Auxin-regulated chromatin switch directs acquisition of flower primordium founder fate
Figures
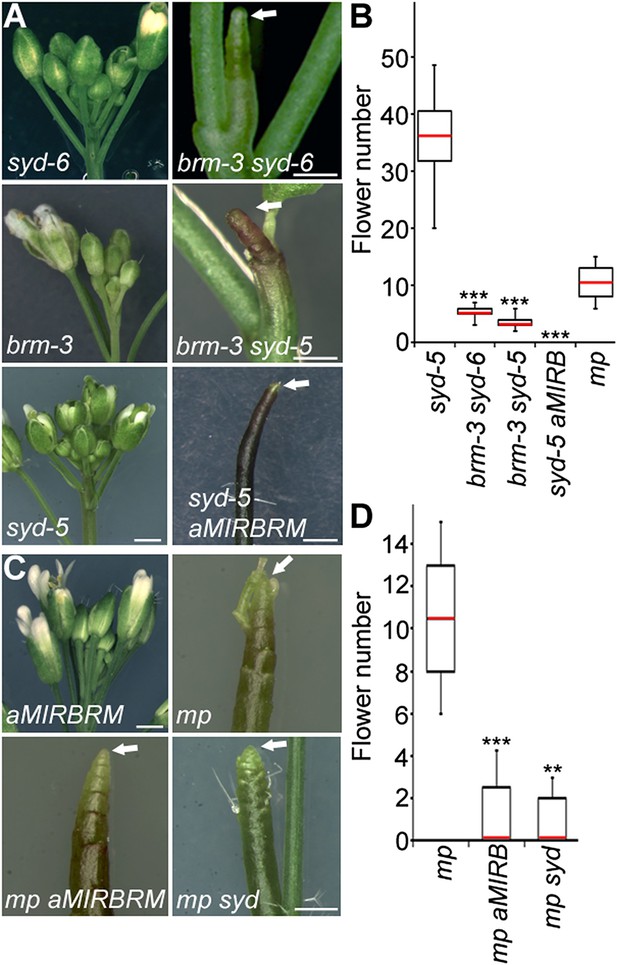
An essential role for SWI/SNF ATPases in flower primordium initiation.
(A) ‘Pin’ inflorescence phenotype (white arrow) of brm syd double mutants. Scale bars = 1 mm. Allelic strength of mutants: syd-6, very weak; brm-3 weak; aMIRBRM, strong; syd-5 null. (B) Quantification of the flower primordia initiated in (A). n > 18. p-value: Mann–Whitney U test. (C) Enhancer tests using hypomorph mp-S319 mutant (Schlereth et al., 2010). White arrows point to ‘pin’ inflorescences. Scale bars = 1 mm. brm-1 null mutant (Hurtado et al., 2006) combined with the mp-S319 hypomorph mutant is seedling lethal like the mp-B4149 null mutant (Weijers et al., 2006) (Figure 1—figure supplement 1) and has developmental defects in the embryo (Figure 1—figure supplement 2). (D) Quantification of the flower primordia initiated in (C). n > 5. p-value: Mann–Whitney U test.
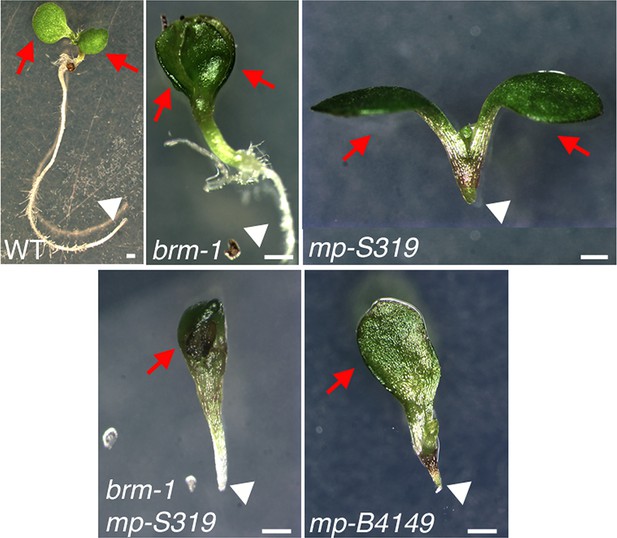

brm-1 null mutant mutants enhance mp-S319 hypomorph seedling phenotypes to phenocopy mp-B4149 null mutants.
Scale bars = 1 mm. Red arrows: number of cotyledons; white arrowhead: root growth arrest.
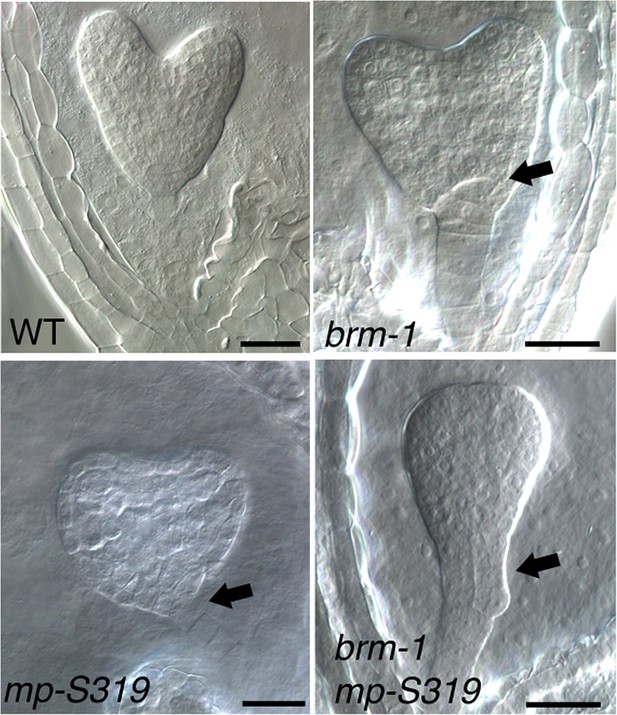
brm-1 null mutant enhance mp-S319 hypomorph embryo phenotypes.
