A FYVE zinc finger domain protein specifically links mRNA transport to endosome trafficking
Figures
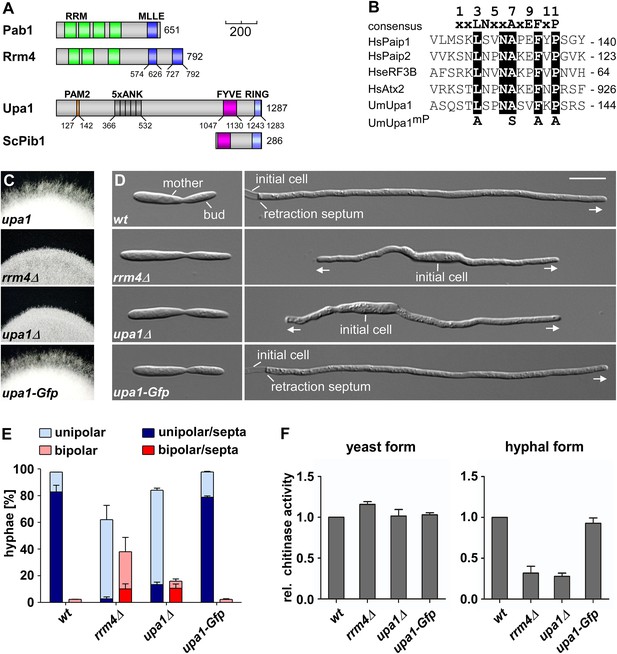
Loss of Upa1 causes defects in hyphal growth.
(A) Schematic representation of proteins drawn to scale (bar, 200 amino acids) using the following colouring: green, RNA recognition motif (RRM); dark blue, MLLE domain (SMART E-values 6.7 and 0.35 for Rrm4, Letunic et al., 2009); red, PAM2 motif; dark grey, Ankyrin repeats; purple, FYVE domain; light blue, RING domain. (B) Comparison of PAM2 sequences found in Upa1 (accession number UMAG_12183) with those of human proteins, such as Paip1 (accession number NP_006442.2), Paip2 (accession number CAG38520.1), eRF3B (accession number CAB91089.1), and Atx2 (accession number NP_002964.3). (C) Edge of colonies growing on charcoal-containing medium under hyphae-inducing conditions (48 h.p.i.). (D) Growth of AB33 derivates in the yeast (left) and hyphal form (right; 8 h.p.i.; size bar, 10 µm). Growth direction is marked by arrows. (E) Percentage of hyphae (8 h.p.i.): unipolarity, bipolarity, and septum formation was quantified (error bars, s.e.m.; n = 3 independent experiments, >100 hyphae were counted per experiment; note that septum formation is given relative to the values of unipolar or bipolar hyphae set to 100%). (F) Relative chitinase activity mainly detecting endochitinase Cts1 (Koepke et al., 2011) in the yeast (left) or hyphal form (right; error bars, s.e.m.; n = 3 independent experiments).
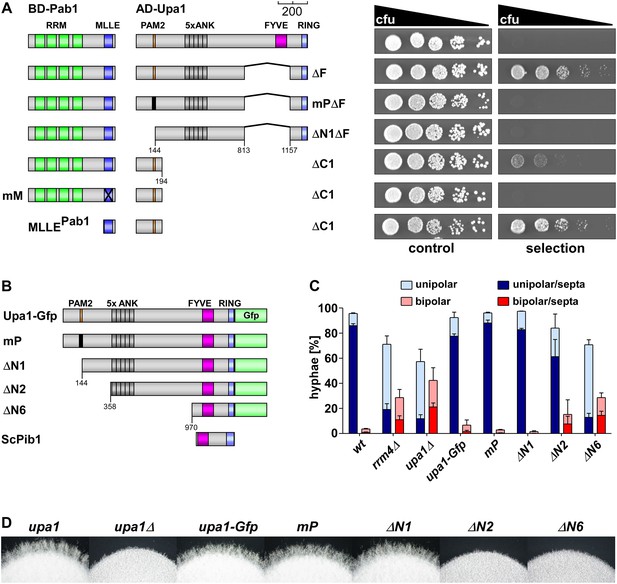
The PAM2 motif of Upa1 interacts specifically with the MLLE domain of Pab1.
(A) Two-hybrid analysis with schematic representation of variants tested (left) and growth plates (right). Yeast cultures were serially diluted 1:5 (decreasing colony forming units, cfu) and spotted on respective growth plates assaying for reporter gene expression (see ‘Materials and methods’). (B) Schematic representation of N-terminal truncated Upa1 variants fused at C-terminus with Gfp, drawn to scale (see Figure 1A; mP, mutation in the PAM2 motif indicated as black bar). (C) Percentage of hyphae (8 h.p.i.): unipolarity, bipolarity, and septum formation was quantified (error bars, s.e.m.; n = 3 independent experiments, >100 hyphae were counted for each strain per experiment; note that septum formation is given relative to the values of unipolar or bipolar hyphae set to 100%). (D) Edge of colonies growing on charcoal-containing medium under hyphae-inducing conditions (48 hr p.i).
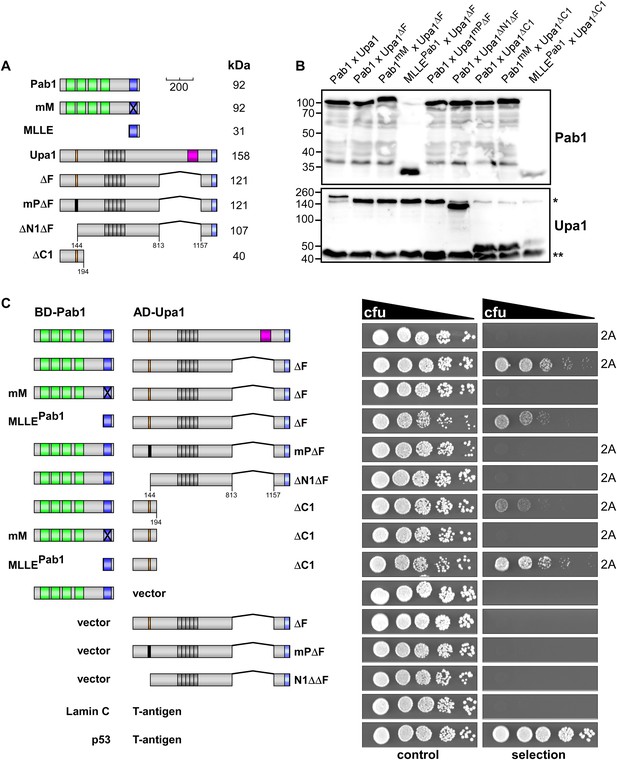
Interaction of Upa1 PAM2/Pab1 MLLE in vivo.
(A) Schematic representation of the constructs used (for details see legend in Figure 2A). (B) Western blot analysis of yeast extracts expressing a Pab1 and Upa1 variants (given above the lanes) carrying a Myc tag and a HA epitope tag, respectively; * and ** mark cross reacting proteins. (C) Two-hybrid analysis as described in Figure 2 (2A marks results already shown in Figure 2A). Positive and negative controls (interaction of p53 with T-Antigen and Lamin C with T-Antigen, respectively) were recommended by the provider of the Matchmaker 3 system (Clontech). Vector indicates the use of a yeast two-hybrid plasmid without insert.
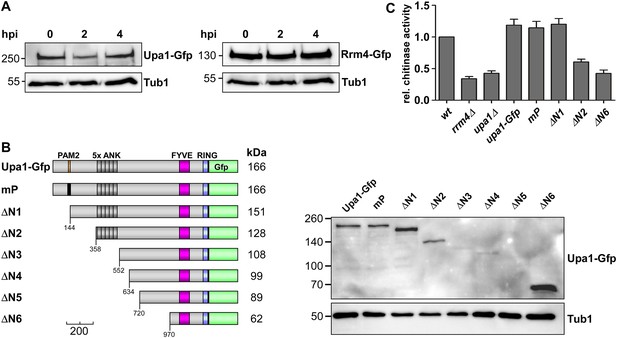
The PAM2 motif is dispensable for Cts1 secretion.
