Expanding xylose metabolism in yeast for plant cell wall conversion to biofuels
Figures
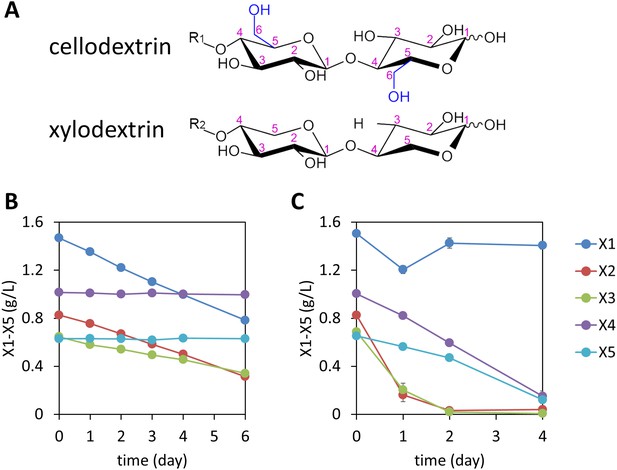
Consumption of xylodextrins by engineered S. cerevisiae.
(A) Two oligosaccharide components derived from the plant cell wall. Cellodextrins, derived from cellulose, are a major source of glucose. Xylodextrins, derived from hemicellulose, are a major source of xylose. The 6-methoxy group (blue) distinguishes glucose derivatives from xylose. R1, R2 = H, cellobiose or xylobiose; R1 = β-1,4-linked glucose monomers in cellodextrins of larger degrees of polymerization; R2 = β-1,4-linked xylose monomers in xylodextrins of larger degrees of polymerization. (B) Xylose and xylodextrins remaining in a culture of S. cerevisiae grown on xylose and xylodextrins and expressing an XR/XDH xylose consumption pathway, CDT-2, and GH43-2, with a starting cell density of OD600 = 1 under aerobic conditions. (C) Xylose and xylodextrins in a culture as in (B) but with a starting cell density of OD600 = 20. In both panels, the concentrations of xylose (X1) and xylodextrins with higher DPs (X2–X5) remaining in the culture broth after different periods of time are shown. All experiments were conducted in biological triplicate, with error bars representing standard deviations.
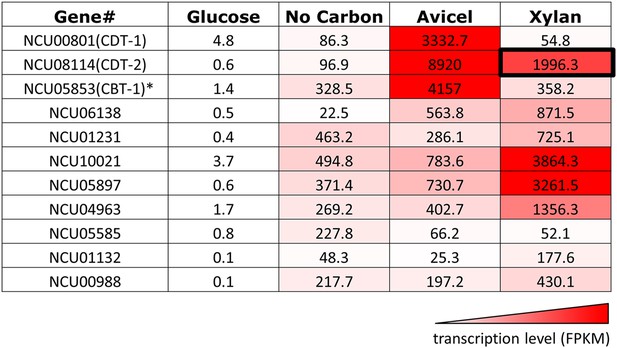
Transcriptional levels of transporters expressed in N. crassa grown on different carbon sources.
Transcript levels reported in fragments per kilobase per million reads (FPKM) are derived from experiments published in Coradetti et al. (2012); Sun et al. (2012). *CBT-1 transports cellobionic acid, the product of lytic polysaccharide monooxygenases (LPMOs, or CaZy family AA9 and AA10) (Xiong et al., 2014).
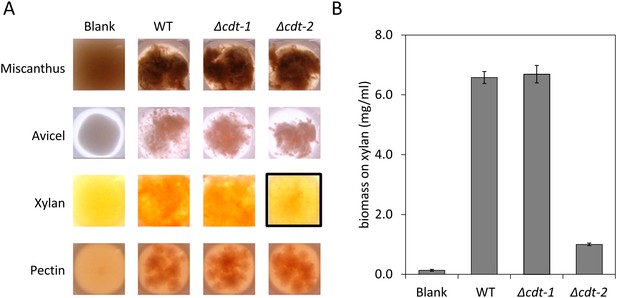
Growth of N. crassa strains on different carbon sources.
(A) Wild-type (WT) N. crassa, or N. crassa with deletions of transporters cdt-1 (Δcdt-1) or cdt-2 (Δcdt-2), were grown on M. giganteus plant cell walls, or purified plant cell wall components. Avicel is a form of cellulose derived from plant cell walls. The black box shows the severe growth phenotype of the Δcdt-2 strain grown on xylan medium. (B) N. crassa biomass accumulation after 3 days of growth on xylan.
