The homo-oligomerisation of both Sas-6 and Ana2 is required for efficient centriole assembly in flies
Figures
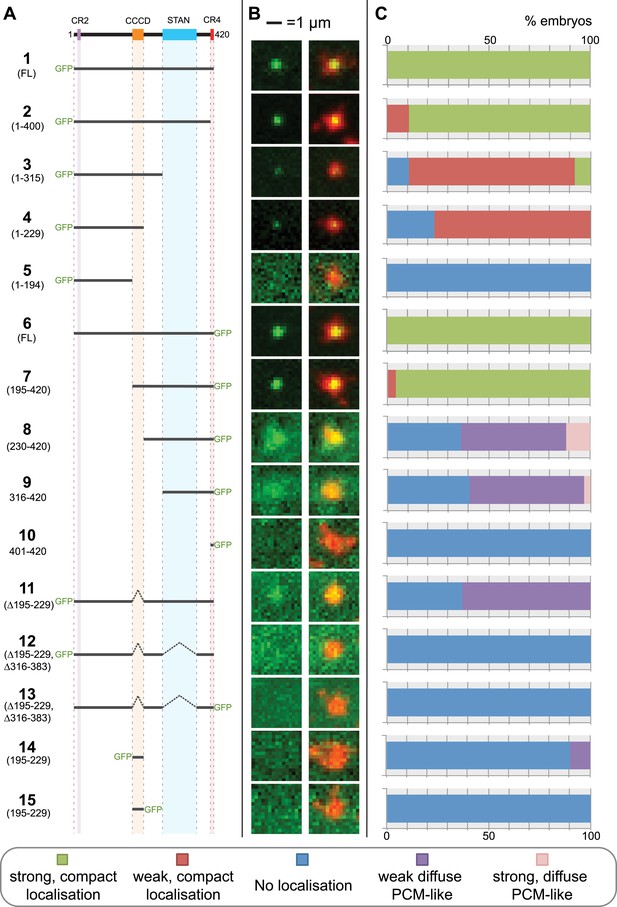
A structure/function analysis of Drosophila Ana2.
(A) A schematic representation of Drosophila Ana2 highlighting the conserved domains and illustrating the GFP constructs analysed in this study. In vitro transcribed mRNA encoding each of these constructs was injected into Drosophila embryos expressing the PCM marker, RFP-Cnn; the distribution of each fusion protein was analysed in living embryos. (B) Micrographs show examples of typical centrosomes in embryos injected with the Ana2 constructs shown in (A). The localisation of the GFP-fusion protein (green) is shown on its own (left panel) and merged with RFP-Cnn (right panel). (C) Bars quantify the localisation behaviour of the various GFP-fusions. Images of 30–80 embryos were analysed for each construct. Images of each embryo were collected and then manually sorted into various categories based on the centrosomal localisation of the GFP-fusion construct (see colour table at bottom of figure). All sorting was performed blind.
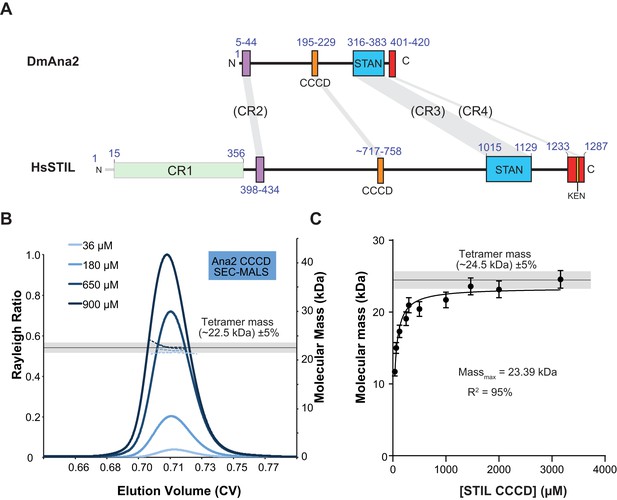
The Ana2 and STIL central coiled-coil domain (CCCD) regions form tetramers in solution.
