Egr-5 is a post-mitotic regulator of planarian epidermal differentiation
Figures
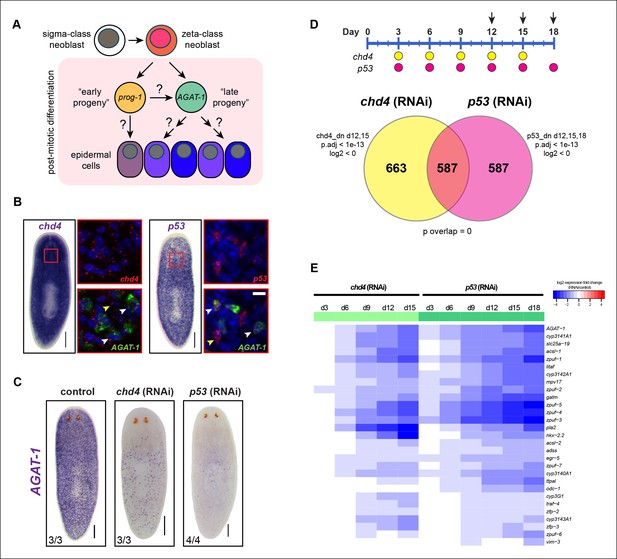
Identification of a common transcriptional down-regulated gene set in chd4 and p53 RNAi animals.
(A) Current model of planarian epidermal lineage specification. Sigma-class neoblasts give rise to zeta-class neoblasts, which in turn generate prog-1 (early progeny) and AGAT-1 (late progeny) expressing cells. The precise molecular relationship between these cell types remains unclear, but collectively give rise to an unknown number of epidermal cell types through an unknown number of transitional states. (B) Whole-mount in situ (WISH) expression patterns of chd4 and p53 in wild-type planaria. Left panels: chd4 colorimetric WISH and double fluorescent in situ (FISH) of chd4 (red), AGAT-1 (green) and DAPI (blue). Right panels: p53 colorimetric WISH and double FISH of p53 (red), AGAT-1 (green) and DAPI (blue). Magnified regions are single confocal planes from boxed regions. White arrowheads highlight cells with co-localized expression of either chd4 or p53 and AGAT-1. Yellow arrowheads highlight additional mesenchymal cells that do not express AGAT-1. Scale bars: 200 μm; 10 μm (zoomed images). (C) chd4(RNAi) and p53(RNAi) result in the loss of AGAT-1 expressing cells. Representative colorimetric WISH images shown at 3Fd18 of RNAi treatment. Scale bar: 200 μm. (D) Venn diagram of genes down-regulated in chd4 and p53 RNAi data sets. Timeline of RNAi treatment (Fed d0, d3, d6) and RNA collected for chd4 (yellow circles) and p53 RNAi (pink circles). Arrows highlight time points used to identify down-regulated gene set. Criteria used for genes to make the cut-off are shown. dn, down-regulated genes; p.adj, adjusted p-value; log2, fold change of RNAi over control (see Materials and methods). For the chd4 and p53 RNAi overlapping data set (587 genes), a hypergeometric distribution and a universe size of 28,668 was used to generate p-value for determining significance of overlap by chance. See also Supplementary file 1. (E) Heat map depicting candidate genes that were selected from the chd4(RNAi) and p53(RNAi) down-regulated data sets for further characterization after in situ hybridization screen. log2 fold changes in RNAi expression relative to each control time point are shown.
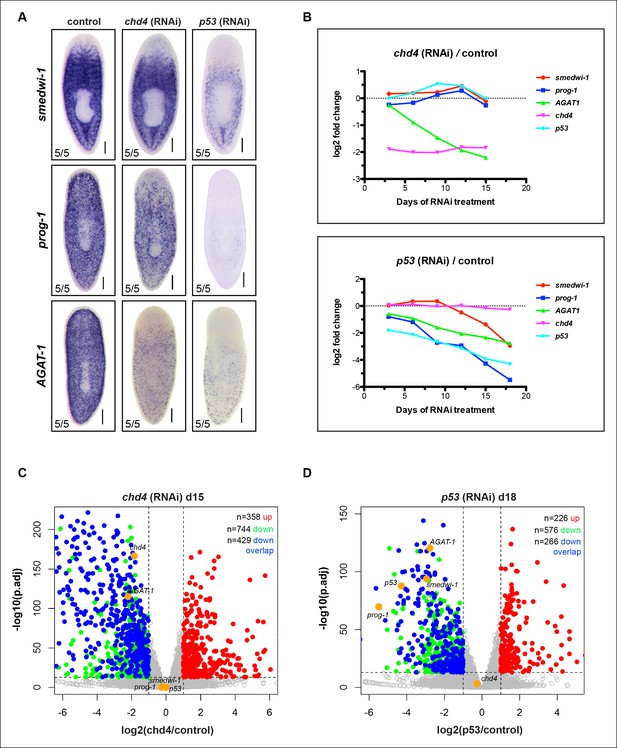
RNA-seq validation of control genes in chd4 and p53 RNAi animals.
(A) Colorimetric WISH was performed on control, chd4 and p53 RNAi animals for smedwi-1 (stem cell marker), prog-1 (early progeny) and AGAT-1 (late progeny) from the same cohort of worms used for RNA-seq analysis. Representative images of 3Fd15 RNAi worms are shown. Scale bar: 200 μm. (B) Plot of log2 ratios of control genes (smedwi-1, prog-1, AGAT-1) and targeted RNAi genes (chd4 and p53) in chd4(RNAi) (top panel) and p53(RNAi) (bottom panel) time course. For each time point, reads mapping to gene models were counted in chd4(RNAi) and p53(RNAi) data sets and were then compared to the unc22(RNAi) control, and log2 ratios were generated. (C-D) Volcano plots illustrating genes significantly down-regulated (green dots; adjusted p-value <1e-13, log2 <-1) and up-regulated (red dots; adjusted p-value <1e-13, log2 >1) at 3Fd15 in chd4(RNAi) data set (C) and at d18 in p53(RNAi) data set (D). Control genes are highlighted in yellow. Blue dots represent genes from the 587 gene overlap data set (Figure 1D) that are also significantly down-regulated in the plotted time points. See also Supplementary file 1. See Supplementary file 2 for up-regulated gene list.
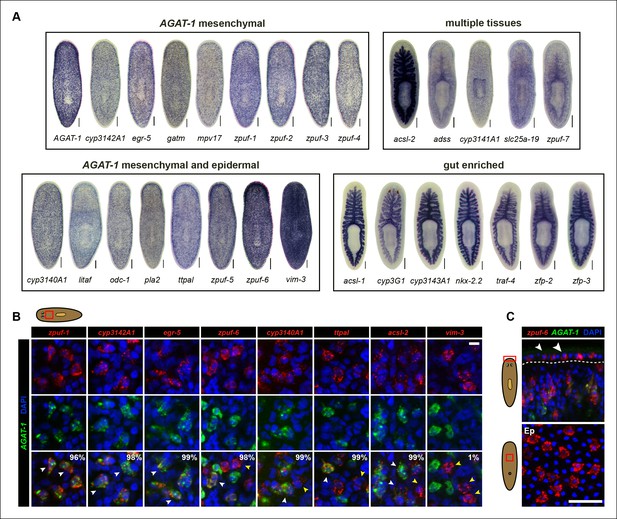
Expression patterns of candidate genes from chd4(RNAi) and p53(RNAi) data sets.