Arrows: irregular cell division in hypophysis. Scale bars = 30 μm.
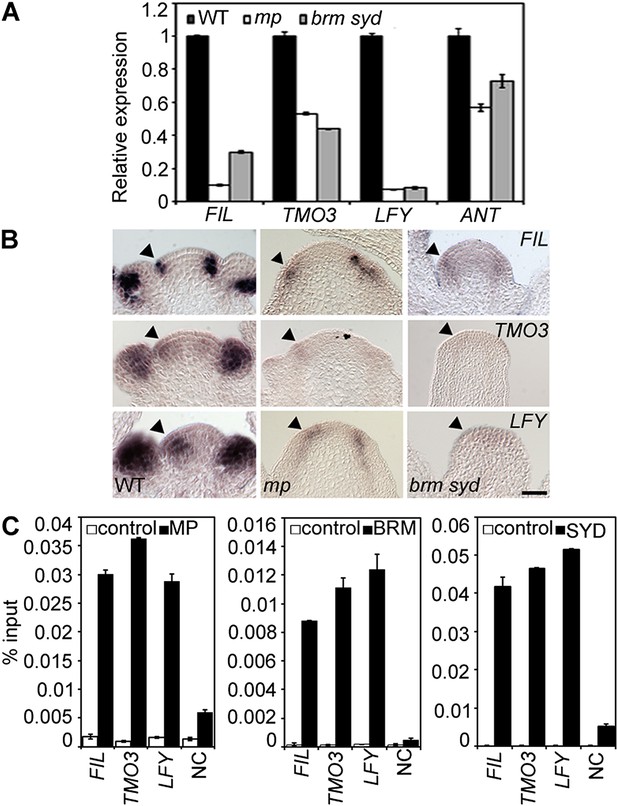
MP and BRM/SYD directly regulate common target genes.
(A) Expression levels of FIL, TMO3, LFY and ANT in wild-type (WT), mp-S319 or brm-3 syd-5 inflorescence shoot apices normalized to that of EIF4A-1. Expression in WT was set to one. (B) In situ hybridization of wild-type, mp-S319 or brm-3 syd-5 inflorescence shoot apices prior to ‘pin’ formation using antisense FIL, TMO3 and LFY probes. Black arrowheads: organogenic region from where flower primordia usually arise. Figure 2—figure supplement 1 shows that MP expression is not visibly reduced in brm-3 syd-5 mutants. Inducible increase or reduction of MP function triggered increased or decreased expression of FIL, TMO3, LFY and ANT, respectively (Figure 2—figure supplement 2). (C) Anti-GFP chromatin immunoprecipitation (ChIP) to test pSYD:GFP-SYD and pBRM:BRM-GFP occupancy at pMP:MP-HA bound sites (as determined by anti-HA ChIP). For MP, BRM and SYD occupancy at the ANT locus see Figure 2—figure supplement 3. For comparison of the binding pattern of BRM, SYD and MP at the FIL, TMO3 or LFY loci see Figure 2—figure supplement 4. Control: anti-GFP or anti-HA ChIP in non-transgenic plants. NC: negative control locus (Ta3 retrotransposon).
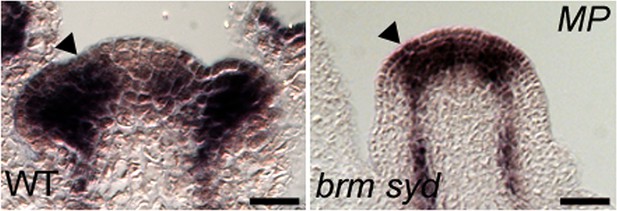
MP expression in wild type and brm-3 syd-5 mutant inflorescences.
Black arrowhead: Site of flower initiation. Sections are from the same experiment and slide. Scale bar = 30 µm.
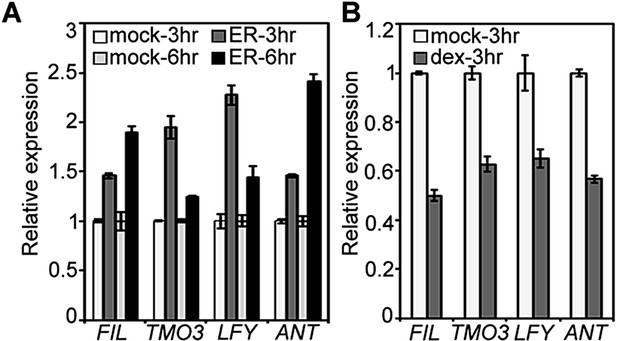
Elevated MP activity leads to increased and reduced MP activity to decreased accumulation of LFY, TMO3, FIL and ANT.
(A) qRT-PCR of estradiol (ER, 10 µM) or mock treated pER>>MP∆C plants. (B) qRT-PCR of dexamethasone (dex, 10 µM) or mock treated 35S:axr3-GR plants. axr3 is an auxin-insensitive MP-interacting Aux/IAA protein. See methods for details.
