A general strategy to construct small molecule biosensors in eukaryotes
Figures
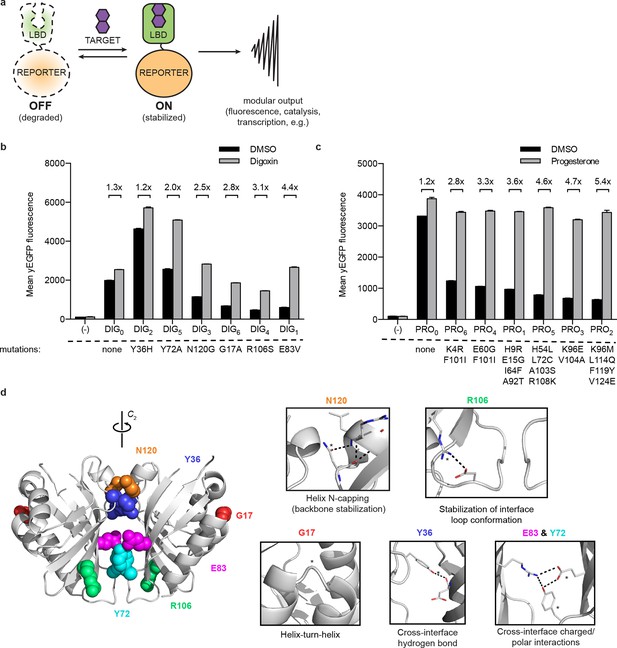
A general method for construction of biosensors for small molecules.
(a) Modular biosensor construction from a conditionally destabilized LBD and a genetically fused reporter. The reporter is degraded in the absence but not in the presence of the target small molecule. (b) yEGFP fluorescence of digoxin LBD-GFP biosensors upon addition of 250 μM digoxin or DMSO vehicle. (c) yEGFP fluorescence of progesterone LBD-GFP biosensors upon addition of 50 μM progesterone or DMSO vehicle. (d) Positions of conditionally destabilizing mutations of DIG0 mapped to the crystal structure of the digoxin LBD (PDB ID 4J9A). Residues are shown as colored spheres and key interactions highlighted in insets. In b-c, fold activation is shown above brackets, (-) indicates cells lacking biosensor constructs, and error bars represent the standard error of the mean (s.e.m.) of three biological replicates.
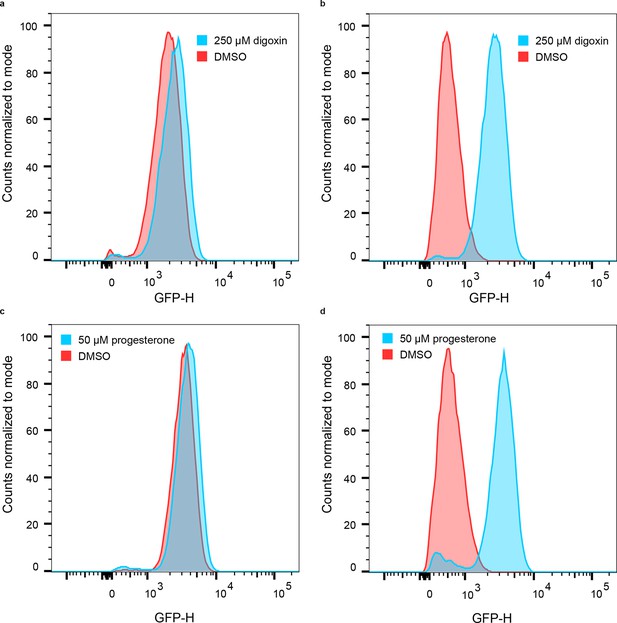
Population responses to cognate ligand for cells bearing LBD-biosensors.
(a) yEGFP fluorescence of DIG0-GFP upon addition of 250 μM digoxin or DMSO vehicle. (b) yEGFP fluorescence of DIG1-GFP upon addition of 250 μM digoxin or DMSO vehicle. (c) yEGFP fluorescence of PRO0-GFP upon addition of 50 μM progesterone or DMSO vehicle. (d) yEGFP fluorescence of PRO1-GFP upon addition of 50 μM progesterone or DMSO vehicle. The data for each condition are presented for a single representative biological replicate with 20,000 events measured by flow cytometry.