(A) A schematic representation of D. melanogaster Ana2 and human STIL highlighting the conserved domains. Note that vertebrate STIL proteins contain a conserved region 1 (CR1) of unknown function that is not present in Ana2. (B) A SEC-MALS analysis of the Drosophila Ana2 CCCD (aa193–229) was performed. Injected protein concentrations are indicated by different shades of blue—solid lines show the relative Rayleigh ratio, dashed lines the observed mass. The black horizontal line indicates the theoretical mass for an Ana2 CCCD tetramer, the grey bar indicates a ±5% tolerance. (C) An analysis of the observed mass of human STIL CCCD (717–758) at various injected protein concentrations obtained from SEC MALS experiments. Error bars represent an estimated ±5% error in the MALS mass measurement, as each data point represents a single injection and mass measurement. The black line and grey bar represent the theoretical tetramer mass ±5% tolerance. The data were fitted to a hyperbolic function in Graphpad Prism 6.01, including a 5% SEM for each mass value, with no extrapolation. This fitting estimated that the STIL CCCD was tending towards a mass of 23.4 kDa (theoretical tetramer mass = 24.5 kDa) with an R2 value of 95%.
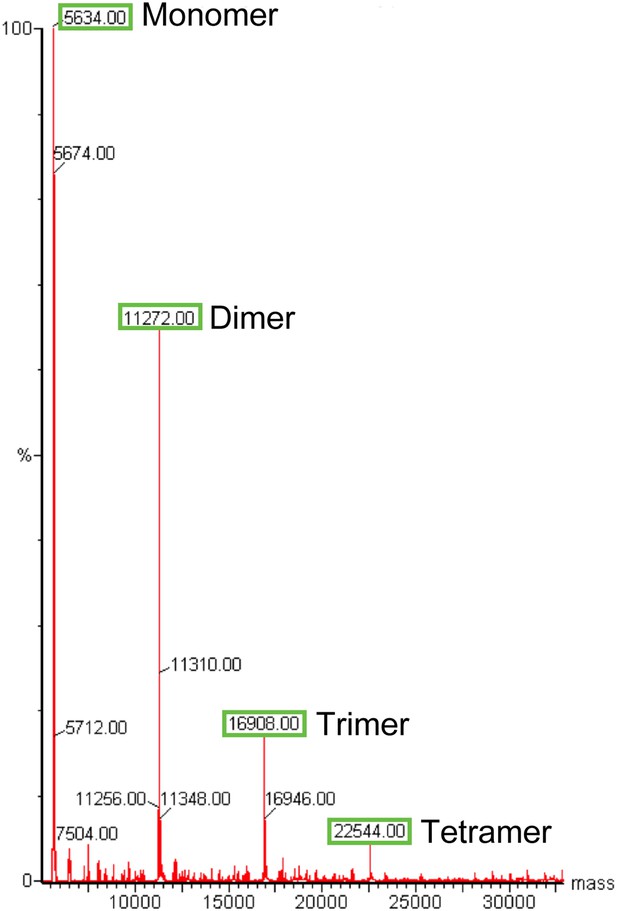

Electrospray-ionisation mass spectrum of the Ana2 CCCD.
The masses (in Da) for dimeric, trimeric and tetrameric Ana2 CCCD species can still be observed, demonstrating that the Ana2 CCCD self-interaction can partially survive these normally denaturing experimental conditions.
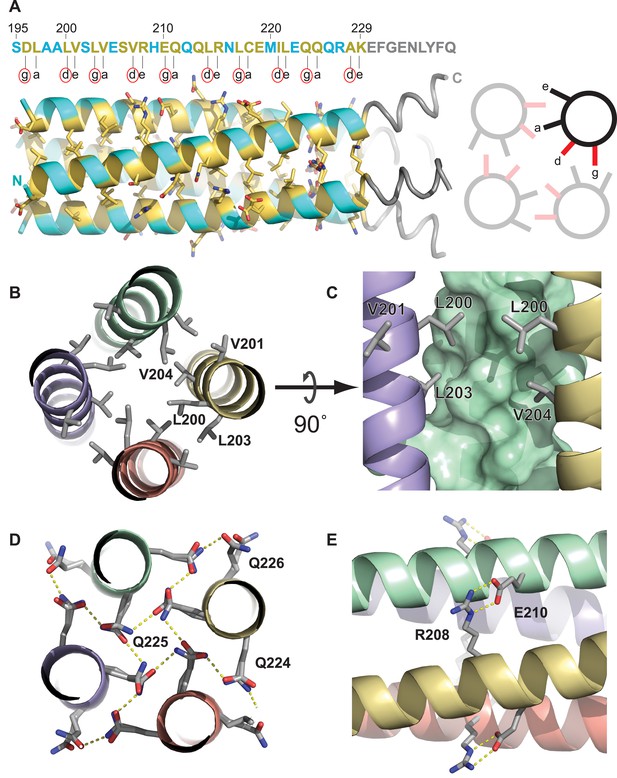
The Ana2 CCCD forms a parallel four helical tetramer.
