The kinetochore prevents centromere-proximal crossover recombination during meiosis
Figures
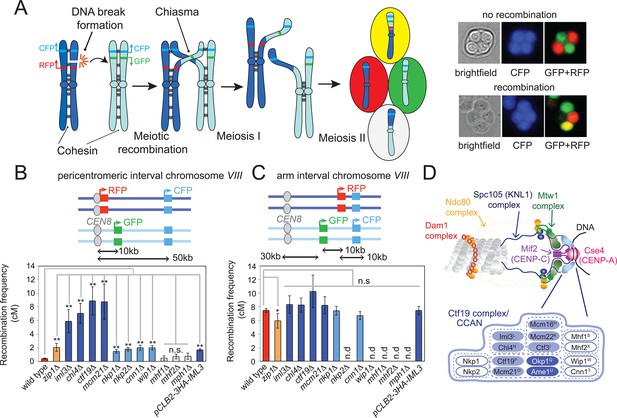
The Ctf19 kinetochore sub-complex represses pericentromeric meiotic recombination.
(A) Scheme of meiosis and the live cell reporter assay to measure CO recombination. Homologous chromosomes are shown in light and dark blue with Green fluorescent protein (GFP), tdTomato (RFP) and m-Cerulean (CFP) reporters represented in green, red and cyan, respectively. Expression of reporters in spores leads to segregation of coloured markers as indicated in the images of live tetrads. (B, C) Map distances (centiMorgans (cM)) and standard error (bars) were determined for a ~10 kb pericentromeric (B) or chromosomal arm (C) interval as described in Materials and methods. p-values were obtained using Fisher’s exact test (* p<0.05; ** p<0.0001). (D) Schematic representation of the kinetochore showing yeast Ctf19 sub-complex components with superscripts indicating the centomere protein (CENP) equivalent in humans. Proteins essential for vegetative growth or proper spore viability after meiosis are shown in dark and light blue, respectively. CO, crossover; n.d., not determined.
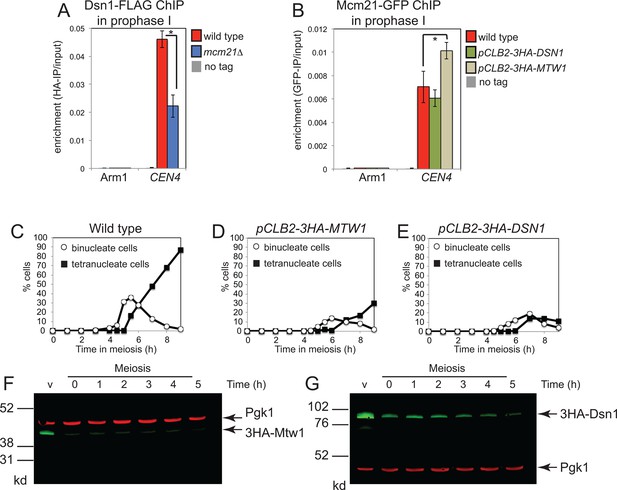
Partial inter-dependence of the Ctf19 complex and MIND subunits.
(A) The Ctf19 component, Mcm21, is required for proper levels of Dsn1 at the centromere during prophase I. Strains AM11633 (no tag), AM20078 (DSN1-6HIS-3FLAG) and AM20080 (DSN1-6HIS-3FLAG mcm21△) all carrying ndt80△ were harvested 5 hr after resuspension in sporulation medium to induce a prophase I arrest and Dsn1 levels at CEN4 and a chromosomal arm site were measured by anti-FLAG ChIP-qPCR. (B) Effect of depleting the MIND components, Mcm21 or Dsn1, on centromeric levels of the Ctf19 component, Mcm21. Strains AM11633 (no tag), AM20296 (MCM21-yeGFP), AM20295 (MCM21-yeGFP pCLB2-3HA-DSN1), AM20294 (MCM21-yeGFP pCLB2-3HA-MTW1) all carrying ndt80△ were treated as in (A) except that anti-GFP ChIP-qPCR was performed. In (A) and (B) the mean of four biological replicates is shown with error bars indicating standard error. p values were obtained using a paired t test. * p<0.05. (C–G) Nuclear division and efficiency of protein depletion in pCLB2-3HA-MTW1 and pCLB2-3HA-DSN1 strains. Wild type (AM1835), pCLB2-3HA-MTW1 (AM20084) and pCLB2-3HA-DSN1 (AM20082) strains were induced to sporulate and the percentages of binucleate and tetranucleate cells were determined after scoring 100 cells at each of the indicated times (C–E). (F, G) The amount of 3HA-Dsn1 (F) or 3HA-Mtw1 (G) was analysed by anti-HA western immunoblot at the indicated times. Pgk1 is shown as a loading control. ChIP, Chromatin immunoprecipitation; MIND, Mtw1 including Nnf1-Nsl1-Dsn1; qPCR, quantitative polymerase chain reaction
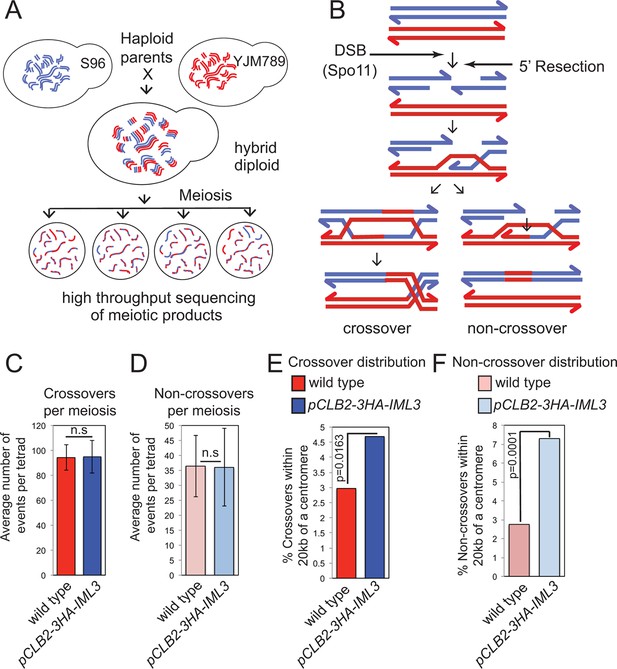
Genome-wide analysis shows that a functional Ctf19 complex is required to prevent both pericentromeric COs and NCOs.
