Evolution of an ancient protein function involved in organized multicellularity in animals
Figures
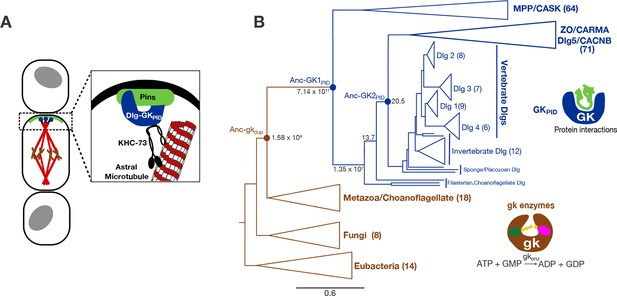
Function and phylogeny of the guanylate kinase (gk) and GKPID protein family.
(A) The GKPID of the protein Discs-large (Dlg, blue) serves as a scaffold for spindle orientation by physically linking the localized cortical protein Pins (green) to astral microtubules (red) via the motor protein KHC-73 (black). (B) Reduced phylogeny of the protein family containing gk enzymes (brown) and protein-binding GKPIDs (blue). Parentheses show the number of sequences in each clade. Reconstructed proteins Anc-gkdup (the preduplication ancestor of gk enzymes and GKPIDs in animals/choanoflagellates), Anc-GK1PID and Anc-GK2PID (the GKPID in the common ancestor of animals and choanoflagellates, and of animals, respectively) are marked as circles with approximate likelihood ratio support. Scale bar indicates number of substitutions per site. For unreduced phylogeny, see Figure 1—figure supplement 1. Characteristics of the reconstructed sequences are found in Figure 1—figure supplement 2. For sequences analyzed, see Figure 1—source data 1. For sequences and posterior probabilities of amino acid states, see Figure 1—source data 2, Figure 1—source data 3.
-
Figure 1—source data 1
Species and identifiers for sequences used in alignment and phylogenetic analysis.
- https://doi.org/10.7554/eLife.10147.004
-
Figure 1—source data 2
Posterior probability distribution of ancestral states for Ancgkdup.
- https://doi.org/10.7554/eLife.10147.005
-
Figure 1—source data 3
Posterior probability distribution of ancestral states for AncGK1PID.
- https://doi.org/10.7554/eLife.10147.006
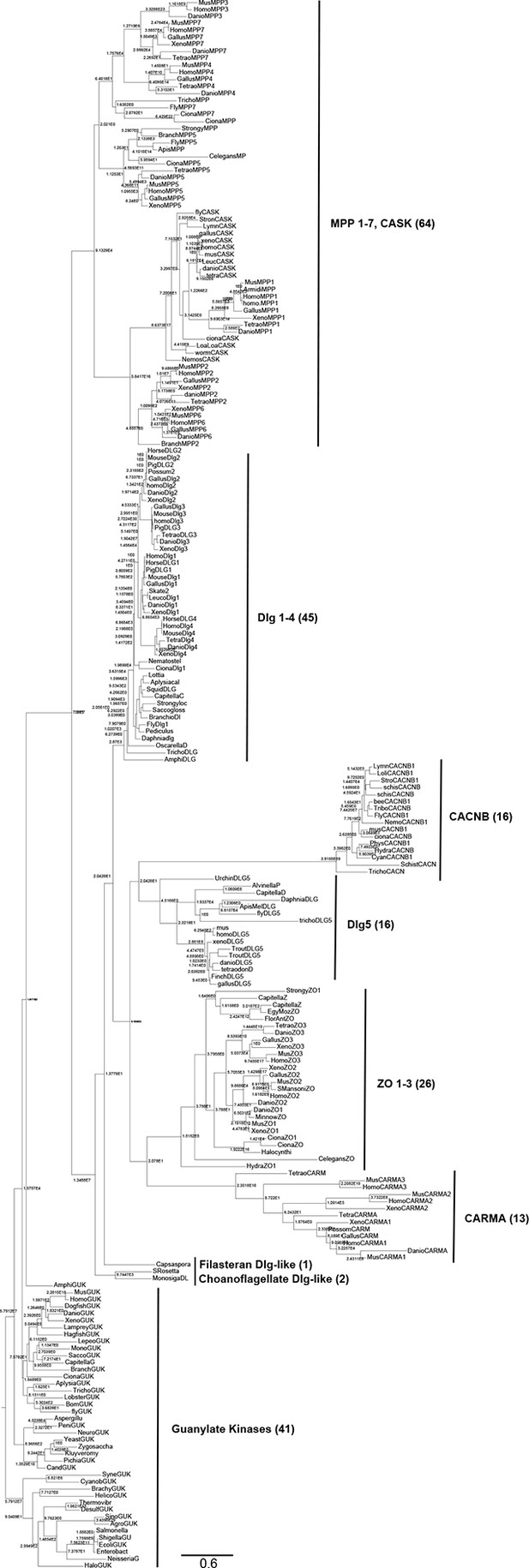
Complete phylogeny of 224 guanylate kinase enzyme and GKPIDs.