(A) Left, the structure of the Ana2 CCCD tetramer generated around the crystallographic fourfold symmetry axis. The primary amino acid sequence is shown above the structure; residues in the g, a, d and e positions of the helical heptad repeat are indicated below the sequence. All these residues were ≥30% buried (according to PISA server analysis) and are coloured in yellow, with side-chains in stick format—other residues are coloured in cyan (side-chains not shown). The TEV cleavage remnant is shown in grey. Right, schematic transverse view of the tetramer indicating how the g, a, d and e residues of the heptad repeat are buried at the tetramer interface. Note that the g and d residues (coloured red, and highlighted with a red circle underneath the primary amino acid sequence) form one side of this interface; these 10 residues were mutated to generate forms of the protein that could no longer form tetramers (see main text). (B–E) Schematics illustrate the molecular determinants of tetramerisation, with interfacing residues shown as grey sticks. (B) A hydrophobic cluster of interface residues. The labelled residues sit at the g, a, d and e positions of the heptad repeat, and pack closely forming a hydrophobic environment. (C) A side on view of the same cluster, with one chain shown as a surface. (D) A transverse N-C view of a QQQ triad which adopts positions g, a and b of the heptad. These polar side-chains form an inward facing hydrogen-bond network. (E) A side-on view showing a salt bridge between adjacent chains of the tetramer.
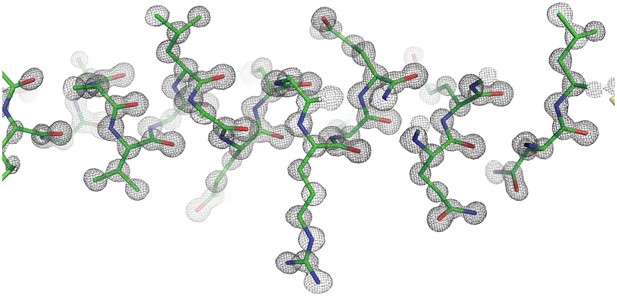
Representative electron density for the Ana2 CCCD crystal structure at 0.8 Å resolution.
The Ana2 CCCD is displayed in stick format, with the 2Fo-Fc map contoured at 1.8 σ.
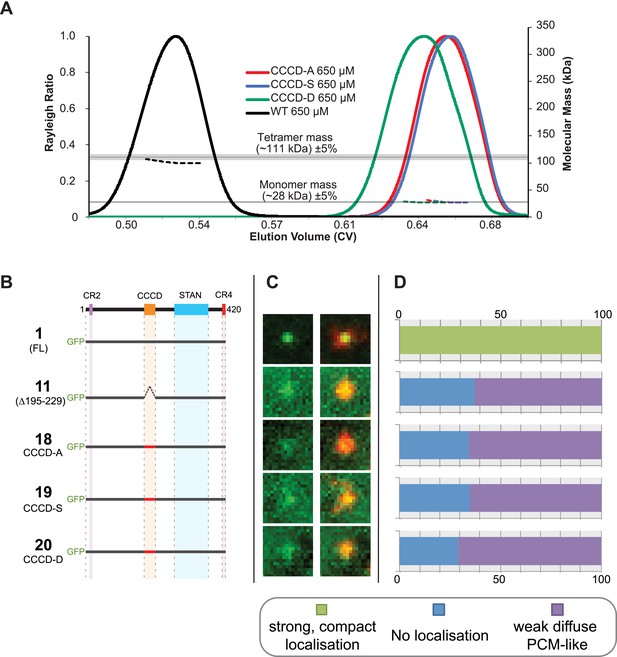
Mutations of the CCCD that perturb tetramer formation in vitro perturb the localisation of Ana2 to centrioles in vivo.