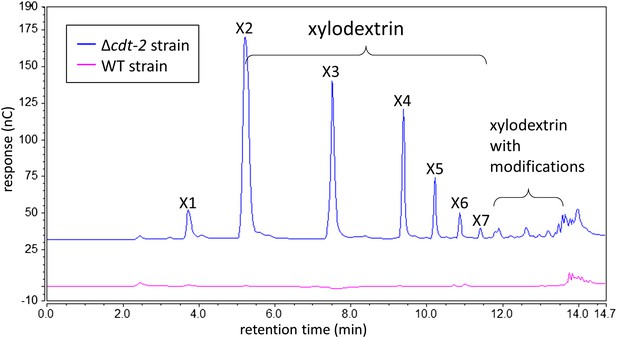
Xylodextrins in the xylan culture supernatant of the N. crassa Δcdt-2 strain.
25 µl of 1:200 diluted N. crassa xylan culture supernantant was analyzed by HPAEC on a CarboPac PA200 column. While no detectable soluble sugars were found in the culture supernatant of the wild-type strain (magenta line), the Δcdt-2 strain (blue line) left a high concentration of unmodified and modified xylodextrins in the culture supernatant. Little xylose was found, indicating xylose was transported by means of different transporters.
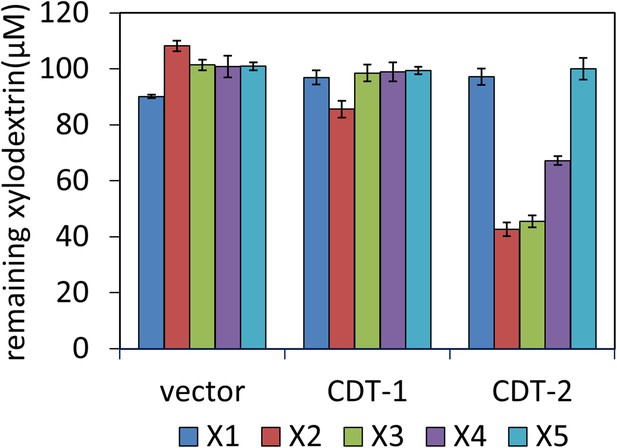
Transport of xylodextrins into the cytoplasm of S. cerevisiae strains expressing N. crassa transporters.
The starting xylodextrin concentration for each purified component was 100 µM. The remaining xylose (X1) and xylodextrins in the culture media are shown for experiments with S. cerevisiae harboring an empty expression plasmid (vector), or with S. cerevisiae individually expressing transporters CDT-1 or CDT-2. Xylodextrins used include xylobiose (X2), xylotriose (X3), xylotetraose (X4), and xylopentaose (X5). Error bars indicate standard deviations of biological triplicates.
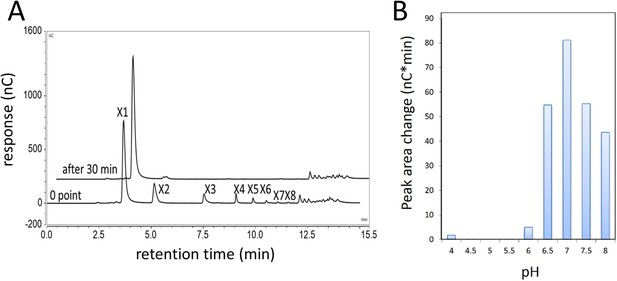
Xylobiase activity of the predicted β-xylosidase GH43-2.
(A) GH43-2 hydrolysis of xylodextrins with degrees of polymerization from at least 2–8 (X2–X8). The 30 min chromatogram is offset for clarity. (B) The pH optimum of GH43-2, determined by measuring the extent of hydrolysis of xylobiose to xylose. The HPAEC chromatogram peak area change for xylose is shown.
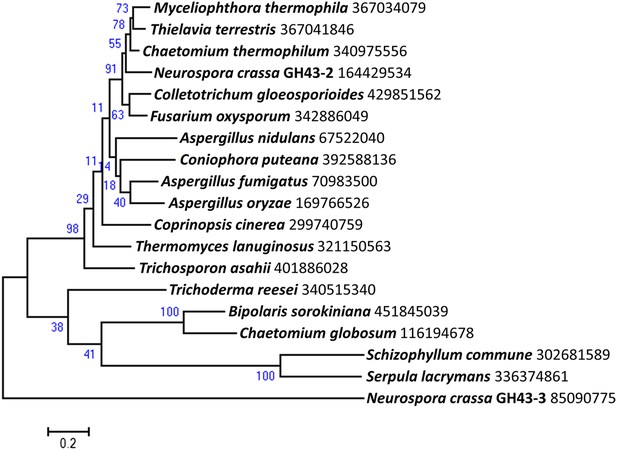
Phylogenetic distribution of predicted intracellular β-xylosidases GH43-2 in filamentous fungi.
