Kinesin Kip2 enhances microtubule growth in vitro through length-dependent feedback on polymerization and catastrophe
Figures
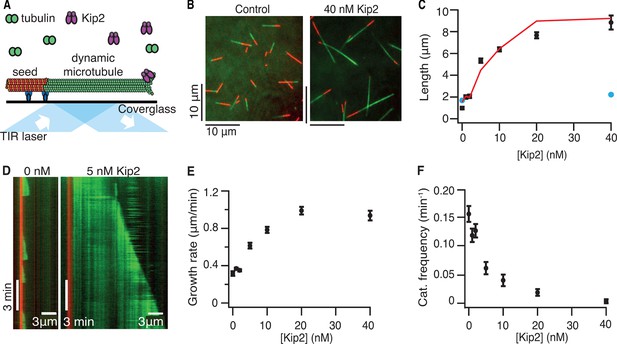
Kip2 is a microtubule polymerase and an anti-catastrophe factor for porcine tubulin.
(A) Schematic of the experimental design: porcine tubulin (green) polymerizes onto stabilized microtubules (red) bound to the coverslip with antibodies (blue), imaged using TIRF microscopy. (B) Microscopy images of dynamic microtubules grown from stabilized seeds without (left) and with 40 nM Kip2 (right) at t = 10 min. (C) Microtubule length as a function of Kip2 concentration in ATP (black circles) or AMP-PNP (blue circles) at t = 10 min. The red line indicates the expected microtubule length at t = 10 min, calculated from the measured growth rates and catastrophe frequencies in Table 1 according to the formula L = (v+/f+-)[1-exp(-tf+-)], where v+ is the growth rate and f+- is the catastrophe frequency (ignoring rescues and assuming that regrowth occurs without delay). (D) Kymographs showing typical microtubule growth without (left) and with 5 nM Kip2 (right) in ATP. (E) Microtubule growth rate as a function of Kip2 concentration in ATP. (F) Catastrophe frequency as a function of Kip2 concentration in ATP. All error bars are standard errors of the mean. Please refer to Table 1 for values. AMP-PNP, adenylyl imidodiphosphate; ATP, adenosine triphosphate; TIRF, total internal reflection fluorescence.
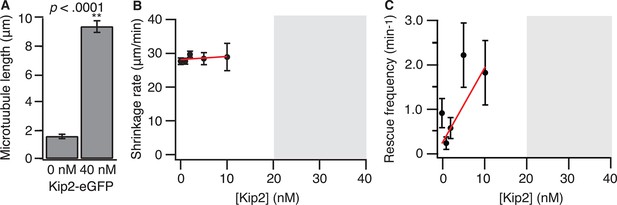
Kip2 has no significant effect on microtubule shrinkage rate or rescue frequency.
(A) Porcine microtubule length at 0 or 40 nM Kip2-eGFP in ATP, measured at t = 10 min. (B) Porcine microtubule shrinkage rate as a function of Kip2 concentration. The gray box indicates the Kip2 concentration regime in which microtubule catastrophe is very rare, and we did not quantify microtubule shrinkage rate. The data were fitted by linear regression, weighted by the SE. Slope = 0.09 ± 0.11, y-intercept = 28.1 ± 0.58 μm/min. (C) Porcine microtubule rescue frequency (shrinkage rate divided by rescue distance) as a function of Kip2 concentration. The gray box indicates the Kip2 concentration regime in which microtubule catastrophe is very rare and we did not measure shrinkage rate or rescue frequency. The data were fitted by linear regression, weighted by the SE. Slope = 0.17 ± 0.07, y-intercept = 0.18 ± 0.11 μm. The slope was not significantly different from zero (p>0.05). Error bars are SE. ATP, adenosine triphosphate; SE, standard error of the mean.
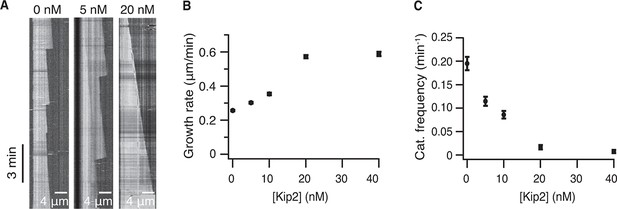
Kip2 is a microtubule polymerase and an anti-catastrophe factor for yeast tubulin.