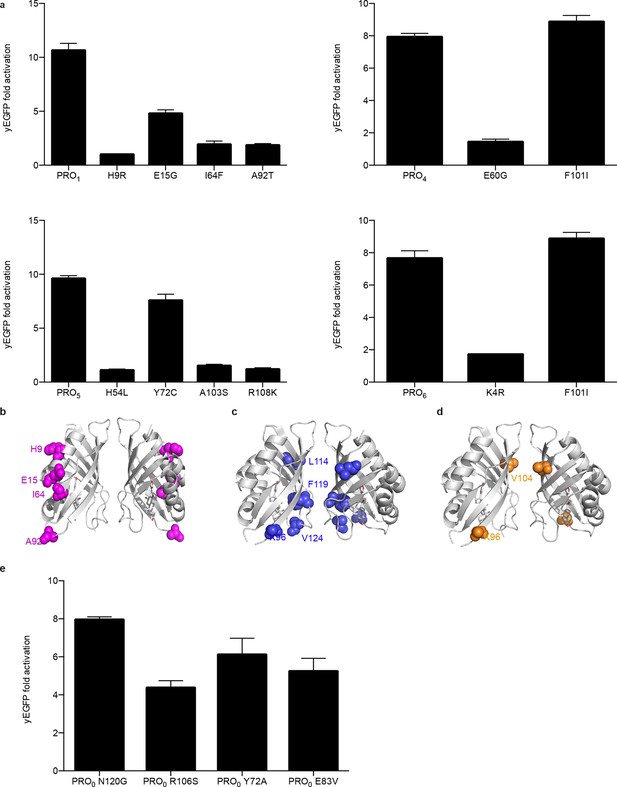
Characterization of mutations conferring progesterone-dependent stability.
(a) Single-mutant deconvolutions of mutations conferring progesterone sensitivity. The parental biosensor appears in the leftmost column of each panel. (b–e) Positions of mutations in PRO1 (b), PRO2 (c), and PRO3 (d) are mapped to the crystal structure of the digoxin LBD (PDB ID 4J9A) and are shown in colored spheres. (e) Fold activation of PRO0-GFP biosensors with digoxin biosensor mutations upon addition of 50 μM progesterone. In a and e, error bars represent s.e.m. of three biological replicates.
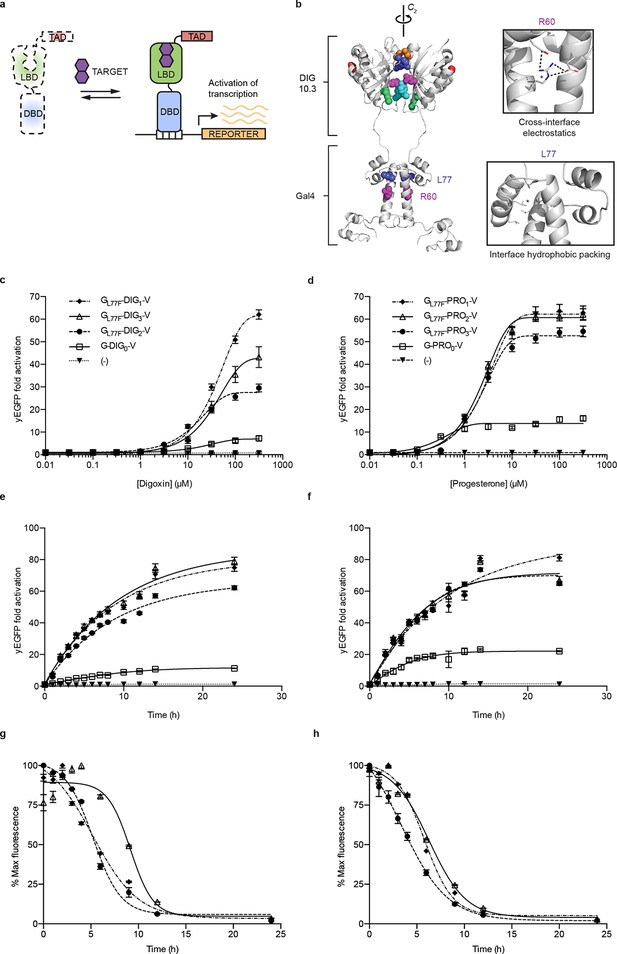
Ligand-dependent transcriptional activation.