Homologs of GH43-2 (NCU01900) were found with BLAST (Altschul et al., 1997) queries of respective sequence against NCBI protein database. Representative sequences from a diversified taxonomy were chosen and aligned with the MUSCLE algorithm (Edgar 2004). A maximum likelihood phylogenetic tree was calculated based on the alignment with the Jones-Taylor-Thornton model by using software MEGA v6.05 (Tamura et al., 2013). Xylan-induced extracellular GH43-3 (NCU05965) was used as an outgroup. The NCBI GI numbers of the sequences used to build the phylogenetic tree were indicated besides the species names. 1000 bootstrap replicates were performed to calculate the supporting values shown on the branches. The scale bar indicates 0.2 substitutions per amino acid residue.
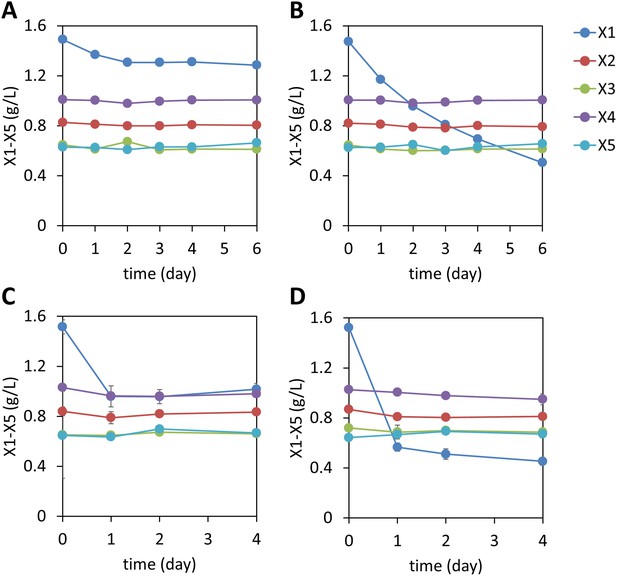
Xylodextrin consumption profiles of S. cerevisiae strains lacking the xylodextrin pathway.
Shown are the concentrations of the remaining sugars in the culture broth after different periods of time of (A) the WT D452-2 strain with starting cell density at OD600 = 1, (B) D452-2 with a S. stipitis xylose utilization pathway (plasmid pLNL78, Table 1) with a starting cell density at OD600 = 1, (C) WT D452-2 strain with a starting cell density at OD600 = 20, and (D) D452-2 with a S. stipitis xylose utilization pathway (plasmid pLNL78) with a starting cell density at OD600 = 20. In all panels, xylose (X1) and xylodextrins of higher DPs (X2–X5) are shown. Error bars represent standard deviations of biological triplicates (panels A–D).
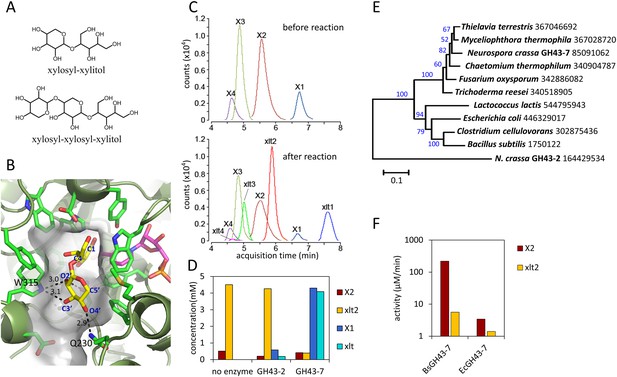
Production and enzymatic breakdown of xylosyl-xylitol.
(A) Structures of xylosyl-xylitol and xylosyl-xylosyl-xylitol. (B) Computational docking model of xylobiose to CtXR, with xylobiose in yellow, NADH cofactor in magenta, protein secondary structure in dark green, active site residues in bright green and showing side-chains. Part of the CtXR surface is shown to depict the shape of the active site pocket. Black dotted lines show predicted hydrogen bonds between CtXR and the non-reducing end residue of xylobiose. (C) Production of xylosyl-xylitol oligomers by N. crassa xylose reductase, XYR-1. Xylose, xylodextrins with DP of 2–4, and their reduced products are labeled X1–X4 and xlt1–xlt4, respectively. (D) Hydrolysis of xylosyl-xylitol by GH43-7. A mixture of 0.5 mM xylobiose and xylosyl-xylitol was used as substrates. Concentration of the products and the remaining substrates are shown after hydrolysis. (E) Phylogeny of GH43-7. N. crassa GH43-2 was used as an outgroup. 1000 bootstrap replicates were performed to calculate the supporting values shown on the branches. The scale bar indicates 0.1 substitutions per amino acid residue. The NCBI GI numbers of the sequences used to build the phylogenetic tree are indicated beside the species names. (F) Activity of two bacterial GH43-7 enzymes from B. subtilis (BsGH43-7) and E. coli (EcGH43-7).