(A) A SEC-MALS analysis of wild type (WT) and mutant forms of the CCCD where the 10 d and g residues important for tetramer formation (circled in red, Figure 3A) have been mutated either to Ala (CCCD-A), Ser (CCCD-S) or Asp (CCCD-D). Horizontal black lines illustrate the theoretical molecular mass of a tetramer and monomer, grey shading represents ±5% tolerance. Note that, in contrast to the SEC-MALS analysis presented in Figure 1A, the diLipoyl domains of the fusion proteins have not been removed in this experiment, so the masses of the monomer and tetramer are higher. (B) A schematic representation of the GFP-Ana2 fusions that contain mutations of the CCCD (constructs #1 and #11 are the same constructs shown in Figure 1A). In vitro transcribed mRNA encoding each of these constructs was injected into Drosophila embryos expressing the PCM marker, RFP-Cnn; the distribution of each fusion protein was analysed in living embryos. (C) Micrographs show examples of typical centrosomes in embryos injected with the Ana2 constructs shown in (A). The localisation of the GFP-fusion protein (green) is shown on its own (left panel) and merged with RFP-Cnn (right panel). (D) Bars quantify the localisation behaviour of the various GFP-fusions. Images of 34–40 embryos were analysed for each construct. Images of each embryo were collected and then manually sorted into various categories based on the centrosomal localisation of the GFP-fusion construct (see colour table at bottom of figure). All sorting was performed blind. The data shown here for constructs #1 and #11 is the same as that presented in Figure 1C.
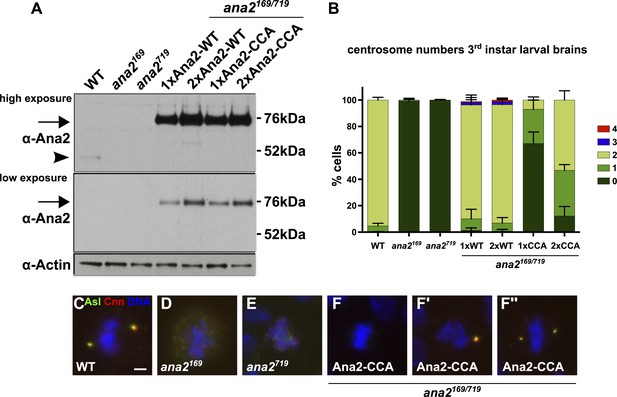
A Mutant form of Ana2 that cannot tetramerise efficiently in vitro cannot support efficient centriole duplication in vivo.
(A) Two exposures of a western blot illustrating the relative expression levels of endogenous Ana2 (arrowhead) and either WT Ana2-GFP or Ana2-CCA-GFP (arrow) (expressed from one (1×) or two (2×) copies of the transgene) in third instar larval brains in either a WT or ana2 mutant background. Actin is shown as a loading control. (B) The bar chart shows the number of centrosomes or centrosome-like structures (CLSs) observed in mitotic third instar larval brain cells (scored by the presence of both the centriole marker Asl and the centrosome marker Cnn) in WT, ana2 mutant and ana2 mutants expressing one (1×) or two (2×) copies of either WT Ana2-GFP or Ana2-CCA-GFP, as indicated. A total of at least 300 mitotic cells from at least five different brains were scored for each genotype; error bars represent the SD. (C–F′′) Micrographs show the distribution of Asl (green) and Cnn (red) in representative mitotic third instar larval brain cells of the indicated genotypes. DNA is in blue. The images in F–F′′ show cells rescued with the Ana2-CCA-GFP construct that have either no centrosomes (F) or one (F′) or two (F′′) CLSs. Scale bar in C: 2 µm.