(A) Assay to measure meiotic recombination genome-wide by analysis of SNPs after high-throughput sequencing of germinated spores resulting from hybrid meiosis. (B) Scheme of meiotic recombination showing CO and NCO outcomes. (C, D) Overall recombination levels are not affected by the pCLB2-3HA-IML3 mutation. The average numbers of COs (C) and NCOs (D) per tetrad are not significantly different between wild type and pCLB2-3HA-IML3 cells. Error bars represent standard deviation. A two-tailed t test indicated non-significance (p>0.05). (E, F) Both COs (E) and NCOs (F) within 20 kb of the pericentromere are increased in pCLB2-3HA-IML3 cells. Data for wild type is from (Oke et al., 2014). Number of meioses scored was 8 for pCLB2-3HA-IML3 and 52 for wild type. p-values were calculated using chi-square test with Yates correction. CO, crossover; DSB, double strand break; NCO, non-crossover; SNPs, single nucleotide polymorphisms.
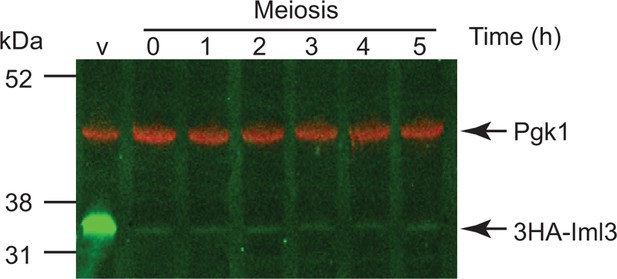
Depletion of Iml3 during meiosis.
Placement of IML3 under the control of the CLB2 promoter results in depletion of Iml3 during meiosis. Cells carrying pCLB2-3HA-IML3 (strain AM17552) were induced to undergo meiosis and samples extracted for anti-HA western immunoblot at the indicated times. A cycling, vegetative (V) sample is also shown. Pgk1 is shown as a loading control.
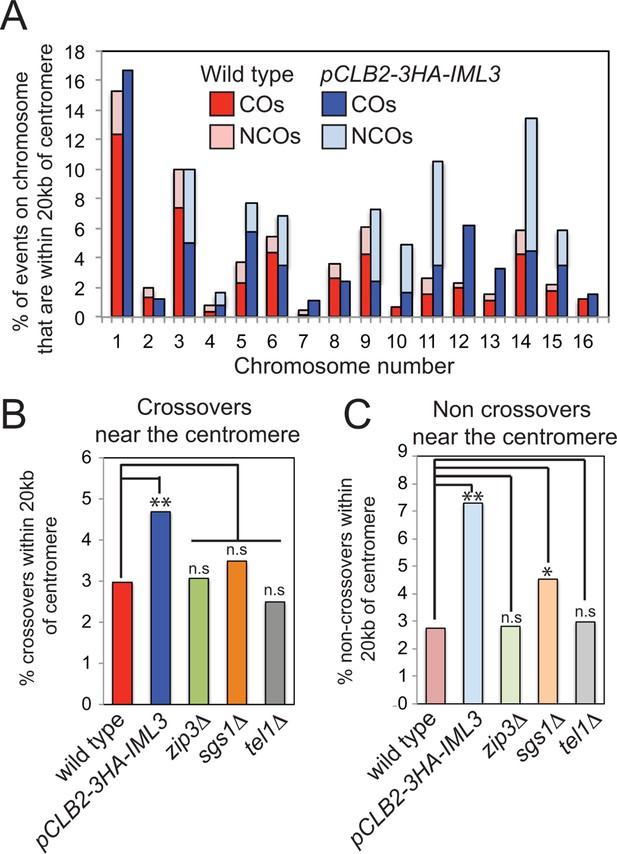
Depletion of Iml3 in meiosis increases the frequency of recombination events within 20 kb of the centromere.
(A) The distribution of the observed events on each chromosome is shown for wild type and pCLB2-3HA-IML3 cells. The percentages of CO or NCO events within 20 kb of the pericentromere were calculated as a fraction of the combined total of CO and NCO events on that chromosome. (B, C) Comparison of the effect of Iml3 depletion on centromeric recombination with other mutants known to affect CO (B) and NCO (C) frequency. Events are shown as a percentage of the combined total of the CO and NCO events per meiosis. Significance was determined using a 2x×2 contingency table and p values were calculated using a Chi square test with Yates correction (** p<0.0001; * p<0.05; n.s., not significant). Data for zip3△, sgs1△ and tel1△ is from (Oke et al., 2014). CO, crossover; NCO, non-crossover.