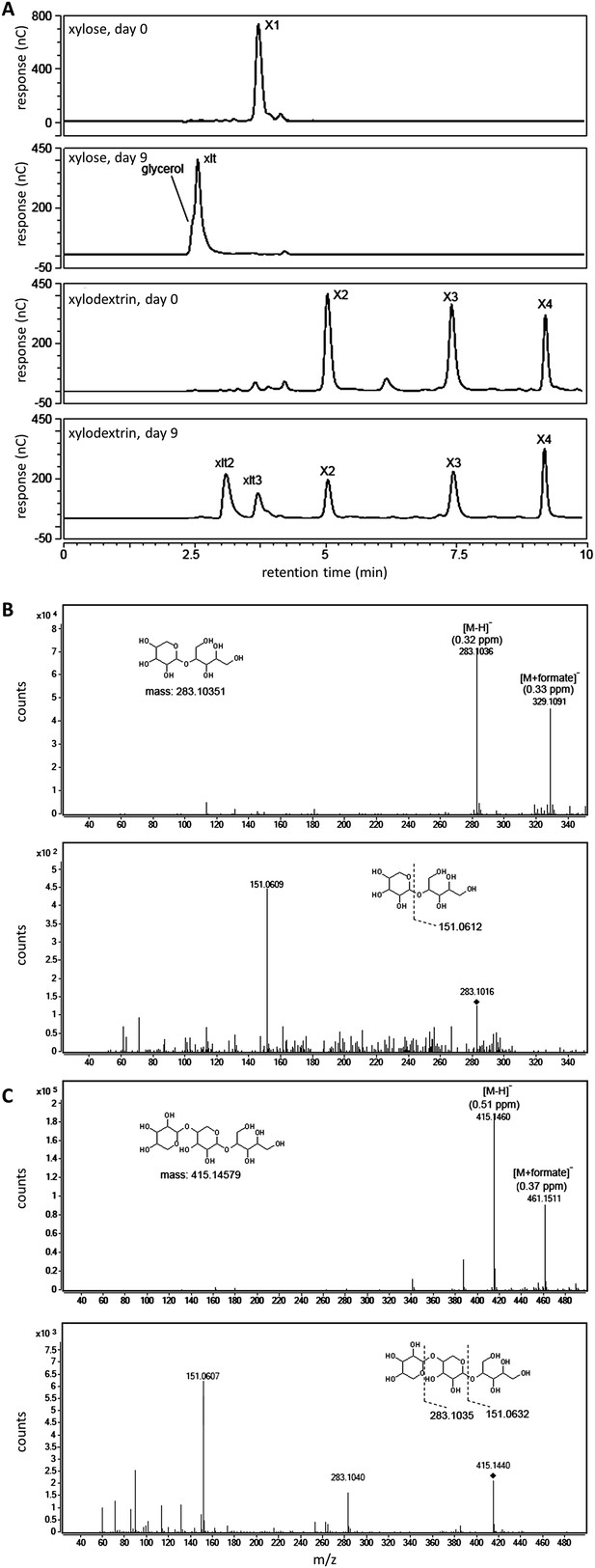
Xylosyl-xylitol oligomers generated in yeast cultures with xylodextrins as the sole carbon source.
(A) Carbohydrates from culture supernatants of strain SR8U expressing CDT-2 and GH43-2 (plasmid pXD8.4), resolved by HPAEC, abbreviated as follows: X1, xylose; X2, xylobiose; X3, xylotriose; X4, xylotetraose; xlt, xylitol; xlt2, xylosyl-xylitol; xlt3, xylosyl-xylosyl-xylitol. (B) LC-MS and LC-MS/MS spectra for xylosyl-xylitol. High-resolution MS spectra show m/z ratios for the negative ion mode. The deprotonated and formate adduct ions were determined with an accuracy of 0.32 and 0.33 ppm, respectively. The MS/MS spectrum in the lower panel shows the product ion matching the predicted fragment. The parental ion, [xylosyl-xylitol + H]−, is denoted with the black diamond mark. (C) LC-MS and LC-MS/MS spectra for xylosyl-xylosyl-xylitol. The deprotonated and formate adduct ions were determined with an accuracy of 0.51 and 0.37 ppm, respectively. The MS/MS spectrum in the lower panel shows the product ions matching the predicted fragments. The parental ion, [xylosyl-xylosyl-xylitol + H]−, is denoted with the black diamond mark.