(a) TF-biosensor construction from a conditionally destabilized LBD, a DNA-binding domain and a transcriptional activation (TAD) domain. (b) Positions of conditionally destabilizing mutations of Gal4 mapped to a computational model of Gal4-DIG0 homodimer. Residues are shown as colored spheres and key interactions are highlighted in insets. The TAD is not shown. (c) Concentration dependence of response to digoxin for digoxin TF-biosensors driving yEGFP expression. (d) Concentration dependence of response to progesterone for progesterone TF-biosensors driving yEGFP expression. (e) Time dependence of response to 250 μM digoxin for digoxin TF-biosensors. (f) Time dependence of response to 50 μM progesterone for progesterone TF-biosensors. (g) Time-dependent response to withdrawal of 250 μM digoxin for digoxin TF-biosensors. (h) Time-dependent response to withdrawal of 50 μM progesterone for progesterone TF-biosensors. In c-f, (-) indicates cells lacking biosensor plasmids and error bars represent s.e.m. of three biological replicates. Marker symbols in e and g are the same as in c. Marker symbols in f and h are the same as in d.
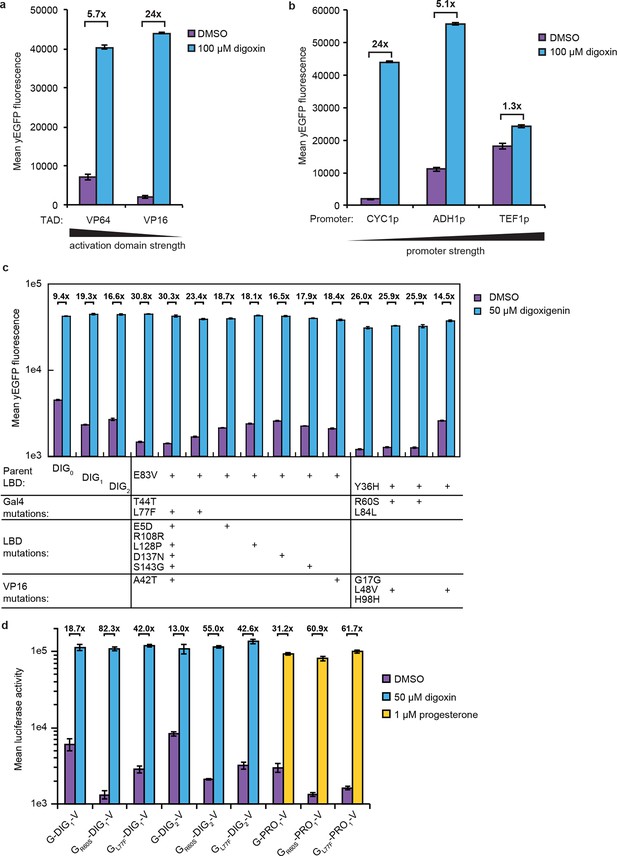
Improvements to TF-biosensor response.
Digoxin-dependent expression of yEGFP by G-DIG1-V TF-biosensors either (a) containing VP64 or VP16 as the TAD and expressed from a CYC1 promoter or (b) containing a VP16 TAD and expressed from a CYC1, ADH1, or TEF1 promoter. (c) Individual mutations identified in a FACS analysis of an error-prone PCR library of G-DIG-V biosensors were tested for their effect on biosensor function using digoxigenin. Transformants were analyzed in an yEGFP yeast reporter strain containing a deletion of PDR5 (PyE14). Improvements in fold activation relative to parental sequences were localized to mutations in Gal4. (d) R60S and L77F mutations found in Gal4 were introduced into G-DIG1-V, G-DIG2-V, and G-PRO1-V. In each case, the Gal4 mutations had the effect of lowering the amount of luciferase activity in the absence of the relevant ligand. In a-d, error bars represent s.e.m. of three biological replicates.