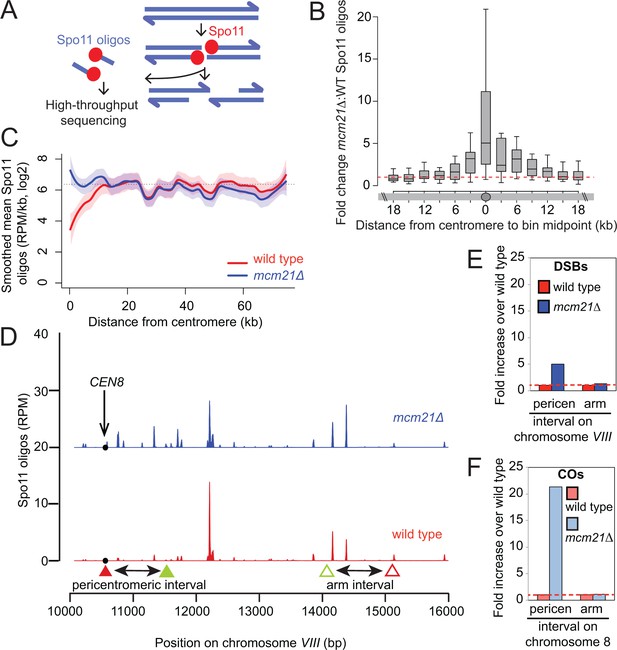
The kinetochore protects the centromere-proximal domain from DSBs.
(A) Sequencing of Spo11-oligos allows DSBs to be mapped genome wide. (B) Fold change in average Spo11-oligo density (RPM per kb) in 3 kb segments in mcm21△ cells compared to wild type (from Zhu and Keeney (2015)), over a 36 kb region surrounding all 16 centromeres. Boxes show median and interquartile range, whiskers enclose data points within 1.5 times the interquartile range; outliers are not shown. Red dashed line, fold change of one. (C) Mean Spo11 signal as a function of distance from the centromere. Spo11 oligo density within 500 bp bins starting from the centromere and moving up to 75 kb away, averaged across the 32 chromosome arms was determined. The horizontal dotted line indicates genome average. The red (wild type) and blue (mcm21△) lines indicate loess smoothing, and the shading indicates the 95% confidence interval. (D) DSBs in the region examined in the CO assay. Spo11 oligo counts smoothed with a 201-bp Hann window are shown. The black circle indicates the centromere, filled triangles indicate the midpoints of coordinates where RFP (red) and GFP (green) cassettes were targeted to for CEN8 analysis in the live cell recombination assay; open triangles indicate the locations where the cassettes were targeted to for ARM8 analysis. (E and F) Fold change in the number of DSBs (E) or COs (F) in mcm21△ vs. wild type within the same pericentromeric or arm intervals on chromosome 8 that were analyzed in the live cell recombination assay (Figure 1B and C). CO, crossover; DSBs, double strand breaks; RPM, reads per million mapped.
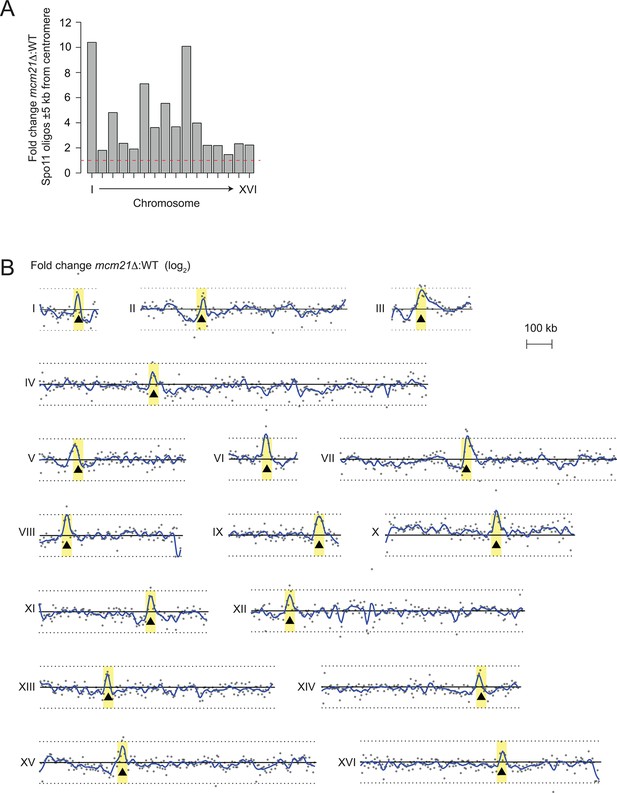
Genome-wide view of meiotic recombination initiation in the mcm21△ mutant.
(A) Fold change in mcm21△ over wild type of Spo11-oligo counts (RPM) in the 10 kb encompassing each centromere. Red dashed line indicates fold change of one. (B) Whole-chromosome view of changes in the Spo11-oligo distribution in mcm21△ . Each point is the fold change (plotted on log2 scale) of mcm21△ over wild type Spo11 oligos (RPM) summed in 5-kb bins. Blue lines, smoothed fit (loess); black triangles, centromeres; yellow shading, centromere ± 20 kb; black solid line, 0 (log2 scale) indicating no change over wild type; black dotted lines, 2-fold change (log2 scale). RPM, reads per million mapped.