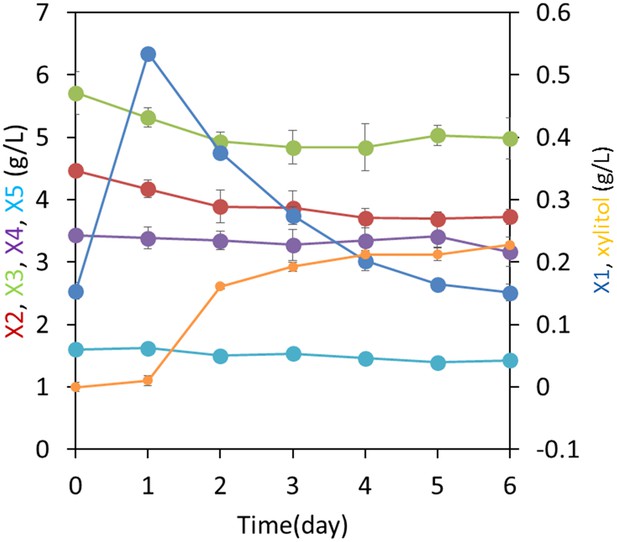
Xylodextrin metabolism by a co-culture of yeast strains to identify enzymatic source of xylosyl-xylitol.
A mixture of a xylose utilizing strain (SR8) with a cell density at OD600 = 1.0 and a xylodextrin hydrolyzing strain (D452-2 expressing CDT-2 and GH43-2 from plasmid pXD8.4) with a cell density at OD600 = 20 was co-cultured in a medium containing 2% xylodextrin. Xylobiose (X2) and xylotriose (X3) decreased, whereas xylose (X1) initially increased. Subsequent X1 consumption correlated with production of xylitol. Notably, xylosyl-xylitol oligomers were not detected, suggesting that the xylodextrin reductase activity was present only in the xylose-fermenting strain expressing XR. Error bars represent standard deviations of biological triplicates.
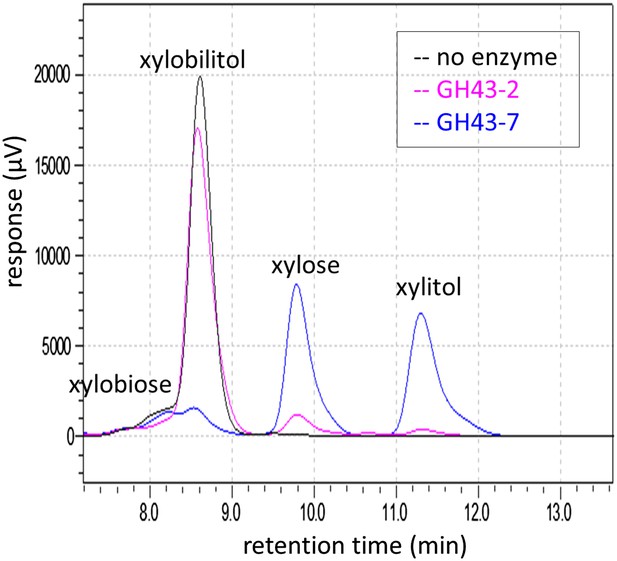
Chromatogram of xylosyl-xylitol hydrolysis products generated by β-xylosidases.
Reaction products from the enzymatic assays in Figure 2D were resolved by ion-exclusion HPLC. Peak areas were used to quantify the concentration of substrates and products at the end of the reaction.
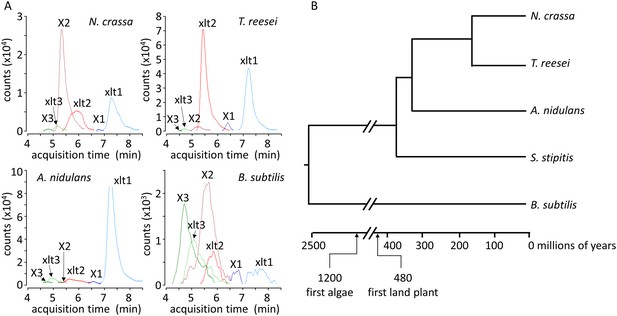
Xylosyl-xylitol and xylosyl-xylosyl-xylitol production by a range of microbes.