(A) Western blot analysis of strains expressing Upa1-Gfp (left) or Rrm4-Gfp (right). Cells were harvested at 0, 2 and 4 h.p.i.; detection of α-tubulin Tub1 served as control for equal protein amounts. (B) Western blot analysis of strains expressing various N-terminal truncations of Upa1 (depicted schematically on the left, see Figure 1). α-Gfp antibodies were used for detection of Upa1 and detection of Tub1 served as control for equal protein amounts. Note that the expression of Upa1ΔN3-Gfp to Upa1ΔN5-Gfp was strongly reduced. Therefore, these variants were not analysed further. (C) Relative chitinase activity mainly detecting endochitinase Cts1 in the hyphal form (error bars, s.e.m.; n = 3 independent experiments; Koepke et al., 2011).
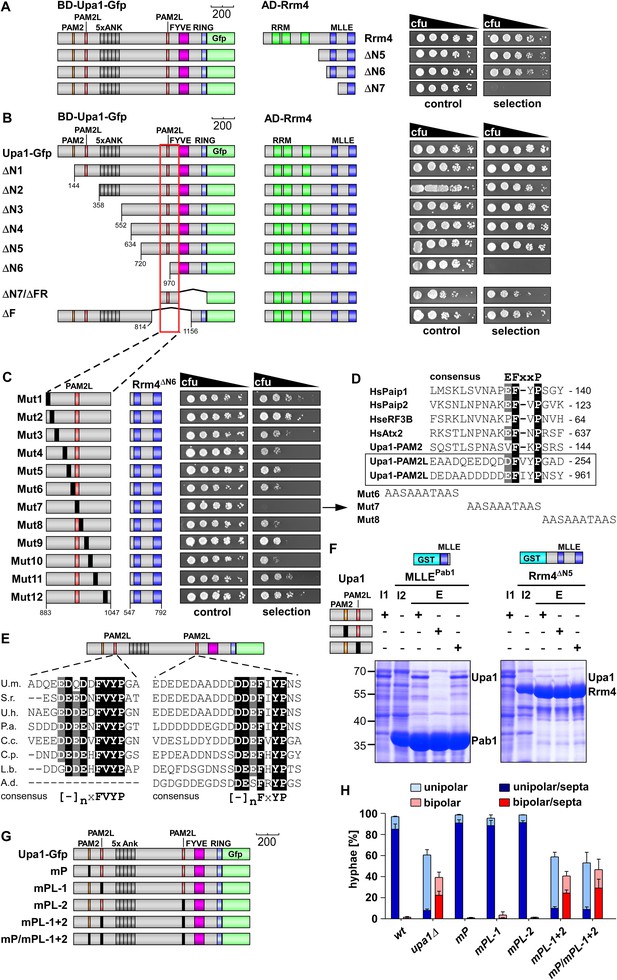
Upa1 contains two PAM2L motives for interaction with Rrm4.
(A) Two-hybrid analysis with schematic representation of variants tested (left) and growth plates (right). Yeast cultures were serially diluted 1:5 (decreasing cfu) and spotted on respective growth plates assaying for reporter gene expression. (B) Two-hybrid analysis as in (A). Red rectangle indicates minimal region in Upa1 interacting with Rrm4. (C) Two-hybrid analysis as in (A). Upa1 region identified in (B) was analysed by linker scanning mutagenesis (mutations indicated as black bar, Mut1-12). (D) Comparison of PAM2 and PAM2L sequences as in Figure 1B. Note, the second PAM2L motif was only mutated in Mut7 (E) PAM2L sequences of Upa1 compared to related sequences from basidiomycetes (U.m., Ustilago maydis UMAG_12183 / XP_758247.1; S.r., Sporisorium reilianum, accession number sr13323 / CBQ72642.1; U.h., Ustilago hordei accession number UHOR_03,485 / CCF52210.1; P.a. Pseudozyma antarctica GAK65366.1; C.c. Coprinopsis cinerea CC1G_00,427 / XP_001837291.2; C.p. Coniophora putanea XP_007767511.1; L.b. Laccaria bicolor XP_001876756.1; A.d. Auricularia delicate XP_007337909.1). (F) GST co-purification experiments with components expressed in E. coli N-terminal His6-tagged versions of Upa1, Upa1mP, and Upa1mPL (amino acids 1–363) were expressed to the same level (first input lane, I1; see Figure 3—figure supplement 5B). MLLE domains of Pab1 or Rrm4 (MLLEPab1 or Rrm4ΔN5, respectively) were expressed as GST fusion proteins (second input lane, I2). After GST affinity chromatography proteins were eluted (lanes marked with "E"). Interaction studies were performed with whole protein extracts from E. coli to demonstrate specific binding. (G) Schematic representation of Upa1 variants carrying mutations (black boxes) in the PAM2 and PAM2L regions. (H) Percentage of hyphae (8 h.p.i.): unipolarity, bipolarity, and septum formation was quantified (error bars, s.e.m.; n = 3 independent experiments, >100 hyphae were counted per experiment; note that septum formation is given relative to the values of unipolar or bipolar hyphae set to 100%).
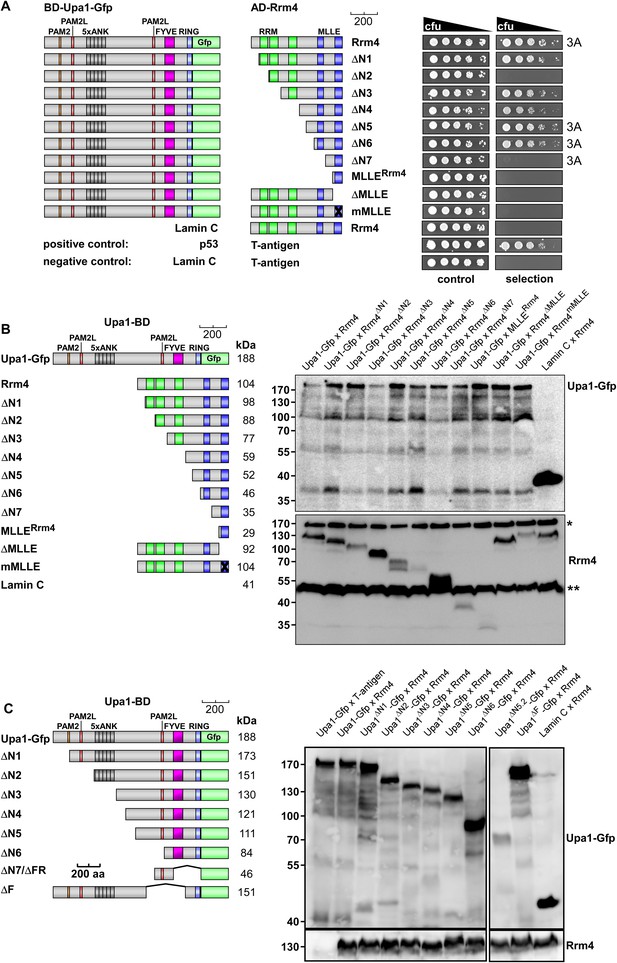
Upa1 interacts with Rrm4 in vivo.
Control experiments for the detailed two-hybrid analysis given in Figure 3. (A) Two-hybrid analysis mapping the domain in Rrm4 that interacts with Upa1 was carried out as described in Figure 3. Positive and negative controls (interaction of p53 with T-Antigen and Lamin C with T-Antigen, respectively) were recommended by the provider of the Matchmaker 3 system (Clontech). Note that Rrm4ΔN2 failed to interact with Upa1-Gfp for unknown reasons. (B) Western blot analysis of yeast extracts expressing Upa1-Gfp and Rrm4 variants (given above the lanes) carrying a Myc tag and a HA epitope tag, respectively. * and ** mark cross reacting proteins. Schematic representation of variants is given on the left. (C) Western blot analysis of yeast extracts expressing Upa1-Gfp variants (schematically shown on the left) and Rrm4 (given above the lanes). Upa1-Gfp was detected with αc-Myc-antibody and Rrm4 with αHA-antibody.