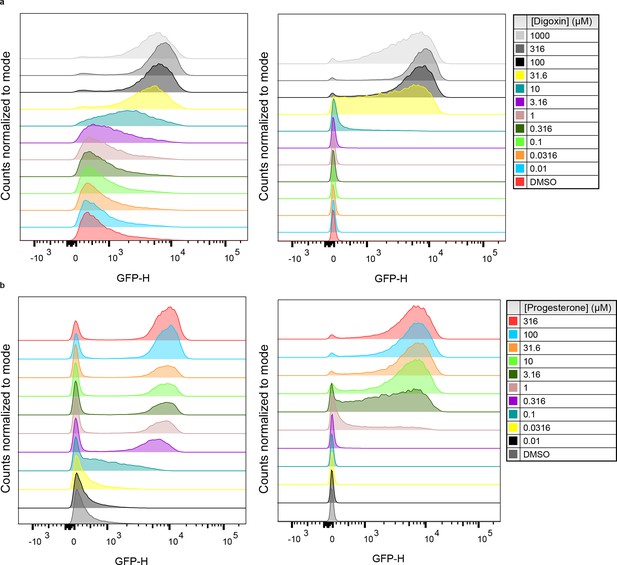
Population responses to cognate ligand for cells bearing TF-biosensors.
(a) yEGFP fluorescence of DIG0-GFP upon addition of digoxin or DMSO vehicle (left) and yEGFP fluorescence of DIG1-GFP upon addition of digoxin or DMSO vehicle (right). (b) yEGFP fluorescence of PRO0-GFP upon addition of progesterone or DMSO vehicle (left) and yEGFP fluorescence of PRO1-GFP upon addition of progesterone or DMSO vehicle (right). Data for each condition are presented for a single representative biological replicate with 20,000 events measured by flow cytometry.
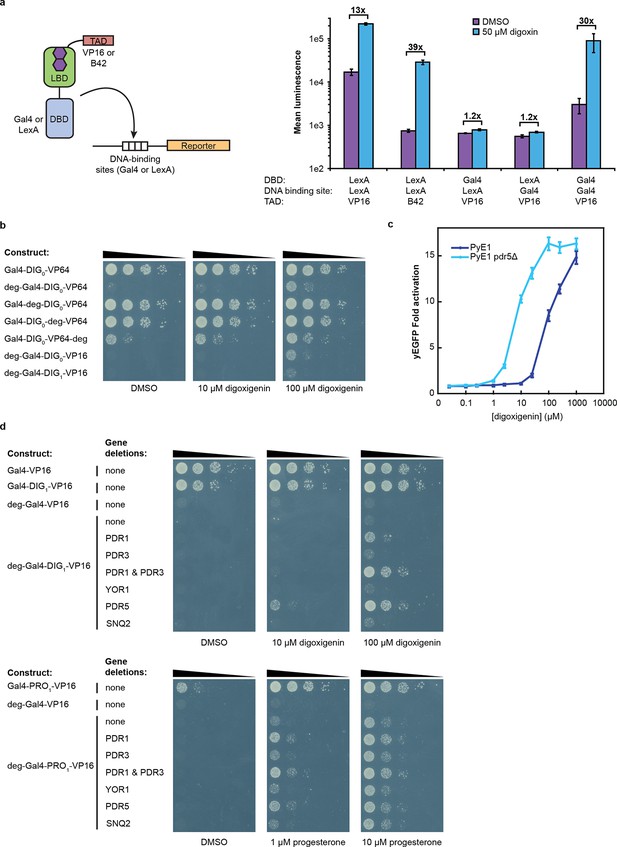
Tuning TF-biosensors for different contexts.
(a) The TAD and DBD of the TF-biosensor and the corresponding binding site for the DBD in the reporter promoter can be swapped for a different application. Expression of a plasmid-borne luciferase reporter was driven by TF-biosensors containing either a LexA or Gal4 DBD and either a VP16 or B42 TAD. Promoters for the reporter contained DNA-binding sites for either Gal4 or LexA. (b) TF-biosensors were transformed into the yeast strain PJ69-4a and tested for growth on this minimal media containing 1 mM 3-aminotriazole (3-AT) and the indicated steroid. To determine the effect of including an additional destabilization domain, the degron from Matα2 was cloned into one of four positions. (c) G-DIG1-V biosensor response to digoxigenin in yEGFP reporter strain PyE1 either with or without a deletion to the ORF of PDR5. (d) Ligand and TF-biosensor-dependent growth on this media in yeast strains containing deleted ORFs for efflux-related transcription factors (PDR1 and PDR3) or ABC transporter proteins (YOR1, PDR5, SNQ2). In a and c, error bars represent s.e.m. of three biological replicates.