(A) Xylodextrin-derived carbohydrate levels seen in chromatograms of intracellular metabolites for N. crassa, T. reesei, A. nidulans and B. subtilis grown on xylodextrins. Compounds are abbreviated as follows: X1, xylose; X2, xylobiose; X3, xylotriose; X4, xylotetraose; xlt, xylitol; xlt2, xylosyl-xylitol; xlt3, xylosyl-xylosyl-xylitol. (B) Phylogenetic tree of the organisms shown to produce xylosyl-xylitols during growth on xylodextrins. Ages taken from Wellman et al. (2003); Galagan et al. (2005); Hedges et al. (2006).
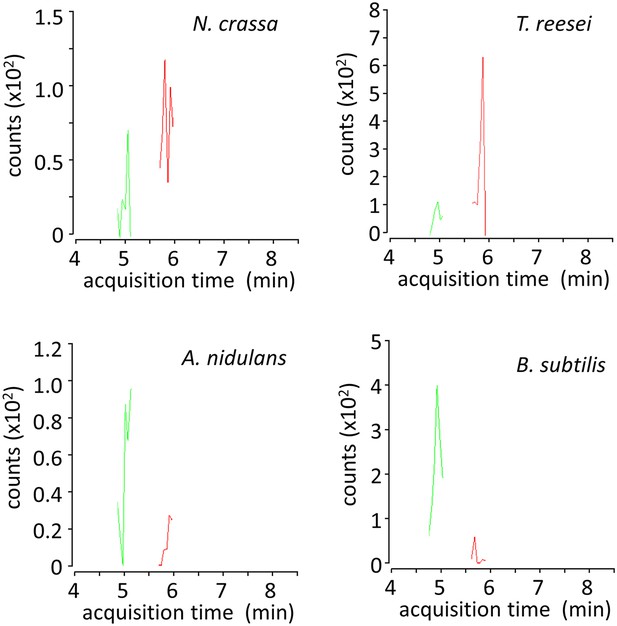
LC-MS/MS multiple reaction monitoring chromatograms of xylosyl-xylitols from cultures of microbes grown on xylodextrins.
Shown are MS/MS transitions for xylosyl-xylitol (in red, m/z 283.1035 → 151.0612 transition) and xylosyl-xylosyl-xylitol (in green, m/z 415.1457 → 151.0612 transition) analyzed from intracellular metabolites of N. crassa, T. reesei, A. nidulans and B. subtilis grown on xylodextrins, after separation by liquid chromatography.
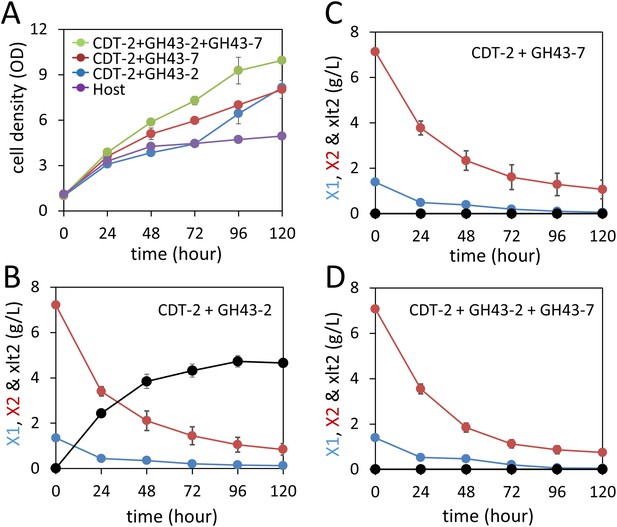
Aerobic consumption of xylodextrins with the complete xylodextrin pathway.
(A) Yeast growth curves with xylodextrin as the sole carbon source under aerobic conditions with a cell density at OD600 = 1. Yeast strain SR8U without plasmids, or transformed with plasmid expressing CDT-2 and GH43-2 (pXD8.4), CDT-2 and GH43-7 (pXD8.6) or all three genes (pXD8.7) are shown. (B–D) Xylobiose consumption with xylodextrin as the sole carbon source under aerobic conditions with a cell density of OD600 = 20. Xylosyl-xylitol (xlt2) accumulation was only observed in the SR8U strain bearing plasmid pXD8.4, that is, lacking GH43-7. Error bars represent standard deviations of biological triplicates (panels A–D).
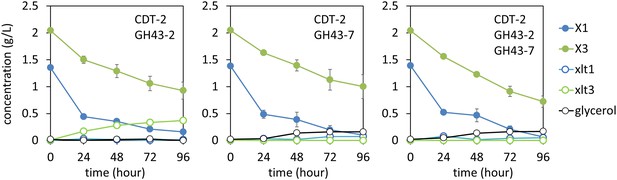
Culture media composition during yeast growth on xylodextrin.