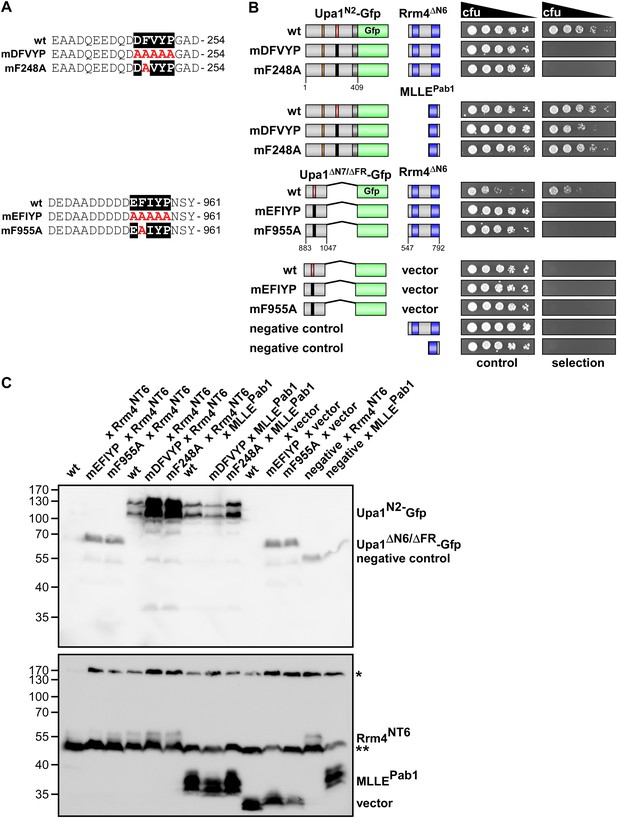
The evolutionarily conserved core of both PAM2L motifs is essential for interaction with Rrm4.
(A) Sequences of the PAM2L motifs from Upa1 (conserved core in white and alanine mutations are given in red). (B) Two-hybrid analysis with schematic representation of variants tested (left) and growth plates (right). Yeast cultures were diluted 1:5 (decreasing colony forming units, cfu) and spotted on respective growth plates assaying for reporter gene expression. (C) Western blot analysis of yeast extracts expressing Upa1-Gfp and Rrm4 variants (given above the lanes) carrying a Myc tag and a HA epitope tag, respectively. * and ** mark cross reacting proteins. Schematic representation of variants is given in B.

Conserved PAM2L motif in the Upa1 N-terminal region.
Sequence comparison of Upa1 homologues of various basidiomycetes (names and accession numbers are given in Figure 3). Remarkably, the FVYP sequence of PAM2L (red box) is the only conserved region at the N termini.
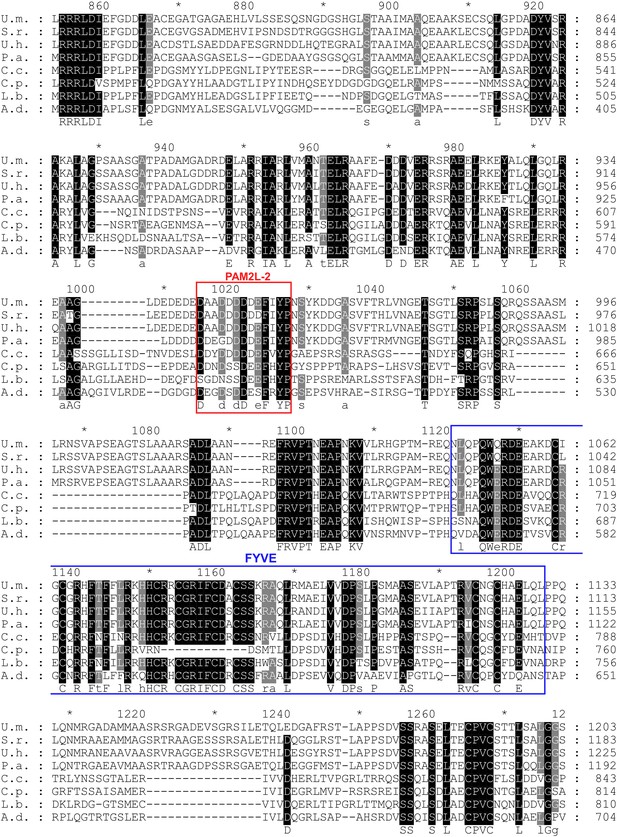

Conserved PAM2L motif in the central region of Upa1.
Sequence comparison of Upa1 homologues from various basidiomycetes (names and accessions numbers given in Figure 3). The FxYP sequence of PAM2L (red box) is conserved in the central part. The computationally predicted FYVE zinc finger domain is given in blue (SMART; Letunic et al., 2009).
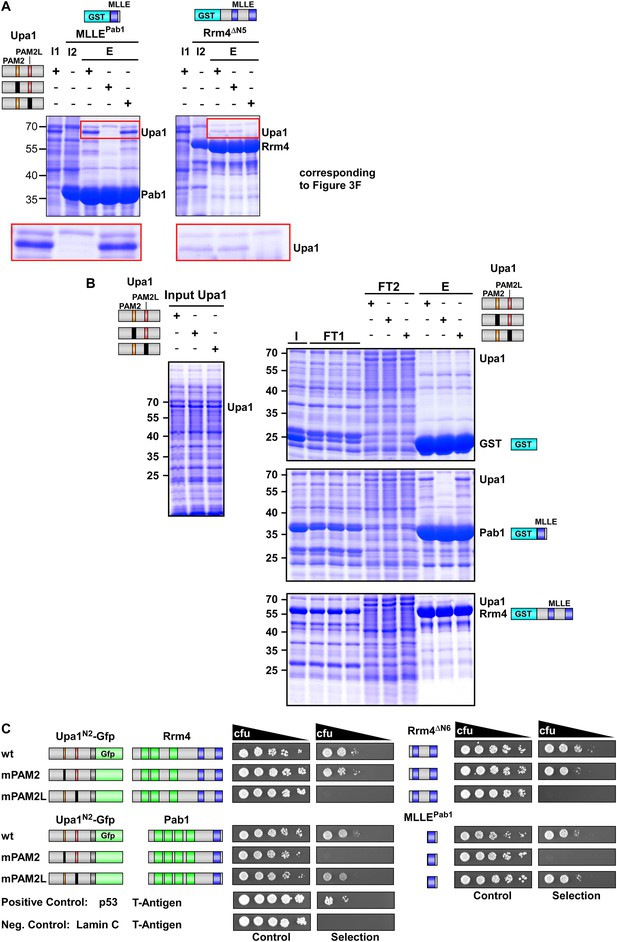
Sequence specific recognition of the PAM2 and PAM2L sequence with the MLLE domains of Pab1 and Rrm4, respectively.
(A) Results of Figure 3F are given with the important region enlarged showing the co-purifying Upa1 variants (red boxes). (B) Coomassie stained SDS-PAGE gels of protein fractions analysed by GST-pulldown assays shown in Figure 3F. On the left side, protein extracts of E. coli expressing all variants of His6-Upa1 are shown. The corresponding band is labelled. Please note, that these proteins—like all variants of Upa1—exhibit a band at a higher kDa size than predicted. On the right side, experimental steps of the pulldown experiment are shown (I = Input; FT1 = flow through 1; FT2 = flow through 2; E = elute of bound proteins). The band height of each protein is indicated on the right. (B) Two-hybrid analysis with schematic representation of variants tested and growth plates as described in Figure 2. Variants of a N-terminal region of Upa1 (Upa1N2; amino acid 1 to 408) carrying no mutation (wt), a mutation in the PAM2 motif (mPAM2) or a mutation in the PAM2L motif (mPAM2L), were tested against full length as well as MLLE-containing versions of Rrm4 and Pab1.
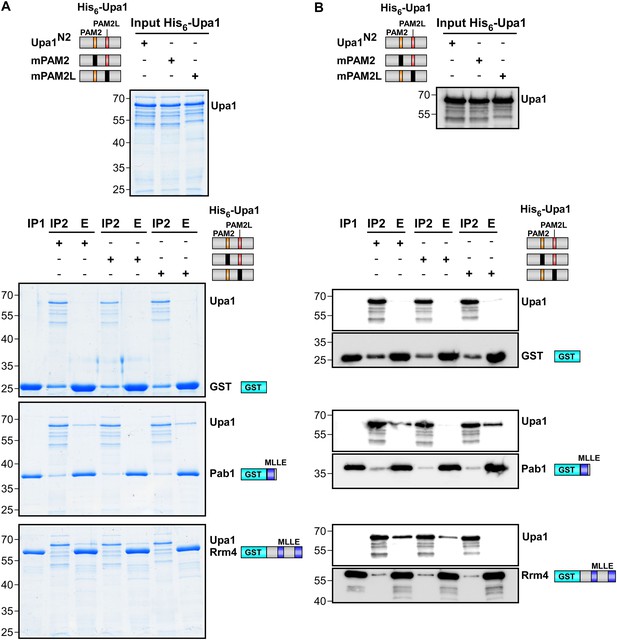
Sequence specific recognition of the PAM2 and PAM2L sequence with the MLLE domains using purified components.
