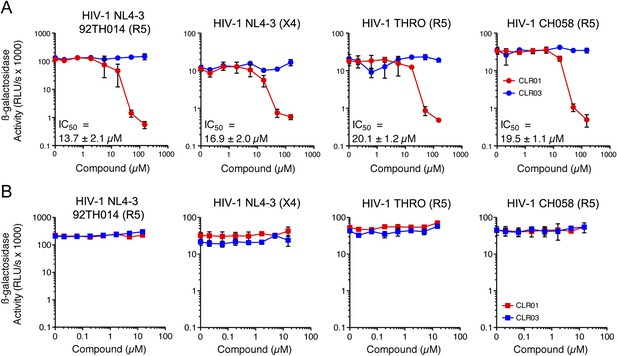
CLR01 has direct anti-HIV-1 activity.
(A) CLR01 blocks HIV-1 infection by targeting virions. The CXCR4 (X4)-tropic lab-adapted HIV-1 NL4-3 strain, a CCR5 (R5)-tropic V3 loop recombinant thereof, and two R5-tropic transmitted/founder viruses (THRO and CH058) were incubated with CLR01 or CLR03 for 10 min and then used to infect TZM-bl cells. Infection rates were determined 3 days post infection. Values represent mean β-galactosidase activities derived from triplicate infections ±SD (RLU/s: relative light units per second). (B) The antiviral activity of CLR01 is not directed against the cell. TZM-bl cells were exposed to CLR01 or CLR03 for 1 hr. Thereafter, cell culture medium was removed and cells were infected with the indicated viruses. Infection rates were determined 3 day post infection. Values represent mean β-galactosidase activities derived from triplicate infections ±SD (RLU/s: relative light units per second).
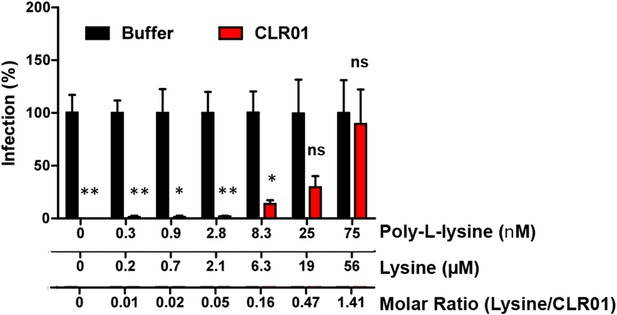
Poly-L-lysine counteracts the antiviral activity of CLR01.
CLR01 (40 µM) was titrated with poly-L-lysine. After a 10 min incubation with HIV-1 these mixtures were used to infect TZM-bl cells. Infection rates were determined 3 days post infection via a β-galactosidase assay. Values represent normalized mean infection rates ±SEM. Unpaired t-tests were used to compare the buffer control to the CLR01 condition at each poly-L-lysine concentration (ns denotes not significant; * denotes p < 0.05; ** denotes p < 0.01).
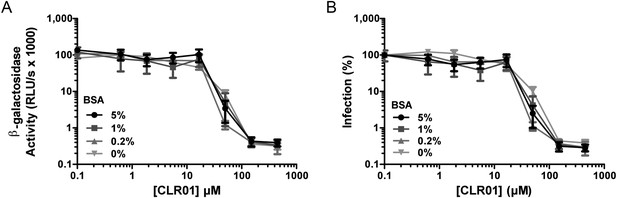
Coated BSA does not antagonize the antiviral activity of CLR01.
Microtiter plates were coated over night with the indicated amounts of BSA. After a 20-min incubation at room temperature, serial dilutions of CLR01 were added and incubated with HIV-1 for another 10 min at room temperature. With these mixtures, TZM-bl cells were infected. Infection rates were determined 3 days post infection. (A) Values represent mean β-galactosidase activities derived from triplicate infections ±SD (RLU/s: relative light units per second). (B) Values represent normalized mean infection rates derived from triplicate infections ±SD.
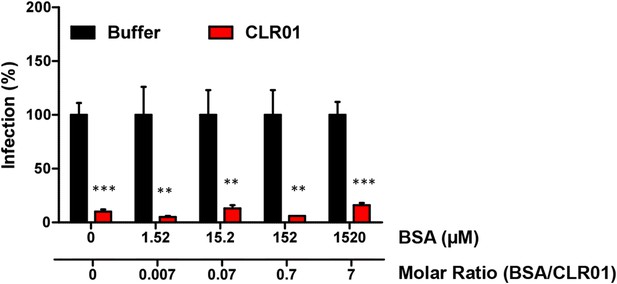
BSA does not antagonize the antiviral activity of CLR01.