(A) Kymographs from DIC microscopy showing typical microtubule growth with 4-μM unlabeled yeast tubulin without (left) and with Kip2 (5 nM, center, 20 nM, right) in ATP. (B) Yeast microtubule growth rate as a function of Kip2 concentration in ATP. (C) Catastrophe frequency as a function of Kip2 concentration of yeast microtubules in ATP. Error bars are SE. Please refer to Table 2 for values. ATP, adenosine triphosphate; DIC, differential interference contrast; SE, standard error of the mean.
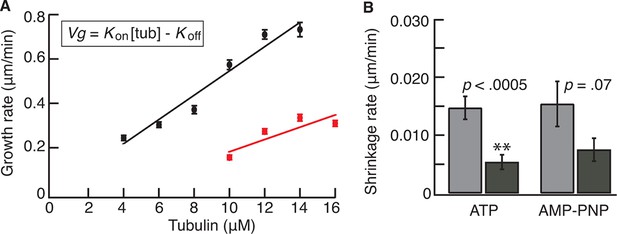
Kip2 increases the growth rate in GTP-tubulin and lowers the off rate of GMPCPP tubulin.
(A) Porcine microtubule growth rate from GMPCPP seeds as a function of tubulin concentration in ATP without Kip2 (red) and with 40 nM Kip2 (black) in solution. The data were fit using linear regression, weighted by the SE. The slope corresponds to a second-order association rate for free GTP-tubulin dimers of 0.70 ± 0.30 μM-1·s-1 without Kip2 and 1.48 ± 0.16 μM-1·s-1 with 40 nM Kip2. The y-intercepts were –1.9 ± 4.0 s-1 without Kip2 and –0.21 ± 1.6 s-1 with 40 nM Kip2; the intercepts did not differ significantly from zero. (B) Shrinkage rates of GMP-CPP microtubules at 0 nM Kip2 (light gray) and 40 nM Kip2 (dark gray) in ATP and AMP-PNP. Error bars are SEs. AMP-PNP, adenylyl imidodiphosphate; ATP, adenosine triphosphate; GTP, guanosine-5'-triphosphate; SE, standard error of the mean.

SDS–PAGE gels of Kip2 and Kip2-eGFP.
Lane 1: Pooled Kip2 fractions after gel filtration (78 kDa). Lane 2: Pooled Kip2-eGFP fractions after gel filtration (105 kDa). Molecular weight markers (Seeblue Plus2) are indicated by horizontal lines. eGFP, enhanced green fluorescent protein; SDS–PAGE, sodium dodecyl sulfate polyacrylamide gel electrophoresis.
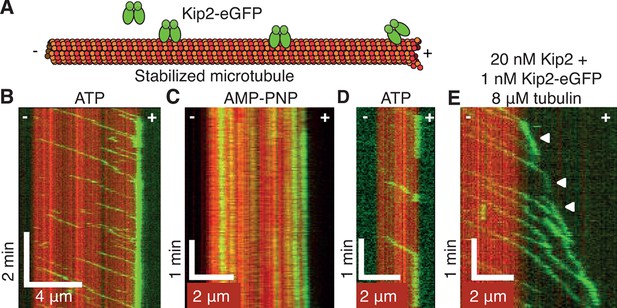
Kip2 is a highly processive motor that dwells at plus ends.
(A) Schematic of the experimental design. (B) Kymograph showing processive motility and plus end accumulation of individual Kip2-eGFP molecules on GMPCPP-stabilized microtubules in 1 mM ATP. The concentration of Kip2-eGFP was 0.085 nM. (C) Kymograph showing tightly bound Kip2-eGFP molecules in AMP-PNP. (D) Kymograph showing end residence of individual Kip2-eGFP molecules in ATP. (E) Kymograph showing end residence of 1 nM Kip2-eGFP spiked into 20 nM unlabeled Kip2 in the presence of 8 μM unlabeled tubulin in ATP. Arrow heads indicate microtubule plus end tracking events. ATP, adenosine triphosphate; AMP-PNP, adenylyl imidodiphosphate; eGFP, enhanced green fluorescent protein; GMPCPP, guanylyl (a,ß)methylene-diphosphonate.