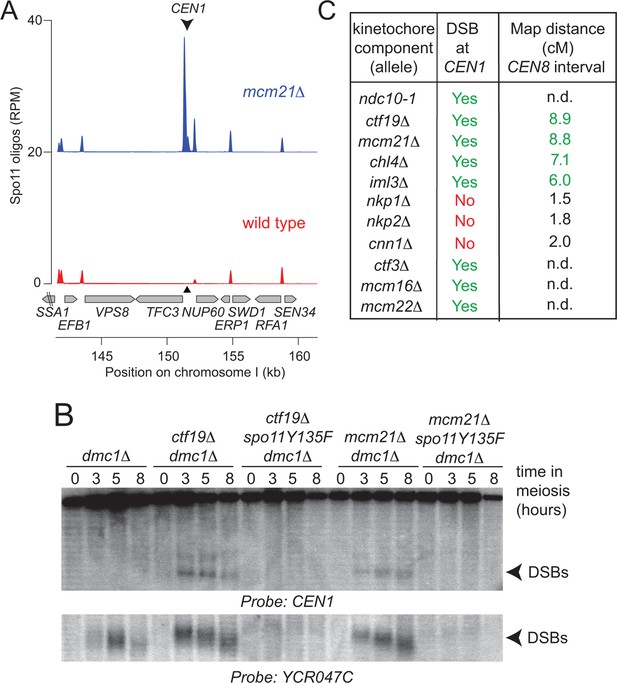
Analysis of DSB formation in Ctf19 complex mutants.
(A) Appearance of a strong DSB hotspot proximal to CEN1 in mcm21△ cells. Spo11-oligo density in a 20 kb region surrounding the centromere of chromosome I in mcm21△ (top) and wild type (bottom). Spo11-oligo counts (RPM) were smoothed with 201-bp Hann window. (B) Detection of CEN1-proximal DSBs in Ctf19 complex mutants by Southern blotting and their dependence on SPO11 catalysis (using a catalytic dead mutant allele of SPO11, spo11-Y135F). Repair-deficient (dmc1△) cells were harvested at defined times after inducing sporulation ((t=0, 3, 5, 8 hr) and faster migrating DNA species (indicative of DSBs) were detected using a probe to CEN1 or the control YCR047C locus. Arrowheads, Spo11-dependent DSBs. Strains used were GV48 (dmc1△), GV1912 (dmc1△ ctf19△), GV2128 (dmc1△ ctf19△ spo11-Y135F-HA), GV2050 (dmc1△ mcm21△) and GV2205 (dmc1△ mcm21△ spo11-Y135F-HA). (C) Summary of tested mutants and their importance for suppression of DSBs and inhibition of COs close to centromeres. n.d., not determined. CO, crossover; DSB, double strand break; RPM, reads per million mapped.
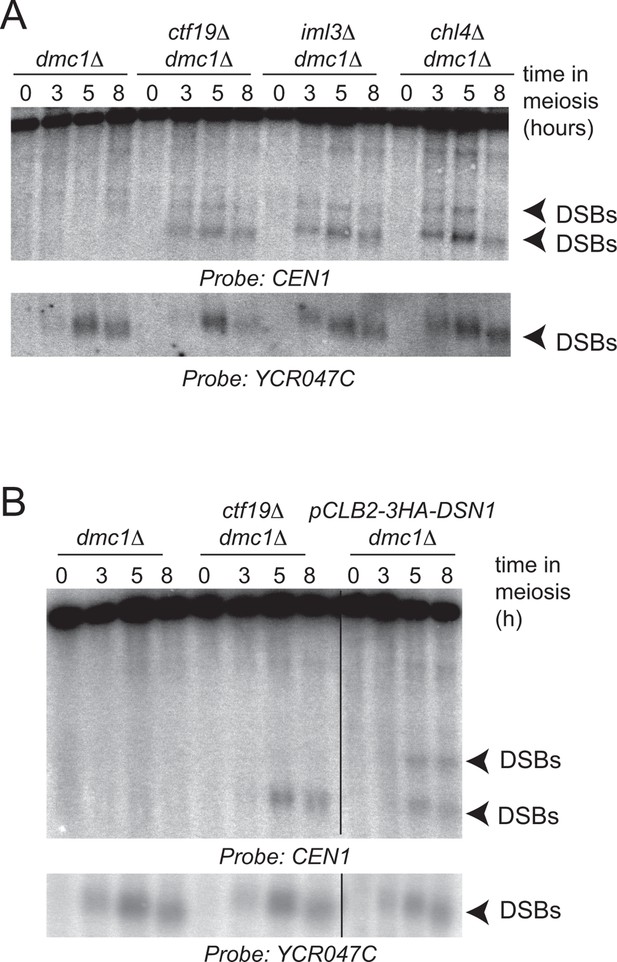
Ctf19 complex components are required to prevent DSB formation close to CEN1.