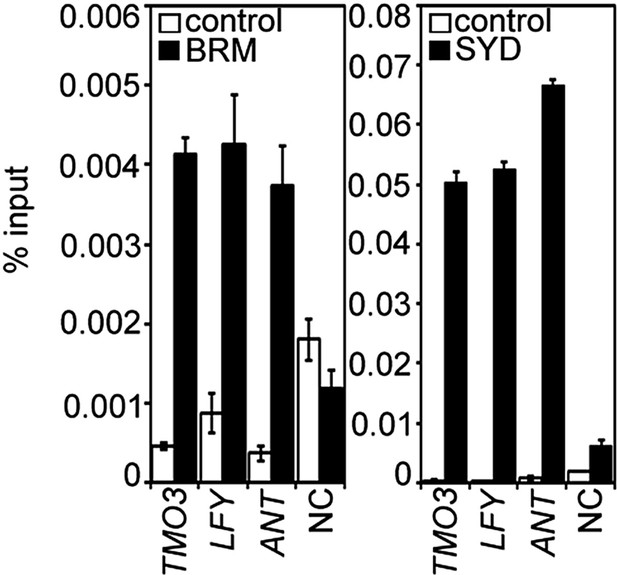
BRM and SYD occupancy at known MP target loci.
https://doi.org/10.7554/eLife.09269.009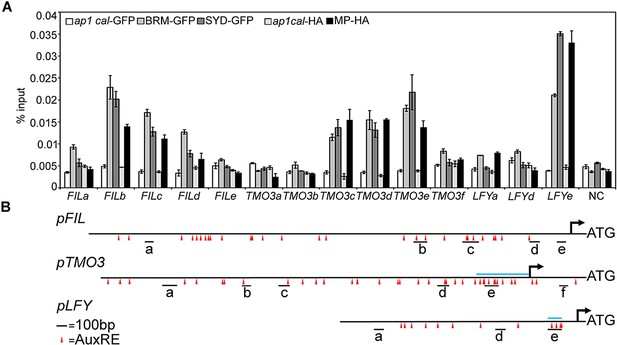
BRM, SYD, or MP occupancy at different regions of the FIL, TMO3 and LFY loci.
(A) ChIP-qPCR. (B) Diagram of the upstream intergenic regions. Red triangles: core AuxREs, blue line above locus: previously defined MP bound regions at the TMO3 and LFY loci (Schlereth et al., 2010; Yamaguchi et al., 2013). Black line below loci: PCR amplicons. Amplicon TMO3e was used for all other MP, BRM and SYD ChIP experiments (Figure 2C and Figure 2—figure supplement 3). Amplicons FILb and LFYe were used for all further ChIP experiments (MP, BRM, SYD, HDA19, TPL) as well as for FAIRE.
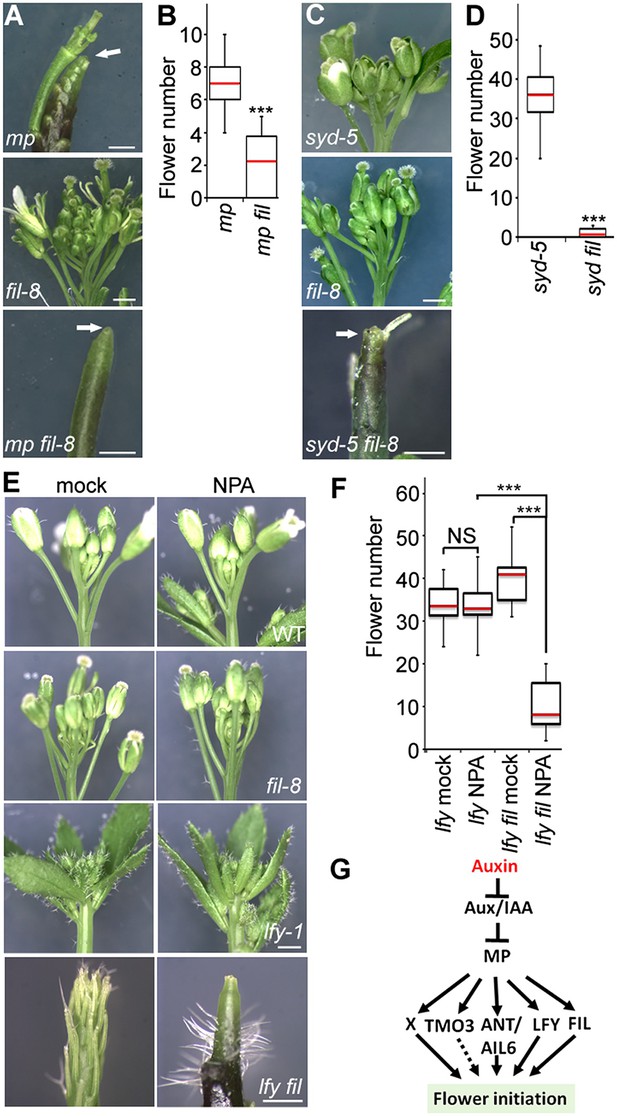
The direct MP and BRM/SYD target FIL plays a role in flower primordium initiation.
(A) Enhancer test using the hypomorph mp-S319 and the null fil-8 mutant. Scale bars = 1 mm. White arrows point to pin inflorescences. (B) Quantification of flower primordia initiated in (A). n > 10. p-value: Mann–Whitney U test. (C) Enhancer test using null syd-5 mutant and fil-8. White arrow points to pin-like inflorescence. Scale bars = 1 mm. (D) Quantification of flower primordia initiated in (C). n > 5. p-value: Mann–Whitney U test. (E) ‘Pin’ inflorescence phenotype of lfy-1 fil-8 double mutant treated with the auxin transport inhibitor N-1-naphthylphthalamic acid (NPA). Scale bars = 1 mm. (F) Quantification of the flower primordia initiated in (E). n > 12. p-value: Mann–Whitney U test. (G) Updated model for auxin/MP-mediated flower primordium initiation together with BRM/SYD. Dashed arrow: role not yet proven. X: additional MP target(s) with a role in flower initiation.