Yeast growth with xylodextrin as the sole carbon source (concentration g/l) under aerobic conditions with a cell density at OD600 = 20. Yeast strain SR8 transformed with plasmid expressing CDT-2 and GH43-2 (pXD8.4), CDT-2 and GH43-7 (pXD8.6), or all three genes (pXD8.7). All growth experiments were performed in biological triplicate, and error bars indicate the standard deviation between experiments.
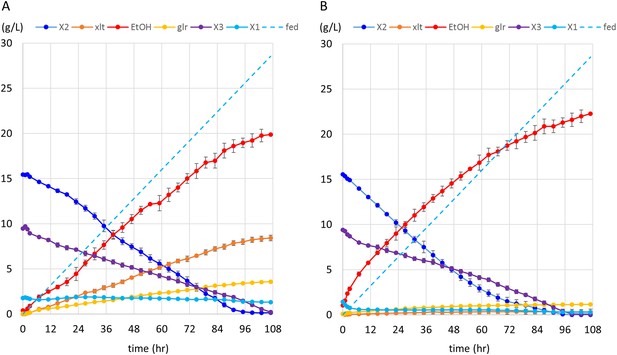
Anaerobic fermentation of xylodextrins in co-fermentations with xylose or glucose.
(A) Anaerobic fermentation of xylodextrins and xylose, in a fed-batch reactor. Strain SR8U expressing CDT-2, GH43-2, and GH43-7 (plasmid pXD8.7) was used at an initial OD600 of 20. Solid lines represent concentrations of compounds in the media. Blue dotted line shows the total amount of xylose added to the culture over time. Error bars represent standard deviations of biological duplicates. (B) Anaerobic fermentation of xylodextrins and glucose, in a fed-batch reactor. Glucose was not detected in the fermentation broth. Error bars represent standard deviations of biological duplicates.
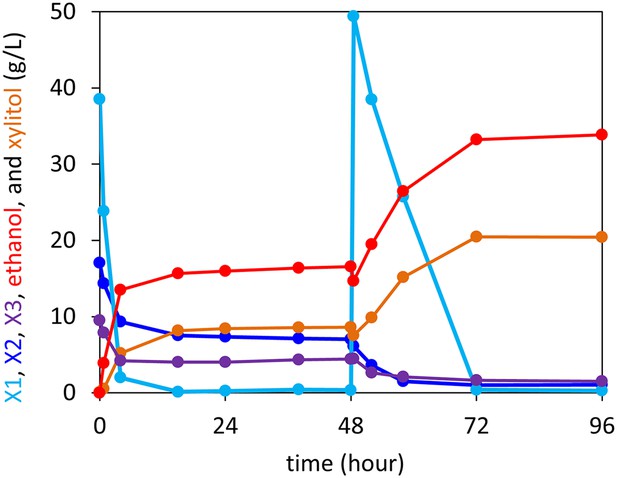
Anaerobic xylodextrin utilization in the presence of xylose.
Strain carrying the complete xylodextrin pathway (CDT-2, GH43-2, GH43-7, XR/XDH) grown under anaerobic conditions in oMM media (Lin et al., 2014) containing 4% xylose and 3% xylodextrin. The consumption of xylobiose (X2) and xylotriose (X3) stalled when xylose (X1) was depleted and resumed after supplying additional xylose at hour 48. This experiment is representative of those carried out with different xylose to xylodextrin ratios.
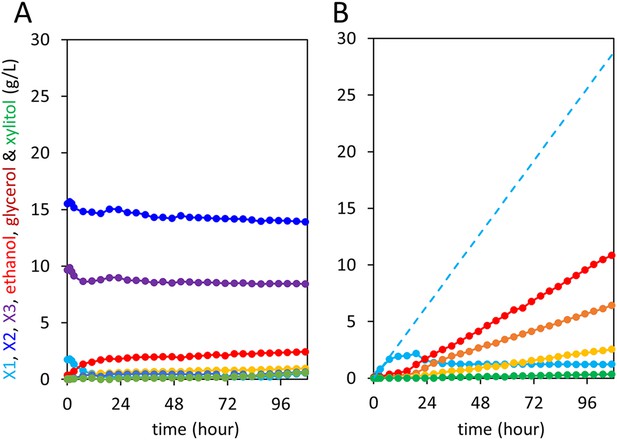
Control anaerobic fermentations with S. cerevisiae strain expressing the complete xylodextrin utilization pathway.
Strain SR8U with plasmid pXD8.7 expressing CDT-2, GH43-2, and GH43-7 was used at an initial OD600 of 20. Solid lines represent concentrations of compounds in the media. Blue dotted line shows the total amount of xylose added to the culture over time. (A) Fermentation profile of the strain in oMM medium containing 4% xylodextrin in the reactor without feeding xylose. (B) Fermentation profile of the strain in oMM medium without xylodextrin in the reactor but with continuous xylose feeding.