Purified protein fractions (His6-tagged Upa1 versions and GST-tagged MLLE domains of Pab1 and Rrm4) analysed by GST-pulldown assays as shown in Figure 3F. Coomassie-stained gels are shown in (A) and results of Western blot analysis in (B) using αHis and αGST antibodies.
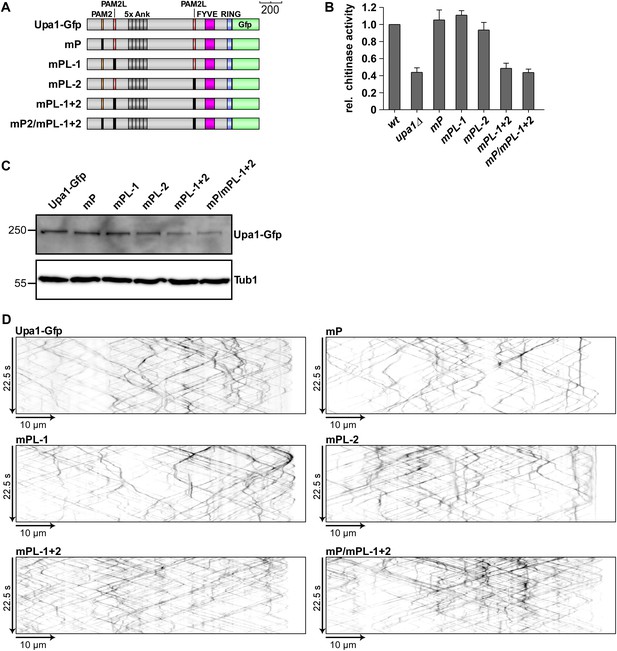
The PAM2L motifs are functionally important for efficient secretion of Cts1.
(A) Schematic representation of Upa1 variants carrying mutations (black boxes) in the PAM2 and PAM2L regions. (B) Relative chitinase activity detecting endochitinase Cts1 (Koepke et al., 2011) in the hyphal form (error bars, s.e.m.; n = 3 independent experiments). (C) Western blot analysis of strains expressing Upa1-Gfp versions (depicted in A; α-Gfp antibodies were used for detection of Upa1 and detection of Tub1 served as control for equal protein amounts). (D) Kymographs of hyphae expressing Upa1-Gfp versions showing bidirectional movement of signals as diagonal lines and indicating that the mutations in the PAM2L motives did not cause drastic differences in endosomal movement of Upa1-Gfp versions.
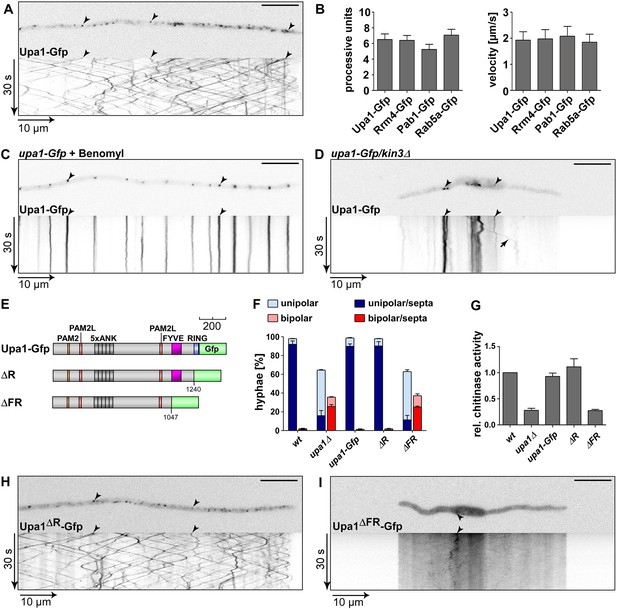
Endosomal targeting of Upa1 is functionally important.
(A) Micrograph (size bar, 10 μm) and corresponding kymograph of hyphae expressing Upa1-Gfp showing bidirectional movement of signals as diagonal lines (arrowheads, Video 1). (B) Bar diagrams depicting amount of processive Upa1-Gfp signals (left, processive units per 10 μm hyphal length to accommodate for size differences between individual hyphae; error bars, s.d.; more than 30 hyphae per strain) and their velocity (right; velocity of tracks showing >5 μm processive movement; error bars, s.d.; 10 to 12 hyphae and more than 900 tracks per strain). (C) Hyphae treated with microtubule inhibitor benomyl. Micrograph (size bar, 10 μm) and corresponding kymograph showing static signals as vertical lines (arrowheads; Video 4). (D) Hyphae expressing Upa1-Gfp and carrying deletion in kin3. Micrograph (size bar, 10 μm) and corresponding kymograph showing static signals as vertical lines (arrowheads; Video 5). Arrow points towards residual movement. (E) Schematic representation of Upa1 fused at C-terminus with Gfp drawn to scale (see Figure 1A). (F) Percentage of hyphae (8 h.p.i.): unipolarity, bipolarity, and septum formation was quantified (error bars, s.e.m.; n = 3 independent experiments, >100 hyphae were counted per experiment; note that septum formation is given relative to the values of unipolar or bipolar hyphae set to 100%). (G) Relative chitinase activity mainly detecting endochitinase Cts1 in the hyphal form (Koepke et al., 2011; error bars, s.e.m., n = 3 independent experiments). (H, I) Micrographs (size bar, 10 μm) and corresponding kymographs of hyphae expressing Upa1ΔR-Gfp (H) or Upa1ΔFR-Gfp (I) (Videos 10,11).
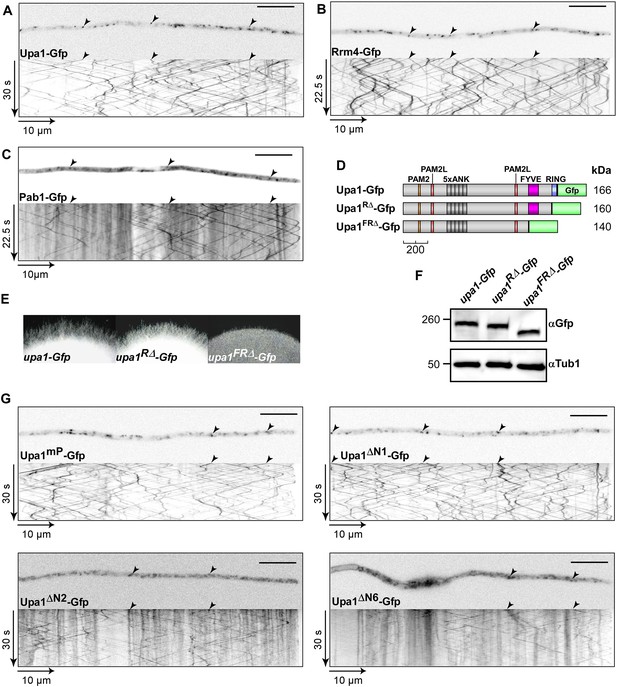
The FYVE domain is crucial for function.
(A–C) Micrographs (size bar, 10 μm) of hyphae expressing Upa1-Gfp (A), Rrm4-Gfp (B), or Pab1-Gfp (C) and corresponding kymographs showing bidirectional movement of signals as diagonal lines (arrowheads; Videos 1–3). Note that the cytoplasmic background signals of Upa1-Gfp resemble Rrm4-Gfp rather than Pab1-Gfp suggesting that Upa1 does only interact with Pab1-Gfp on the cytoplasmic surface of endosomes. (D) Schematic representation of C-terminal truncated Upa1 variants fused at its C-terminus with Gfp drawn to scale (see Figure 1A). (E) Edge of colonies growing on charcoal-containing medium under hyphae-inducing conditions (48 h.p.i.). (F) Western blot analysis of strains expressing various C-terminal truncations of Upa1 (depicted schematically in D). α-Gfp antibodies were used for detection of Upa1-Gfp variants and detection of Tub1 served as control for equal protein amounts. (G) Micrographs (size bar, 10 μm) and corresponding kymographs of hyphae expressing Upa1-Gfp variants shown in Figure 4 (Videos 6–9).