Repair-deficient (dmc1△) cells (strains GV48 (dmc1△), GV1912 (dmc1△ ctf19△), GV1870 (dmc1△ iml3△) and GV2139 (dmc1△ chl4△)) were harvested at defined times after inducing sporulation (t=0, 3, 5, 8 hr) and faster migrating DNA species (indicative of DSBs) were detected using a probe to CEN1 or the control YCR047C locus. Arrowheads, Spo11-dependent DSBs. DSBs, doublestrand breaks

Ctf19-dependent cohesin enrichment prevents pericentromeric COs.
(A–D), The Ctf19 complex enriches meiotic cohesin in the pericentromere during prophase I. (A–C) Wild type (AM4015), iml3△ (AM4016) and mcm21△ (AM13833) strains carrying REC8-3HA and ndt80△ were harvested 5h after resuspension in sporulation medium and Rec8 association was analyzed by ChIP-Seq. Rec8 association with chromosome V and a close up of the 50 kb pericentromeric interval is shown (A). The median Rec8 level for all 16 pericentromeric regions is shown over a 25 kb region on each side of the centromere for iml3△ (B) and mcm21△ (C) compared to wild type. (D) Strains as in (A) together with chl4△ (AM4017), ctf19△ (AM20086) and a no tag control (AM11633) carrying ndt80△ were arrested in prophase I by harvesting 5h after being induced to sporulate. The level of Rec8 at the indicated sites was determined by anti-HA ChIP-qPCR. Primer sets used corresponded to sites on the arm of chromosome IV (arm1, 2, 3), within the 20 kb pericentromere (pericen1, 2) or ~150bp from CEN4 (CEN4) and sequences and coordinates are given in Supplementary file 4B. Error bars represent standard error (n=4 biological replicates for iml3△, chl4△, ctf19△ and mcm21△; n=8 for no tag and wild type). *p<0.05, unpaired t test. (E) Chromosomal Rec8 levels are reduced in scc4-m35 cells. Wild type (AM4015), scc4-m35 (AM18211) cells and a no tag control (AM11633) carrying ndt80△ were arrested in prophase I by harvesting 5 hr after being induced to sporulate. The level of Rec8 at the indicated sites was determined by anti-HA ChIP-qPCR. Error bars represent standard error (n=4 biological replicates). *p<0.05, paired t test. (F, G) Map distances (in cM) in the pericentromere (F) or a chromosome arm (G) interval in wild type, a control SCC4 replacement strain, or the scc4-m35 mutant were determined and their significance analysed as described in Figure 1. ChIP-Seq, chromatin immunoprecipitation with sequencing.
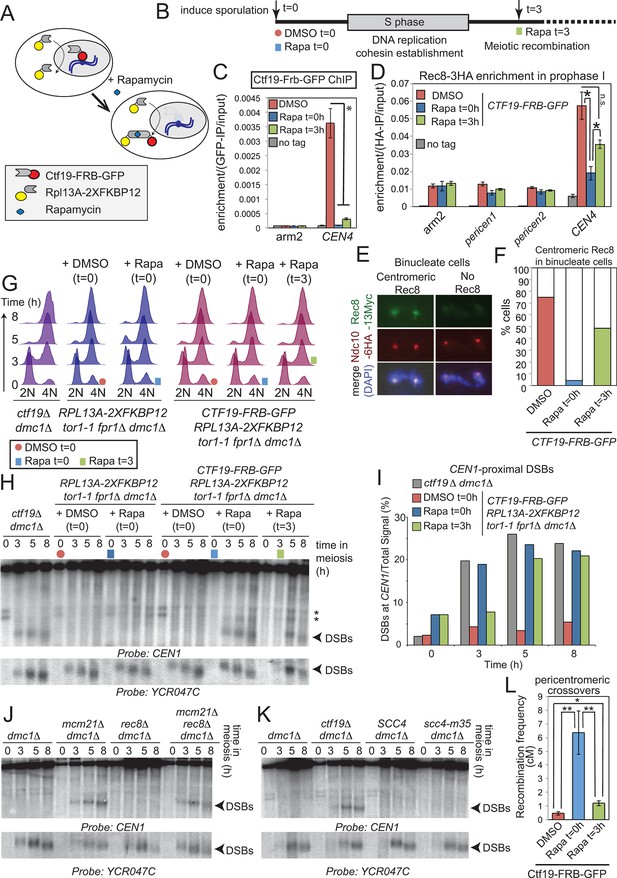
Pericentromeric cohesin does not prevent DSBs, but ensures their repair does not form COs.