Nodes are labeled with approximate likelihood ratio supports, and branch lengths are in substitutions per site (see scale bar). The tree is rooted on the bacterial GK enzymes. Major paralogs in the GKPID family, with the number of sequences included in each, are labeled. See Figure 1—source data 1 for species and accessions used. See Supplementary files 1 and 2 for alignment and tree file.
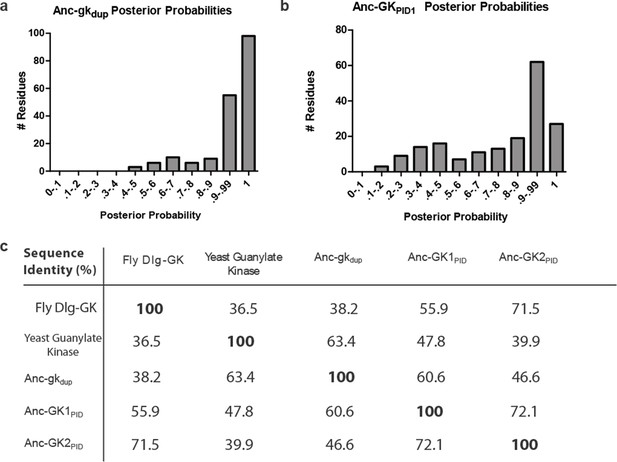
Sequence characteristics of maximum likelihood reconstructions of Anc-gkdup and Anc-GK1PID.
(A,B) The histogram shows the distribution over sites of posterior probability support for maximum likelihood amino acid states (see Figure 1—source data 2, Figure 1—source data 3 for full sequences and support). (C) Sequence similarity of ancestral sequences to extant gk enzymes and GKPIDs. The table shows the percent of residues identical between each pair of sequences.
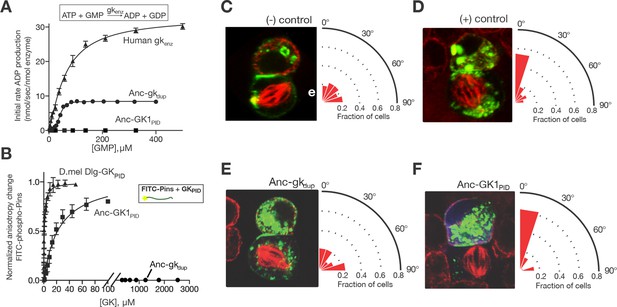
Evolution of a novel spindle-orientation function in ancestral GKPID.
(A) Anc-gkdup (circles) is an active nucleotide kinase in a coupled enzyme assay for the reaction shown; Anc-GK1PID (boxes) is inactive. Activity of the human gk enzyme (triangles) is shown for reference. Error bars show SEM for three replicates. (B) The more recent ancestral protein Anc-GK1PID (boxes) binds a 20 amino-acid peptide (see methods) from the Pins protein in a fluorescent anisotropy assay, but Anc-gkdup (cirlcles) does not. Pins binding by the GKPID of the Drosophila melanogaster Dlg protein (triangles) is shown for reference. Error bars show SEM for three replicates. (C–F) Evolution of spindle orientation function as assayed in cultured S2 cells that do not express endogenous Dlg protein. Cells were transfected with a GK construct (C, –control: empty transfection vector; D, + control: GKPID from extant Drosophila Dlg) and scored for alignment of the mitotic spindle (red, tubulin, visualized immunocytochemically) relative to the Pins cortical crescent (green, a GFP-tagged Pins-Ecd fusion). In the example images for each experiment, two cells are shown, the bottom one of which is dividing. The angle of the mitotic spindle relative to a line bisecting the Pins crescent (from 0°, precisely aligned, to 90°) was recorded in many dividing cells; the radial histogram (right) shows the distribution of observed angles among all cells scored with a given genotype. Cells transfected with Anc-gkdup (E) do not display robust spindle orientation, but those transfected with Anc-GK1PID do (F). SEM: Standard error of the mean.