(A) WISH expression of representative candidate gene set from Figure 1E and grouped by expression pattern (see text for details). Scale bars: 200 μm. See also Supplementary file 3. (B) Expression pattern of various candidate genes in (A), analyzed by double FISH with AGAT-1. Images represent single confocal planes from anterior regions. Percentages represent fraction of AGAT-1+ cells that co-express the candidate gene (∼200-400 cells were quantified; low but detectable expression was counted as co-localized). White arrowheads highlight co-localization; yellow arrowheads highlight additional cells that have no detectable AGAT-1 expression (AGAT-1neg). Scale bar: 10 μm. (C) Double FISH of zpuf-6 and AGAT-1 highlighting zpuf-6 expression in discrete cells in the planarian epidermis (Top panel, white arrowheads). Bottom panel, ventral epidermal (Ep) view. DapI used for nuclear staining. Scale bar: 50 μm.
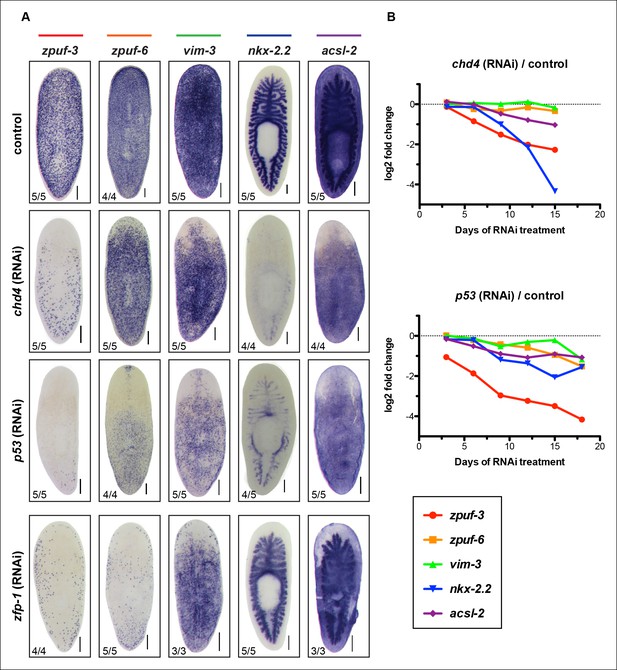
Validation of select candidate genes in chd4, p53 and zfp-1 RNAi conditions by WISH.
(A) Representative WISH of candidate genes encompassing the full range of expression patterns in control, chd4 and p53 RNAi animals fixed at 3Fd21 (For nkx-2.2, 3Fd18). Genes expressed in mesenchyme, epidermis and gut are reduced in chd4 and p53 RNAi animals. Genes expressed in mesenchyme and epidermis are also reduced in zfp-1(RNAi) animals (3Fd21) but gut expression is not affected. Scale bar: 200 μm. (B) Plot of log2 ratios of candidate genes from (A) in chd4(RNAi) (top panel) and p53(RNAi) (bottom panel) RNA-seq time course.
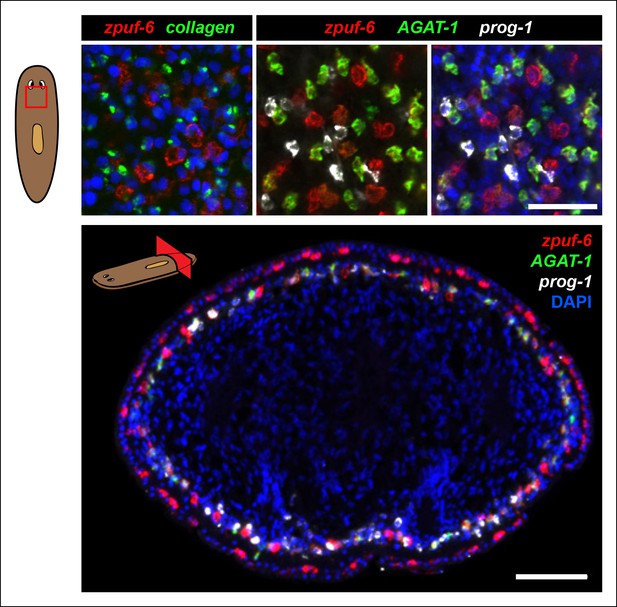
Detailed characterization of zpuf-6 expression pattern by whole-mount FISH.
Zpuf-6 is not expressed in muscle cells or in prog-1 cells. Top left panel: Double FISH of zpuf-6 and a muscle marker (collagen). Top right panels: Triple FISH of zpuf-6, AGAT-1 and prog-1. Bottom panel: transverse tissue section of triple FISH highlighting the spatial distribution of zpuf-6, AGAT-1 and prog-1. zpuf-6 is expressed in both sup-epidermal and epidermal layers. All images shown are single confocal planes. Scale bars: 50 μm.
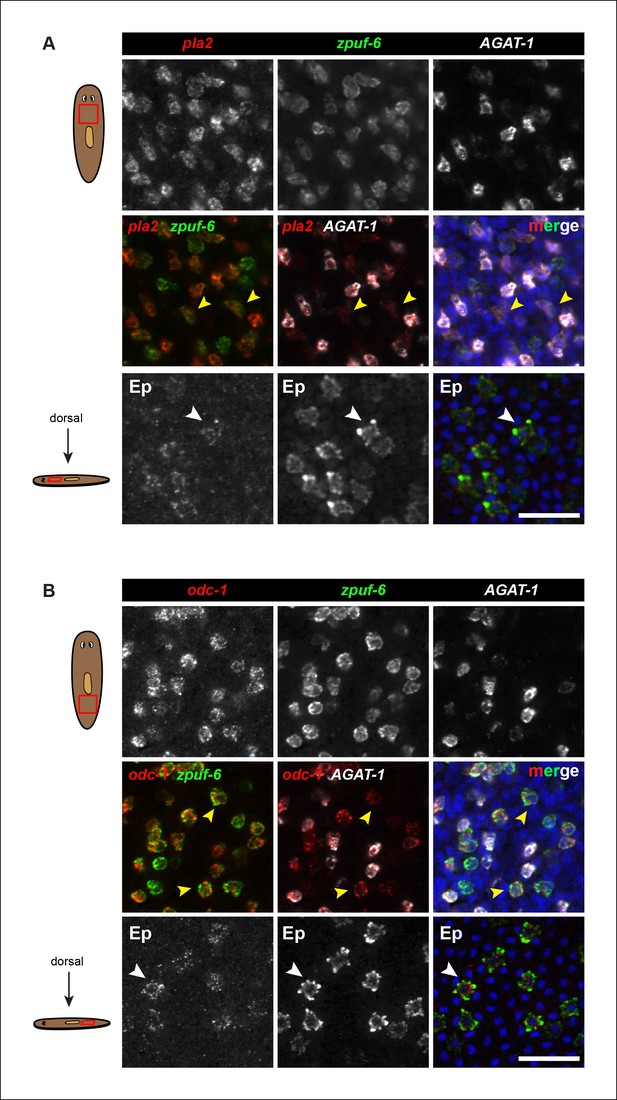
Expression of pla2 and odc-1 in mesenchyme and epidermis.
