SMC condensin entraps chromosomal DNA by an ATP hydrolysis dependent loading mechanism in Bacillus subtilis
Figures
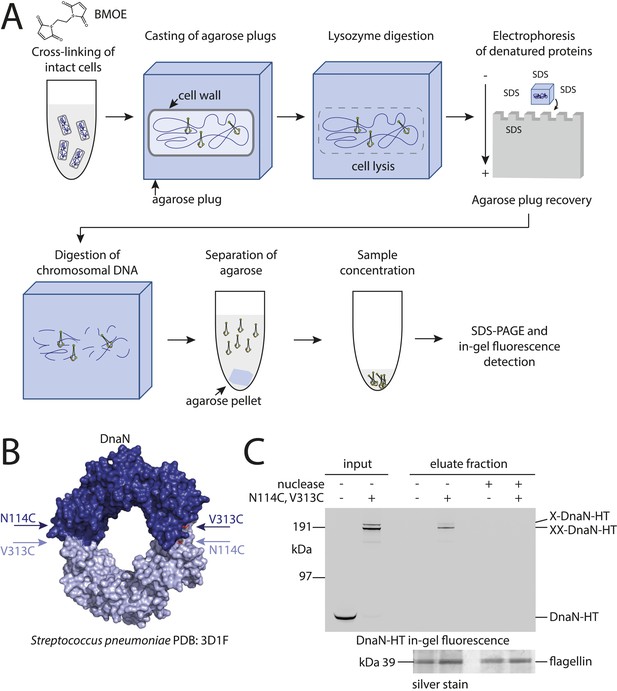
Development of the chromosome entrapment assay using DnaN.
(A) Scheme for the chromosome entrapment assay. Cells are incubated with the cysteine cross-linker BMOE, lysed in agarose plugs and subjected to an electric field in the presence of SDS buffer. Proteins stably bound to chromosomal DNA are re-isolated from nuclease treated agarose plugs, concentrated and analyzed by SDS-PAGE. (B) Crystal structure of S. pneumoniae DnaN (PDB: 3D1F) in surface representation. The monomers of DnaN are shown in dark and light blue colours, respectively. The positions of an engineered pair of cysteine residues (N114C and V313C) at the monomer–monomer interface of B. subtilis DnaN are indicated by arrows. (C) Chromosome entrapment by DnaN. Cells of strains BSG1449 (dnaN-HT) and BSG1459 (dnaN(N114C, V313C)-HT) were cross-linked with BMOE and subjected to the chromosome entrapment assay. Input and eluate fractions were analysed by in-gel detection of fluorescently labeled HT fused to DnaN (top panel). Eluate fractions of samples treated with or without nuclease during cell lysis are indicated as nuclease ‘+’ or ‘−’, respectively. Eluate fractions were further analyzed by silver staining revealing that another protein was consistently co-isolated during the chromosome entrapment assay (bottom panel). This protein—identified as flagellin by mass spectrometry—was retained independently of the integrity of the chromosome. The following figure supplement is available: Figure 1—figure supplement 1: DNA entrapment by DnaN.
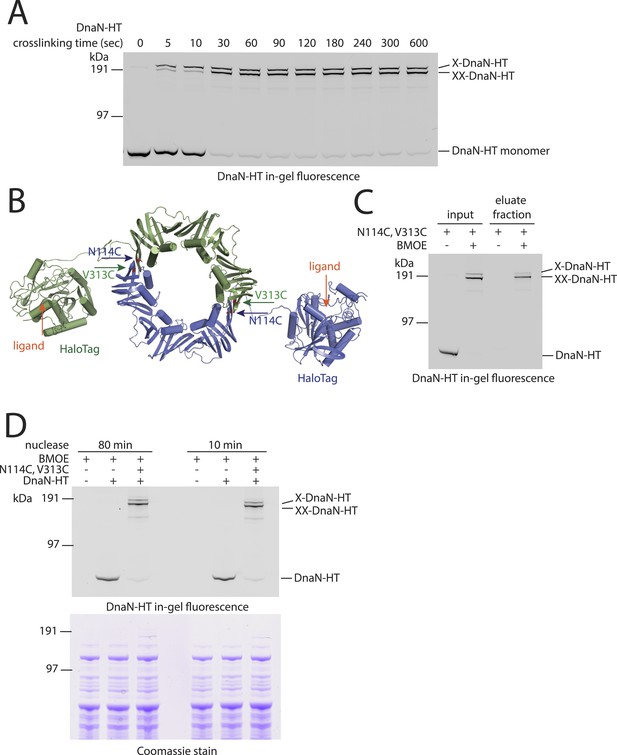
DNA entrapment by DnaN.