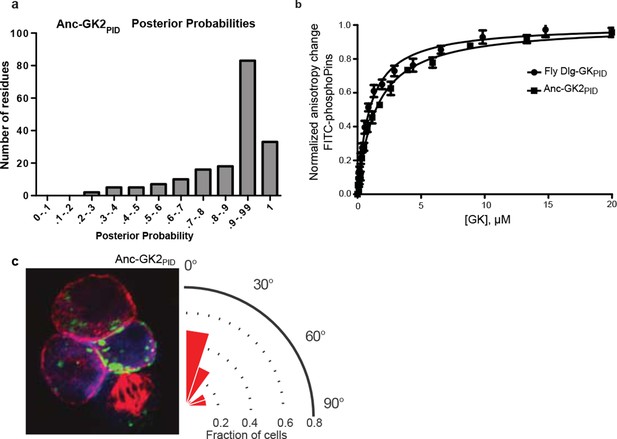
Properties of ancestral protein AncGK2-PID.
(A) Distribution over sites of posterior probabilities of ML amino acid states in AncGK2PID . Mean posterior probability = 0.87. (B) Anc-GK2PID(squares) binds the Pins peptide ligand with high affinity in a fluorescence anisotropy assay. Binding by the Drosophila melanogaster Dlg GKPID is shown for comparison (circles). Error bars who standard error of the mean of three replicates. (C) Anc-GK2PID is capable of orienting the mitotic spindle.
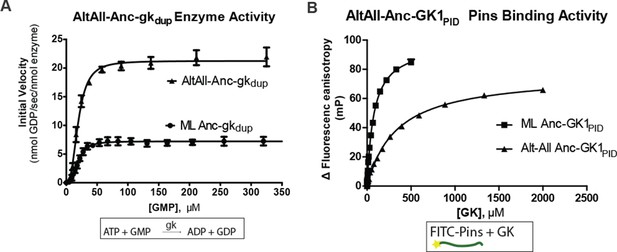
Robustness of functional inferences about ancestral proteins to uncertainty about the sequence reconstruction.
For each of Anc-gkdup and Anc-GK1PID, an alternate reconstruction (Alt-All) was synthesized, containing the next-best amino acid state at all sites with multiple plausible states ((PP>0.2). The enzyme activity of Anc-gkdup in a coupled enzymatic assay for cofactor turnover (panel A) and the Pins-binding activity of Anc_GK1PID (panel B) in a fluorescence anisotropy assay are shown for both maximum likelihood and Alt-All reconstructions. Affinities and maximal velocities differ quantitatively, but the presence/absence of each property is robust to incorporation of uncertainty about the ancestral sequence (see Figure 1F).
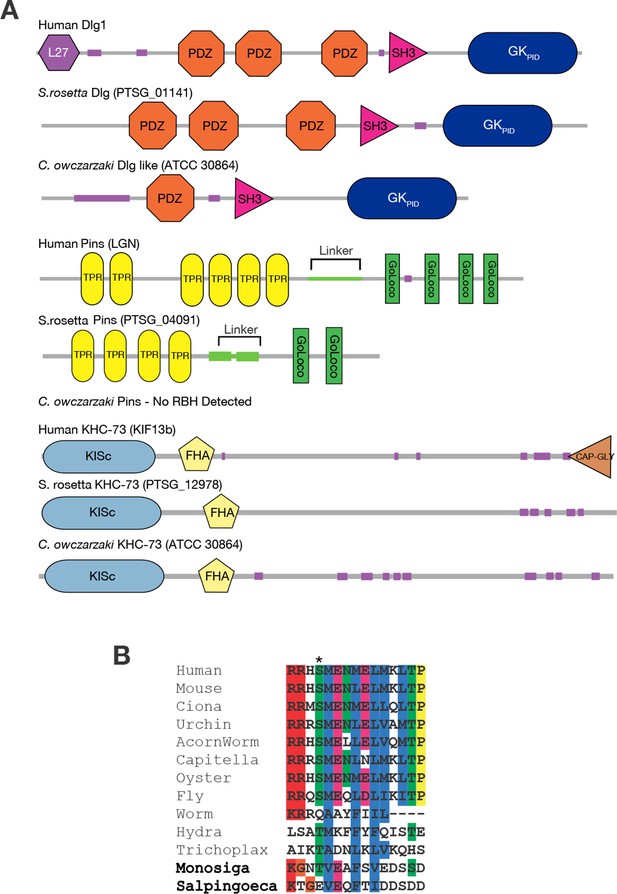
Taxonomic distribution of proteins in the spindle orientation complex.