(A) Triple FISH of pla2, zpuf-6 and AGAT-1 in the mesenchyme. Top row: single channel of each individual gene. Middle row: Left panel, double FISH of pla2 and zpuf-6. Middle panel, double FISH of pla2 and AGAT-1. Right panel, merge. Yellow arrowheads highlight cells that co-express pla2 and zpuf-6 and are AGAT-1neg. Bottom row: Double FISH of pla2 and zpuf-6 in the epidermis (Ep). White arrowheads highlight co-localization. Scale bars: 50 μm. (B) Triple FISH of odc-1, zpuf-6 and AGAT-1 in the mesenchyme. Top row: single channel of each individual gene. Middle row: Left panel, double FISH of odc-1 and zpuf-6. Middle panel, double FISH of odc-1 and AGAT-1. Right panel, merge. Yellow arrowheads highlight cells that co-express odc-1 and zpuf-6 and are AGAT-1neg. Bottom row: Double FISH of odc-1 and zpuf-6 in the epidermis. White arrowheads highlight co-localization. Scale bars: 50 μm.
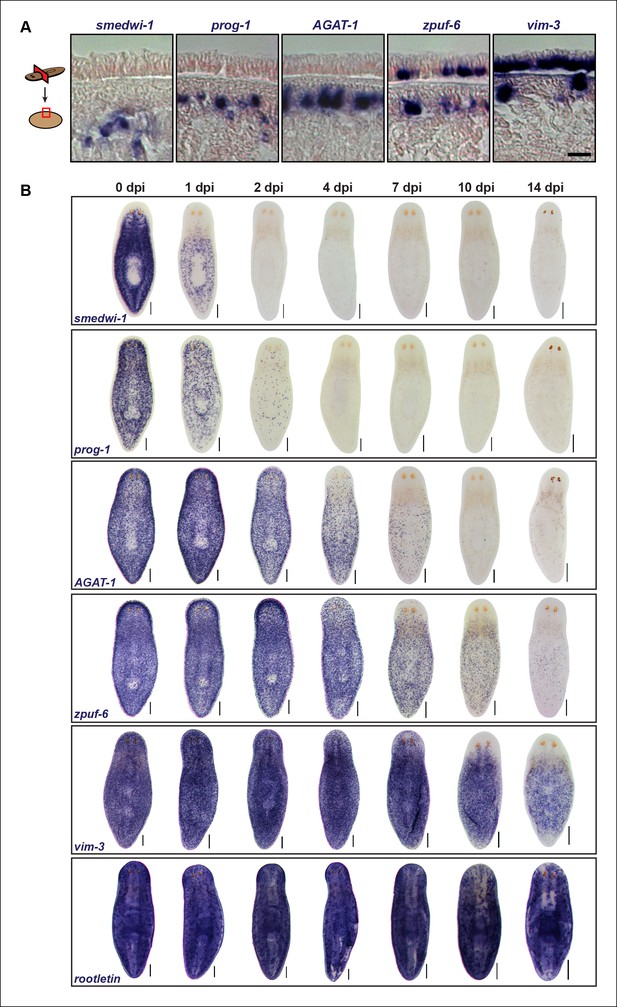
Epidermal progeny markers are expressed in distinct spatiotemporal domains.
(A) Transverse tissue sections of colorimetric WISH-stained animals expressing putative markers in the epidermal lineage. Sections were counterstained with nuclear fast red. Scale bar: 10 μm. (B) Colorimetric WISH of animals after 6000 Rad irradiation exposure for markers of neoblasts (smedwi-1), early progeny (prog-1), late progeny (AGAT-1), and other genes identified in this study (zpuf-6 and vim-3) marking transitions into the epidermis. Rootletin marks differentiated cells of the ciliated epidermis as well as protonephridia. Expression patterns of epidermal progeny markers are lost in a ventral-to-dorsal, anterior-to-posterior manner. A representative image from 4–6 worms for each gene and time point are shown. dpi, days post-irradiation. Dorsal views. See Figure 3—figure supplement 1 for close-up ventral views. Scale bars: 200 μm.
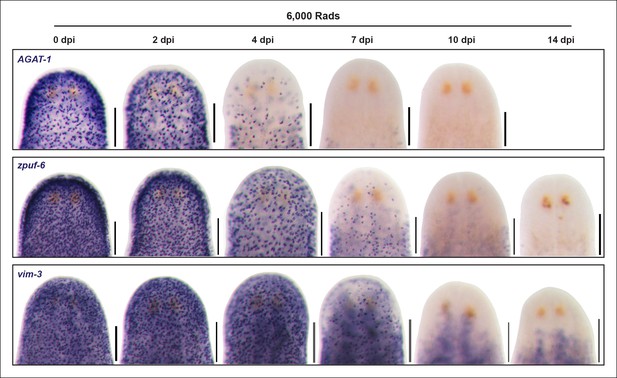
Ventral plane sections of epidermal progeny markers after irradiation.
Colorimetric WISH of animals after 6,000 Rad irradiation exposure for markers of AGAT-1, zpuf-6 and vim-3, highlighting loss of ventral expression prior to loss of dorsal expression. A representative image from 4-6 worms for each gene and time point are shown. dpi, days post-irradiation. Scale bars: 200 μm.
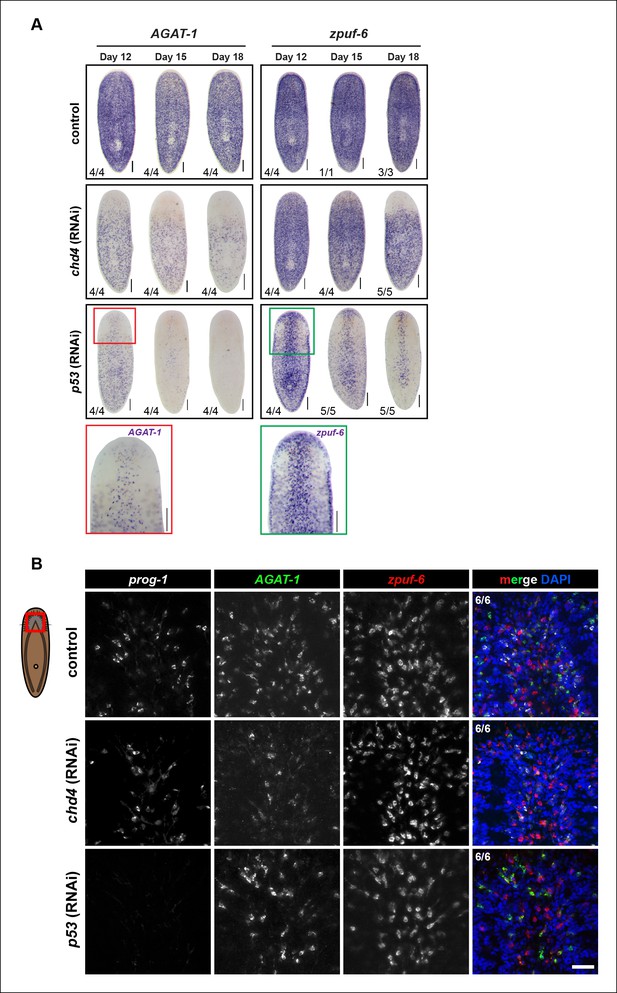
Correlated spatial expression patterns of prog-1, AGAT-1 and zpuf-6.
(A) Expression of AGAT-1 and zpuf-6 after chd4 and p53 RNAi knockdowns. Colorimetric WISH staining is lost in a ventral-to-dorsal, anterior-to-posterior fashion. Expression of markers in p53(RNAi) animals display a striped pattern of cells (zoom). Ventral views. Scale bars: 200 μm. (B) Triple FISH of prog-1, AGAT-1 and zpuf-6 at 3Fd12 in control, chd4 and p53 RNAi knockdown conditions. The spatial patterns of remaining AGAT-1 and zpuf-6 cells are highly correlated. Images shown are single confocal planes of the ventral anterior region. Scale bar: 50 μm.