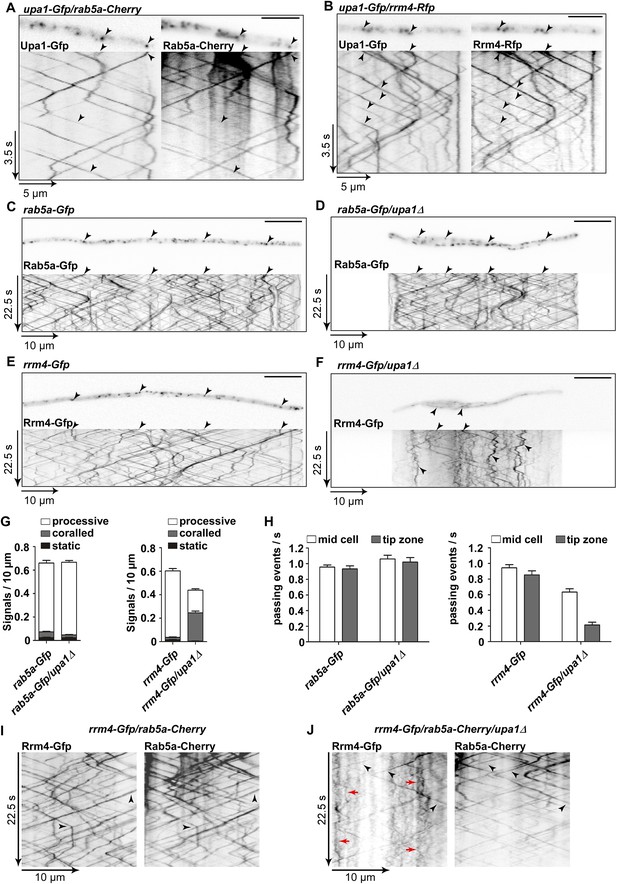
Upa1 is crucial for Rrm4 movement on Rab5a-positive endosomes.
(A) Dynamic co-localisation studies of Upa1-Gfp (left) and Rab5a-Cherry (right) using dual view and msALEX microscopy (see ‘Materials and methods’; arrowheads indicate co-localising signals). Micrographs (size bar, 10 μm) and corresponding kymographs of hyphal tip (Video 12). (B) Dynamic co-localisation studies of Upa1-Gfp (left) and Rrm4-Rfp (right) as in (A) (Video 13). (C–F) Micrographs (size bar, 10 μm) and corresponding kymographs of hyphae expressing Rab5a-Gfp (C), Rab5a-Gfp/upa1Δ (D), Rrm4-Gfp (E), or Rrm4-Gfp/upa1Δ (F) (Videos 14–17). (G) Bar diagrams depicting amount of Rab5a-Gfp signals per 10 μm hyphae in wt and upa1Δ cells (left, error bars, s.d.; >22 hyphae), as well as amount of Rrm4-Gfp signals per 10 μm hyphae in wt and upa1Δ cells (right, error bars, s.d.; >15 hyphae). (H) Number of Rab5a-Gfp (left) and Rrm4-Gfp (right)—signals passing zones in the middle of the hyphae and 10 μm from the apical pole in wt and upa1Δ cells, respectively (passing events of signals/s, error bars, s.d.; more than 15 hyphae). (I) Dynamic co-localisation studies of Rrm4-Gfp (left) and Rab5a-Cherry (right) using dual view and msALEX microscopy (arrowheads indicate co-localising signals). (J) Same analysis as in (I) using a strain carrying a deletion in upa1. Corralled movement of Rrm4 signals not found associated with Rab5a is highlighted by red arrows.
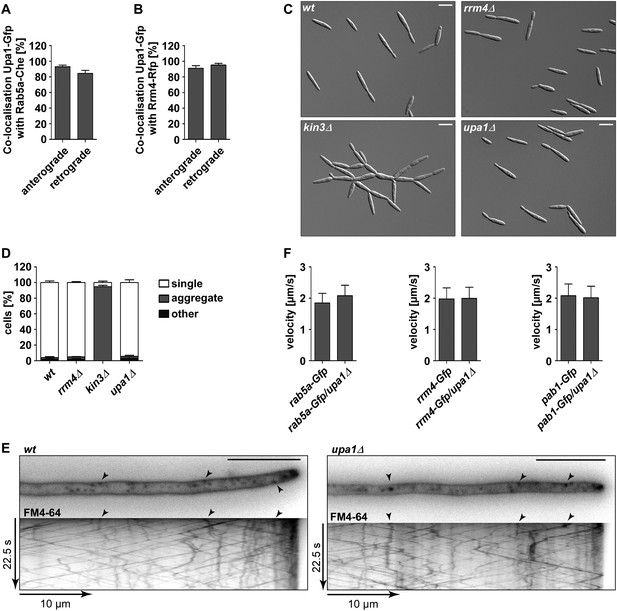
Upa1 co-localises with Rab5a and Rrm4, but loss of Upa1 does not affect long-distance transport of endosomes.
(A) Bar diagram showing the percentage of mobile Upa1-Gfp signals that co-localise with Rab5a-Cherry in the retrograde and anterograde direction (error bars s.e.m.; n = 11 hyphae). (B) Bar diagram showing the percentage of mobile Upa1-Gfp signals that co-localise with Rab5a-Cherry in the retrograde and anterograde direction (error bars s.e.m.; n = 10 hyphae). (C) DIC images of loss of function mutants in kin3, upa1, and rrm4 showing that cytokinesis is not disturbed. (D) Quantification of sporidia appearance in single cell form or cell aggregates (error bars s.e.m.; n = 3 experiments with >100 cells each). (E) FM4-64 uptake assays showing wild-type and upa1Δ mutants. FM4-64 uptake is not disturbed. (F) Bar diagram showing the velocity of processive units (>5 μm/30 s) of Rab5a-Gfp (left), Rrm4-Gfp (middle) and Pab1-Gfp (right) in wt and upa1Δ background strains (error bars, s.d.; 10 hyphae each strain with >1000 tracks [Upa1-Gfp], >550 tracks [Rrm4-Gfp] and >260 tracks [Pab1-Gfp] analyzed).
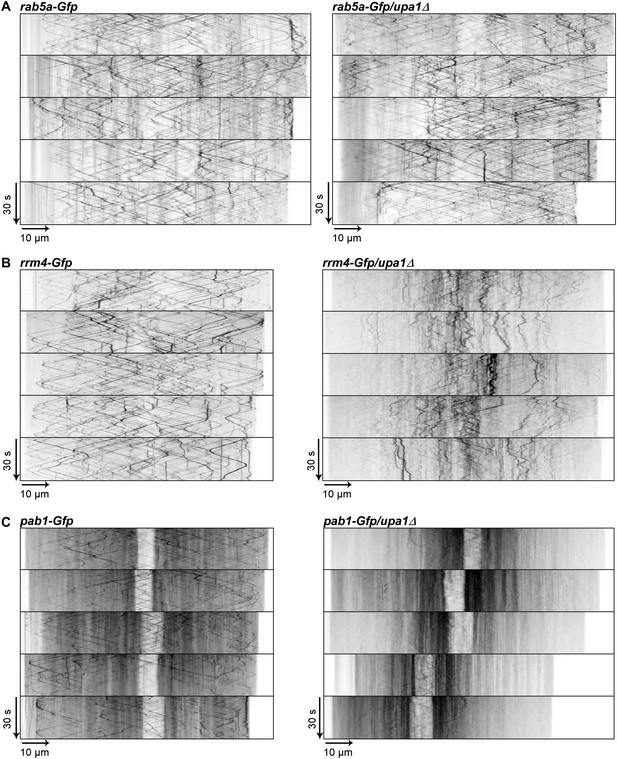

Rrm4 and Pab1 movement is altered in the absence of Upa1.