(A) The genomes of the choanoflagellate Salpingoeca rosetta and the filasterean Capsaspora owczarzaki contain orthologs of human Dlg, Pins, and KHC-73. The domain architecture of each protein is shown, as inferred using the SMART database. Each group of proteins are reciprocal best BLAST hits (RBH) to the human query protein shown. For details, see Figure 3—source data 1. (B) Aligned sequences from the linker portion of Pins (see panel A), which binds to Dlg. Colors highlight identical or biochemically conservative residues. Asterisk, phosphorylated or negatively charged residue 436, which in the Drosophila melanogaster Pins protein anchors Dlg binding. For complete species names and accessions, see Figure 3—source data 2.
-
Figure 3—source data 1
Results of reciprocal Blast search of metazoan and nonmetazoan genomes for orthologs of GKPID, Pins and KHC-73.
Each row shows, for a query sequence from Drosophila melanogaster, the accession and e-score for the best-hit sequence in the target species, as well as the reciprocal best-hit and e-score when the target best-hit is used as a query against D. melanogaster.
- https://doi.org/10.7554/eLife.10147.013
-
Figure 3—source data 2
Identifiers and species for Pins sequences shown in Figure 3B.
- https://doi.org/10.7554/eLife.10147.014
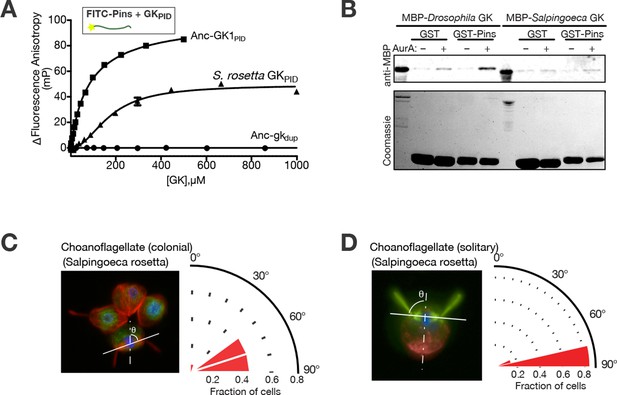
Spindle orientation and its molecular components in choanoflagellates.
(A) Purified GKPID from the Salpingoeca rosetta Dlg ortholog (triangles) binds fluorescently labeled Drosophila melanogaster Pins. Binding by Anc-GK1PID (squares) and lack of binding by Anc-gkdup (circles) are shown for comparison. Error bars, SEM for three replicates. (B) Purified GKPID of S. rosetta does not interact with Pins of S. rosetta; under the same conditions, the D. melanogaster orthologs of these proteins do bind. A GST-fusion of each Pins linker regions (visible on the coomassie stained gel) was incubated with an MBP-fusion of each species’ GKPID and detected by anti-MBP western blot. As previously found, binding of D. melanogaster Pins requires phosphorylation by Aurora A kinase; S. rosetta GKPID does not bind its own Pins whether or not the kinase is present. (C, D) Spindle orientation in colonial ( panel C, n = 12) and noncolonial (panel D, n = 7) S. rosetta. For each condition, the image shows one representative colony; the cell at bottom center is mitotic, as evidenced by condensed DNA (blue, DAPI) without a defined nuclear envelope (green, visualized using anti-nuclear pore complex). Red, mitotic spindle visualized using anti-tubulin. In colonial cells, the angle of the mitotic spindle (solid white line) was measured relative to a line perpendicular to the plane of the colony extending through the colony’s center (dashed line). The histogram shows the distribution of spindle angles among all dividing S. rosetta cells measured, with 90° representing perfect alignment relative to the colony ring. In noncolonial cells, the spindle angle was measured relative to a line extending from the midpoint between the bases of the two duplicate flagella to the cell body’s centroid (dotted line). An angle of 90° represents perfect alignment relative to the flagella. SEM: Standard error of the mean.