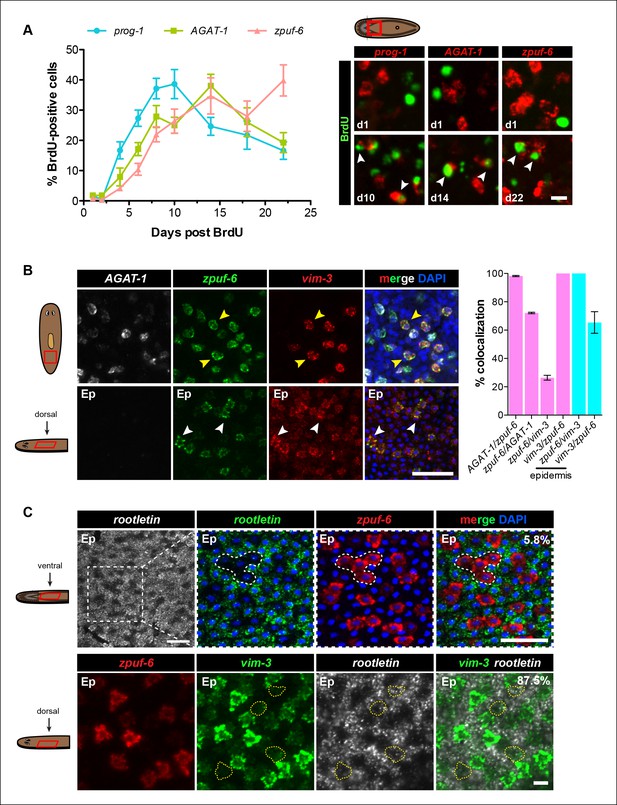
Zpuf-6+ epidermal cells are not terminally differentiated.
(A) Turnover dynamics for prog-1, AGAT-1 and zpuf-6 expressing cell populations. Animals were soaked with BrdU for 24 hr and chased for the indicated time periods. Quantification of the percentage of prog-1+, AGAT-1+ or zpuf-6+ cells analyzed that are BrdU+ are plotted. Error bars: SEM (see Materials and methods). Representative images of BrdU+ cells, are maximum intensity projections over 1 cell diameter of a subset of the quantified images, of the minimum and maximum time points for each gene are shown. White arrowheads highlight double-positive cells. Scale bar: 10 μm. (B) Lineage relationship between AGAT-1, zpuf-6 and vim-3. Top row: Triple FISH showing overlapping expression between AGAT-1, zpuf-6 and vim-3 in the mesenchyme. Yellow arrowheads highlight cells that are zpuf-6+ vim3+ AGAT-1neg. AGAT-1 and vim-3 exhibit little to no co-expression (Figure 2B). Bottom row: Dorsal epidermal (Ep) view of zpuf-6 and vim-3. White arrowheads highlight cells that co-express zpuf-6 and vim-3 but there are additional cells expressing vim-3. Images are single confocal planes. Scale bar: 50 μm. Quantifications of the percent colocalizations of combinations of AGAT-1, zpuf-6 and vim-3 mesenchymal (magenta) and zpuf-6 and vim-3 dorsal epidermal cells (cyan) are shown (∼200-400 cells were quantified over >3 animals; error bars: SD). Gene1/gene2 notation signifies percentage of gene1+ cells that are also positive for gene2 expression. Notable percentages: zpuf-6+/AGAT-1+ (72%), zpuf-6+/vim-3+ (26%). (C) zpuf-6+ epidermal cells express very low levels of rootletin. Top row: double FISH of rootletin and zpuf-6. 5.8%=percentage of strong zpuf-6+ epidermal cells that co-express rootletin (~300 cells). Scale bars: 50 μm. Bottom row: triple FISH of zpuf-6, vim-3 and rootletin in dorsal epidermis. Only overlay of vim-3 and rootletin is shown (far right panel). Yellow-dashed shapes outline cells that express very low levels of vim-3 that also express rootletin. 87.5%=percentage of weak vim-3+ dorsal epidermal cells that co-express rootletin (~200 cells). Images are single confocal planes. Scale bar: 10 μm.
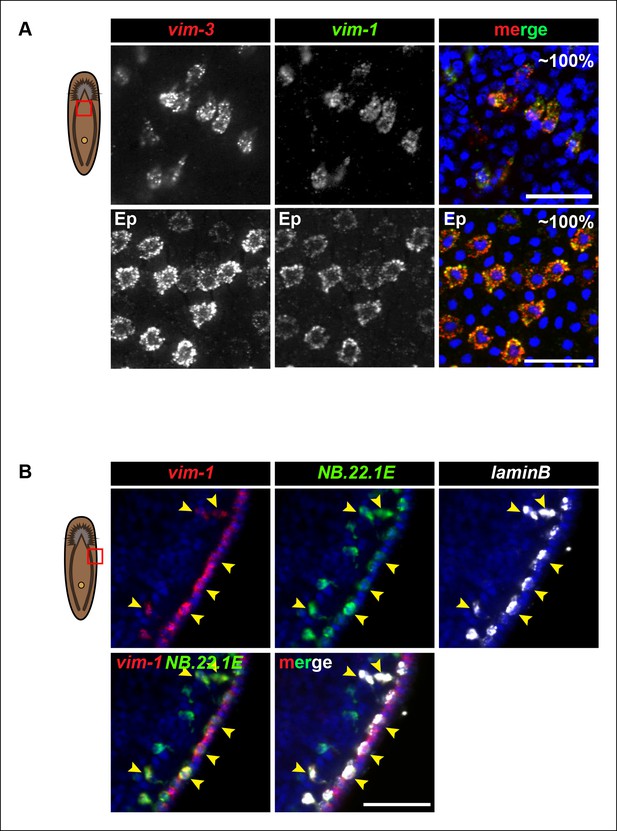
Vim-3 and vim-1 are co-expressed in the same cell types.
(A) Images represent single confocal planes from ventral anterior regions of the mesenchyme (top panels) and epidermis (Ep) (bottom panels). Percentages represent fraction of vim-3+ cells that co-express vim-1 (∼200 mesenchymal cells and 400 epidermal cells were quantified). Scale bars: 50 μm. (B) Vim-1/vim-3 is expressed in NB.22.1E+ and laminB+ cells at the animal body margin (yellow arrowheads). Images represent single confocal planes. Scale bar: 50 μm.
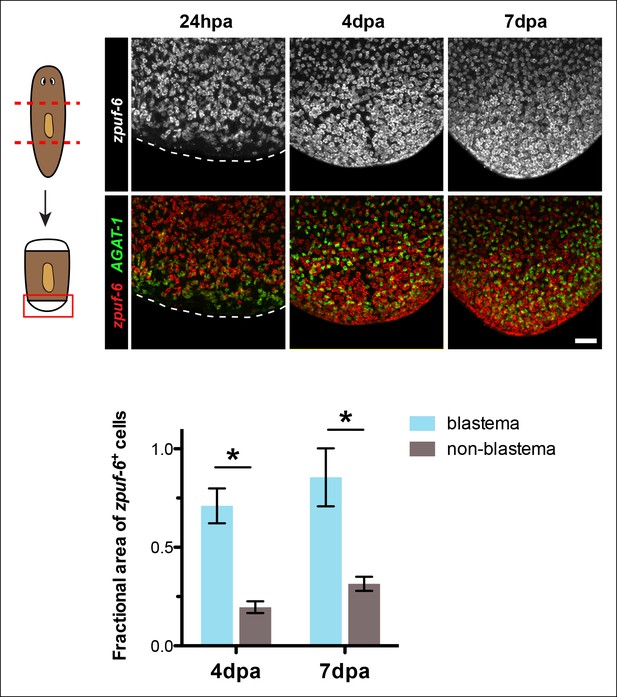
zpuf-6+ cells are enriched at regenerating blastemas.