(A–D) Anchoring away Ctf19 after DNA replication and cohesion establishment is not sufficient to prevent the appearance of centromere-proximal DSBs. (A, B) Scheme of the anchor away system and experimental setup used to deplete Ctf19 during meiosis. (C) Addition of Rapamycin leads to Ctf19 removal from the pericentromere. Three cultures of strain AM18978 (CTF19-FRB-GFP RPL13A-2XFKBP12 tor1-1 fpr1△ ndt80△ REC8-3HA) were induced to sporulate. Either DMSO or Rapamycin were added to two of the cultures (t=0). Rapamycin was added 3 hr after inducing sporulation to the third culture (t=3). A fourth culture was a no tag ndt80△ control (AM11633) to which DMSO was added. All cultures were harvested 5 hr after inducing sporulation (prophase I arrest) and Ctf19 levels were analyzed by anti-GFP ChIP-qPCR at the indicated sites. Error bars represent standard error (n=4 biological replicates). *p<0.05, paired t-test. See Figure 5 and Supplementary file 4B for details of primer sets used. (D–F) Anchoring away Ctf19 after DNA replication allows establishment of centromeric Rec8. (D) Cells treated as in (C) were processed for anti-HA ChIP-qPCR at the indicated sites. Error bars represent standard error (n=4 biological replicates). *p<0.05, paired t-test. (E and F) Three cultures of strain AM20138 (CTF19-FRB-GFP NDC10-6HA pGAL-NDT80 pGPD1-GAL4(848)-ER RPL13A-2XFKBP12 tor1-1 fpr1△ ndt80△ REC8-3HA) were resuspended in sporulation medium (t=0). DMSO or Rapamycin were immediately added to the first and second cultures (t=0), while the third culture received Rapamycin after 3 hr incubation (t=3). After 6 hr total, β-estradiol was added to release cells from the prophase I arrest, samples were harvested at 15 min intervals and chromosome spreads were prepared and stained with anti-HA and anti-Myc antibodies. (E) Examples of binucleate cells with centromeric or no Rec8. (F) Percentages of binucleate cells with centromeric Rec8 are shown for indicated conditions. (G–I) Timed depletion of Ctf19 to test DSB formation dependencies. (G) DNA replication is largely complete prior to anchoring Ctf19 away in cultures where Rapamycin was added at 3 hr. A control strain GV2367 (RPL13A-2XFKBP12 tor1-1 fpr1△ dmc1△) and equivalent experimental strain carrying CTF19-FRB-GFP (GV2354) were induced to undergo meiosis together with a ctf19△ dmc1△ mutant (GV1912). Rapamycin or DMSO were added at the indicated times (red circles, addition of DMSO at t=0; blue squares, addition of Rapamycin at t=0; green squares, addition of Rapamycin at t=3) and samples were processed for FACS analysis (timepoints t=0, 3, 5, 8 hr). (H) Analysis of DSB formation in the experiment shown in (B). Southern blot shows that DSB formation close to CEN1 occurs either when Ctf19 is anchored away early (t=0) or after DNA replication and cohesin enrichment (t=3). Red circles, addition of DMSO at t=0; blue squares, addition of Rapamycin at t=0; green squares, addition of Rapamycin at t=3; Arrowheads, Spo11-dependent DSBs; asterisks, cross-hybridizing species. (I) Quantification DSBs shown in (H). (J, K) Inhibition of centromere-proximal DSBs does not depend on cohesin. (J) CEN1-proximal DSBs are observed in rec8△ cells only upon deletion of Ctf19 complex components. Strains GV48 (dmc1△), GV2050 (dmc1△ mcm21△), GV2403 (dmc1△ rec8△), GV2286 (dmc1△ mcm21△ rec8△) were analyzed by Southern blotting as described in Figure 4B. (K) Cohesin impairment does not allow CEN1-proximal DSB formation. Strains used were GV48 (dmc1△), GV1912 (dmc1△ ctf19△), GV2305 (dmc1△ SCC4) and GV2533 (dmc1△ scc4-m35). (L) Anchoring away Ctf19-FRB after DNA replication and cohesin establishment leads to only a modest increase in pericentromeric COs. Three cultures of strain AM19543 [carrying heterozygous pericentromeric RFP/GFP reporters separated by ~10 kb, homozygous chromosomal arm CFP reporters (Figure 1B), estradiol-inducible Ndt80 to allow release from prophase I arrest (pGAL-NDT80 pGPD1-GAL4(848)-ER) and the Ctf19 anchor away system (CTF19-FRB RPL13A-2XFKBP12 tor1-1 fpr1△)] were resuspended in sporulation medium (t=0). DMSO or Rapamycin were immediately added to the first and second cultures (t=0), while the third culture received Rapamycin after 3 hr incubation (t=3). All cultures were incubated for 6 hr total before being released from the prophase I arrest by addition of β-estradiol and tetrads were scored after incubation for 48 hr total. ChIP, chromatin immunoprecipitation; DMSO, dimethyl sulfoxide; DSBs, double strand breaks; FACS, fluorescence-activated cell sorting; qPCR, quantitative polymerase chain reaction.
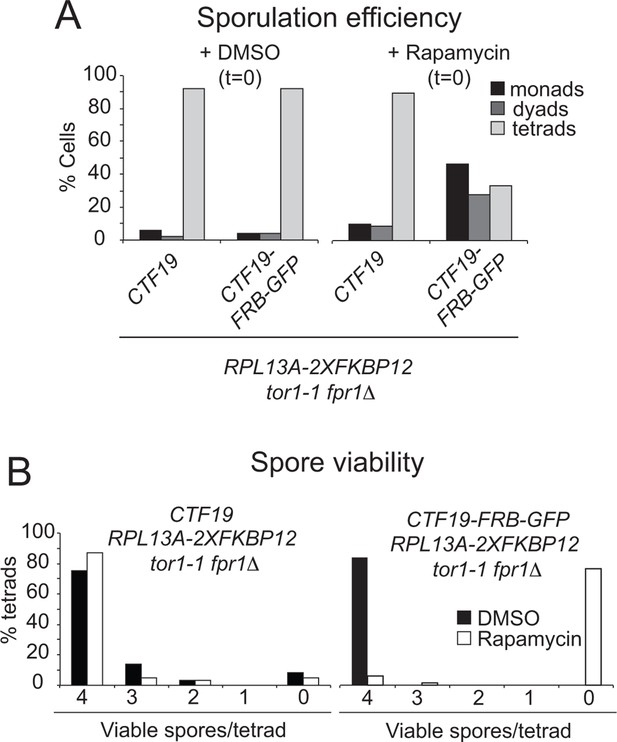
Anchoring Ctf19 away early in meiosis results in reduced sporulation efficiency and spore viability.