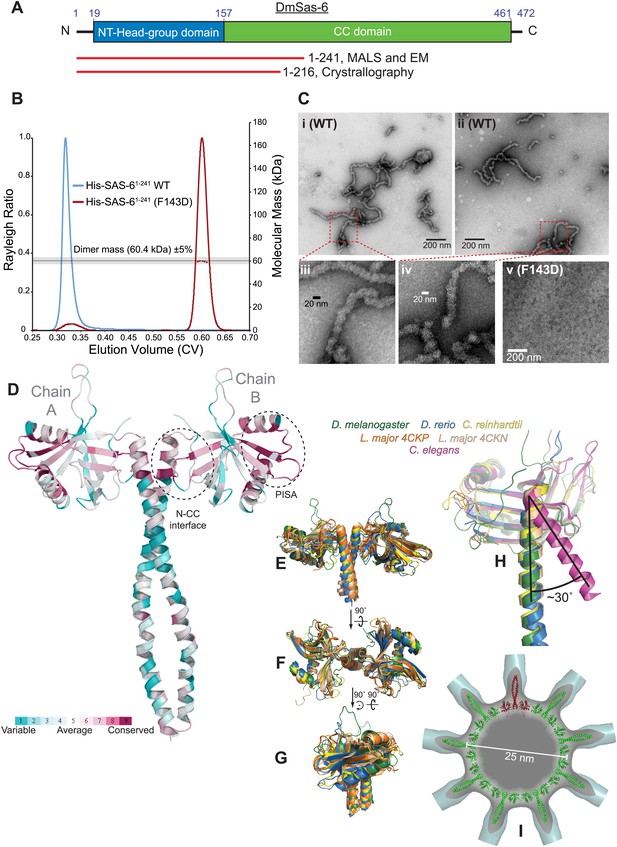
A biochemical and structural analysis of Drosophila Sas-6.
(A) A schematic representation of Drosophila Sas-6 highlighting the position of the N-terminal head domain (blue) and C-terminal coiled-coil (CC) domain (green). Red lines below represent the constructs used in SEC-MALS and EM studies (top) and in X-Ray Crystallography studies (bottom). (B) A SEC-MALS analysis of WT (blue trace) and F143D mutant (red trace) Sas-61–241 proteins, injected at 33 µM. The horizontal black line and grey bar represent the theoretical dimer mass ±5% tolerance. The WT protein could not be analysed by MALS as it eluted in the void volume and appeared to form a range of higher-order oligomers. (C) Negative-stain EM analysis of purified WT ([i]–[iv]) Sas-61–241 protein, showing the chain-like structures formed ([iii] and [iv] show magnified views of the red boxed areas in [i] and [ii]); these structures are not detectable in preparations of the mutant Sas-6-F143D1–241 protein ([v]). (D) The structure of the Sas-6 dimer, coloured according to Consurf conservation scores (Glaser et al., 2003) from cyan (variable) to burgundy (conserved). The conserved PISA domain and the N-CC interface regions are highlighted with dashed circles. (E–G) Superimposed structures from D. melanogaster, D. rerio, Chlamydomonas and Leishmania (as indicated) of the Sas-6 N-terminal head-group with a short stretch of the coiled-coil domain. (H) Superimposed structures of the N-CC interface in D. melanogaster, D. rerio, Chlamydomonas and C. elegans. Note how the interface is rotated by ∼30° in C. elegans (purple) compared to the other structures. (I) The DmSas-6 structure modelled into a ninefold symmetric flat ring (green, single dimer shown in red), similar to that observed in crystallo for LmSAS-6. This ring structure was docked into the EM density of the Triconympha cartwheel structure (Guichard et al., 2013) (cyan surface, cut away to reveal the DmSAS-6 ring).
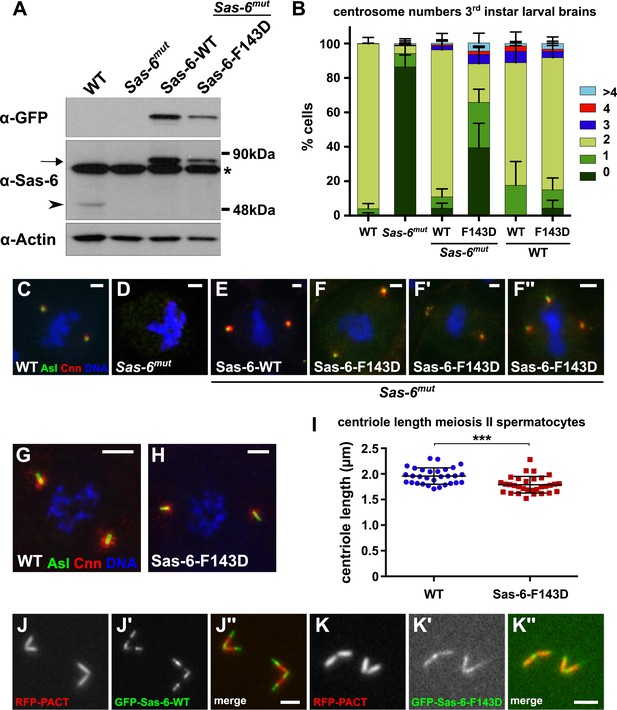
A Mutant form of Sas-6 that cannot oligomerise efficiently in vitro cannot support efficient centriole duplication in vivo.