(A–C) Representative kymographs of the movement of Rab5a-Gfp (A), Rrm4-Gfp (B) and Pab1-Gfp (C) in wt and in upa1Δ background strains. Each movie was taken with an exposure time of 200 ms and consisted of 150 frames.
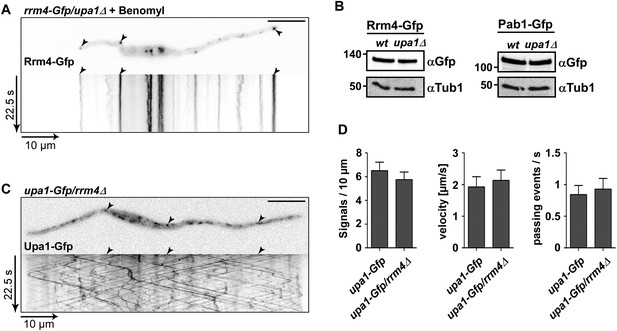
Rrm4 does not influence endosomal localisation of Upa1.
(A) Kymograph of Rrm4-Gfp in upa1Δ background strain treated with benomyl showing that residual movement is microtubule-dependent. (B). Western blots of Rrm4-Gfp and Pab1-Gfp in wt and upa1Δ background strains 6 hr after induction of hyphal growth. α-Gfp antibodies were used for detection of Upa1-Gfp variants and detection of Tub1 served as control for equal protein amounts. (C) Kymograph of Upa1-Gfp in rrm4Δ background showing that Rrm4 is not needed for Upa1 movement. (D) Bar diagrams comparing the amount of Upa1-Gfp signals (left; error bars, s.d.; n = 37/23 cells), the velocity of processive Upa1-Gfp units (middle; >5 μm/30 s; error bars, s.d.; 10 hyphae each strain with >800 tracks) and the number of Upa1-Gfp signals (passing a line 10 μm from the apical pole) in wt and rrm4Δ filaments (right; error bars, s.d.; more than 20 filaments).
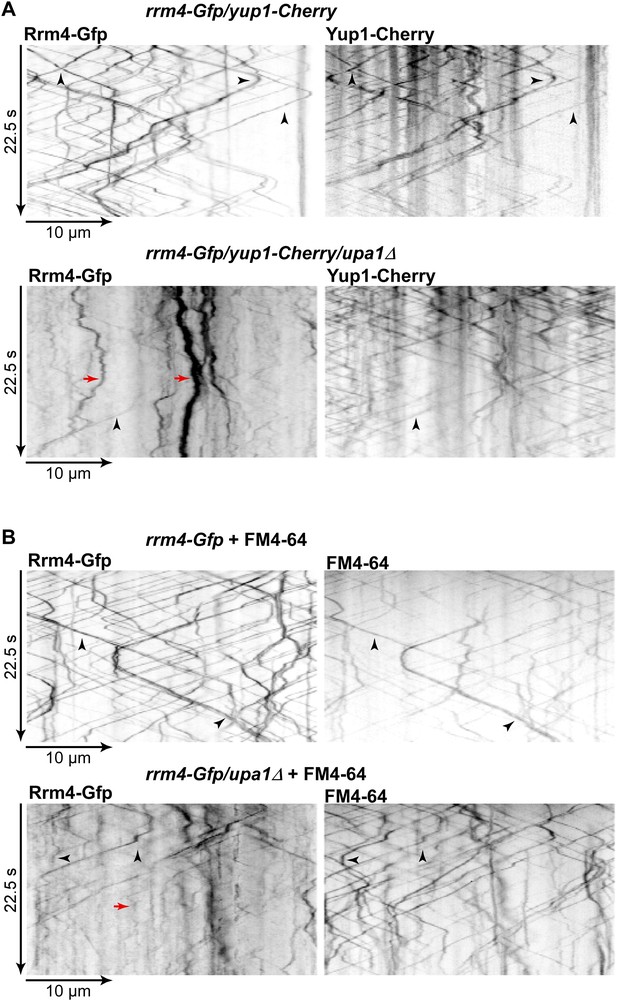
Residual processive movment of Rrm4-Gfp takes place on endosomes.
Dynamic co-localisation studies of Rrm4-Gfp (left) and (A) Yup1-Cherry (right) or (B) FM4-64-stained signals (right) using dual view and msALEX microscopy. Arrowheads indicate co-localising processive signals and corralled movement of Rrm4-Gfp signals is highlighted by red arrows.
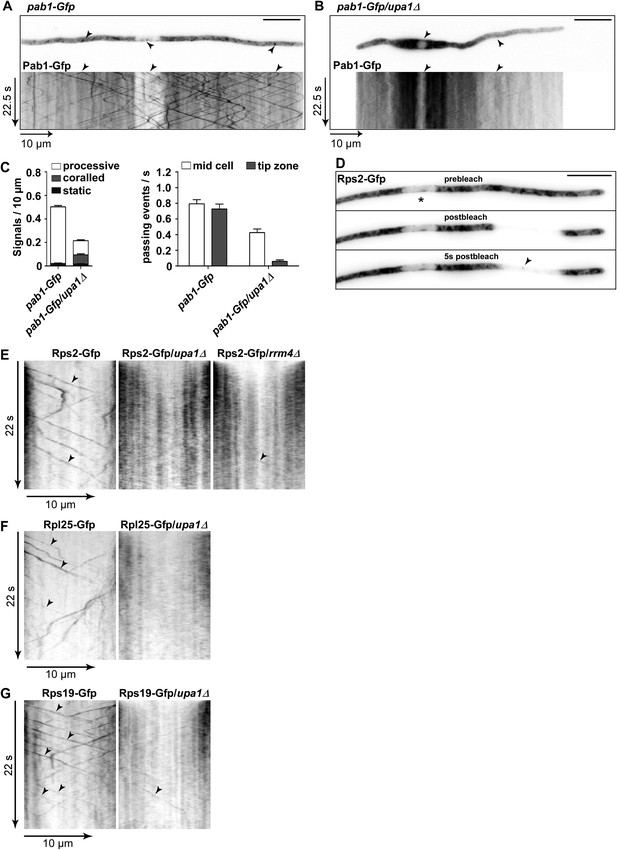
Upa1 functions specifically in mRNP function of endosomes.
(A, B) Micrographs (size bar, 10 μm) and corresponding kymographs (Videos 18, 19) of hyphae expressing Pab1-Gfp (A) or Pab1-Gfp/upa1Δ (B). (C) Bar diagrams depicting amount of Pab1-Gfp signals per 10 μm hyphae in wt and upa1Δ cells (left; error bars, s.d.; more than 13 hyphae) and number of Pab1-Gfp signals passing two zones in the middle of the hyphae and 10 μm from the apical pole in wt and upa1Δ cells (right; passages of signals/s, error bars, s.d.; >13 hyphae). (D) Analysing subcellular localisation of ribosomal protein Rps2-Gfp as an example. Micrographs (size bar, 10 μm) of hyphae expressing Rps2-Gfp (nucleus indicated by asterisk). 20-μm area was bleached by laser irradiation about 10 μm from the hyphal tip. Arrowhead indicates processive signal entering the bleached area. (E, F, G) Kymographs of hyphal areas bleached with laser irradiation, as shown in (D). Arrowheads indicate processive signals.
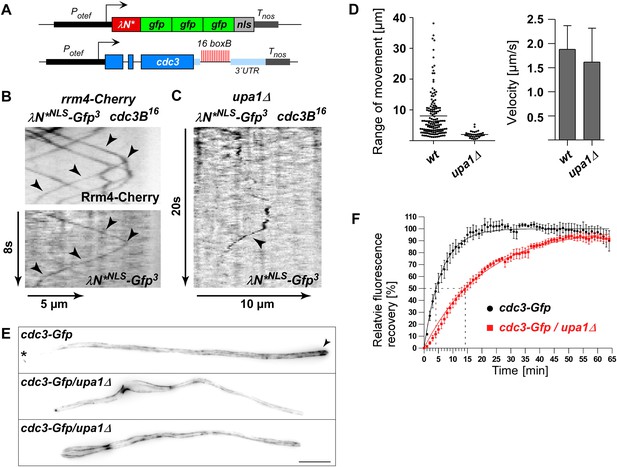
Loss of Upa1 disturbs Rrm4-dependent transport of cdc3 septin mRNA and protein.