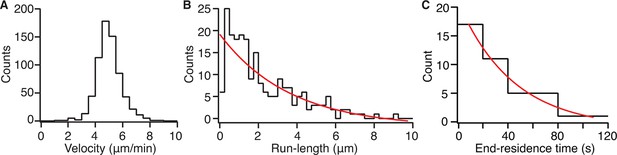
Kip2-eGFP velocity, run length and end residence time distributions.
(A) Histogram showing velocities of single Kip2-eGFP molecules on GMPCPP-stabilized porcine microtubules in 1 mM ATP. (B) Histogram showing the distribution of Kip2-eGFP run lengths on GMPCPP-stabilized porcine microtubules in 1 mM ATP. The run length of Kip2 is predicted to be exponentially distributed, as dissociation from the microtubule lattice is expected to be a random process. The red line depicts a single exponential: f(x) = A exp(-x/x0), where A = 21.1 ± 2.1 and x0 = 3.6 ± 1.0 μm; run lengths between 0 and 0.5 μm are under-represented, likely due to the limited temporal resolution. (C) Histogram showing end residence times of Kip2-eGFP on GMPCPP-stabilized porcine microtubules. End residence times were included in the analysis only if single Kip2-eGFP molecules could be observed to arrive at, and dissociate from, a microtubule plus end. The red line depicts a single exponential: f(t) = A exp(-t/t0), where A = 18.2 ± 2.4 and t0 = 44.1 ± 14.5 s. ATP, adenosine triphosphate; eGFP, enhanced green fluorescent protein; GMPCPP, guanylyl (a,ß)methylene-diphosphonate.
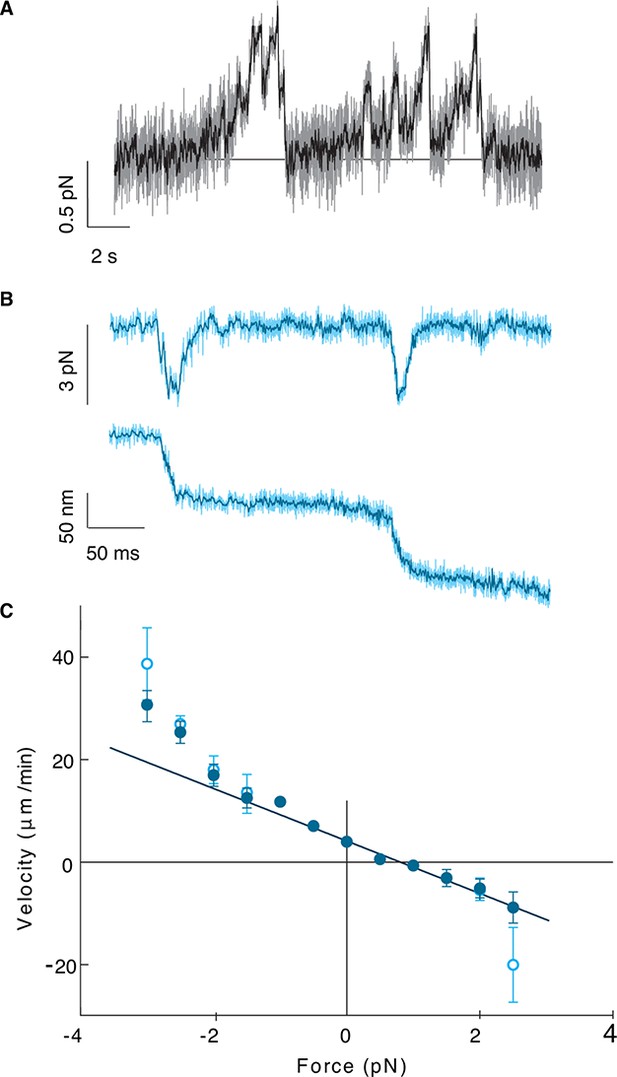
Kip2 is a low-force motor.
