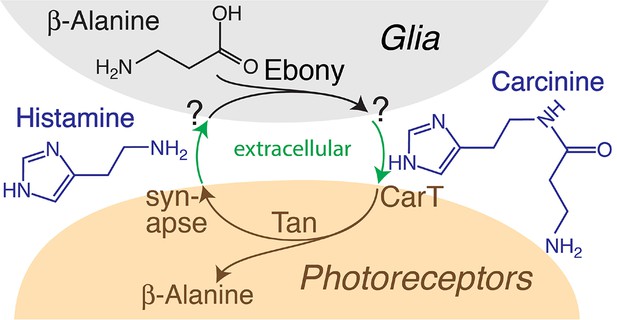
The carcinine transporter CarT is required in Drosophila photoreceptor neurons to sustain histamine recycling
Figures
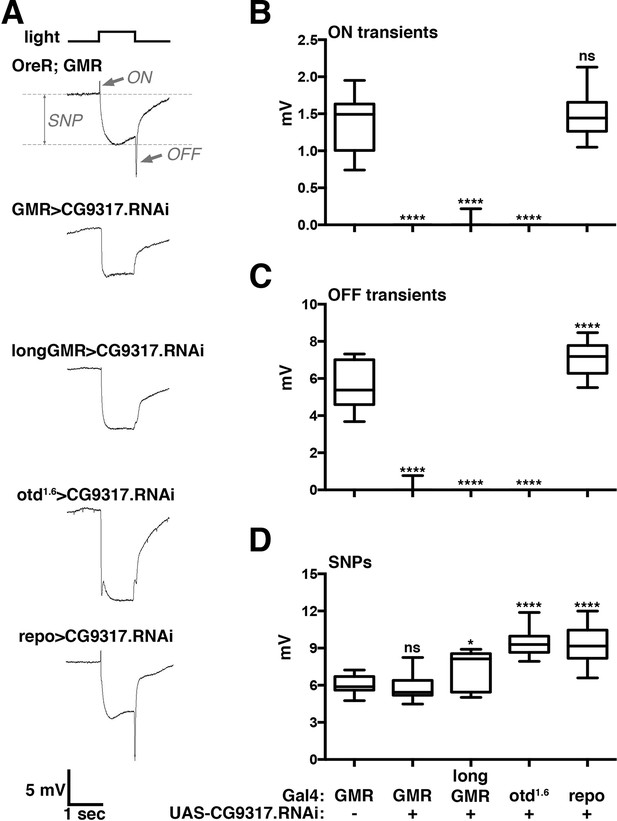
Photoreceptor specific knockdown of CG9317 blocks visual neurotransduction.
(A) ERGs recorded from female flies expressing a UAS-CG9317 RNAi transgene (VDRC 101145) targeting CG9317 under the control of the indicated Gal4 driver specific for photoreceptors (GMR-Gal4, longGMR-gal4 and otd1.6-Gal4) or glia (repo-Gal4). Quantifications of (B) ON transients, (C) OFF transients, and (D) sustained negative photoreceptor potentials were averaged from three replicate experiments, including at least 45 traces from 15 flies. Arrows indicate SNP, ON and OFF transient of the control recording. Graphs report upper and lower quartiles (box) and minimum and maximum values (whiskers). ns, not significant; *p < 0.05, ****p < 0.0001 compared to OreR;GMR-Gal4 control. ERGs, electroretinograms; SNPs, sustained negative potentials.
-
Figure 1—source data 1
List of transporter genes tested by GMR-Gal4 driven knockdown.
- https://doi.org/10.7554/eLife.10972.004
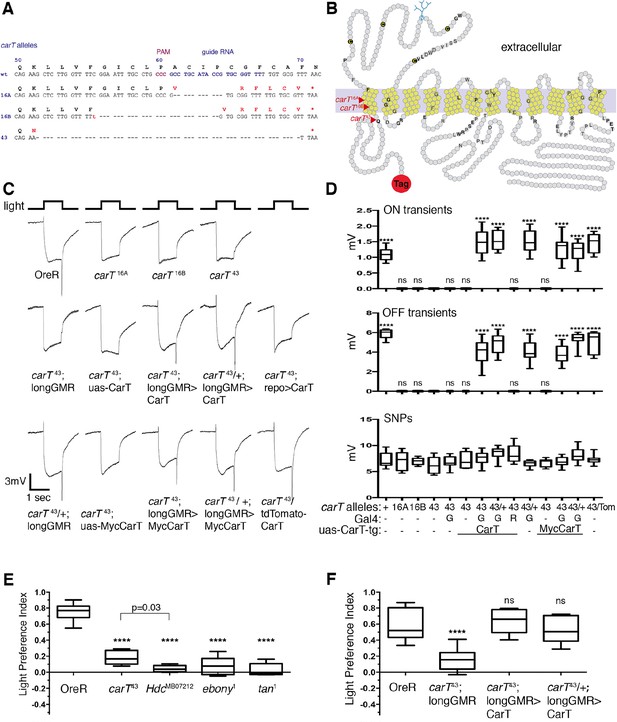
CarT is required in photoreceptors but not glia for normal cellular and behavioral visual responses.