CLR01 (220 µM) was titrated with the indicated concentrations of BSA. HIV-1 was incubated with these mixtures for 10 min and then used to infect TZM-bl cells. Infection rates were determined 3 days post infection via a β-galactosidase assay. Values represent normalized mean infection rates derived from triplicate measurements ±SD. Unpaired t-tests were used to compare the buffer control to the CLR01 condition at each BSA concentration (** denotes p < 0.01; *** denotes p < 0.001).
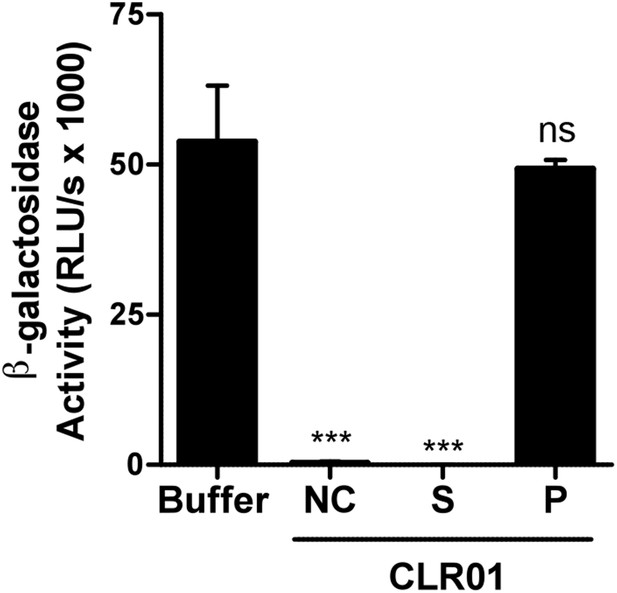
Preclearing CLR01 via centrifugation does not affect antiviral activity.
CLR01 (50 µM) was centrifuged at 20,000×g and 20°C for 20 min. HIV-1 92TH014 was incubated for 10 min with the supernatant (S) or the pellet fraction (P). As a control, virus was also incubated either with PBS (Buffer) or 50 µM CLR01 that was not centrifuged (NC). After infection of TZM-bl cells with these mixtures infection rates were measured 2 days post infection. Values represent mean β-galactosidase activities derived from triplicate infections ±SD (RLU/s: relative light units per second). Unpaired t-tests were used to compare the buffer control to each CLR01 condition at each poly-L-lysine concentration (ns denotes not significant; ** denotes p < 0.01; *** denotes p < 0.001).
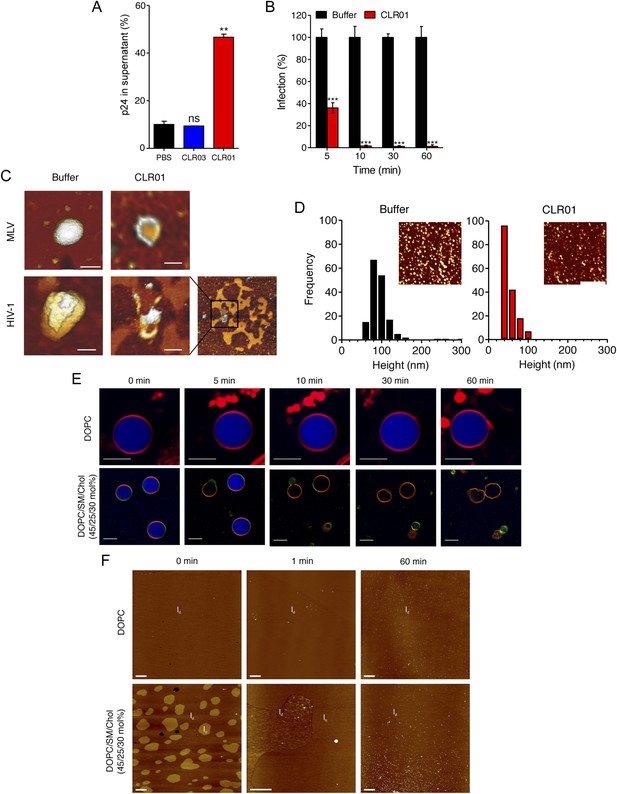
CLR01 destroys retroviral particles and selectively disrupts raft-rich membranes.