(A) Cross-linking time course of DnaN-HT. Cells of strain BSG1459 were incubated with the cross-linker BMOE and quenched at the indicated times. Extracts were analysed by in-gel fluorescence detection of DnaN-HT. At later time points the double cross-linked DnaN species (‘XX-DnaN-HT’) is enriched over non-cross-linked and single cross-linked DnaN-HT. (B) Cartoon represenation of the DnaN-HT construct based on the crystal structure of S. pneumoniae DnaN (PDB: 3D1F) and Rhodococcus Haloalkane dehalogenase (PDB: 1BN6). The structure of the DnaN dimer (monomers in green and blue colours, respectively) was juxtaposed to the structure of the Haloalkane dehalogenase linked by a flexible peptide. The binding pockets for HT ligand binding are indicated by orange arrows. The positions of the engineered pair of cysteine residues (N114C and V313C) at the DnaN–DnaN interface are indicated by blue and green arrows. (C) Chromosome entrapment assay of strain BSG1459 (dnaN(N114C, V313C)-HT) in presence or absence of the cross-linker BMOE. (D) Effect of benzonase (‘nuclease’) on protein extracts of strains BSG1001, BSG1449 and BSG1459. Cell extracts were analysed by in-gel fluorescence detection (upper panel) of DnaN-HT after long-time (‘80 min’) or short-time (‘10 min’) incubation with benzonase. Lower panel shows Coomassie staining of the same SDS-PAGE gel to control for equal protein extraction efficiency.
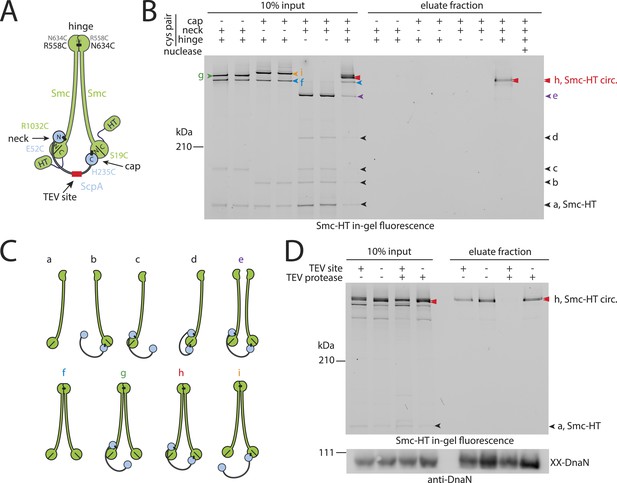
Prokaryotic condensin entraps the chromosome.