(A) Stall force measurement tace. Sampling rate: 10 kHz, raw data (gray), boxcar filtered to 50 Hz (black). A force of 0.5 N corresponds to a displacement of about 17 nm. (B) Time trace for a slip event under 3 pN assisting force. Sampling rate: 20 kHz, raw data (light cyan), boxcar filtered to 400 Hz (dark cyan). (C) Kip2 force-velocity curve: positive is a hindering (load) force and negative is an assisting force. Open symbols include slip events. Error bars are standard errors of the mean.
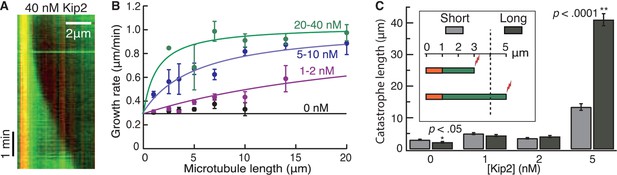
Kip2 promotes porcine microtubule growth in a length-dependent manner.
(A) Kymograph showing acceleration of microtubule growth with increasing length at 40 nM Kip2. (B) Porcine microtubule growth rate as a function of length without Kip2 (black) and binned for 1–2 nM Kip2 (purple), 5–10 nM Kip2 (blue) and 20–40 nM Kip2 (green). Lengths are binned for 0–2 μm, 2–3 μm, 3–4 μm, 4–6 μm, 6–8 μm, 8–12 μm, 12–16 μm and 16–24 μm. The data were fit with the equation, where v0 = 0.294 ± 0.009 μm/min is the initial growth rate; vmax = 1.03 ± 0.03 μm/min is the maximum growth rate; L is microtubule length and A = 39.8 ± 5.4 μm·nM. (C) Mean catastrophe length at various Kip2 concentrations for short (light gray) and long (dark gray) microtubules. In the short microtubule bins, we summed the total distance that microtubules grew while shorter than 4 μm and divided by the number of catastrophes that occurred at lengths < 4 μm. In the long microtubule bin, we summed the total distance that microtubules grew while longer than 4 μm and divided by the number of catastrophes that occurred at lengths > 4 μm (Figure 4C, inset). The number of catastrophes was 120 (0 nM Kip2), 102 (1 nM Kip2), 111 (2 nM Kip2) and 23 (5 nM Kip2). The number of catastrophes at higher Kip2 concentrations was too small to make statistically significant comparisons. Error bars are standard errors of the mean.
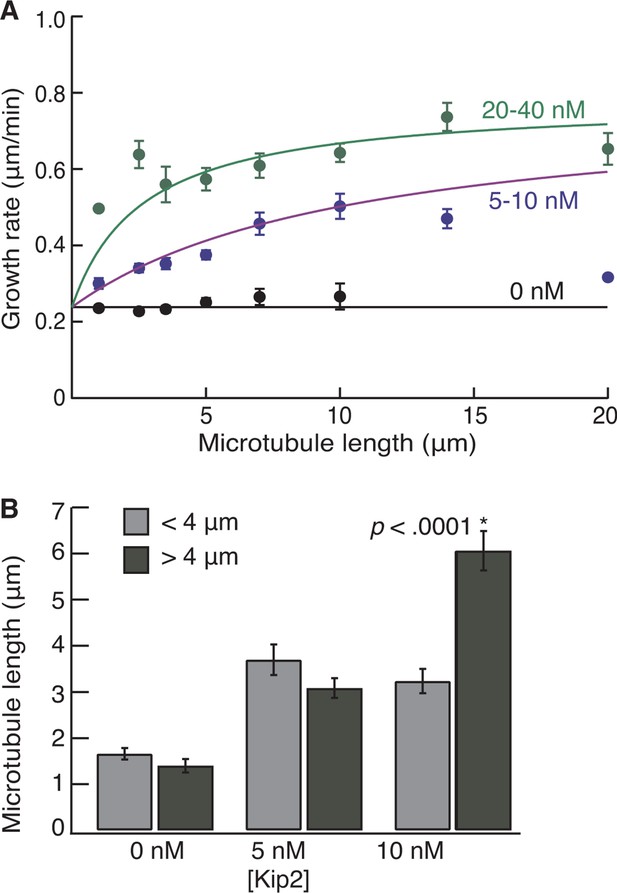
Length dependence of growth and catastrophe for yeast tubulin.