(A) CLR01 releases p24 capsid antigen from HIV particles. HIV-1 was incubated with PBS, 100 µM CLR03, or 100 µM CLR01 and centrifuged at 20,000×g and 4°C for 1 hr. The p24 content of the supernatant was determined via p24 ELISA. Values represent means ±SD. Unpaired t-tests were used to compare the buffer control to the CLR03 or CLR01 condition (ns denotes not significant; ** denotes p < 0.01). (B) HIV-1 was incubated at 37°C with 10 µM CLR01 or buffer control. Aliquots were taken after different time points and analyzed regarding their infectivity using TZM-bl reporter cells. Values represent normalized mean infection rates derived from triplicate measurements ±SD compared to the buffer control (100%). Unpaired t-tests were used to compare the buffer control to the CLR01 condition at each time point (*** denotes p < 0.001). (C) CLR01 destroys retroviral particles. Images obtained by atomic force microscopy (AFM) show single MLV and glycoprotein-deficient HIV particles before and after treatment with 100 µM CLR01. Scale bar: 100 nm. (D) CLR01 destroys MLV particles. Height distribution of MLV particles after treatment with buffer (left panel) or 100 µM CLR01 (right panel). Values were derived from AFM images shown in the insets. Scale bar: 2 µm. (E) CLR01 selectively destroys membranes with high lipid raft content. Giant unilamellar vesicles (GUVs) consisting of pure DOPC were labeled with N-Rh-DHPE (red). GUVs containing a mixture of DOPC, SM and Chol (45/25/30 mol%) were labeled with N-Rh-DHPE (red) and Bodipy-Chol (green). Both types of GUVs were filled with buffer containing the fluorophore ATTO 647 (blue) and treated with 150 µM CLR01 for the indicated times before images were taken by confocal microscopy. Note that ATTO 647 remains inside the DOPC GUVs treated with CLR01, but escapes the DOPC/SM/Chol GUVs treated with CLR01. Scale bar: 10 μm. (F) Upper panel: AFM images (10 µm scans) of a pure DOPC lipid membrane on mica before injection (0 min) and 1 min and 60 min after injection of 800 µl of 150 µM CLR01 in 10 mM NaH2PO4, pH 7.6 into the AFM fluid cell. The whole scan area is shown with a vertical color scale from dark brown to white corresponding to an overall height of 8 nm. The thickness of the hydrated membrane is 3.7 nm. Lower panel: AFM image (10 µm scan) of a DOPC/SM/Chol (45/25/30 mol%) lipid membrane on mica before injection (0 min) and 1 min and 60 min after injection of 800 µl of 150 µM CLR01 in 10 mM NaH2PO4, pH 7.6 into the AFM fluid cell. The whole scan area is shown with a vertical color scale from dark brown to white corresponding to an overall height of 8 nm and indicating a homogeneous lipid bilayer with coexisting domains in lo (liquid-ordered) and ld (liquid-disordered domain) phase. The height difference between domains is 1 nm; the ld phase has a thickness of 4.0 nm.
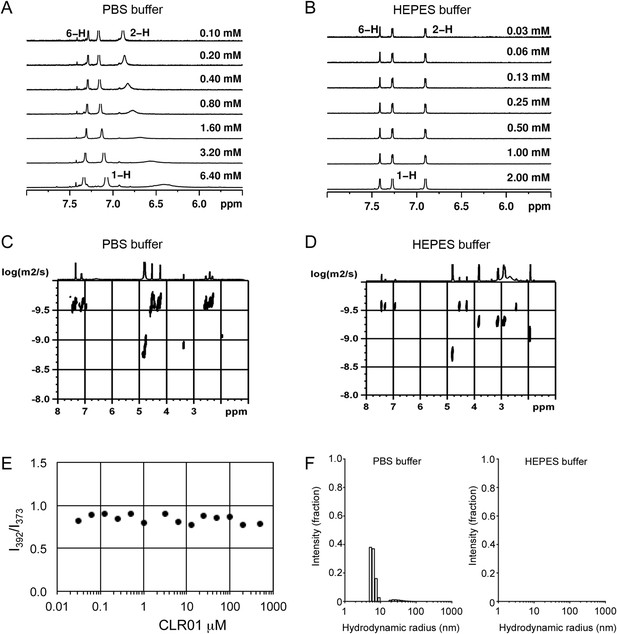
CLR01 does not form colloid or micelle aggregates.