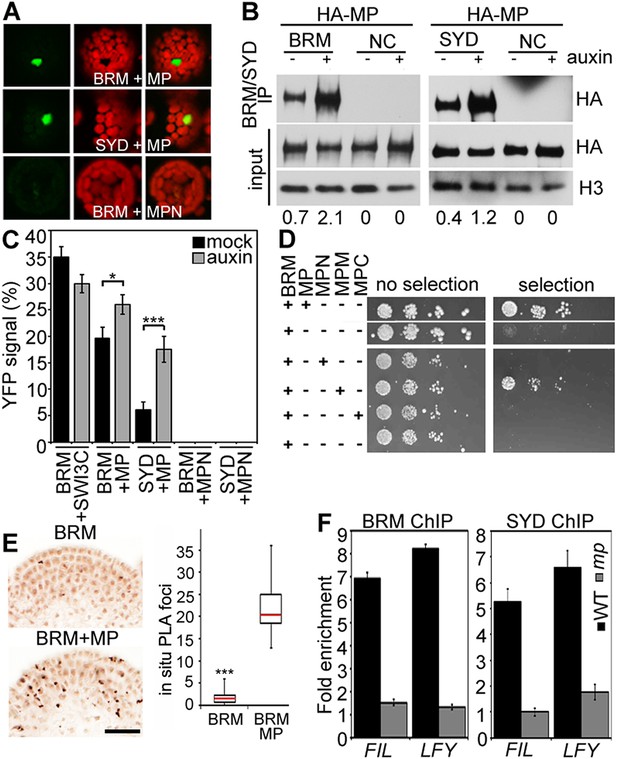
MP physically interacts with and recruits BRM and SYD to target loci.
(A) Bimolecular fluorescence complementation (BiFC) test of MP and BRM or SYD protein interaction in plant cells. Green: BiFC signal in the nucleus, red: chloroplast auto-fluorescence. MPN: N-terminal domain of MP. (B) Co-immunoprecipitation using anti-FLAG antibody in plant cells expressing HA-MP with or without FLAG-BRM or FLAG-SYD. Western blot is probed with anti-HA or anti-histone H3 antibody. Below: Amount of precipitated HA-MP (% of input). See also Figure 4—figure supplement 1. (C) Quantification of BiFC events in the absence or presence of auxin. The error bars are proportional to the standard error of the pooled percentage computed using binomial distribution. n = 3. p-value; Mann–Whitney U test. SWI3C: BRM chromatin remodeling complex component (positive control). (D) Yeast-two-hybrid test of interaction between BRM and MP or MP domains: N: N-terminus, M: middle region, C: C-terminus. See Figure 4—figure supplement 2 for domains of the MP protein. Growth was assayed minus (left) or plus (right) 3-amino-1,2,4-triazole. Thin white line: cropped image from one plate. (E) In situ proximity ligation assay (PLA) with anti-GFP and anti-HA antibodies in pBRM:BRM-GFP or pMP:MP-HA pBRM:BRM-GFP shoot apices. Left: individual sections, right: quantification of interaction foci. n > 12. p-value: Student's t-test. (F) BRM and SYD ChIP enrichment at the FIL and LFY loci relative to the control locus (Ta3 retrotransposon) in wild-type and mp-S319 mutant inflorescences.
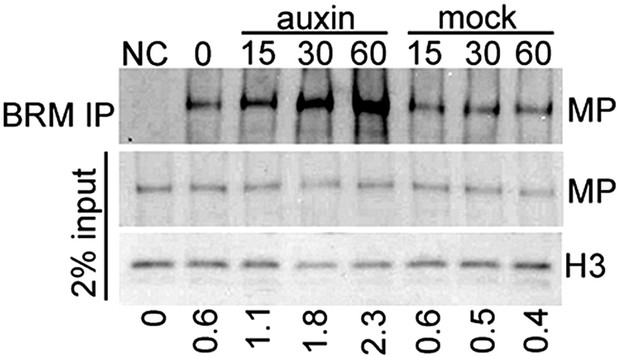
Auxin treatment enhanced the physical interaction between BRM and MP.
IP: epitope tagged BRM (FLAG). Western blot: epitope-tagged MP (HA) and histone H3 (loading control). Negative control sample (NC): plants expressing YFP-tagged BRM and HA-tagged MP. Below: Amount of MP immunoprecipitated (percent of input).
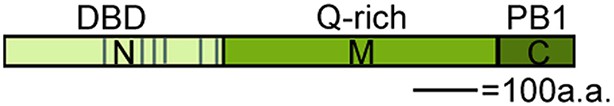
Domains of MONOPTEROS.
N: N-terminal DNA binding (DBD) and dimerization domain, M: Glutamine (Q) rich middle region linked to transcriptional activation, C: C-terminal Phox and Bem1 (PB1)-like domain required for interaction with Aux/IAA proteins (Boer et al., 2014; Korasick et al., 2014; Tiwari et al., 2003). Grey lines in N-terminal domain: nuclear localization signals.