(A) Schematic representation of components of the modified λN RNA reporter system (Potef, constitutively active promoter; Tnos, heterologous transcriptional terminator; cdc3B16 carries 16 copies of boxB hairpin in its 3′ UTR; λN*NLS-Gfp3, modified λN peptide fused to triple Gfp; and NLS; Baumann et al., 2014). (B) Dynamic co-localisation of strain expressing Rrm4-Cherry, λN*NLS-Gfp3, and cdc3B16. Kymograph with directed particles (arrowheads). (C) Kymograph of upa1Δ strain expressing λN*NLS-Gfp3 protein and cdc3B16 mRNA. Occasionally directed particles are observed that exhibit altered processive movement. (D) Diagram showing range of movement of λN*NLS-Gfp3-labelled mRNAs (vertical bar = mean, 151 mRNA particles for wt and 33 particles for upa1Δ) on the left and velocity λN*NLS-Gfp3-labelled mRNAs on the right (error bars, SD; 54 and 96 hyphae for wt and upa1Δ, respectively). (E) Micrographs of Cdc3G or Cdc3G/upa1Δ expressing hyphae (maximum projection of z-stacks with 0.27 μm steps; size bar, 10 μm). Arrowhead marks gradient of septin filaments emanating from the hyphal tip. (F) FRAP analysis of Cdc3-Gfp or Cdc3-Gfp/upa1Δ expressing hyphae 7–10 h.p.i. (about 10 μm from the hyphal tip; data were fitted to uniphasic exponential equation, dashed lines indicate half time of recovery; n = 3 independent experiments with 4–6 hyphae per experiment; error bars represent s.e.m.). Fluorescence is normalised to plateau (Baumann et al., 2014).
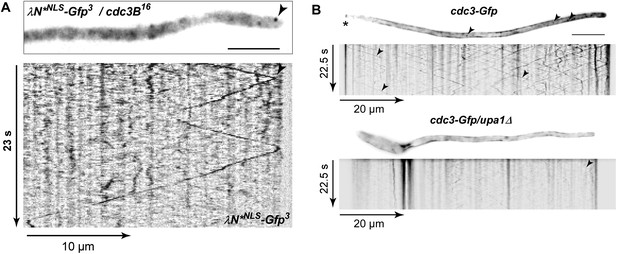
Endosome-dependent movement of cdc3 mRNA and protein.
(A) Hyphal tip of a strain expressing the λN*NLS-Gfp3 protein and cdc3B16 mRNA. Micrograph (size bar, 10 μm) and corresponding kymograph show directed particles (arrowheads; Video 20). (B) Micrographs (size bar, 10 μm; asterisk marks retraction septum) and corresponding kymographs of hyphae expressing Cdc3-Gfp (top) or Cdc3-Gfp/upa1Δ (bottom). Arrowheads indicate Cdc3-Gfp-positive endosomes.
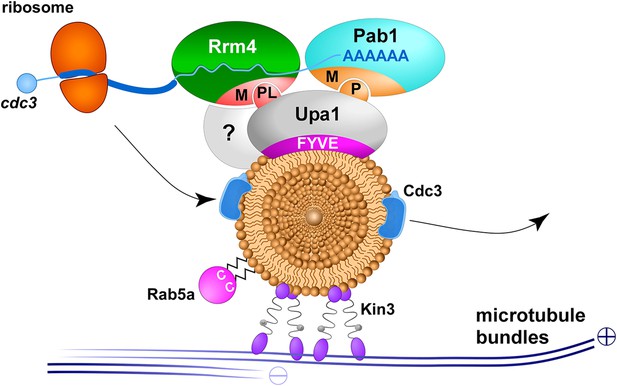
Upa1 functions specifically in endosomal mRNA transport.
Model proposing Upa1 function during endosomal mRNA transport. Microtubules are given in blue, kinesin-3 type motor Kin3 transports endosomes in the plus-end direction. The small GTPase Rab5a (magenta) marks this specific endosomal compartment, described as early endosomes (Higuchi et al., 2014). Upa1 binds endosomes via FYVE domain and the MLLE domain (M) of Pab1 via PAM2 motif (P), as well as the MLLE domain of Rrm4 via PAM2L motif (PL; for simplicity only one motif is shown). Note that the interaction of Upa1 with Pab1 is dispensable, whereas the interaction of Upa1 with Rrm4 is crucial for the endosomal localisation of septin mRNA (blue line with poly[A]-tail), septin protein (blue) and ribosomes (orange). A currently unknown adaptor protein is highlighted with a question mark.
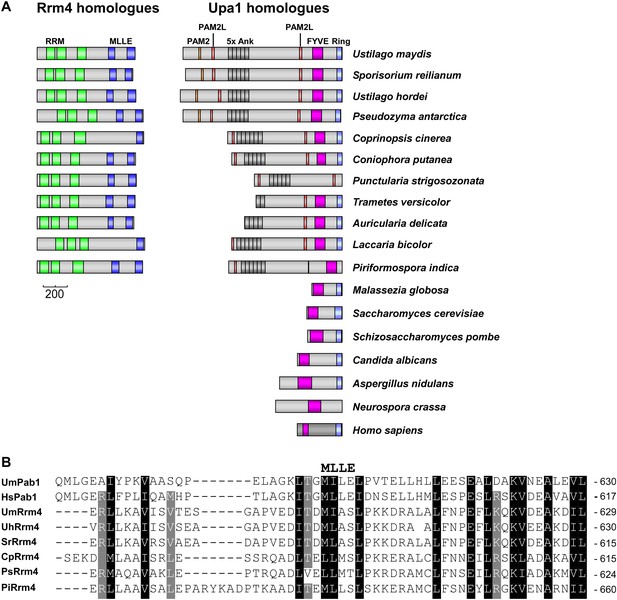
Rrm4 and Upa1 homologues in fungi.
(A) Domain architecture of proteins was predicted by analysing amino acid sequence of the proteins using SMART (http://smart.embl-heidelberg.de/smart/set_mode.cgi?NORMAL=1) and NCBI conserved domain search (http://www.ncbi.nlm.nih.gov/Structure/cdd/wrpsb.cgi). Rrm4-homologues: U. maydis (UMAG_10836); Sporisorium reilianum (sr14312); Ustilago hordei (UHOR_05154); Pseudomonas antarctica (GAK64672.1); Coprinopsis cinerea (XP_001832566.2); Coniophora putanea (XP_007771597.1), Punctularia strigosozonata (XP_007384300.1); Tinea versicolor (XP_008042363.1), Acaulospora delicata (XP_007341926.1); L. bicolor (XP_001881076.1); Piriformospora indica (CCA67340.1); Upa1 homologues: U. maydis (UMAG_10836); S. reilianum (sr13323); U. hordei (UHOR_03485); P. antarctica (GAC72163); C. cinerea (XP_001837291.2); C. putanea (XP_007767511.1), P. strigosozonata (XP_007382070.1); T. versicolor (XP_008035292.1), A. delicata (XP_007337909.1); L. bicolor (XP_001876756.1); P. indica (CCA71703-CCA71704), Malassezia globosa (XP_001732453.1); S. cerevisiae (YDR313C); Schizosaccharomyces pombe (NP_595987.1); Candida albicans (XP_719309.1); Aspergillus nidulans (ANID_11,932.1); Neurospora crassa (CU08360); Homo sapiens (NP_001017368.1). (B) Alignment of the protein sequence of MLLE domains of the Poly(A)-binding proteins from U. maydis (UmPab1; UMAG_03,449) and humans (HsPab1; AAH23520), as well as the N-terminal situated MLLE domains of Rrm4 (UmRrm4; UMAG_10836) and homologues.
Videos
Upa1-Gfp moves bidirectionally in a hypha of AB33upa1-Gfp.
Video corresponds to Figure 4A (size bar = 10 μm, timescale in seconds, 200 ms exposure time, 150 frames, 5 frames/s display rate; QuickTime format, 6275 kB).
Rrm4-Gfp moving bidirectionally in a hypha of AB33rrm4-Gfp.