(A) 1H NMR dilution titration of CLR01 from 6 mM to 100 μM in PBS leads to very weak dimerization; (B) in HEPES buffer CLR01 remains monomeric over the entire concentration range. (C) DOSY spectrum of 6 mM CLR01 in PBS buffer; cross peaks at 4.8 ppm correspond to aqueous solvent. (D) DOSY spectrum of 2 mM CLR01 in HEPES buffer; cross peaks between 2 and 4 ppm correspond to HEPES buffer, cross peaks at 4.8 ppm correspond to the aqueous solvent. (E) Attempted cmc determination by analysis of the I392/I373 emission intensity ratio of 10 μM PBS-buffered pyrene solutions containing CLR01 from 30 nM to 500 μM produces a straight line, as opposed to the SDS control (data not shown). (F) DLS measurements of CLR01 (1 mM) in PBS or HEPES buffer show formation of minute amounts of particles with RH ∼5–8 nm or ∼20–40 nm in PBS. No particles were detected in HEPES buffer. The vast majority of CLR01 molecules are too small to be detected and are presumably monomeric.
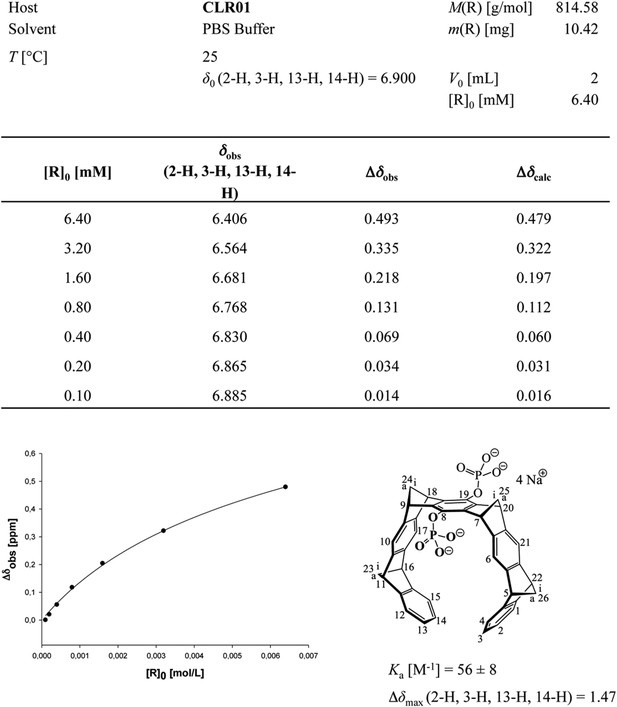
Aromatic proton shifts during dilution of PBS-buffered CLR01.
Upfield shifts of the inner aromatic protons (2-H, 3-H, 13-H, 14-H) document their mutual inclusion inside the cavity of the other host molecule, leading to specific anisotropic shielding exclusively in the dimer. Thus, the NMR shift changes prove formation of weak dimers (Ka ∼60 M−1) and exclude unspecific aggregation, which would manifest as smaller chemical shift changes evenly distributed among the CH protons of CLR01.
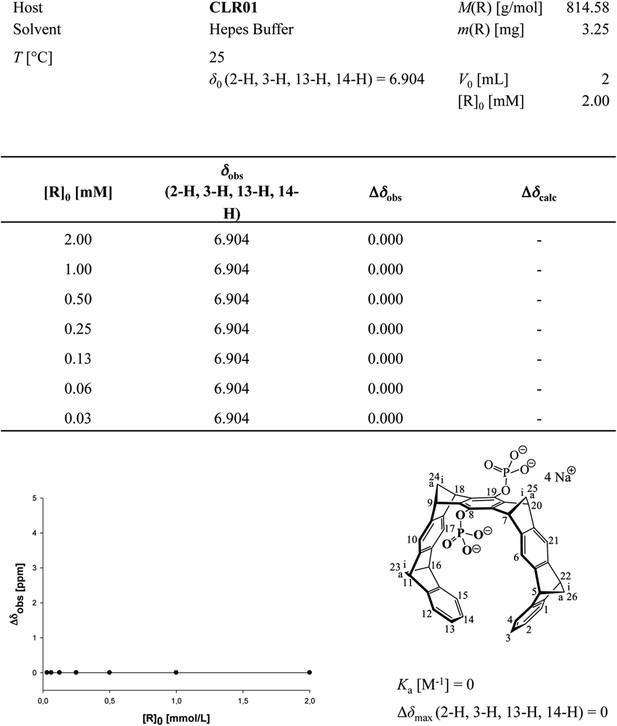
Aromatic proton shifts during dilution of HEPES-buffered CLR01.