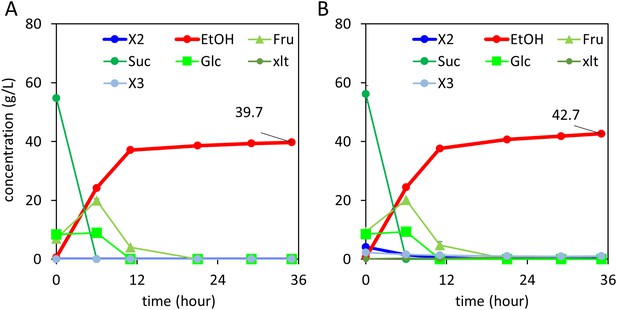
Xylodextrin and sucrose co-fermentations.
(A) Sucrose fermentation. Vertical axis, g/l; horizontal axis, time in hours. (B) Xylodextrin and sucrose batch co-fermentation using strain SR8U expressing CDT-2, GH43-2, and GH43-7 (plasmid pXD8.7). Vertical axis, g/l; horizontal axis, time in hours. The xylodextrins were supplied at 10 g/l which containing xylobiose (4.2 g/l) and xylotriose (2.3 g/l). Not fermented in the timeframe of this experiment, the xylodextrin sample also included xylotetraose and xylopentaose, in addition to hemicellulose modifiers such as acetate.
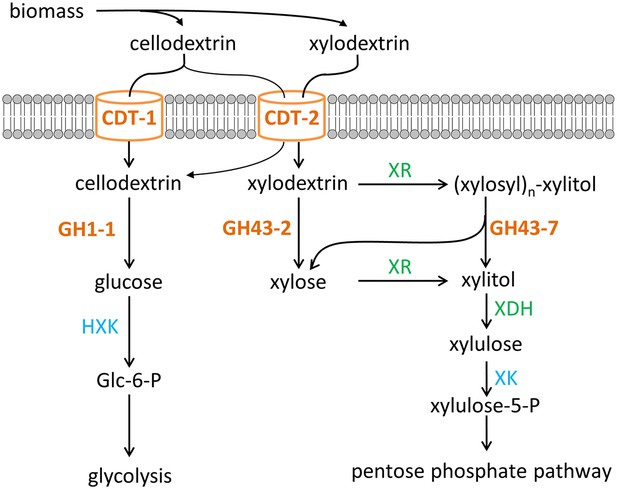
Two pathways of oligosaccharide consumption in N. crassa reconstituted in S. cerevisiae.
Intracellular cellobiose utilization requires CDT-1 or CDT-2 along with β-glucosidase GH1-1 (Galazka et al., 2010) and enters glycolysis after phosphorylation by hexokinases (HXK) to form glucose-6-phosphate (Glc-6-P). Intracellular xylodextrin utilization also uses CDT-2 and requires the intracellular β-xylosidases GH43-2 and GH43-7. The resulting xylose can be assimilated through the pentose phosphate pathway consisting of xylose/xylodextrin reductase (XR), xylitol dehydrogenase (XDH), and xylulokinase (XK).
Tables
A list of plasmids used in this study
| Plasmid | Genotype and use | Use | Ref. |
|---|---|---|---|
| pRS426_NCU08114 | PPGK1-CDT-2 | transport assay | (Galazka et al., 2010) |
| pRS423_GH43-2 | PTEF1-GH43-2 | enzyme purification | this study |
| pRS423_GH43-7 | PTEF1-GH43-7 | enzyme purification | this study |
| pRS313_NcXR | PCCW12-NcXR | enzyme purification | this study |
| pET302_EcGH43-7 | EcGH43-7 | enzyme purification | this study |
| pET302_BsGH43-7 | BsGH43-7 | enzyme purification | this study |
| pLNL78 | PRNR2-SsXK::PTEF1-SsXR::PTEF1-SsXDH | fermentation | (Galazka et al., 2010) |
| pXD2 | PRNR2-SsXK::PTEF1-SsXR::PTEF1-SsXDH::PPGK1-CDT-2::PTEF1-GH43-2 | fermentation | this study |
| pXD8.4 | PCCW12-CDT-2::PCCW12-GH43-2 | fermentation | this study |
| pXD8.6 | PCCW12-CDT-2::PCCW12-GH43-7 | fermentation | this study |
| pXD8.7 | PCCW12-CDT-2::PCCW12-GH43-7::PCCW12-GH43-7 | fermentation | this study |