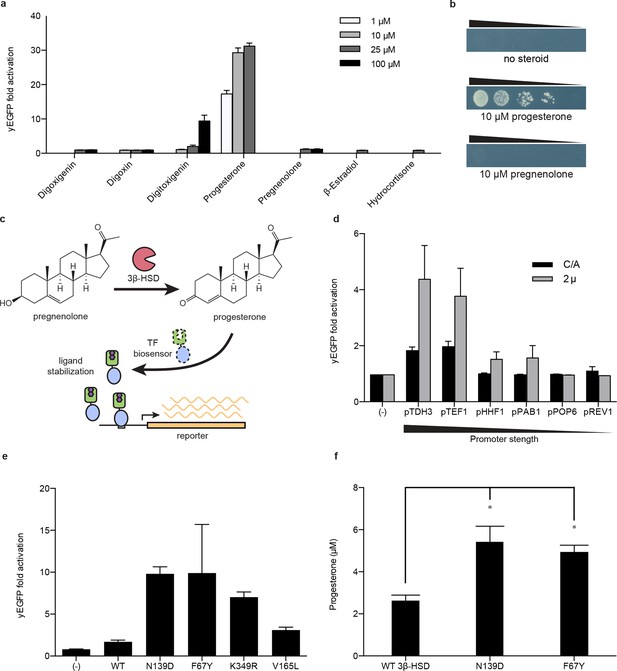
Application of biosensors to metabolic engineering in yeast.
(a) Fold activation of GL77F-PRO1-V by a panel of steroids in yEGFP reporter strain PyE1. Data are represented as mean ± SEM. (b) Growth of degron-G-PRO1-V in HIS3 reporter strain PJ69-4a is stimulated by progesterone but not pregnenolone. (c) Schematic for directed evolution of 3β-HSD using TF-biosensors for the conversion of pregnenolone to progesterone. (d) Fold activation of GL77F-PRO1-V by a panel of plasmids expressing wild-type 3β-HSD under varying promoter strengths in yEGFP reporter strain PyE1 when incubated in 50 μM pregnenolone. Data for plasmids containing CEN/ARS and 2 μ (2 micron) origins are shown. (e) Fold activation of GL77F-PRO1-V by a panel of evolved 3β-HSD mutants expressed under the TDH3 promoter on a CEN/ARS plasmid and incubated in 50 μM pregnenolone. (f) Progesterone titer in 1 OD of cells produced by strains expressing 3β-HSD mutants. Progesterone became toxic at levels of 100 μM and above, leading to substantial cell death. β-estradiol and hydrocortisone were not soluble in yeast growth media at levels above 25 μM. In a and d-f, data are presented as mean ± s.e.m. of three biological replicates. In d and e, (-) indicates cells lacking 3β-HSD. *indicates significance with a threshold of p < 0.05 using 2-tailed Student’s t-test.
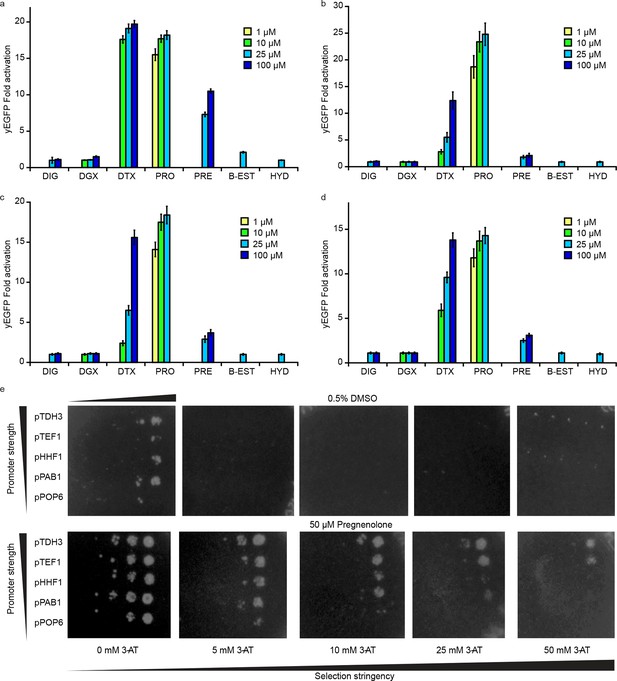
Specificity of PRO biosensors enables selection for auxotrophy complementation.