RPL13A-2xFKBP12 tor1-1 fpr1△ strains carrying FRB-tagged (CTF19-FRB-GFP; GV2275) or untagged Ctf19 (CTF19; GV1853) were treated with Rapamycin or DMSO, as a control, upon resuspension in sporulation medium (t=0). (A) The percentages of cells producing monads (1 spore), dyads (2 spores) or tetrads (4 spores) was scored (n=200). (B) The number of spores germinating per tetrad was scored after dissection. At least 60 spores were analysed per condition.
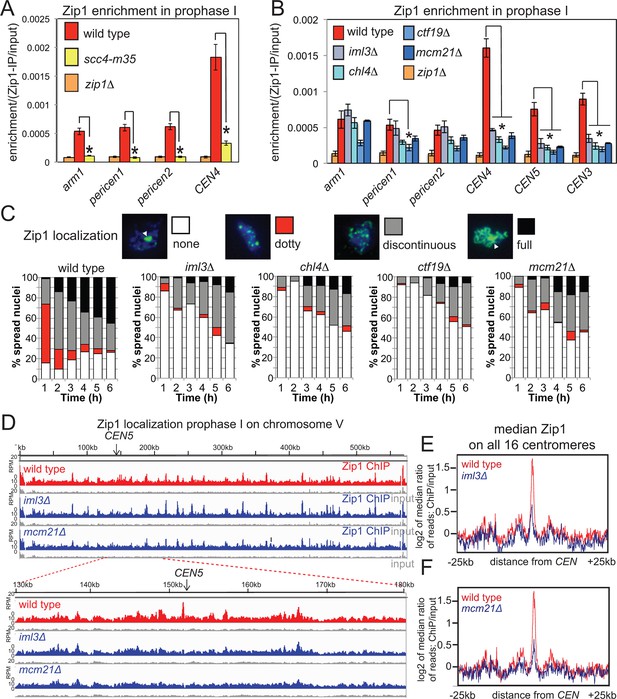
Cohesin enables centromeric Zip1 recruitment.
(A) Zip1 enrichment on chromosomes is reduced in scc4-m35 mutants. Wild type (AM11633), scc4-m35 (AM18881) and zip1△ (AM10913) cells carrying ndt80△ were induced to sporulate and harvested after 5 hr (prophase I) arrest for anti-Zip1 ChIP-qPCR. Error bars represent standard error (n=4 biological replicates). p<0.05, paired t test. See Figure 5 and Supplementary file 4B for details of primer sets used. (B-–F) The Ctf19 complex is required for Zip1 localization at centromeres. (B) ChIP-qPCR analysis of Zip1 localization in prophase I in Ctf19 complex mutants. Wild type (AM11633), iml3△ (AM10686), chl4△ (AM10658), ctf19△ (AM10660), mcm21△ (AM10664) and zip1△ (AM10913) cells carrying ndt80△ were induced to sporulate and harvested after 5 hr for anti-Zip1 qPCR. Error bars represent standard error (n=3 biological replicates). p<0.05, paired t test. See Figure 5 and Supplementary file 4B for details of primer sets used. (C) Analysis of Zip1 localization on chromosome spreads as cells progress into prophase I. Examples of Zip1 localization on chromosome spreads are shown with Zip1 in green and DNA in blue. Categories of Zip1 localization were scored in 100 spread nuclei at each of the indicated times after resuspension in sporulation medium. Strains used carried NDC10-6HA, pGAL-NDT80, pGPD1-GAL4-ER and were AM8769 (wild type), AM8772 (iml3△), AM8770 (chl4△), AM9049 (ctf19△) and AM8861 (mcm21△). (D–F) ChIP-Seq analysis of Zip1 localization during prophase I in wild type, iml3△ and mcm21△ strains carrying ndt80△ (B) and harvested 5 hr after resuspension in sporulation medium. (D) Zip1 localization along chromosome V is shown as an example with the 50 kb region around the centromere amplified. (E, F) Median Zip1 localization over a 50 kb domain surrounding all 16 centromeres is shown compared to wild type for iml3△ (E) and mcm21△ (F). ChIP, chromatin immunoprecipitation; qPCR, quantitative polymerase chain reaction.
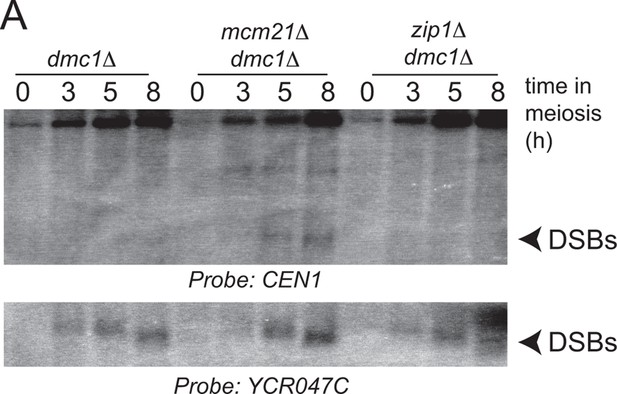
Zip1 does not induce pericentromeric DSB formation.