Shown are posterior trunk fragments cut from wild-type animals and regenerated for the indicated times: hours or days post-amputation (hpa and dpa, respectively) and subjected to double FISH with zpuf-6 and AGAT-1. Dotted lines demarcate the posterior end of the animal. Maximum intensity projections are shown. Scale bar: 100 μm. Quantification of the fractional area of zpuf-6+ cells in the regenerating blastema compared to non-blastema (old tissue). The overall positive signal of zpuf-6+ cells was measured in a fixed region in the blastema compared to the same area in the non-blastema region. Data represent means from 2 regenerating trunk fragments for each time point; error bars: SD. Student's t test: *, p < 0.05.
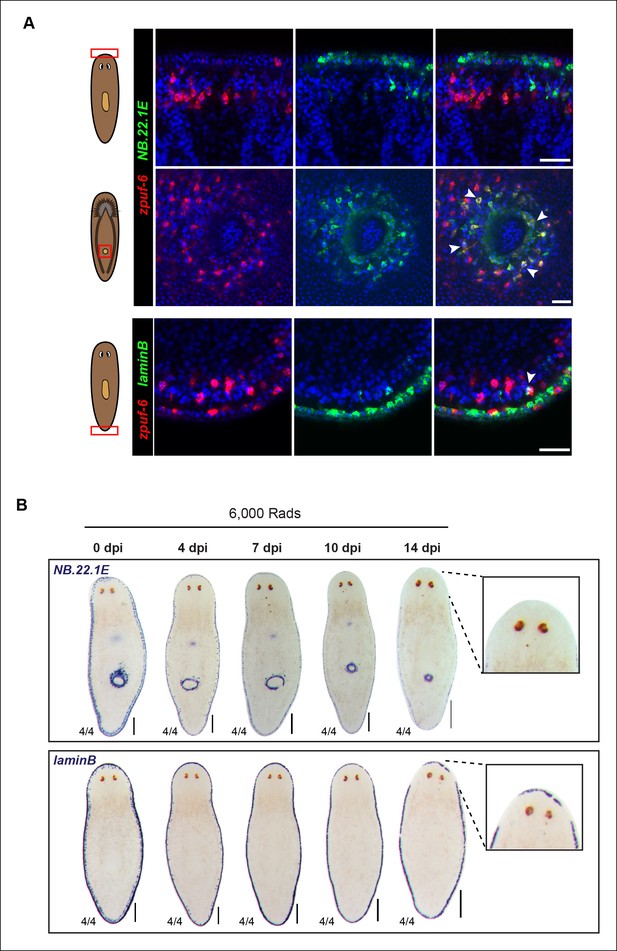
NB.22.1E+ and laminB+ cells exhibit little overlap with zpuf-6 and display slow cell turnover kinetics after irradiation.
(A) Whole-mount FISH of zpuf-6 and markers of epidermal edge cells (NB.22.1E and laminB). zpuf-6 exhibits little to no overlap with NB.22.1E in edge cells (top row) but overlaps with NB.22.1E+ epidermal cells surrounding the ventral mouth opening (middle row, white arrowheads). zpuf-6 shows very little overlap with laminB+ edge cells (bottom row, white arrowheads). Scale bars: 50 μm. (B) Colorimetric WISH of animals after 6,000 Rad irradiation exposure for markers NB.22.1E and laminB. By 14dpi (days post-irradiation), cells around the anterior have disappeared, suggesting that these cells do not turnover as quickly as zpuf-6+ epidermal cells. For NB.22.1E marker, sub-epithelial mesenchymal cells and those around the ventral mouth opening disappear by 7dpi, suggesting they turn over more rapidly than the cells at animal margin. Ventral views. Scale bars: 200 μm.
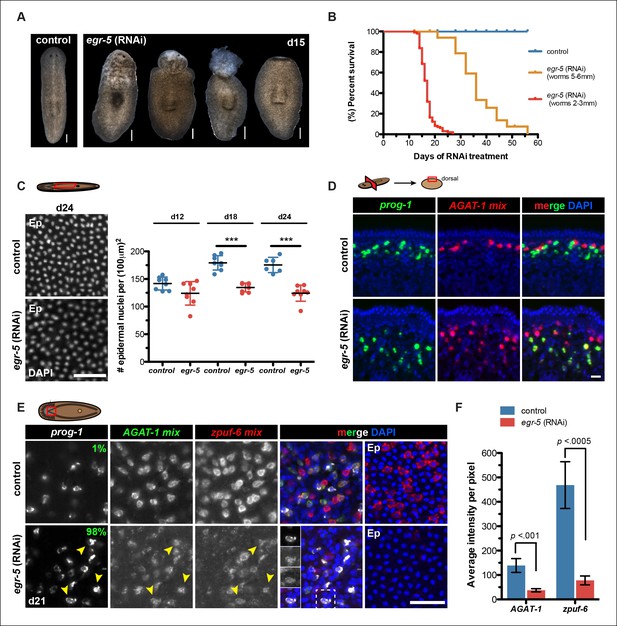
Egr-5 is required for the proper differentiation of epidermal progeny cells.
(A) Intact phenotypes for representative control and egr-5(RNAi) animals. Various stages of phenotypic progression of egr-5 RNAi knockdown (4Fd15) are shown. Scale bars: 200 μm. (B) Survival curves for control and egr-5(RNAi) animals. The efficacy of RNAi and resulting gross phenotype is dependent on size of animals at the start of RNAi feedings. Animals starting around 2-3 mm in size (red lines) were fed 5 times (5F) (n = 99); animals starting around 5-6 mm in size (yellow lines) were fed 6 times (6F) (n = 65). Death was marked by complete animal lysis. (C) Effects of egr-5(RNAi) on epidermal cell density. DapI was used to quantify epidermal (Ep) nuclear density in the ventral mid-sections; representative images for 4Fd24 are shown. Scale bar: 50 μm. Quantification of epidermal cell density are plotted for d12, d18 and d24 for control (blue) and egr-5(RNAi) (red) animals. Each symbol represents individual animals (average of 2 regions/animal). Black lines and error bars (colored) represent mean and SD, respectively. Student's t test: ***, p <0.00005. (D) Spatial domain expansion of prog-1 and AGAT-1 cells in egr-5(RNAi). Tissue transverse sections of control and egr-5(RNAi) animals stained for markers of early epidermal progeny cells (8Fd30). To help improve visualization of AGAT-1-expressing cells, an RNA probe mix of other AGAT-1 co-expressed genes were used (AGAT-1 mix = AGAT-1, zpuf-1, zpuf-3, zpuf-4). Scale bar: 10 μm. (E) Misexpression of epidermal progeny markers in egr-5(RNAi) animals. Triple FISH of prog-1, AGAT-1 and zpuf-6 markers in control and egr-5(RNAi) animals (4Fd21). Percentages shown in prog-1 panel represent percentage of prog-1+ cells that also express AGAT-1 (400-–700 cells were counted between 3-–4 worms per condition). Yellow arrowheads highlight cells that are prog-1+ AGAT-1+ zpuf-6+ (also shown in inset panels in merge panel). Rightmost column: epidermal (Ep) view highlighting loss of zpuf-6 expression in epidermis. AGAT-1 mix = AGAT-1, zpuf-1, zpuf-3, zpuf-4; zpuf-6 mix = zpuf-6, zpuf-5 and zpuf-8. Scale bar: 50 μm. (F) Average fluorescence intensity for AGAT-1 mix and zpuf-6 mix probes in control and egr-5(RNAi) animals from (E). Error bars: SD. p-values are results of unpaired Student's t test.