(A) Yeast microtubule growth rate as a function of length, without Kip2 (black) and binned for 5–10 nM Kip2 (blue) and 20–40 nM Kip2 (green). Lengths are binned for 0–2 μm, 2–3 μm, 3–4 μm, 4–6 μm, 6–8 μm, 8–12 μm, 12–16 μm and 16–24 μm. The data were fit with the equation, where v0 = 0.238 ± 0.005 μm/min is the initial growth rate; vmax = 0.78 ± 0.02 μm/min is the maximum growth rate; L is microtubule length and A = 79.4 ± 8.7 μm·nM. (B) Mean length at catastrophe for yeast microtubules at 0 nM Kip2 (n = 170 catastrophes), 5 nM Kip2 (n = 101) and 10 nM Kip2 (n = 80). Bars depict dynamic microtubule lengths below 4 μm (light gray) or above 4 μm (dark gray). Error bars are SE. SE, standard error of the mean.
Tables
Parameters of microtubule dynamics for 12 μM porcine tubulin (mean ± SE).
| [Kip2] (nM) | Length (μm) | Growth rate (μm/min) | Catastrophe frequency (min-1) | Catastrophe distancea (μm) | Shrinkage rate (μm/min) | Rescue frequency (min-1) | Rescue distanceb (μm) |
|---|---|---|---|---|---|---|---|
| 0 | 1.0 ± 0.1 (n = 54) | 0.32 ± 0.02 (n = 172) | 0.166 ± 0.015 (n = 126) | 1.9 ± 0.2 | 27.6 ± 1.0 (n = 130) | 0.88 ± 0.33 (n = 7) | 32 ± 12 |
| 1 | 1.0 ± 0.2 (n = 75) | 0.37 ± 0.01 (n = 152) | 0.126 ± 0.012 (n = 104) | 2.9 ± 0.3 | 27.7 ± 0.9 (n = 85) | 0.20 ± 0.14 (n = 2) | 140 ± 100 |
| 2 | 2.1 ± 0.2 (n = 88) | 0.35 ± 0.01 (n = 159) | 0.135 ± 0.013 (n = 115) | 2.6 ± 0.3 | 29.7 ± 0.9 (n = 110) | 0.54 ± 0.24 (n = 5) | 55 ± 25 |
| 5 | 5.3 ± 0.2 (n = 82) | 0.62 ± 0.03 (n = 77) | 0.065 ± 0.011 (n = 33) | 10 ± 2 | 28.4 ± 1.8 (n= 45) | 2.2 ± 0.7 (n = 9) | 13 ± 4 |
| 10 | 6.4 ± 0.2 (n = 75) | 0.78 ± 0.03 (n = 38) | 0.043 ± 0.011 (n = 16) | 18 ± 5 | 29 ± 4 (n = 18) | 1.8 ± 0.7 (n = 6) | 16 ± 6 |
| 20 | 7.7 ± 0.3 (n = 68) | 0.99 ± 0.04 (n = 36) | 0.020 ± 0.006 (n = 10) | 50 ± 16 | - | -. | - |
| 40 | 8.8 ± 0.6 (n = 26) | 0.94 ± 0.05 (n = 18) | 0.004 (n = 1) | 235 | - | - | - |
-
aThe catastrophe distance is the growth rate divided by the catastrophe frequency.
-
bThe rescue distance is the shrinkage rate divided by the rescue frequency.
Parameters of microtubule dynamics for 4 μM yeast tubulin (mean ± SE).
| [Kip2] (nM) | Growth rate (μm/min) | Catastrophe frequency (min-1) |
|---|---|---|
| 0 | 0.257 ± 0.004 (n = 300) | 0.234 ± 0.017 (n = 191) |
| 5 | 0.302 ± 0.005 (n = 263) | 0.137 ± 0.012 (n = 141) |
| 10 | 0.353 ± 0.008 (n = 146) | 0.103 ± 0.010 (n = 116) |
| 20 | 0.572 ± 0.01 (n = 57) | 0.020 ± 0.006 (n = 13) |
| 40 | 0.589 ± 0.012 (n = 48) | 0.009 ± 0.004 (n = 5) |