Repair-deficient (dmc1△) cells (strains GV48 (dmc1△), GV2050 (dmc1△ mcm21△), GV2734 (dmc1△ zip1△) were harvested at defined times after inducing sporulation ((t=0, 3, 5, 8 hr) and faster migrating DNA species (indicative of DSBs) were detected using a probe to CEN1 or the control YCR047C locus. Arrowheads, Spo11-dependent DSBs. DSBs, double strand breaks.
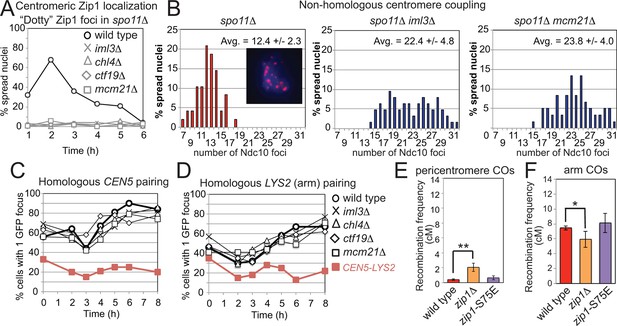
Recruitment of Zip1 to centromeres by the Ctf19 complex independently promotes centromere coupling and suppression of pericentromeric COs.
(A, B) Analysis of Zip1 and kinetochore foci (Ndc10-6HA) on chromosome spreads in spo11△ cells progressing into meiotic prophase I. The percentages of cells with “dotty” Zip1 foci, representing kinetochores, were scored in 100 spread nuclei at each of the indicated times after resuspension of strains spo11△ (AM9018), spo11△ iml3△ (AM9288), spo11△ chl4△ (AM9287), spo11△ ctf19△ (AM9017) and spo11△ mcm21△ (AM8861) in sporulation medium (A). The number of Ndc10 kinetochore foci per spread nucleus was scored in the indicated strains at the 6 hr time point with the average and standard deviation indicated. n=48 (spo11△), 63 (spo11△ iml3△) and 60 (spo11△ mcm21△). (B) The inset shows Ndc10-6HA staining (red) in an example nucleus. DNA is shown in blue. (C) The Ctf19 complex is not required for SPO11-dependent homologous pairing. Pairing of homozygous CEN5-GFP (C) or LYS2-GFP (D, arm) foci was scored for 100 cells at each of the indicated times after resuspension in sporulation medium in ndt80△ cells. Heterologous CEN5-GFP and LYS2-GFP foci labels were used as a control for spurious interactions (strain AM12823, red filled squares). Strains used in (C) were AM12829 (wild type), AM13348 (iml3△), AM12466 (chl4△), AM13346 (ctf19△) and AM12837 (mcm21△). Strains used in (D) were AM12469 (wild type), AM12978 (iml3△), AM12980 (chl4△), AM12831 (ctf19△) and AM12825 (mcm21△). A representative experiment is shown. (E, F) The centromere-coupling function of Zip1 is separable from its role in suppression of recombination in the pericentromere. Recombination frequency in the pericentromere (E) or chromosomal arm (F) interval on chromosome VIII for the centromere-coupling defective zip1-S75E mutant is shown together with data for wild type and zip1△ reproduced from Figure 1B,C. COs, crossovers.
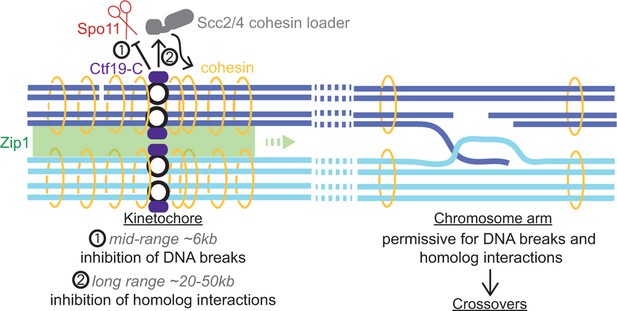
Schematic diagram indicating dual repression of pericentromeric recombination by the Ctf19 kinetochore sub-complex.
(1) Mid-range (~6 kb) suppression of DSBs. (2) Long range (~20-50 kb) inhibition of COs by cohesin and Zip1, directing repair of rare centromere-proximal breaks to the sister chromatid to avoid deleterious pericentromeric inter-homolog chiasmata.
Additional files
-
Supplementary file 1
Raw data and analyses of map distances in the pericentromeric interval using the fluorescence live cell reporter assay.
- https://doi.org/10.7554/eLife.10850.019
-
Supplementary file 2
Raw data and analyses of map distances in the arm interval using the fluorescence live cell reporter assay.
- https://doi.org/10.7554/eLife.10850.020
-
Supplementary file 3
Analysis of genome-wide recombination.
Source data used to generate Figure 2 and Figure 2—figure supplement 2.
- https://doi.org/10.7554/eLife.10850.021
-
Supplementary file 4
Yeast strains and qPCR primers used in this study.
(A) List of yeast strains. (B) List of qPCR primers.
- https://doi.org/10.7554/eLife.10850.022