Specificity for progesterone (PRO) over digoxigenin (DIG), digoxin (DGX), digitoxigenin (DTX), pregnenolone (PRE), β-estradiol (B-EST), and hydrocortisone (HYD) for (a) G-PRO0-V (b) G-PRO1-V (c) G-PRO2-V and (d) G-PRO3-V. (e) Growth response of yeast strain PyE1 transformed with 3β-HSD on CEN/ARS plasmids under various promoters and plated on SC –his (and –ura –leu for plasmid maintenance) containing titrations of 3-AT and either 0.5% DMSO (upper panels) or 50 μM pregnenolone (lower panels). Progesterone became toxic at levels of 100 μM and above, leading to substantial cell death. Beta-estradiol and hydrocortisone were not soluble in yeast growth media at levels above 25 μM. In a-d, error bars represent s.e.m. of three biological replicates.
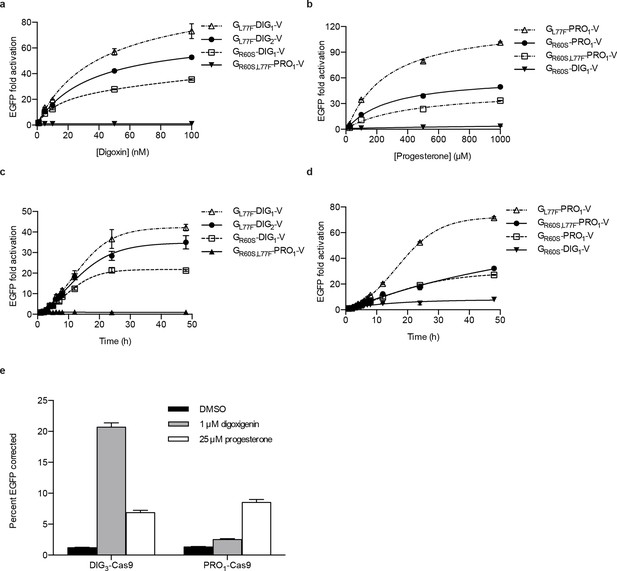
Activation of biosensors in mammalian cells and regulation of CRISPR/Cas9 activity.
(a) Concentration dependence of response to digoxin for constructs containing digoxin TF-biosensors and Gal4 UAS-E1b-EGFP reporter individually integrated into K562 cells. GR60S,L77F-PRO1-V serves as a digoxin insensitive control. (b) Concentration dependence of response to progesterone for constructs containing progesterone TF-biosensors and Gal4 UAS-E1b-EGFP reporter individually integrated into K562 cells. GR60S-DIG1-V serves as a progesterone insensitive control. (c) Time dependence of response to 100 nM digoxin for constructs containing digoxin TF-biosensors and Gal4 UAS-E1b-EGFP reporter individually integrated into K562 cells. GR60S,L77F-PRO1-V serves as a digoxin insensitive control. (d) Time dependence of response to 25 μM progesterone for constructs containing progesterone TF-biosensors and Gal4 UAS-E1b-EGFP reporter individually integrated into K562 cells. GR60S-DIG1-V serves as a progesterone insensitive control. (e) DIG3 and PRO1 fused to the N-terminus of S. pyogenes Cas9 were integrated into a K562 cell line containing a broken EGFP. EGFP function is restored upon transfection of a guide RNA and donor oligonucleotide with matching sequence in the presence of active Cas9. The data are presented as mean fluorescence ± s.e.m. of three biological replicates.
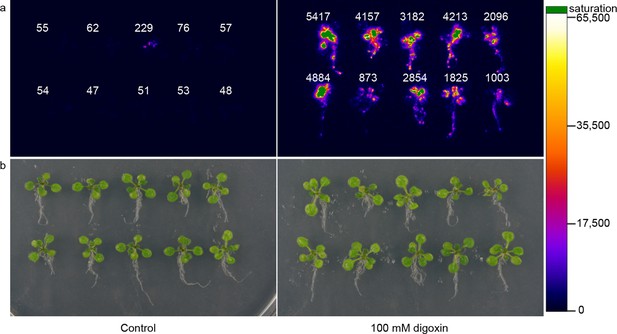
Application of biosensors in plants.