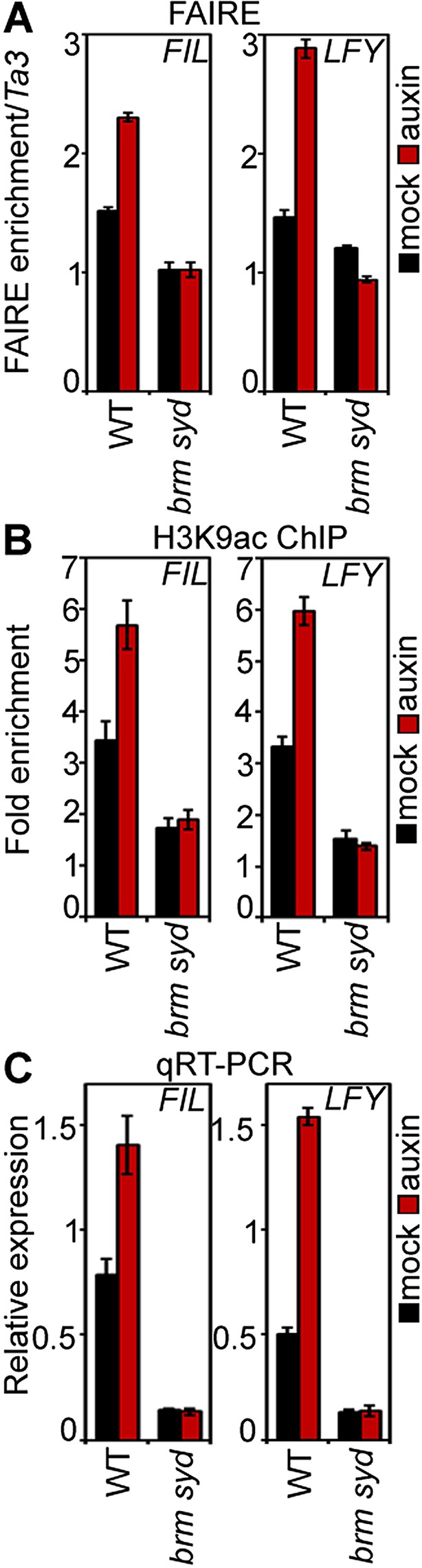
SWI/SNF chromatin remodeling ATPases are required for increased accessibility of MP target loci in response to auxin.
(A) DNA accessibility at FIL and LFY loci in the context of chromatin assayed by Formaldehyde Assisted Isolation of Regulatory Elements (FAIRE) in response to auxin treatment in wild-type (WT) and brm-3 syd-5 inflorescences. The ratio of FAIRE enrichment at the locus of interest was normalized over that at the Ta3 retrotransposon. (B) Anti-histone 3 lysine 9 acetylation (H3K9ac) ChIP at the FIL (left) and LFY (right) locus normalized over that at Ta3 in genotypes and treatments shown in (A). (C) FIL and LFY RNA accumulation relative to EIF4A-1 in genotypes and treatments shown in (A).
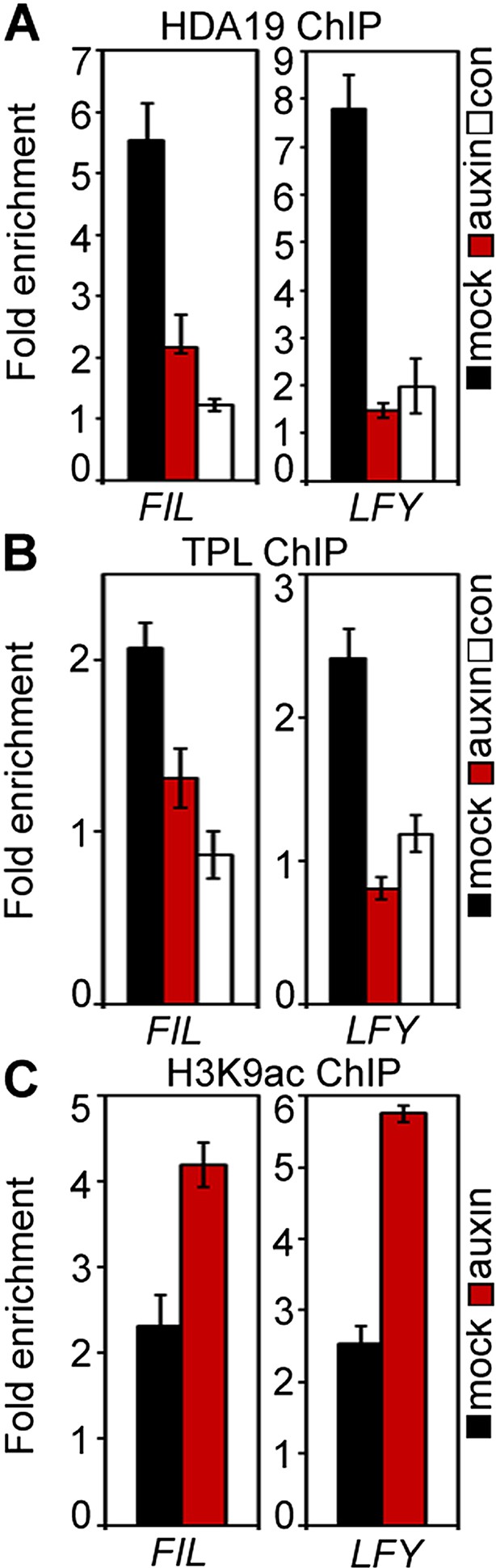
TPL/HDA occupancy and H3K9ac levels at the FIL and LFY target loci with and without auxin application.