(A) Scheme for the cross-linking of Smc-HaloTag (‘HT’) and ScpA into a covalent Smc–ScpA–Smc ring. (B) Chromosome entrapment of covalent Smc2–ScpA rings. Cells of strains BSG1782, BSG1809-1813 and BSG1831 were cross-linked and subjected to the chromosome entrapment assay. Cross-linked Smc-HT species were visualized by in-gel fluorescence detection. The presence or absence of cysteine pairs at each of the three ring interfaces are indicated by ‘+’ and ‘−’, respectively. An aliquot of cells of strains BSG1782 was incubated with benzonase during cell lysis (nuclease ‘+’). The positions of uncross-linked Smc-HT and fully cross-linked, circular Smc–ScpA–Smc species are indicated by ‘Smc-HT’ and ‘Smc-HT circ.’; all species are labelled by colour-coded arrowheads (see panel C for legend). Circular species (‘h’) are labeled by a double pointed arrowhead. (C) Schematic depiction of the structure of cross-linked Smc–ScpA species (‘a’–‘i’). (D) TEV cleavage of ScpA prevents entrapment of Smc–ScpAB in agarose plugs. In-gel fluorescence detection of Smc-HT derived from strains BSG1807 and BSG1832. The presence or absence of TEV sites in ScpA and of TEV protease during cell lysis is indicated by ‘+’ and ‘−’, respectively. Cleavage of ScpA(TEVs) by TEV protease creates new species of cross-linked Smc-HT (see ‘input’ samples) and prevents entrapment of Smc-HT in agarose plugs (see ‘eluate fraction’) (top panel). ‘XX-DnaN’ serves as internal assay control visualized by immunoblotting of cross-linked species of DnaN protein (bottom panel). The following figure supplement is available: Figure 2—figure supplement 1: DNA entrapment by wild-type Smc–ScpAB (I) and Figure 2—figure supplement 2: DNA entrapment by wild-type Smc–ScpAB (II).
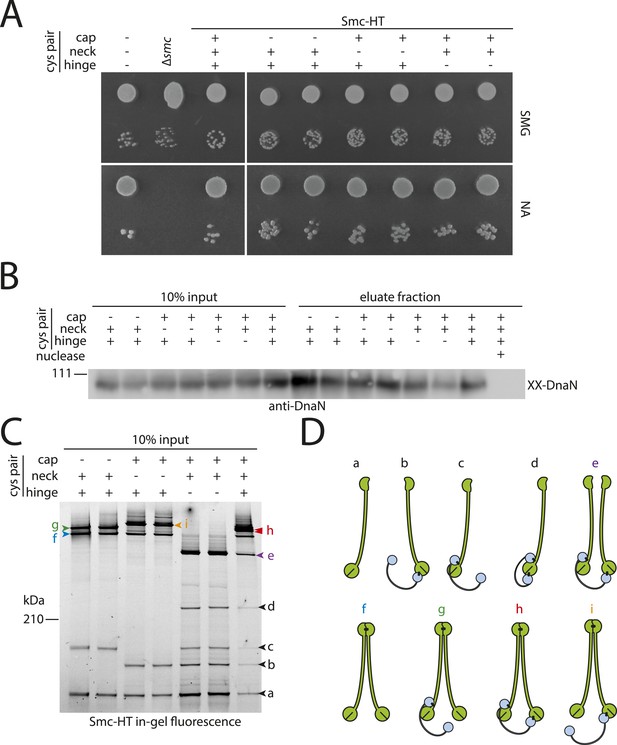
DNA entrapment by wild-type Smc–ScpAB (I).
(A) Colony formation assays of strains BSG1002, BSG1007, BSG1782, 1809–1813 and BSG1831 were performed on minimal medium agar (SMG) or nutrient agar (NA). Cultures were diluted 9^2-fold (top row) and 9^5-fold (bottom row) and grown at 37°C for 24 hr (SMG) and 14 hr (NA), respectively. The presence and absence of cysteine pairs for cross-linking of the hinge, cap and neck interface are indicated by ‘+’ and ‘−’, respectively. (B) Immunoblot of SDS-PAGE gels shown in Figure 2B using anti-DnaN antibody. All strains carry cysteine mutations for DnaN cross-linking. Doubly cross-linked DnaN (‘XX-DnaN’) serves as internal assay control as visualized by immunoblotting. (C) Identification of cross-linked species of Smc-HT and ScpA. The gel image is identical to Figure 2A with higher contrast settings to visualize low-intensity species. All cross-linked species are labelled (with a colour-coded letter) (‘a’–‘i’). Circular species (‘h’) are labeled by a double pointed arrowhead. (D) Same as Figure 2, panel C. Schematic depiction of the structure of cross-linked Smc–ScpA species (‘a’–‘i’).