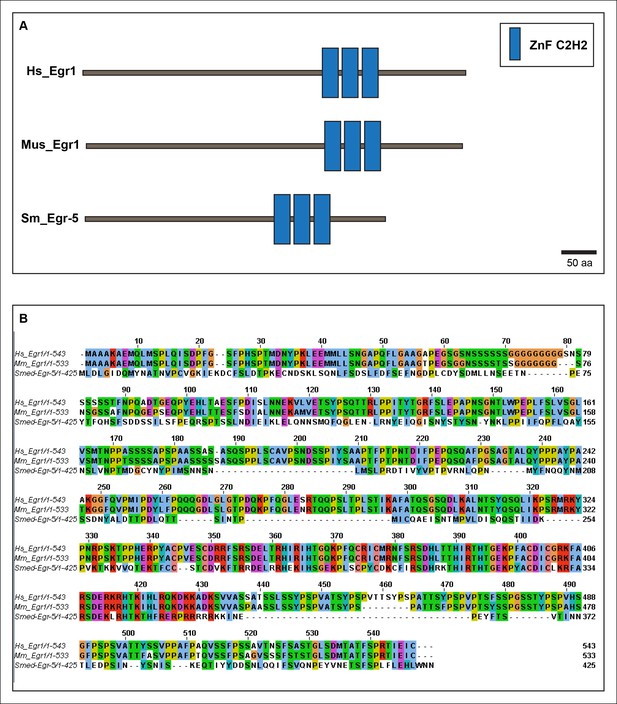
Smed-Egr-5 is a conserved member of the early growth response family of transcription factors.
(A) Schematic of predicted domain structures of Smed-Egr-5 (Sm) with Egr1 from humans (Hs) and mouse (Mus), adapted from the Smart Modular Architecture Research Tool (SMART) (http://smart.embl-heidelberg.de). (B) Alignment of planarian Egr-5 protein sequence (Smed-Egr-5) with Egr1 sequences from human (Hs) and Mus musculus (Mm). Predicted Sm ZnF_C2H2 domains are residues 268-346.
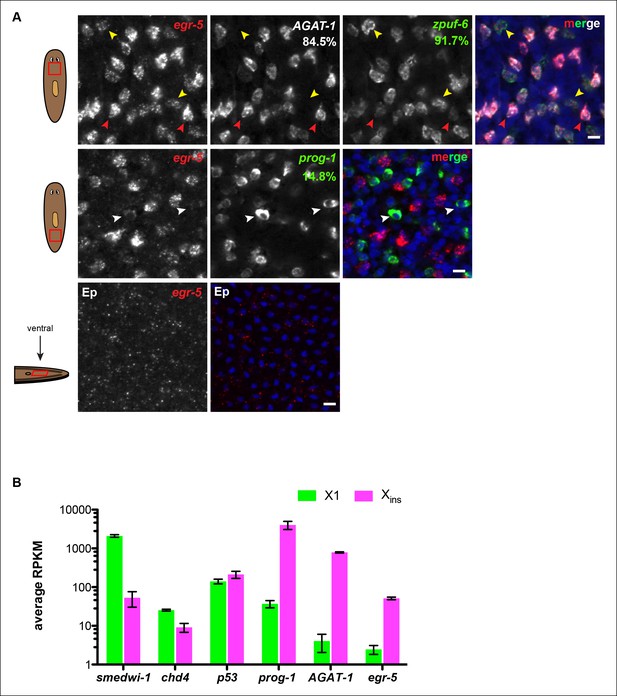
Egr-5 is expressed in multiple post-mitotic epidermal progeny cells.
(A) Whole-mount FISH of egr-5 with various markers in the epidermal lineage. Top panel: Cells expressing the strongest levels of egr-5 exhibit strong AGAT-1 expression and very weak zpuf-6 expression (red arrowheads). Cells expressing lower levels of egr-5 exhibit strong zpuf-6 expression and little to no detectable AGAT-1 expression (yellow arrowheads). Middle panel: Some cells expressing low levels of egr-5 are prog-1+ (white arrowheads). Bottom panel: egr-5 is very weakly expressed in some epidermal cells, which overlap with zpuf-6+ epidermal cells (not shown). Percentages represent fraction of egr-5+ cells (low but detectable expression was counted) that co-express the candidate gene (∼200-400 cells were quantified). Images are single confocal planes. Ep, epidermis. Scale bars: 10 μm. (B) egr-5 transcripts are enriched in the post-mitotic, differentiated cell population (Xins). Average RPKM values are plotted for various known neoblast and non-neoblast genes from wild-type planarian cells dissociated by FACS (data are deposited under GEO accession number: GSE73027). X1 cells are irradiation-sensitive and comprise of genes that are expressed in stem cells (smedwi-1). Xins cells are irradiation-insensitive and represent a population of various post-mitotic cells. chd4 and p53 are expressed in both stem cells and post-mitotic cells and are therefore present in both X1 and Xins cell population. prog-1 and AGAT-1 are post-mitotic epidermal lineage markers and show enrichment in the Xins population. egr-5 has no detectable expression in stem cells based on FISH (not shown), and consistent with this, few egr-5 transcripts are detected in the X1 population.
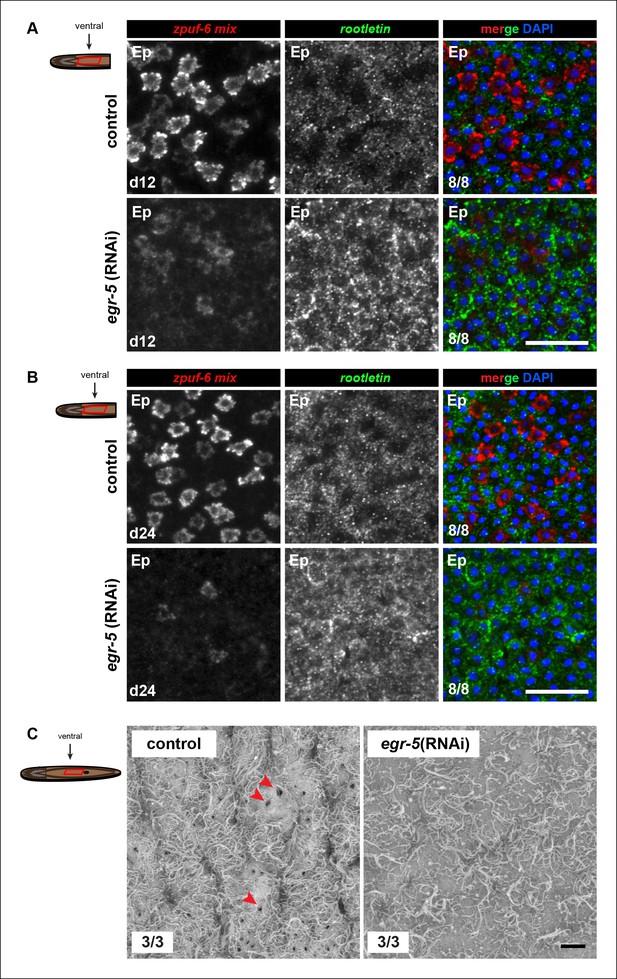
Molecular and ultrastructural analysis of egr-5(RNAi) epidermis.