(A) Histone deacetylase 19 (HDA19)-GFP ChIP. (B) TOPLESS (TPL-GFP) ChIP. (C) Anti-histone 3 lysine 9 acetylation (H3K9ac) ChIP.
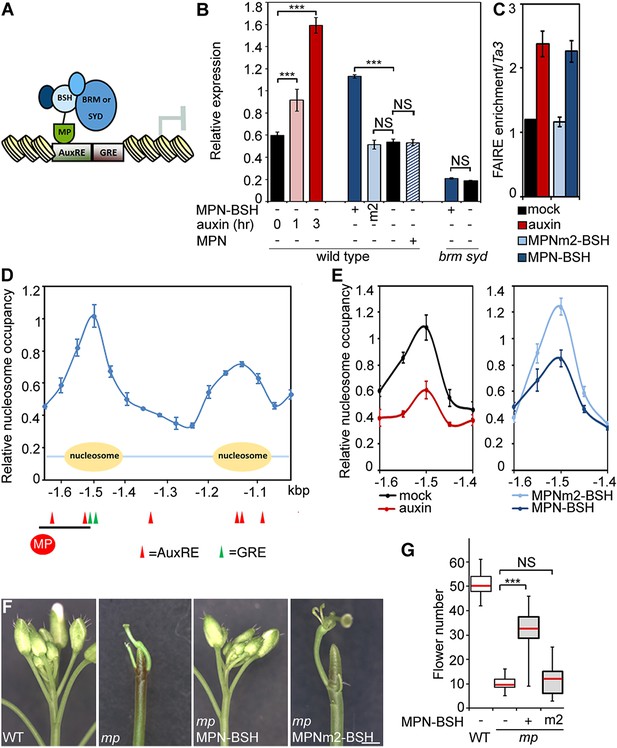
Tethering the BRM or SYD complex to MP target loci mimics MP function.
(A) Tethering of BRM or SYD-containing SWI/SNF complexes to the MP target loci. The shared BRM and SYD complex subunit BUSHY (BSH) (Han et al., 2015) is translationally fused to the MP DNA binding domain (MPN-BSH). (B) Transcriptional activation of the FIL locus by auxin treatment or BRM/SYD tethering via MPN-BSH in isolated plant cells. MPNm1-BSH carries a mutation (G279E; Figure 6—figure supplement 1) that blocks MP dimerization (Boer et al., 2014). MPNm2-BSH carries a second mutation (R215A) that causes loss of DNA binding specificity. Controls: MPN, mock treatment or no plasmid. n > 3. p-value: Student's t-test. (C) DNA accessibility at the FIL locus in response to auxin treatment or BRM/SYD tethering assayed by FAIRE in isolated plant cells. (D) Nucleosome positioning at the FIL locus. Top: MNase digestion followed by tiled oligo qPCR (MNase-qPCR) to monitor nucleosome positioning at the FIL promoter in 3-week-old plants. X-axis: distance from the start codon. Middle: diagram of nucleosome positions. Bottom: red circle: MP protein. Red triangles: core MP binding sites (AuxREs) (Ulmasov et al., 1997; Boer et al., 2014). Black line: region probed in all ChIP or FAIRE experiments (FILb in Figure 2—figure supplement 4). Green triangle: Evolutionarily conserved cis elements. (E) Nucleosome occupancy at the FIL locus in response to auxin treatment or BRM/SYD tethering via MPN-BSH in isolated plant cells by MNase-qPCR. X-axis: distance from the start codon. Figure 6—figure supplement 1 shows the nucleosome occupancy in brm syd mutant plant cells in response to auxin treatment. (F, G) Rescue of mp-S319 flower primordium initiation defect by tethering of BRM or SYD complexes to MP binding sites. Figure 6—figure supplement 2 shows the effect of additional rescue constructs on flower initiation in mp-S319 mutants. Figure 6—figure supplement 3 shows rescue of mp-S319 mutant leaf developmental defects. (F) Representative inflorescence images. Scale bars = 1 mm. (G) Quantification of flower primordium initiation. n > 18. Grey shading: T1 population of transgenic plants. p-value: Mann–Whitney U test.
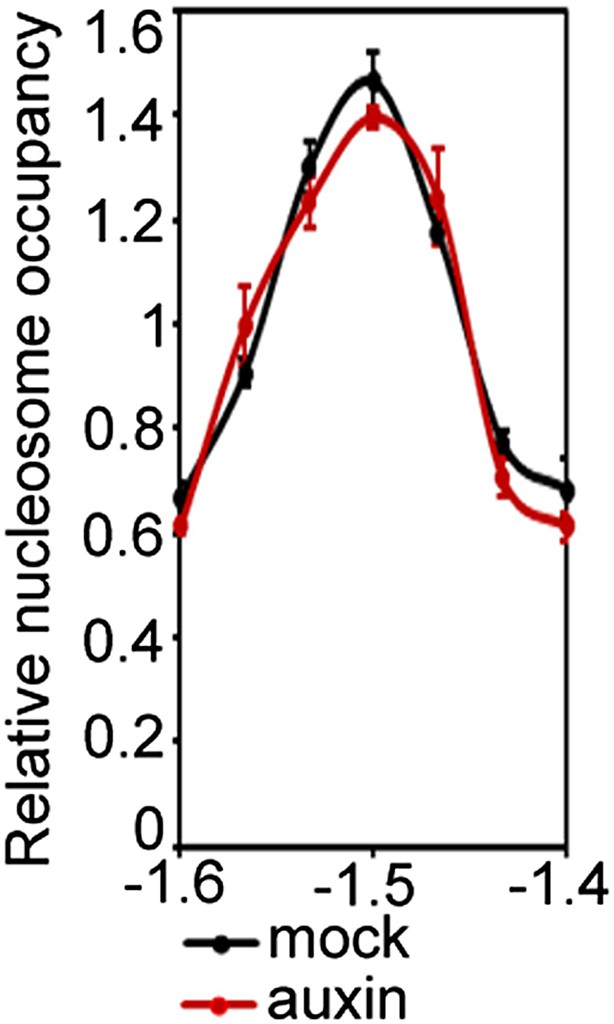
Auxin treatment fails to destabilize the well-positioned nucleosome at the FIL locus in syd brm mutant plant cells.