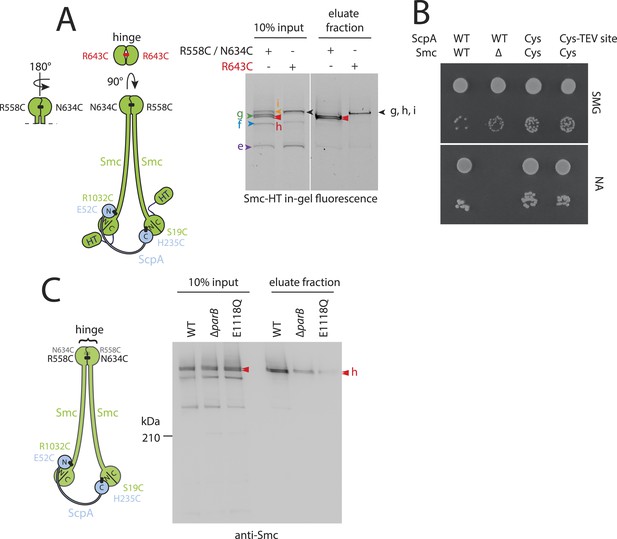
DNA entrapment by wild-type Smc–ScpAB (II).
(A) Cross-linking of the Smc hinge interface with different cysteine residues. Cross-linking of the Smc hinge using cysteine residue (‘R643C’) instead of the ‘R558C’ and ‘N643C’ cysteine pair results in different migration pattern of cross-linked Smc-ScpA species (see input fractions). In this case species ‘g’ and ‘i’ have the same migration behavior as the circular form ‘h’. Only a single species of Smc-ScpA is retained in the agarose plug after the entrapment assay (see eluate fractions). (B) Colony formation assay of strains BSG1002, BSG1007, BSG1807 and BSG1832 were performed as described before. The smc and scpA loci lacking mutations (‘WT’), harbouring cysteine mutations for cross-linking alone (‘Cys’) or in combination with a TEV cleavage site (‘Cys-TEV site’) are denoted. (C) Chromosome entrapment of Smc–ScpAB lacking fusion tags (see scheme on the left). The right panel shows immunoblotting of input and eluate fractions using an anti-Smc antibody. All strains harbour cysteine residues for crosslinking of Smc-ScpA in an otherwise wild-type background (‘WT’) in BSG680, combined with a deletion of the parB gene (‘ΔparB’) in BSG1991 or with the E1118Q mutation at the smc locus in BSG1995. The position of fully cross-linked, circular Smc2-ScpA species is indicated by double pointed arrowhead labelled ‘h’ (see Figure 2—figure supplement 1C), respectively.
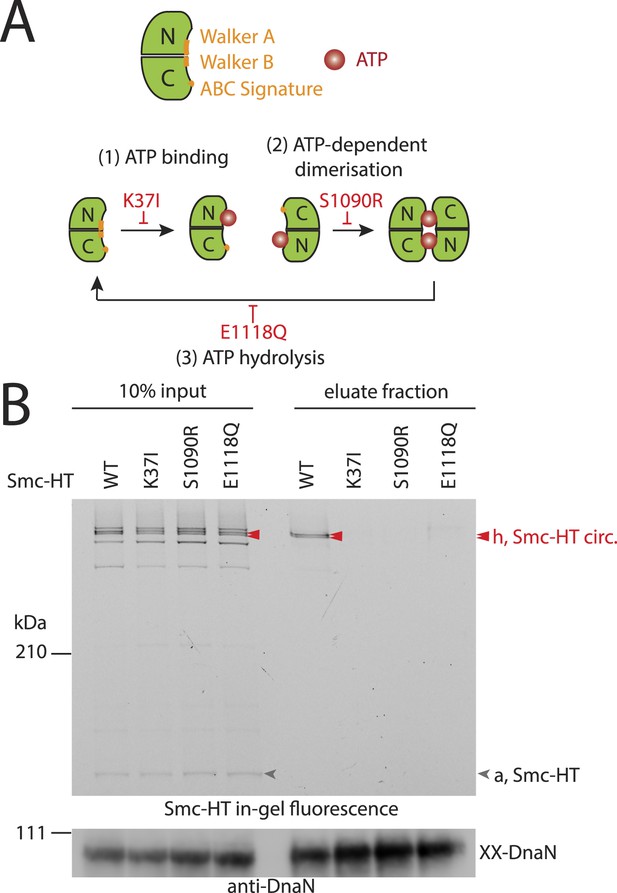
The Smc ATPase is required for loading of DNA into Smc–ScpAB.