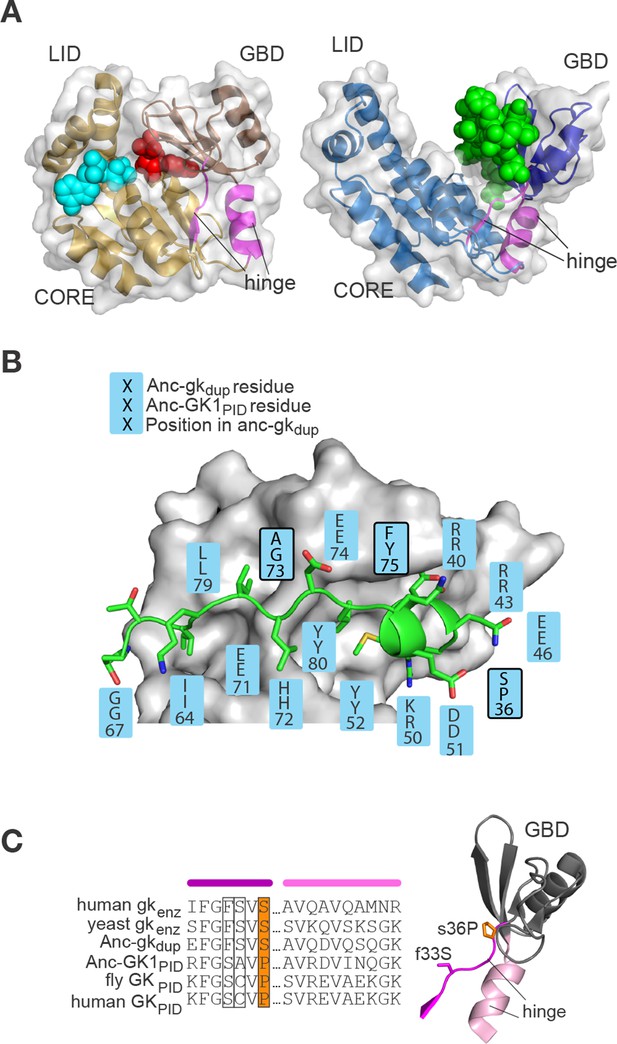
Evolution of the binding interface and hinge during the evolution of AncGKPID spindle orientation functions.
(A) All gk/GKPID family members share a common structural architecture, comprising a catalytic core, two binding lobes (the GMP-binding domain, GBD, shown in dark hue, and the ATP-binding lid), and a flexible hinge region, which connects the GBD to the core and comprises two segments of contiguous residues (magenta). Left: in gk enzymes bound to GMP (red spheres), the lobes adopt a closed conformation, bringing GMP and ATP (cyan spheres) adjacent to each other in the core. Right: the GKPID has an open conformation, in which Pins (green spheres) binds to the surface of the GBD in the cleft between the two binding lobes. Structures shown are mouse gk enzyme (brown, PDB 1LVG) and the GKPID from rat Dlg1 (blue, 3UAT). (B) Most residues in Anc-GK1PID that bind Pins (blue boxes) are unchanged from the ancestral state in Anc-gkdup. White surface, D. melanogaster Dlg GK1PID (3TVT). Green, Pins peptide. Ancestral and derived amino acid states are shown; residues with historical amino acid replacements between the two ancestral proteins are outlined. (C) In the hinge region, two historical substitutions (outlined and colored, with side-chains shown as sticks) were conserved in the ancestral state in extant enzymes and in a different state in extant GKPIDs. Colored bars above the sequence indicate position in the protein structure (right). Hinge segments are shown in pink and the GMP-binding lobe in gray.