https://doi.org/10.7554/eLife.09269.018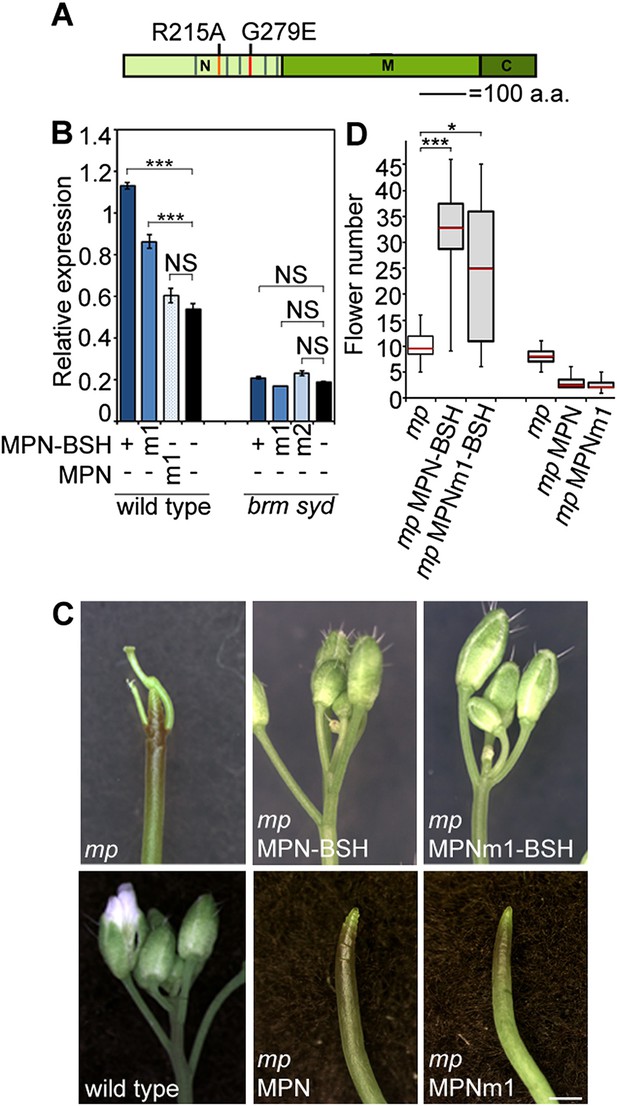
Rescue of mp-S319 by tethering the BRM or SYD complex to MP target loci.
(A) Diagram MONOPTEROS domains. N: N-terminal DNA binding/dimerization domain. M: middle region. C: C-terminal Aux/IAA interaction domain. Red lines: Amino acid mutations that disrupt DNA binding (R215A) and dimerization (G279E). Grey lines: nuclear localization signals. (B) Effect of dimerization deficient MPN-BSH [MPNm1-BSH (G279E)] or of MPN [MPNm1 (G279E)] on FIL expression. (C, D) Effect of MPNm1-BSH (G279E) or of MPN and MPNm1 on flower initiation in the mp-S319 mutant. (C) Representative images. (D) Quantification. n > 15. Grey shading: T1 population of transgenic plants. p-value: Mann–Whitney U test.
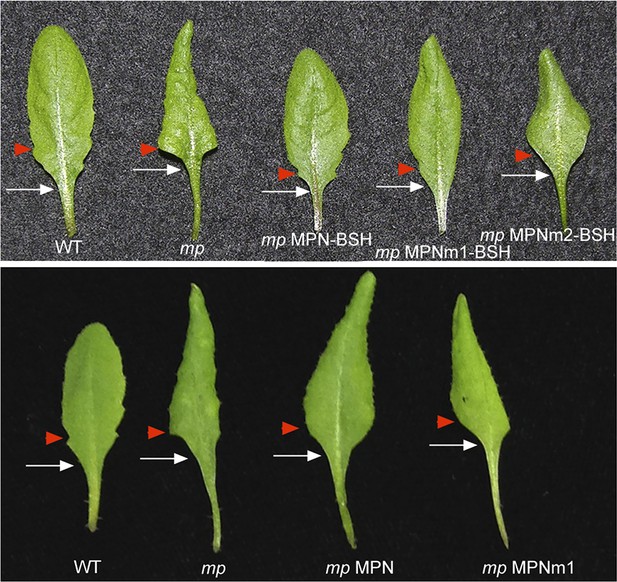

MPN-BSH and MPNm1-BSH rescue mp-S319 mutant leaf phenotypes.
mp-S319 MPN-BSH and mp-S319 MPNm1-BSH rescue the loss in leaf serration (red arrowhead) and elongated petiole (white arrow) phenotypes of mp-S319 mutants. mp-S319 MPNm2-BSH, which cannot bind DNA, displays similar defects as mp-S319 mutants. Likewise, MPN or MPNm1 alone, which cannot recruit BRM/SYD do not rescue the leaf shape defects of the mp mutants.