(A) A western blot illustrating the relative expression levels of endogenous Sas-6 (arrowhead) and either WT GFP-Sas-6 or GFP-Sas-6-F143D (arrow) in third instar larval brains in either a WT or Sas-6 mutant background. Actin is shown as a loading control, and an (*) marks a non-specific band. (B) The bar chart shows the number of centrosomes or CLSs observed in mitotic third instar larval brain cells in WT or Sas-6 mutants (first and second bars); in Sas-6 mutants expressing either WT GFP-Sas-6 or GFP-Sas-6-F143D (third and fourth bars); or WT brains expressing either WT GFP-Sas-6 or GFP-Sas-6-F143D (fifth and sixth bars). At least 600 mitotic cells from at least five different brains were scored for each genotype; error bars represent the SD. (C–F′′) Micrographs show the distribution of Asl (green) and Cnn (red) in representative mitotic third instar larval brain cells of the indicated genotypes. DNA is in blue. The images in F, F′ and F′′ show Sas-6 mutant cells rescued by Sas-6-F143D showing examples of the CLSs. (G, H) Micrographs show the distribution of Asl (green) and Cnn (red) in either a WT primary spermatocyte (G) or a WT primary spermatocyte overexpressing GFP-Sas-6-F143D (H). (I) Graph shows the quantification of centriole length (as measured by Asl staining) in WT primary spermatocytes (blue circles) or WT primary spermatocytes overexpressing GFP-Sas-6-F143D (red boxes); at least 1500 centrioles from at least 30 different testes were scored for each genotype, and each circle or box represents the mean from an individual testes. Statistical significance was assessed using an unpaired two-tailed t-test: (***) indicates p-value < 0.001. (J–K) Micrographs show the distribution of the centriole marker RFP-PACT (red) and either WT GFP-Sas-6 (green) (J–J′′) or GFP-Sas-6-F143D (K–K′′) in WT primary spermatocytes. Scale bars: 2 µm in C–F′′ and J–K′′ and 5 µm in H–I.
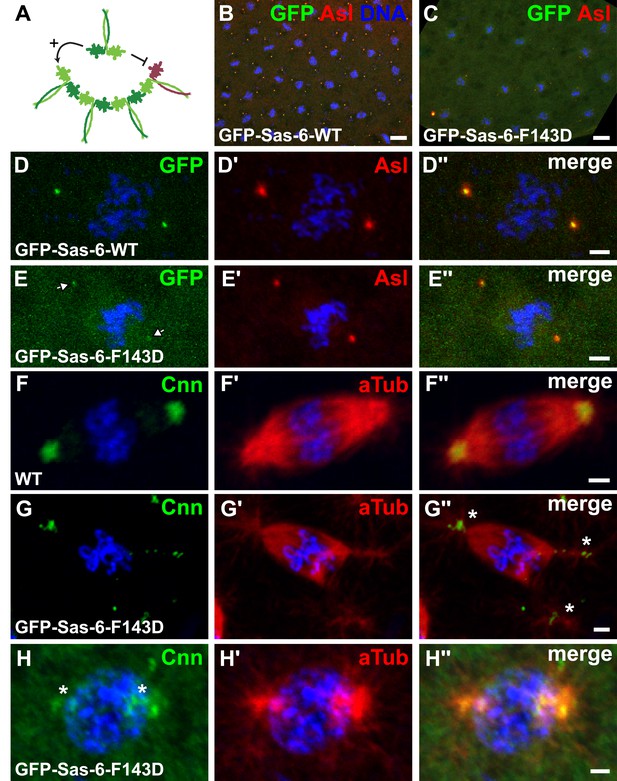
GFP-Sas-6-F143D dominantly suppresses centrosome assembly in early embryos.