(A) A scheme for the ATP hydrolysis cycle of Smc. Schematic positions for Walker A, Walker B and ABC-signature motifs on the Smc head domain are shown (top row). ATP binding to the Walker A domain is blocked in Smc(K37I) ‘(1)’. ATP-dependent engagement of two Smc heads is abolished in the Smc(S1090R) mutant ‘(2)’. The E1118Q mutation strongly reduces ATP hydrolysis ‘(3)’. (B) Smc ATPase mutations abolish chromosomal loading of Smc–ScpAB. In-gel fluorescence detection of Smc-HT of input and eluate fractions from a representative chromosome entrapment assay performed with strains BSG1782 and BSG1784-6. Protein extracts (10% of input) were loaded next to samples subjected to the entrapment assay. Selected cross-linked species of Smc-HT are labeled (top panel). Detection of cross-linked species of DnaN by immunoblotting was used as internal assay control (bottom panel). The following figure supplement is available: Figure 3—figure supplement 1: ATPase mutants of Smc–ScpAB.
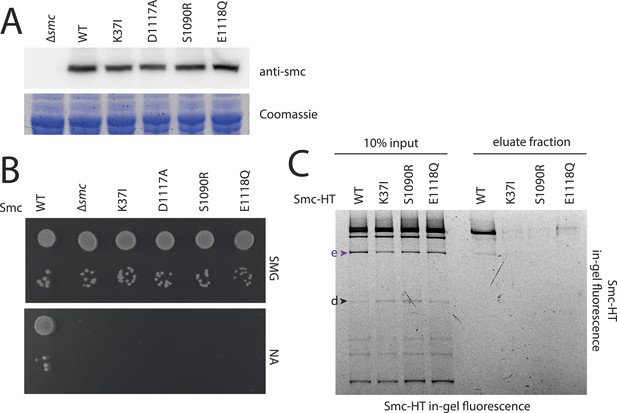
ATPase mutants of Smc-ScpAB.
(A) Upper panel shows Smc expression levels of strains with mutant smc alleles (BSG1002, 1007, 1008, 1045, 1046, 1047 and 1074) determined by immunoblotting using anti-Smc antibodies. Lower panel shows Coomassie staining of a SDS-PAGE gel loaded with the same whole cell extract samples to control for equal protein extraction efficiency. (B) Overnight cultures of strains (BSG1002, 1007, 1008, 1045, 1046, 1047) were spotted on SMG and NA as described before (see Figure 2—figure supplement 1). Smc locus was without mutation (‘WT’), a smc deletion (‘Δsmc’) or mutations in the Smc ATPase domain. (C) Smc ATPase mutations show slightly decreased levels of cross-linking species ‘e’ and increased levels of ‘d’. Image is identical to Figure 3B, with higher contrast.
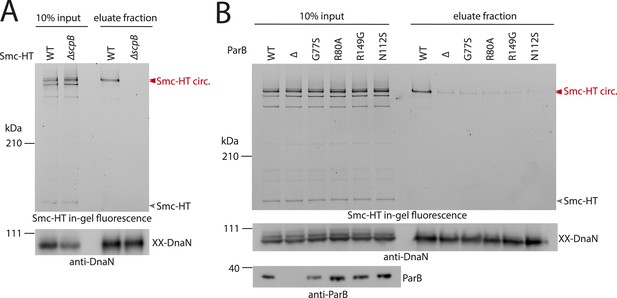
ScpB and ParB are essential for efficient DNA entrapment by Smc complexes.