Video corresponds to Figure 4—supplement figure 1B (size bar = 10 μm, timescale in seconds, 150 ms exposure time, 150 frames, 6 frames/s display rate; QuickTime format, 1536 kB).
Pab1-Gfp moving bidirectionally in a hypha of AB33pab1-Gfp.
Note, that in contrast to Upa1-Gfp and Rrm4-Gfp (Videos 1 and 2) a higher background signal can be attributed to non-transported poly-adenylated mRNAs. Video corresponds to Figure 4—supplement figure 1C (size bar = 10 μm, timescale in seconds, 150 ms exposure time, 150 frames, 6 frames/s display rate; QuickTime format, 473 kB).
Upa1-Gfp movement is dependent on the microtubule cytoskeleton.
Treating AB33upa1-Gfp for one hour with 50 μM microtubule destabilising drug benomyl, inhibits the bidirectional movement of Upa1-Gfp. Video corresponds to Figure 4C (size bar = 10 μm, timescale in seconds, 200 ms exposure time, 150 frames, 5 frames/s display rate; QuickTime format, 2862 kB).
Loss of the plus-end directed kinesin-3 Kin3 disturbs Upa1-Gfp movement in AB33upa1-Gfp/kin3Δ.
Immobile Upa1-Gfp accumulations can be seen in the middle of the cell. Residual movement might be due to dynein activity. Video corresponds to Figure 4D (size bar = 10 μm, timescale in seconds, 200 ms exposure time, 150 frames, 5 frames/s display rate; QuickTime format, 2382 kB).
Upa1mP-Gfp moving bidirectionally in a hypha of AB33upa1mP-Gfp.
Mutating the PAM2 motif by amino acid exchanges does not inhibit movement of Upa1mP-Gfp. Video corresponds to Figure 4—supplement figure 1G top left (size bar = 10 μm, timescale in seconds, 200 ms exposure time, 150 frames, 5 frames/s display rate; QuickTime format, 2901 kB).
Upa1ΔN1-Gfp moving bidirectionally in a hypha of AB33upa1ΔN1-Gfp.
Loss of the first 143 amino acids including the PAM2 motif does not inhibit movement of Upa1ΔN1-Gfp. Video corresponds to Figure 4—supplement figure 1G top right (size bar = 10 μm, timescale in seconds, 200 ms exposure time, 150 frames, 5 frames/s display rate; QuickTime format, 2695 kB).
Upa1ΔN2-Gfp moving bidirectionally in a hypha of AB33upa1ΔN2-Gfp.
Loss of the first 337 amino acids including the PAM2 motif does not inhibit movement of Upa1ΔN2-Gfp although an increase in stationary background signals can be seen. Video corresponds to Figure 4—supplement figure 1G bottom left (size bar = 10 μm, timescale in seconds, 200 ms exposure time, 150 frames, 5 frames/s display rate; QuickTime format, 5614 kB).
Upa1ΔN6-Gfp moving bidirectionally in a hypha of AB33upa1ΔN6-Gfp.
The C-terminal part of Upa1 containing the FYVE and RING domains is sufficient for movement although an increase in stationary background signals can be seen. Video corresponds to Figure 4—supplement figure 1G bottom right (size bar = 10 μm, timescale in seconds, 200 ms exposure time, 150 frames, 5 frames/s display rate; QuickTime format, 4388 kB).
Upa1ΔR-Gfp moving bidirectionally in a hypha of AB33upa1ΔR-Gfp.
The C-terminal part comprising aa 1241–1287 including the RING domain is not necessary for shuttling of Upa1. Video corresponds to Figure 4H (size bar = 10 μm, timescale in seconds, 200 ms exposure time, 150 frames, 5 frames/s display rate; QuickTime format, 4069 kB).
Upa1ΔFR-Gfp shows an even distribution in a bipolar growing hypha of AB33upa1ΔFR-Gfp.
Thus, the C-terminal part comprising of aa 1048–1287 including the FYVE and RING domains is necessary for localising Upa1 to endosomes. Video corresponds to Figure 4I (size bar = 10 μm, timescale in seconds, 200 ms exposure time, 150 frames, 5 frames/s display rate; QuickTime format, 5201 kB).
Upa1-Gfp and Rab5a-Cherry (upper and lower part, respectively) co-localise in shuttling units in hyphae of AB33upa1-Gfp/rab5a-Cherry.
Videos were recorded simultaneously using dual-colour detection and correspond to Figure 5A (size bar = 5 μm, timescale in seconds, 70 ms alternating exposure time, 200 frames, 15 frames/s display rate; QuickTime format, 561 kB)
Upa1-Gfp and Rrm4-Rfp (upper and lower part, respectively) co-localise in shuttling units in hyphae of AB33upa1-Gfp/rrm4-Rfp.
Videos were recorded simultaneously using dual-colour detection and correspond to Figure 5B (size bar = 10 μm, timescale in seconds, 70 ms alternating exposure time, 100 frames, 15 frames/s display rate; QuickTime format, 132 kB).
Rab5a-Gfp moving bidirectionally in a hypha of AB33rab5a-Gfp.
Video corresponds to Figure 5C (size bar = 10 μm, timescale in seconds, 150 ms exposure time, 150 frames, 6 frames/s display rate; QuickTime format, 248 kB).
Rab5a-Gfp moving bidirectionally in a hypha of AB33rab5a-Gfp/upa1Δ.
Loss of Upa1 results in bipolar growing cell, but does not affect Rab5a-Gfp shuttling. Video corresponds to Figure 5D (size bar = 10 μm, timescale in seconds, 150 ms exposure time, 150 frames, 6 frames/s display rate; QuickTime format, 219 kB).
Rrm4-Gfp moving bidirectionally in a hypha of AB33rrm4-Gfp.
Video corresponds to Figure 5E (size bar = 10 μm, timescale in seconds, 150 ms exposure time, 150 frames, 6 frames/s display rate; QuickTime format, 2324 kB).
Rrm4-Gfp moving bidirectionally in a hypha of AB33rrm4-Gfp/upa1Δ.
Loss of Upa1 disturbs shuttling of Rrm4-Gfp, as seen by increased corraled movement. Video corresponds to Figure 5F (size bar = 10 μm, timescale in seconds, 150 ms exposure time, 150 frames, 6 frames/s display rate; QuickTime format, 741 kB).
Pab1-Gfp moving bidirectionally in a hypha of AB33pab1-Gfp.
Video corresponds to Figure 6A (size bar = 10 μm, timescale in seconds, 150 ms exposure time, 150 frames, 6 frames/s display rate; QuickTime format, 729 kB).
Pab1-Gfp moving bidirectionally in a hypha of AB33pab1-Gfp/upa1Δ.
Loss of Upa1 disturbs shuttling of Pab1-Gfp, as seen by the drastically decreased number of processive Pab1-Gfp signals. Video corresponds to Figure 6B (size bar = 10 μm, timescale in seconds, 150 ms exposure time, 150 frames, 6 frames/s display rate; QuickTime format, 353 kB).
Video of cdc3B16 particle visualised by λN*NLS-Gfp3.
Arrowhead at the beginning of the video marks the starting point of the moving particle (size bar = 5 μm, timescale in seconds, 150 ms exposure time, 150 frames, 6 frames/s display rate, QuickTime format, 5495 kB).
Additional files
-
Supplementary file 1
Potential PAM2-containing proteins from U. maydis.
- https://doi.org/10.7554/eLife.06041.047
-
Supplementary file 2
Description of U. maydis strains used in this study.
- https://doi.org/10.7554/eLife.06041.048
-
Supplementary file 3
Generation of U. maydis strains used in this study.
- https://doi.org/10.7554/eLife.06041.049
-
Supplementary file 4
Description of plasmids used for U. maydis strain generation.
- https://doi.org/10.7554/eLife.06041.050
-
Supplementary file 5
Description of plasmids used for yeast two-hybrid analyses.
- https://doi.org/10.7554/eLife.06041.051
-
Supplementary file 6
Description of plasmids used for pulldown experiments.
- https://doi.org/10.7554/eLife.06041.052
-
Supplementary file 7
DNA oligonucleotides used in this study.
- https://doi.org/10.7554/eLife.06041.053