(A) A schematic illustration of how GFP-Sas-6-F143D could act as a dominant-negative in cartwheel assembly. WT Sas-6 (light and dark green) can form WT–WT homodimers or WT-mutant heterodimers with GFP-Sas-6-F143D (red). The homodimers can support cartwheel assembly while the heterodimers can incorporate into the growing cartwheel (through the WT headgroup), but cannot support further cartwheel assembly. The heterodimer must dissociate before a WT homodimer can incorporate into the cartwheel, so allowing cartwheel assembly to proceed. (B–H′′) Micrographs show images from WT embryos expressing no transgene (F) or expressing either WT GFP-Sas-6 (B, D) or GFP-Sas-6-F143D (C, E, G, H) stained to reveal the distribution of GFP, Asl, Cnn or α-tubulin, as indicated. (B, D) Embryos expressing WT GFP-Sas-6 develop normally, and the fusion protein strongly localises to a bright spot in the centre of the centrosomes. (C, E) Embryos expressing GFP-Sas-6-F143D arrest during the early syncytial stages; some of these embryos are reasonably well organized and centrioles are observed at the spindle poles, but these contain very little detectable GFP-Sas-6-F143D. (F–G) Most embryos are less well organized and contain abnormal microtubule (MT) arrays organized by fragmented centrosomes (G, H) when compared to WT (F). Scale bars: 10 µm in B, C, 2 µm in D–D′′ and F–H′′ and 3 µm in E–E′′.
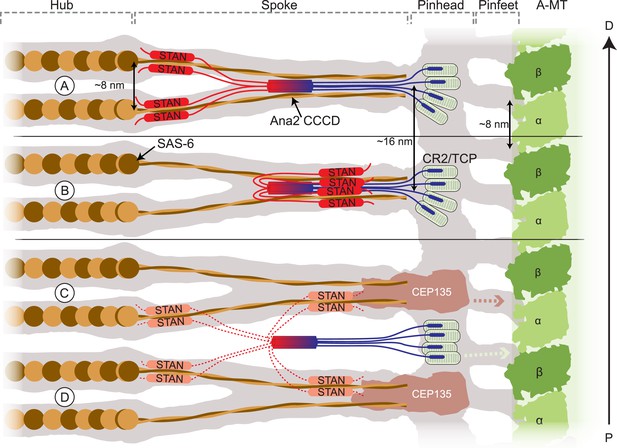
Schematic models of how Ana2 tetramers might contribute to cartwheel assembly.
The grey area shows a representation of the electron density map derived from the Triconympha cartwheel structure (emd-2329/2330) (Guichard et al., 2013). A protofilament of the centriole A-MT is shown in green, and full-length Sas-6 molecules (brown/orange) have been placed into the cartwheel density. The Ana2 CCCD tetramer is coloured blue/red, extending out to the N-terminus in blue and to the C-terminus in red. The N-terminal Ana2 CR2 domain (blue) is shown bound to the Sas-4 C-terminal TCP domain (pale green) (Cottee et al., 2013; Hatzopoulos et al., 2013); Sas-4 is located towards the periphery of the cartwheel (Mennella et al., 2012) so we place the CR2/TCP interaction in the peripheral pinhead region. The STAN domain (red) binds Sas-6 (Ohta et al., 2014); as the C–C domain of Sas-6 is required for targeting to the centriole (Keller et al., 2014) (and so presumably for binding to Ana2). We present two alternative models where the STAN domain binds to the Sas-6 C–C region either at the N-terminal head-linker region (A) or at a more C-terminal region towards the end of the spoke (B). (A, B) In these models, the Ana2 tetramer interacts with two Sas-6 dimers, one that is incorporated into the top Sas-6 ring, and one that is incorporated into the bottom Sas-6 ring. An attractive feature of this model is that it can explain why the fundamental building block of the cartwheel seems to comprise two Sas-6 rings rather than one. Making cartwheel assembly dependent on the co-assembly of two rings could dramatically reinforce the tendency to form a ninefold symmetric ring (see main text). In these models, the density of the Ana2 molecules would fit well into the existing density model. (C, D) In these models, Ana2 forms the same interactions in the same stoichiometry, but the tetramer bridges adjacent paired spoke layers. This model is less attractive as the Ana2 molecules span two of the basic building blocks of the cartwheel (rather than contributing to the stability of the basic building block), and the Ana2 molecules would not fit into the existing density models (although this is possible if they are held in a flexible manner, so their density is not detectable by EM). In these latter models we show Cep135 tethering the SAS-6 spoke to the pinhead (Lin et al., 2013). This interaction likely also occurs in the models shown in (A, B) (although, for ease of presentation, it is not shown); in the models shown in (C, D) this putative interaction between Sas-6 and Cep135 would function as the key interaction stabilising the two ring structure. Models where the Ana2 tetramer interacts with Sas-6 molecules in only one layer of the Sas-6 ring structure are also possible, but are not illustrated here.