(A) Deletion of scpB eliminates loading of chromosomal DNA into Smc complexes. In-gel fluorescence detection of Smc-HT in input and eluate fractions is shown from chromosome entrapment assays performed with strains BSG1782 (‘WT’) and BSG1850 (‘ΔscpB’) (top panel). DnaN was used as internal control (bottom panel). (B) Several parB mutations interfere with efficient chromosomal loading of Smc–ScpAB. Input and eluate fractions from chromosome entrapment assays with strains BSG1782, BSG1783 and BSG1960-3 were analysed by in-gel fluorescence detection of Smc-HT (top panel). DnaN was used as internal control (middle panel). Immunoblotting using polyclonal rabbit anti-ParB antiserum confirms near-normal expression of mutant ParB proteins (bottom panel). The following figure supplement is available: Figure 4—figure supplement 1: Growth of smc, parB double mutants.
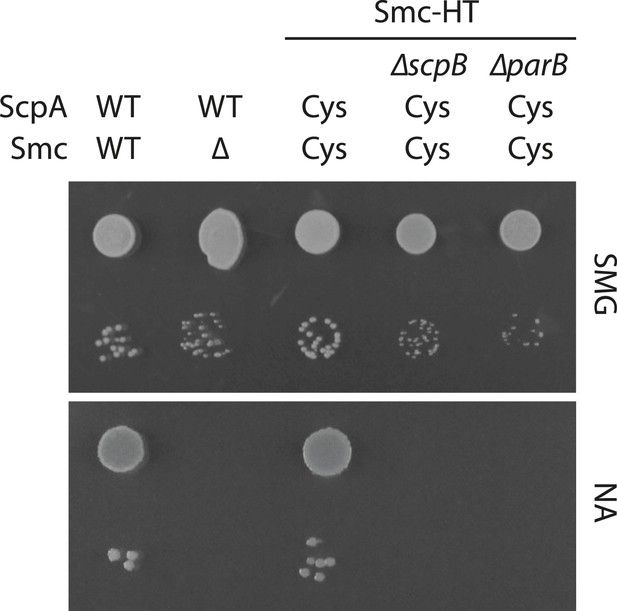
Growth of smc(Cys) mutants.
Overnight cultures of strains (BSG1002, BSG1007, BSG1782, BSG1850 and BSG1783) were serially diluted and spotted as described before (Figure 2—figure supplement 1). Strains were wild-type (‘WT’), harboured the smc deletion (‘Δ’) or cysteine mutations for cross-linking the Smc-ScpA ring (‘Cys’). In addition, scpB (‘ΔscpB’) or parB (‘ΔparB’) were deleted where indicated.
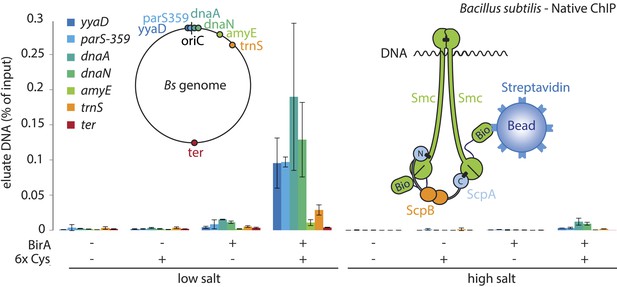
A physical interaction of Smc–ScpAB rings and chromosomal DNA.