(A,B) Expression patterns of zpuf-6 and rootletin cilia gene marker at (A) 4Fd12 and (B) 4Fd24 after initial RNAi treatment for control and egr-5(RNAi) animals. Representative images are single confocal planes of ventral epidermis (Ep). Scale bars: 50 μm. (C) Scanning electron micrograph of images (1500X) of control and egr-5(RNAi) animals (4Fd12) of the ventral epidermis (Ep) right above the mouth opening. Although cilia are still present in egr-5(RNAi) animals, there is an absence of epidermal pores (red arrowheads) that may be a result of epithelial stretching. Scale bar: 10 μm.
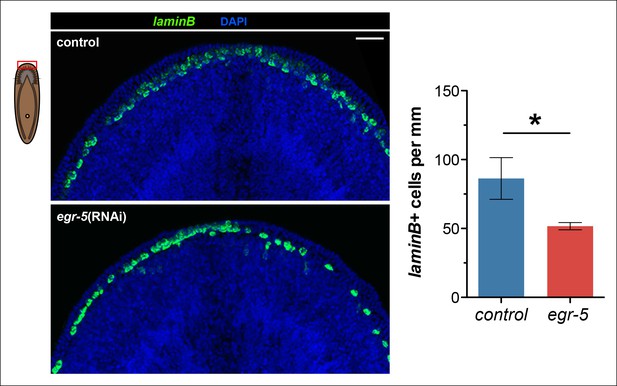
Reduction of laminB+ cells in egr-5(RNAi) animals.
The number of laminB+ cells per mm along the animal margin was quantified in control and egr-5(RNAi) animals. The quantified region was restricted to the anterior because this region exhibits the highest cell turnover of laminB+ cells. Representative images for 4Fd21 are shown. Scale bar: 50 μm. Quantification of laminB+ cell density is plotted from 3 animals for each condition. Data represent means; error bars: SD. Student's t test: *, p < 0.05.
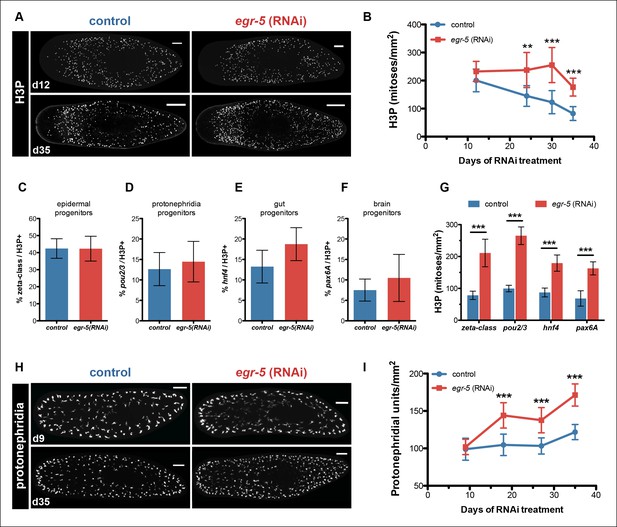
Egr-5 knockdown results in the expansion of multiple progenitor populations.
(A,B) egr-5 knockdown causes an increase in global stem cell proliferation. (A) H3P-positive cells over the surface area in control and egr-5(RNAi) intact animals (8 feedings). Maximum intensity projections of representative H3P patterns for day 12 and day 35 time points are shown. Scale bar: 200 μm. (B) Quantification of H3P-positive cells per surface area from (A). Data represent means, error bars: SD. Student's t test: **, p <0.005, ***, p <0.0001. (C–G) Stem cell proliferation results in the expansion of multiple lineage-committed progenitors. For control and egr-5(RNAi) animals (3Fd27), animals were fixed and stained for H3P and a marker for the following: (C) dividing epithelial progenitors (zeta class: zfp-1, egr-1, fgfr-1), (D) dividing protonephridia progenitors (pou2/3), (E) dividing gut progenitors (hnf4), and (F) dividing brain progenitors (pax6A). Quantifications of percentage of lineage-committed progenitors of total H3P+ cells are shown. Error bars: SD. (G) Quantification of total mitotic cells (H3P) per surface area for control and egr-5(RNAi) animals (3Fd27) from analysis in (C–F). Data represent means, error bars: SD. Student’s t test: ***, p <0.0001. (H,I) egr-5 knockdown causes supernumerary protonephridial units (PU). (H) Control and egr-5(RNAi) intact animals (8 feedings) were stained for the number of proximal units (slc6a-13) to visualize and quantify total PUs. Maximum intensity projections of representative animals for day 9 and day 35 time points are shown. Scale bar: 200 μm. (I) Quantification of PU per surface area. Data represent means, error bars: SD. Student's t test: ***, p <0.0001.
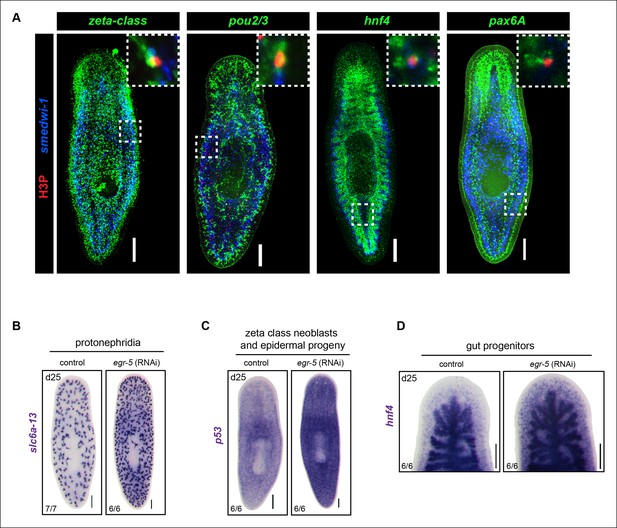
Analysis of multiple dividing tissue progenitors in egr-5(RNAi) animals.
(A) Representative images of egr-5(RNAi) animals from Figure 6C–G fixed and stained for H3P (red), neoblast marker (blue: smedwi-1) and progenitor marker (green). Progenitor markers: zeta-class/epidermal: zfp-1, egr-1, fgfr-1; protonephridia: pou2/3; gut: hnf4; brain: pax6A. Images represent whole-animal maximum intensity projections and zoomed image highlights an example of a positive dividing progenitor cell (i.e. H3P+/smedwi-1+/progenitor marker+). Scale bars: 200 μm. (B-D) Expansion of multiple differentiated tissue markers in egr-5(RNAi) animals. Representative colorimetric WISH images of control and egr-5(RNAi) intact animals (6Fd25) monitoring various lineages/tissues: protonephridia (pou2/3), epidermal progenitors and early/late progeny cells (p53), and gut morphology/gut progenitors (hnf4). Scale bars: 200 μm.