Tables
Ana2 CCCD dataset and refinement statistics
| Dataset statistics | |
| Beamline | Diamond I03 |
| Wavelength (Å) | 0.7293 |
| Spacegroup | I4 |
| Unit cell dimensions (Å/°) | 33.27, 33.27, 74.49/90.00, 90.00, 90.00 |
| Resolution (Å) (overall/inner/outer) | 30.36–0.80/30.36–3.58/0.82–0.80 |
| Completeness (overall/inner/outer) | 97.8/99.9/79.2 |
| Rmerge (overall/inner/outer) | 0.038/0.034/0.547 |
| Rpim (overall/inner/outer) | 0.013/0.011/0.312 |
| CC (1/2) (overall/inner/outer) | 1.00/0.999/0.782 |
| I/σI (overall/inner/outer) | 21.5.71.3/2.2 |
| Mulitiplicity (overall/inner/outer) | 5.8/11.2/3.3 |
| Refinement statistics (parentheses = highest res shell) | |
| Resolution range (Å) | 30.36–0.80 (0.82.0.80) |
| Rwork/Rfree/% test set size | 10.6/11.6/5.06% (21.1/20.3/4.84%) |
| Number of reflections working set/test set | 39,348 (2338)/2099 (119) |
| Number of atoms (non-H) | 499 |
| Waters | 53 |
| Rmsd from ideal values: bond length (Å)/angles (°) | 0.025/2.230 |
| Average B factor (Å2) | 10.70 |
| Ramachandran outliers | 0% |
| Ramachandran favoured | 100% |
| MolProbity score (N number, percentile) | 1.22 (222, 88%) |
-
Ramachandran and Molprobity scores were calculated using MolProbity (Chen et al., 2010).
Sas-61–216 (F143D) dataset and refinement statistics
| Dataset statistics | |
| Beamline | ESRF ID23-2 |
| Wavelength (Å) | 0.8726 |
| Spacegroup | P2 |
| Unit cell dimensions (Å/°) | 47.13, 64.74, 123.73/90.00, 98.91, 90.00 |
| Resolution (Å) (overall/inner/outer) | 41.43–2.92/41.43–13.06/3.00–2.92 |
| Completeness (overall/inner/outer) | 97.4/90.4/96.4 |
| Rmerge (overall/inner/outer) | 0.128/0.035/0.668 |
| Rpim (overall/inner/outer) | 0.060/0.017/0.318 |
| CC (1/2) (overall/inner/outer) | 0.994/0.988/0.799 |
| I/σI (overall/inner/outer) | 12.1/48.4/2.6 |
| Mulitiplicity (overall/inner/outer) | 5.2/4.8/5.1 |
| Refinement statistics (parentheses = highest res shell) | |
| Resolution range (Å) | 41.43–2.92 (3.10–2.92) |
| Rwork/Rfree/% test set size | 18.3/21.5/5.00% (26.2/34.5/5.90%) |
| Number of reflections working set/test set | 14,976 (2456)/788 (154) |
| Number of atoms (non-H) | 3405 |
| Waters | 33 |
| Rmsd from ideal values: bond length (Å)/angles (°) | 0.007/1.055 |
| Average B factor (Å2) | 76.30 |
| Ramachandran outliers | 0% |
| Ramachandran favoured | 94.9% |
| Molprobity score (N number, percentile) | 1.54 (3648, 100%) |
-
Ramachandran and Molprobity scores were calculated using MolProbity (Chen et al., 2010).