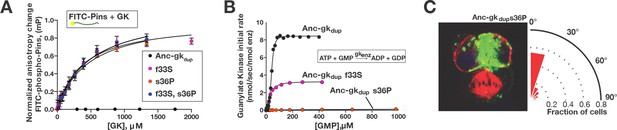

A simple genetic basis for the evolution of spindle orientation functions in the ancestral GKPID.
(A,B) Introducing either of two historical substitutions s36P or f33S into Anc-gkdup confers Pins binding and reduces ancestral guanylate kinase activity. Error bars are SEM of 3 replicates. Combining s36P and f33S does not further affect the derived functions beyond the effects of the single substitutions. (C) Introducing s36P into Anc-gkdup is sufficient to confer the full capacity to drive orientation of the mitotic spindle (compare Figure 1). As shown in Figure 6—figure supplement 1, historical substitutions in the binding interface do not recapitulate the evolution of Pins binding and the loss of gk activity.
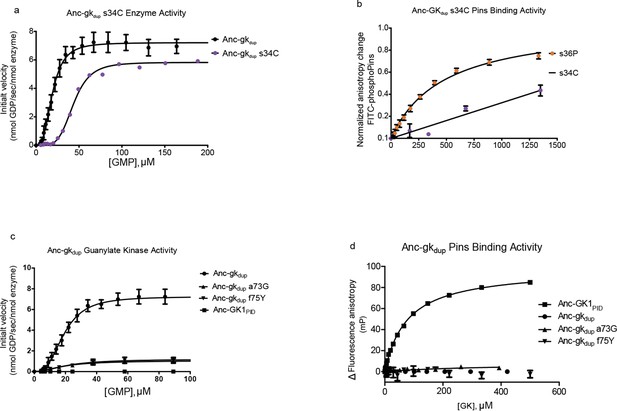
Introducing other historical substitutions into AncGKdu does not confer GKPID-like function.
(A) The hinge substitution s34C (purple) does not abolish guanylate kinase activity in a coupled enzyme activity assay; Anc-gkdup is shown for comparison. (B) s34C (purple) does not confer substantial Pins-binding affinity on Anc-gkdup. Pins-binding activity relative to s36P (orange) in a fluorescence anisotropy assay. (C,D) Anc-GK1PID Pins interface substitutions a73G and f75Y reduce gunaylate kinase enzyme activity (D) but do not confer Pins binding (E) when introduced into Anc-gkdup. Error bars are standard error of the mean with three replicates.
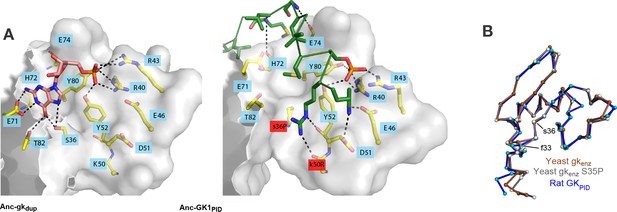
Evolution of GKPID’s new function by unveiling a latent protein-binding site.
(A) The binding surface for Pins in GKPIDs is derived from the GMP-binding surface of gk enzymes. Homology models of Anc-gkdup (left) and Anc-GK1PID (right) are shown as white surface, with all side chains that contact either GMP or Pins as yellow sticks. Pink sticks show GMP; green ribbon shows Pins backbone, with the side chains of all Pins residues that contact the GK protein shown as sticks. The phosphate group on GMP and on Pins residue 436 are shown as orange and red sticks. Black dotted lines, protein-ligand hydrogen bonds. In the AncGK1PID structure , substitutions at sites in the binding interface are shaded red, including key substitution s36P. The binding modes of extant gk enzymes and GKPIDs are similar and support the same conclusions (see Figure 7—figure supplement 1). (B) The structure of the hinge and GMP/Pins-binding lobes is conserved between the Pins-bound GKPID (blue, rat Dlg, 3UAT), the apo-gk enzyme (brown, S. cerevisiae guanylate kinase 1EX6), and the apo-gk-s36P mutant (gray, 4F4J), all in the open conformation.