(a) Activation of luciferase expression in transgenic Arabidopsis plants containing the G-DIG1-V biosensor in the absence (left) or presence (right) of 100 μM digoxin. Luciferase expression levels are false colored according to scale to the right. Relative luciferase units corresponding to 1 min of image pixel integration (to avoid saturating pixels) are shown above each individual plant. (b) Brightfield image of plants shown in a.
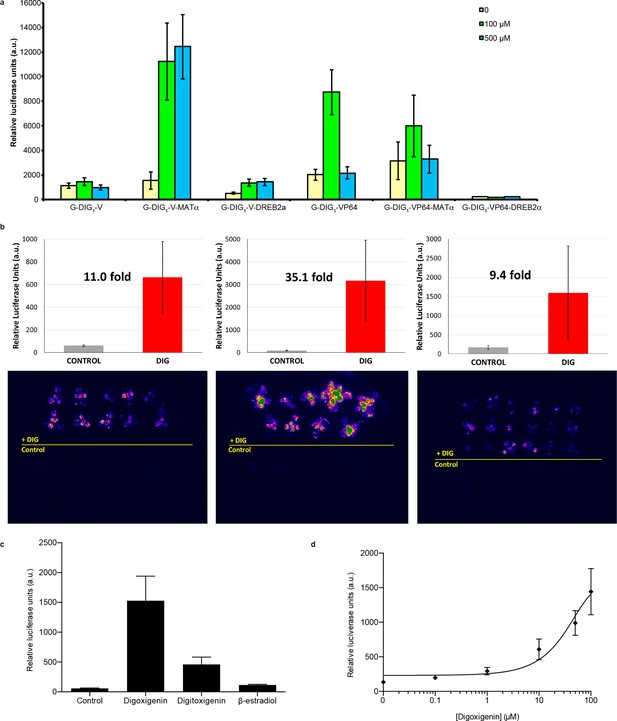
Characterization of DIG biosensor in plants.
(a) Test of DIG1 variants engineered for plant function in Arabidopsis protoplasts. Two transcriptional activation domains, VP16 (V) and VP64 (VP64), as well as two degrons, yeast MATα2 and Arabidopsis DREB2a, were added to DTF-1 (G-DIG1), and the proteins were constitutively expressed from the CaMV35S promoter. The Gal4-activated pUAS promoter controls expression of a luciferase reporter. Transformed protoplasts were incubated with digoxigenin at 0, 100, and 500 μM for 16 hr. (b) Digoxigenin-dependent activation of luciferase expression in three independent transgenic Arabidopsis lines. Plants were incubated in the absence (Control) or presence (DIG) of 100 μM digoxigenin for 42 hr and imaged. Quantification of luciferase expression is presented as mean relative luciferase units ± s.d. of ten plants. (c) Specificity of luciferase activation in transgenic Arabidopsis plants. All inducers were tested at 100 μM concentration. DIG, digoxigenin; DIGT, digitoxigenin; β-EST, β-Estradiol. Data are presented as mean fold activation relative to the control ± s.e.m. of ten technical replicates. (d) Digoxigenin dose response curve in transgenic Arabidopsis plants. Concentrations are expressed in micromolar. The data are presented as mean fold induction relative to the control ± s.e.m. of ten biological replicates.
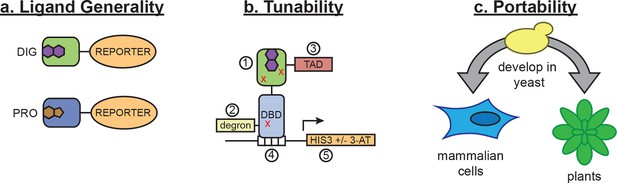
Schematic of biosensor platform.
(a) Biosensors for small molecules are modularly constructed by replacing the LBD with proteins possessing altered substrate preferences. (b) Activity of the biosensor can be tuned by 1) introducing destabilizing mutations (red Xs), 2) adding a degron, 3) altering the strength of the TAD or DNA binding affinity of the TF, 4) changes in the number of TF-binding sites or sequences, and 5) titrating 3-aminotriazole, an inhibitor of His3. (c) Yeast provide a genetically tractable chassis for biosensor development before implementation in more complex eukaryotes, such as mammalian cells and plants.
Additional files
-
Supplementary file 1
Oligonucleotides used in the construction of biosensors are shown.
- https://doi.org/10.7554/eLife.10606.016