Co-purification of chromosomal DNA fragments with native Smc–ScpAB. Cells of strains BSG1104-5 and BSG1107-8 were treated with the cross-linker BMOE prior to cell lysis. Strains carrying (‘+’) or lacking (‘−’) cysteine mutations (‘6xCys’) in the Smc-AviTag construct were expressed in presence (‘+’) or absence (‘−’) of the biotin ligase (‘BirA’). Beads were washed in the presence of either a 150 mM ammonium acetate buffer (‘low salt’) or a 2 M sodium chloride buffer (‘high salt’). The co-purification of DNA fragments with Smc-biotin on streptavidin beads was measured by quantitative PCR using primer pairs specific for genomic positions indicated on a representation of the circular B. subtilis genome. Mean values and standard deviations were calculated from two independent biological replicates. The following figure supplement is available: Figure 5—figure supplement 1: Chromatin immuno-precipitation of Smc.
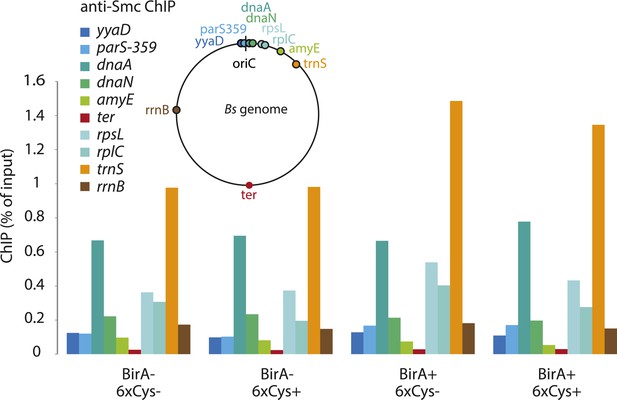
Chromatin immuno-precipitation of Smc.
All Smc variants used in Figure 5 behave similarly in standard ChIP experiments using anti-Smc antibodies. Cells of strains BSG1104-5 and 1107-8 were grown to mid-exponential phase and subjected to chromatin immunoprecipitation (ChIP) using polyclonal rabbit anti-Smc serum. Input and eluate DNA samples were analysed by quantitative PCR using specific primer pairs for the genomic positions indicated on a circular representation of the B. subtilis genome. Pull-down efficiency (ChIP-DNA/input-DNA *100%) was plotted for each primer pair. The lack and presence of the BirA biotin ligase or cysteines for cross-linking of Smc and ScpA proteins are denoted as ‘BirA−’ and ‘BirA+’ or ‘6xCys−’ and ‘6xCys+’, respectively. It is noted that the highly transcribed tRNA locus, trnS, is highly enriched by Smc-Avitag using standard ChIP (but not native ChIP, see Figure 5), being consistent with the notion that enrichment of highly transcribed genes by ChIP is prone to artefacts due to non-uniform formaldehyde cross-linking.
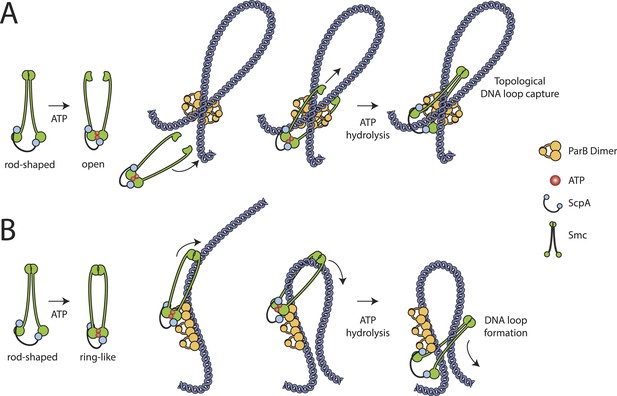
Models for entrapment of chromosomal DNA by Smc–ScpAB.