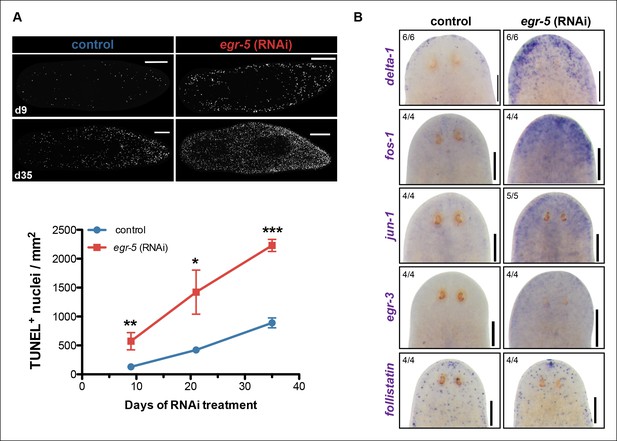
A global stress response is induced by loss of egr-5.
(A) Whole-mount TUNEL assay measuring apoptosis in control and egr-5(RNAi) animals (8 feedings). Quantification of TUNEL-positive nuclei per surface area is plotted. Data represent means; error bars: SEM. Student's t test: *, p <.05, **, p <.005, ***, p <.0001. Representative maximum intensity projection of TUNEL-stained image for day 9 and day 35 are shown. Scale bars: 200 μm. (B) Wound-induced genes are up-regulated in egr-5(RNAi) animals. Representative colorimetric WISH images of delta-1, fos-1, jun-1 egr-3 and follistatin expression in the anterior regions of intact control and egr-5(RNAi) animals at 4Fd21 of RNAi treatment. Scale bars: 200 μm.
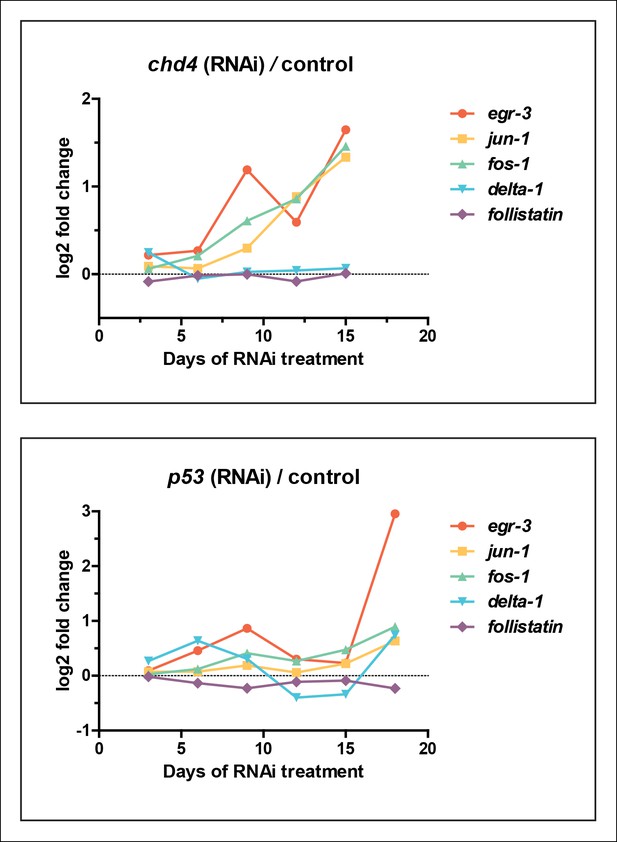
Wound-induced genes are upregulated in chd4 and p53 RNAi datasets.
Plot of log2 ratios of selected wound-induced genes from Figure 7B in chd4(RNAi) (top panel) and p53(RNAi) (bottom panel) whole worm RNA-seq time course.
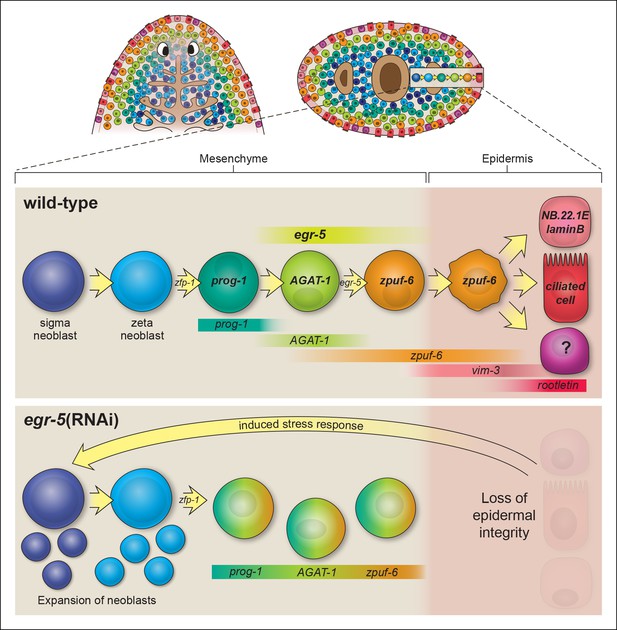
Model of planarian epidermal lineage progression.
Schematic representation of the distribution of neoblasts, epidermal progeny cells in the mesenchyme, and differentiated epidermal cells along the anteroposterior (top left, dorsal view) and dorsoventral (top right, cross section) axes. Gut branches are shown as reference. Below: sigma-class neoblasts give rise to zeta-class neoblasts, which are epidermal progenitors. zfp-1 is required to generate epidermal progeny cells, which begin to express different markers as cells undergo multiple transitions and intercalate into the mature epidermis. Gradient boxes represent domains and relative expression intensity of specified genes. egr-5 is most strongly expressed in AGAT-1+ cells and is required for proper differentiation of epidermal post-mitotic progeny cells. zpuf-6+ epidermal cells likely represent a branching point in the epidermal cell fate decision and can generate multiple different cell types including marginal adhesive cells (NB.22.1E and laminB), multiciliated cells (rootletin), and other unknown cell types. Bottom panel: schematic of egr-5(RNAi) phenotype. Loss of egr-5 disrupts the proper spatiotemporal transition of post-mitotic epidermal cells, resulting in prog-1, AGAT-1 and zpuf-6 to be co-expressed in the same cell, and leading to a depletion of mature epidermal cells. The resulting loss of epidermal integrity is sensed by the neoblasts (induced stress response) and causes an expansion of neoblasts and multiple progenitors before animals eventually lyse due to irreparable loss of an intact epidermal barrier.
Additional files
-
Supplementary file 1
Summary of down-regulated genes in chd4 and p53 RNAi RNA-seq data sets.
Tab1, down-regulated genes in chd4(RNAi), p53(RNAi), and the 587 common down-regulated genes in both chd4 and p53 RNAi data sets. Tab 2, Top BLAST hits of 587 common down-regulated gene list to C. elegans, D. melanogaster, M. musculus, and H. sapiens NCBI Refseq with an e-value cutoff of .001. Tab 3, GO term enrichment analysis for the common down-regulated gene list.
- https://doi.org/10.7554/eLife.10501.026
-
Supplementary file 2
Summary of up-regulated genes in chd4 (d15) and p53 (d18) RNAi RNA-seq data sets.
Genes were classified as up-regulated if the adjusted p value <1e-13 and log2 fold change >1.
- https://doi.org/10.7554/eLife.10501.027
-
Supplementary file 3
In situ expression summary of cloned genes from chd4 and p53 RNAi data sets.
- https://doi.org/10.7554/eLife.10501.028
-
Supplementary file 4
Primer sequences of genes cloned in this study.
- https://doi.org/10.7554/eLife.10501.029