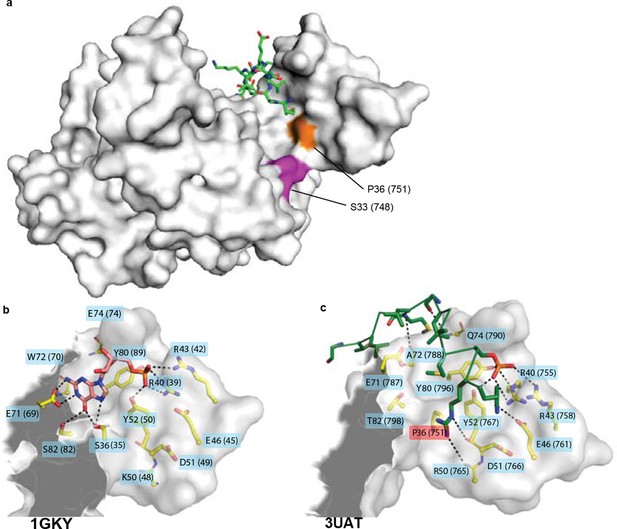
Structural context of key historical mutations.
(A) Location of historical hinge substitutions s36P and f33S. Residues Pro36 (orange) and Ser33 (purple) are shown on the crystal structure of the GKPID from rat Dlg (3UAT, white surface). Pins peptide ligand is shown in green. (B,C) Similarity of GMP binding site in extant guanylate kinase enzyme to Pins binding site in extant GKPID. The guanylate binding domains (GBDs) of yeast gk enzyme (PDB 1GKY, panel B) and of rat Dlg GKPID (PDB 3UAT, panel C) is shown as white surface, with all side chains that contact either GMP or Pins as yellow sticks. Pink sticks show GMP; green ribbon shows Pins backbone, with the side chains of all Pins residues that contact the GKPID protein shown as sticks. The phosphate group on GMP and on Pins residue S436 are indicated. Black dotted lines, protein-ligand hydrogen bonds. Key substitution s36P is highlighted in pink.
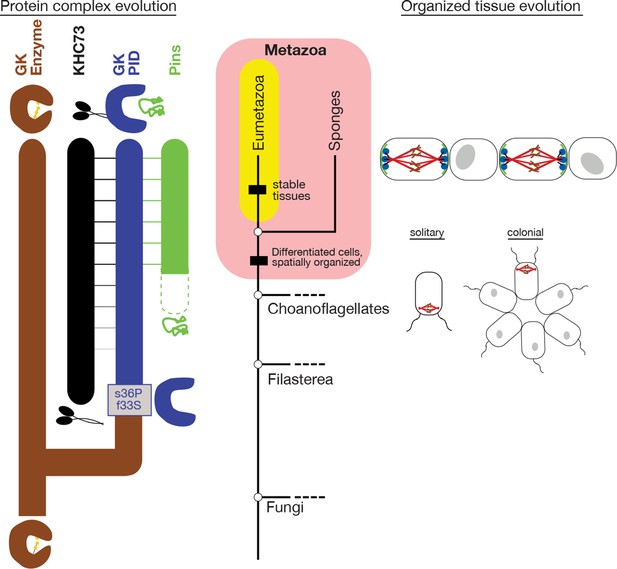
Historical evolution of GKPID-mediated spindle orientation complex.
The center portion shows the phylogeny of Metazoa and closely related taxa. The origin of cell differentiation and spatially organized tissues is marked. The left portion shows major events in the evolution of the components of the spindle orientation complex reconstructed in this study. Duplication of an ancestral gk enzyme (brown) and the key mutations that led to the origin of a GKPID (blue) that could bind other molecules in the complex are shown relative to the phylogeny’s time scale. The apparent date of origin of KHC-73 (black) and Pins (green) are also shown. Dotted green line shows the origin of Pins in a form not yet bound by GKPID. Solid green line shows GKPID-binding form. Horizontal lines indicate binding between proteins. The right portion shows a schematic of the spindle orientation machinery in metazoans, which allows orientation relative to external cues from nearby cells, as well as spindle orientation relative to the internal cell axis as marked by the flagella in both solitary and colonial choanoflagellates.
Additional files
-
Supplementary file 1
Alignment of GK domains used for phylogenetic analysis and reconstruction.
- https://doi.org/10.7554/eLife.10147.022
-
Supplementary file 2
Phylogeny of GK domains in Newick format.
- https://doi.org/10.7554/eLife.10147.023