(A) Loop capture model. DNA loops might be pre-formed within ParB/parS nucleoprotein assemblies. Driven by ATP dependent engagement of Smc head domains Smc–ScpAB adopts a ring-like configuration. Occasional opening of the Smc hinge then allows capture of ParB-DNA loops within Smc–ScpAB. Subsequent ATP hydrolysis by Smc locks the hinge in a closed state and stabilizes the structure. (B) Loop formation model. ParB/parS might serve as a landing platform for Smc–ScpAB allowing Smc–ScpAB in its ring-like conformation to guide DNA into its central cavity. Continuous extrusion of DNA through Smc–ScpAB then drives lengthwise condensation of chromosomes. Ring opening is not required in this model and DNA entrapment by Smc–ScpAB is thus strictly non-topological here. The following figure supplement is available: Figure 6—figure supplement 1: Quantitative Blotting of Smc protein and parB DNA.
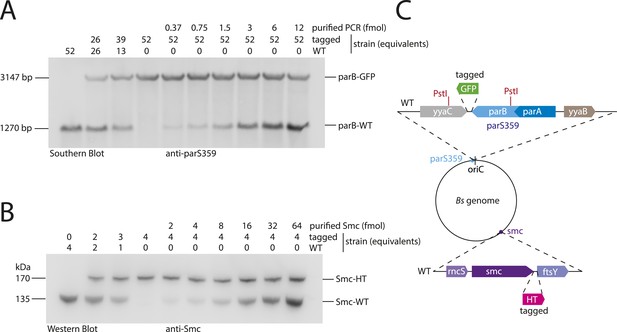
Quantitative Blotting of Smc protein and parB DNA.
To estimate Smc protein and parS-359 DNA abundance in B. subtilis cells, levels of Smc and parS-359 in extracts of wild-type cells were compared to defined amounts of purified Smc protein and parS-359 DNA spiked into equivalent extracts from cell harboring a tagged version of Smc and a modified parS-359 locus (see panel C). The ratio of Smc protein (see panel B) to parS-359 DNA (see panel A) was calculated as average from three independent experiments (biological replicates) to be 61 (±12 SEM, standard error of mean) Smc monomers per parS-359 DNA. Due to potential loss of chromosomal DNA during genomic DNA preparation this might be a slight over-estimation of the real number. (A) Quantitative Southern blotting to estimate the number of molecules (in fmol) of the parS359 locus in a given number of cells. Genomic DNA from 52× cell culture equivalents (1 equivalent equals 0.025 ml*OD600) of wild type (‘WT’, BSG1001) has been digested with the PstI restriction enzyme (see panel C) and loaded into the left lane. DNA loaded into the second and third lane is derived from mixtures of wild-type and tagged (‘parB-GFP’, BSG2058) cells. Defined amounts of a purified PCR product covering the wild-type parS-359 locus have been spiked into cell extracts of the tagged strain. Digested DNA was detected using a Southern probe specific for the parS-359 locus. (B) Quantitative Western blot to estimate the number of molecules of the Smc protein (in fmol) in a given number of cells. Cultures were identical to the ones used in (A). Whole cell protein extract corresponding to 4 culture equivalents of wild-type cells (‘WT’, BSG1001) was loaded into the left lane. Other lanes were loaded with samples derived from wild-type and tagged cells or mixtures thereof with or without spiked-in purified Smc protein as indicated in the figure panel. Smc protein was detected by immunoblotting using a polyclonal rabbit anti-Smc antibody. (C) Protein and DNA spike-in was performed in extracts derived from cells (BSG2058) with modifications at the parS-359 locus (blue filled circle) and the smc gene (purple filled circle). PstI restriction sites (in red) in the parB gene (light blue box) and the neighbouring yyaC gene (grey box) generate a fragment of 1270 bp in size in case of the wild-type locus. From parB-gfp cells (BSG2058), however, a bigger DNA fragment of 3147 bp in size is generated. Strain BSG2058 also harboured a modified smc gene, which is fused to a halotag cassette (pink box).
Additional files
-
Supplementary file 1
Genotypes of Bacillus subtilis strains. All strains are derivatives of either Bacillus subtilis 1A700 (Bacillus Genetic Stock Centre) or Bacillus subtilis 168 ED.
- https://doi.org/10.7554/eLife.06659.016