Dilution titration of CLR01 in HEPES buffer did not produce any upfield shift of the aromatic protons; on the contrary, the monomer spectrum appeared at all concentrations, with sharp signals for all three protons.
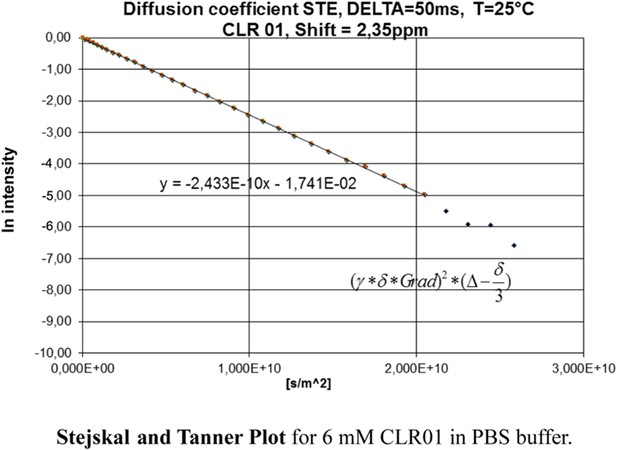
Stejskal and Tanner Plot.
Signal intensity in diffusion NMR experiments obeys the following law: ln(I/I0) = −γ2δ2G2(∆ − δ/3) D, where I = intensity at a given G, I0 = Intensity at G = 0, γ = Gyromagnetic ratio (for 1H: γ = 2.675 × 108T−1s−1), δ = Length of diffusion gradient, ∆ = Diffusion shift, G = Gradient field intensity, and D = Diffusion coefficient. Signal intensities are used to generate a Stejskal Tanner-Plot lg(I/I0) vs G2. Its slope yields the diffusion coefficient. Diffusion coefficient for CLR01 (6.4 mM in PBS): 2.433 × 10−10 m2s−1. By means of the Stokes Einstein equation one calculates the hydrodynamic radius: r(s) = (k*T)/(6*pi*η*D). The hydrodynamic radius for CLR01 (6.4 mM) was determined at 1.0075 × 10−9 m ∼10 Å in PBS buffer; for CLR01 (0.2 mM) the diffusion coefficient dropped to 2.702 × 10−10 m2s−1, corresponding to a hydrodynamic radius of 9.072 × 10−9 m ∼9 Å. In HEPES buffer, the diffusion coefficient for CLR01 (2.0 mM) was determined at 2.449 × 10−10 m2s−1, corresponding to hydrodynamic radius r(s) of 1.001 × 10−9 m ∼10 Å; for CLR01 (0.5 mM) the diffusion coefficient dropped to 2.579 × 10−10 m2s−1, corresponding to a hydrodynamic radius of 9.504 × 10−9 m ∼9 Å. In summary, almost identical r(s) values were found for PBS and HEPES buffer. Thus, at various concentrations even above those used in experiments, the tweezers only produced a hydrodynamic radius slightly above the monomeric species.
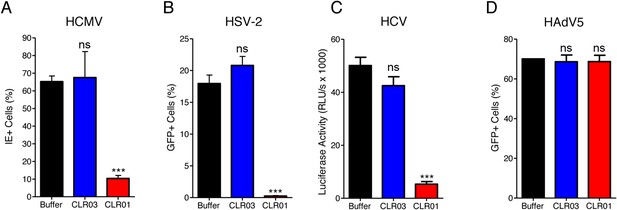
CLR01 is a broad-spectrum inhibitor of enveloped viruses.