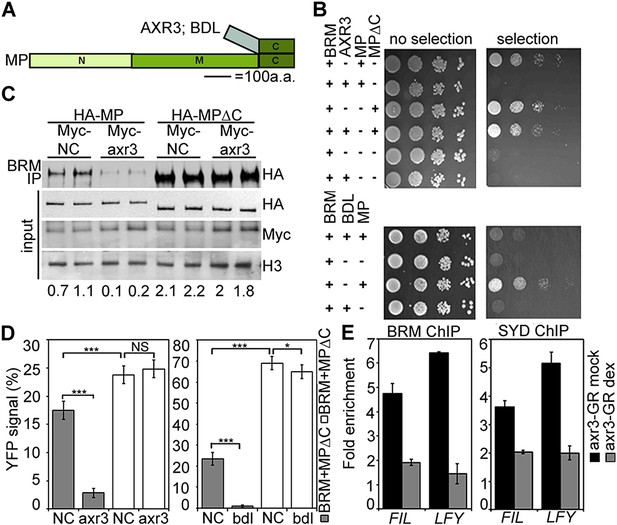
Aux/IAA proteins prevent BRM and SYD recruitment by MP.
(A) Diagram of MP domains. N: N-terminal DNA binding/dimerization domain, M: middle BRM/SYD interacting region, C: C-terminal Aux/IAA interacting domain. (B) Yeast-three-hybrid test of BRM interaction with MP or MP lacking the C-terminal domain (MP∆C) in the presence of the Aux/IAA protein AXR3 (top) or BDL (bottom). Growth was assayed with (right) or without (left) 3-amino-1,2,4-triazole. (C) Co-immunoprecipitation of FLAG-BRM with HA-MP or HA-MP∆C in the presence of the stabilized Myc-axr3. NC: Myc-tagged unrelated protein of similar molecular mass as axr3. Below: Amount of precipitated HA-MP/HA-MP∆C (% input). (D) Quantification of BiFC test of interaction between BRM and MP or BRM and MP∆C in the presence of axr3 (left) or bdl (right) compared to a NC protein. The error bars are proportional to the standard error of the pooled percentage computed using binomial distribution. n = 3. p-value: Mann–Whitney U test. (E) ChIP to assess BRM and SYD association with MP target gene loci before (mock) or after (dex) nuclear entry of axr3-GR. Shown is fold-enrichment relative to a control locus (Ta3 retrotransposon).
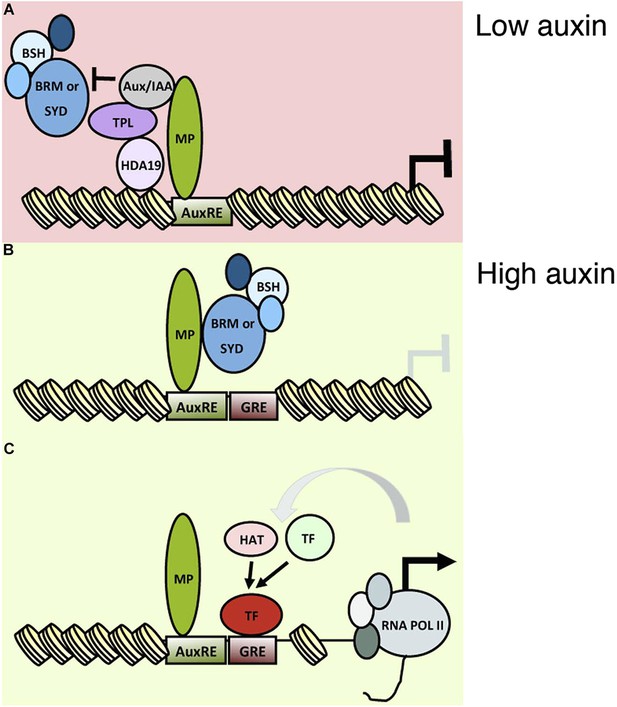
An auxin triggered chromatin state switch.
(A) In conditions of low auxin, Aux/IAA proteins bind to MP transcription factor associated with target loci and prevent gene expression in two ways: by recruiting the co-repressor TOPLESS (TPL) and histone deacetylase HDA19 and by preventing recruitment of the BRM or SYD chromatin remodeling complexes. (B) Upon establishment of a local auxin maximum, Aux/IAA proteins are degraded, this leads to eviction of HDA19 and TPL. Aux/IAA degradation also frees MP to recruit BRM or SYD complexes. The chromatin remodeling complexes open up the compacted chromatin by reducing nucleosome occupancy, thus increasing the accessibility of the genomic DNA near MP bound sites. (C) The ‘chromatin unlocking’ allows additional transcription factors access to their cis elements. This, possibly via additional steps, leads to recruitment of the general transcriptional machinery and initiation of transcription. HAT: histone acetyl transferase. GRE: binding site for transcription factor (TF). Figure 8—figure supplement 1 shows evolutionarily conserved cis elements near the midpoint of the well-positioned nucleosome at the FIL locus.

Two evolutionarily conserved cis elements close to the midpoint of the well-positioned nucleosome at the FIL locus.
Magenta: nucleosome midpoint; turquoise: a bZIP transcription factor bound G-box Related Element (GRE) (Berendzen et al., 2012; Weiste and Dröge-Laser, 2014); green: conserved cis element of unknown function.

MP message abundance relative to that of EIF4A.
MP abundance is similar in mp-S319 and mp-S319 transformed with MPN or MPNm1.
Additional files
-
Supplementary file 1
Primers used in this study.
- https://doi.org/10.7554/eLife.09269.024