(A) Human cytomegalovirus was incubated with PBS, 100 µM CLR03, or 100 µM CLR01. Afterwards, HFF cells were infected and immediate early (IE) antigen positive cells were counted 1 day post infection as a measure for infectivity. Values are means ±SD (n = 3). Unpaired t-tests were used to compare the buffer control to the CLR03 or CLR01 condition (ns denotes not significant; *** denotes p < 0.001). (B) Herpes simplex virus type 2 comprising a GFP reporter gene was treated with PBS, 100 µM CLR03 or 100 µM CLR01 and added to Vero cells. GFP-positive cells were counted using flow cytometry 2 days post infection. Values represent means ±SD (n = 3). Unpaired t-tests were used to compare the buffer control to the CLR03 or CLR01 condition (ns denotes not significant; *** denotes p < 0.001). (C) A luciferase encoding hepatitis C virus was treated with 150 µM CLR01 or 150 µM CLR03 and used for infection of Huh-7.5 reporter cells. Infection was measured 3 days post infection. Values represent means ±SEM (n = 3). Unpaired t-tests were used to compare the buffer control to the CLR03 or CLR01 condition (ns denotes not significant; *** denotes p < 0.001). (D) A GFP-reporter adenovirus type 5 was added to A549 cells after treatment with 158 µM CLR01 or 158 µM CLR03. GFP positive cells were counted using flow cytometry 1 day post infection. Values represent means ±SD (n = 3). Unpaired t-tests were used to compare the buffer control to the CLR03 or CLR01 condition (ns denotes not significant).
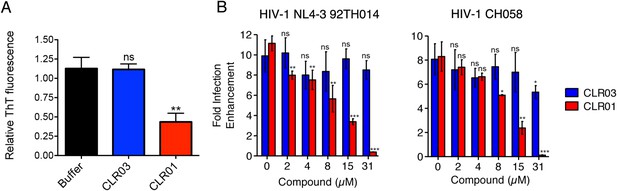
CLR01 diminishes the infection enhancing property of semen.
(A) SEVI fibrils were formed in an artificial semen simulant (AS) (Owen and Katz, 2005). The resulting fibrils were then diluted to 20 µM in AS and treated with 200 µM CLR01 or CLR03, or with buffer for 2 hr. Fibril integrity was assessed using ThT fluorescence. Values represent means ±SEM (n = 4). A one-way ANOVA with the post hoc Dunnett's multiple comparisons test was used to compare the buffer alone control to the other conditions (ns denotes not significant; ** denotes p < 0.01). (B) CLR01 abrogates semen-mediated enhancement of HIV infection. Seminal plasma (10%) or cell culture medium containing CLR01 or CLR03 was mixed with CCR5-tropic HIV-1 or transmitter/founder HIV-1 CH058. After 10 min, TZM-bl cells were infected and infectivity was measured 3 day post infection. Shown are the n-fold increased infection rates obtained for semen-treated virus relative to those of medium-treated virus. Values represent means ±SD (n = 4). Unpaired t-tests were used to compare the buffer control (0 µM compound) to the CLR03 or CLR01 condition at each concentration (ns denotes not significant; * denotes p < 0.05; ** denotes p < 0.01; *** denotes p < 0.001).
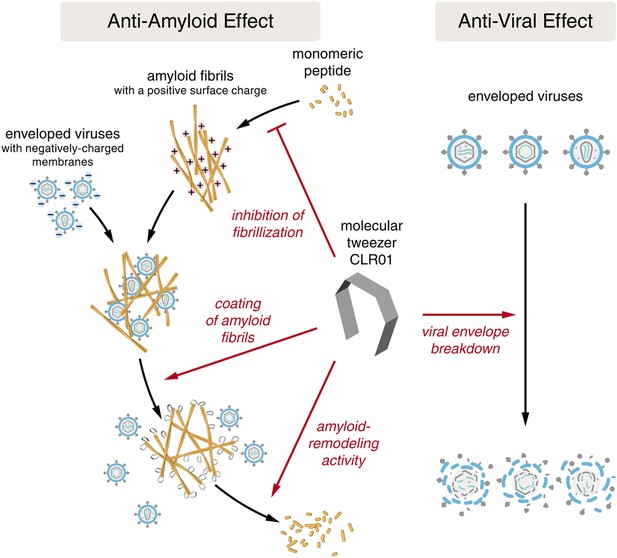
CLR01 acts as a dual-function inhibitor of viral infection.
Schematic overview of the anti-amyloid and antiviral effects of CLR01.