(A) CRISPR targeted site in the carT gene and resulting mutations. (B) Primary structure and predicted transmembrane domains (yellow) of CarT. Red arrows indicate location of premature stops within mutant alleles. Gray and black letters indicate conserved and highly conserved amino acids among SLC22 transporters (Eraly et al., 2004; Koepsell, 2013), respectively. The SLC22 family is characterized by a large extracellular loop containing four cysteines (C) and a glycosylation site (blue) at conserved positions. (C) ERGs recorded from female flies carrying wild type (+) or the indicated CarT alleles (16A, 16B, 43, or tdTomato-CarT) with or without expression of a wild-type or a Myc-tagged UAS-CarT transgene driven by longGMR-Gal4 or repo-Gal4 as indicated. (D) Quantifications of ON and OFF transients, and SNPs of all genotypes in panel C averaged from three replicate experiments, including at least 45 traces from 15 flies (G: longGMR-Gal4; R: repo-Gal4; -: not present). (E) Phototactic behavior of OreR and carT43 mutant flies compared with other mutants that disrupt the histamine–carcinine cycle (HdcMB07212, ebony1, or tan1) presented as a light preference index. (F) Phototactic behavior of flies expressing a UAS-CarT transgene in photoreceptor neurons under control of the longGMR-Gal4 driver in carT43 mutants compared with controls shows the restoration of wild type behavior. For each graph, the box outlines the upper and lower quartiles, and the whiskers show minimum and maximum recorded values. ns, not significant; ****p < 0.0001 compared with carT43 (D) or OreR (E and F). CRISPR, clustered regularly-interspaced short palindromic repeats; ERG, electroretinograms; SNPs, sustained negative potentials.
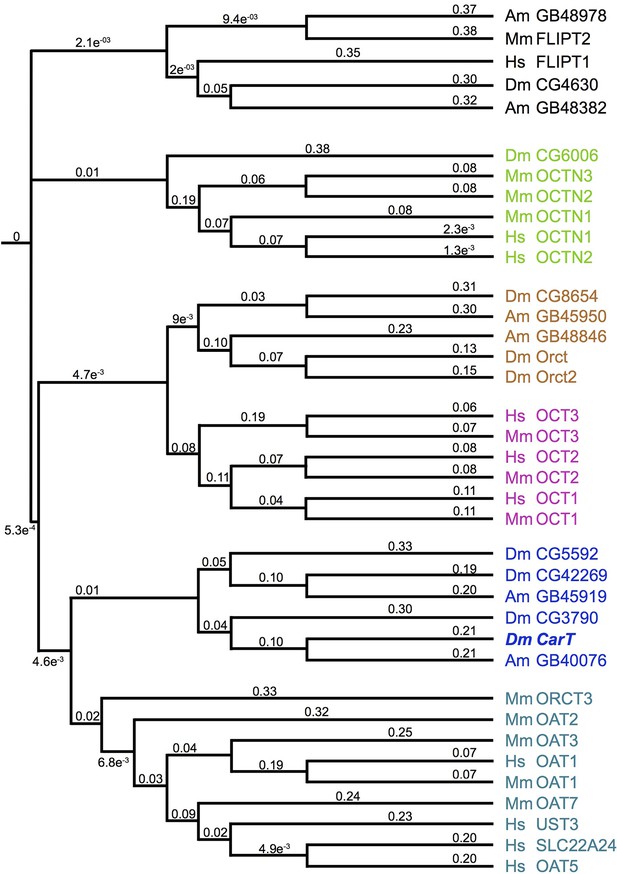
Phylogeny of the Drosophila SLC22 transporters.
Drosophila and bee SLC22 transporters were identified using the Basic Local Alignement Search Tool Protein (BLASTp) provided by the National Center for Biotechnology Information (NCBI) based on similarity to known mammalian and insect family members (Koepsell, 2013; Zhu et al., 2015). A phylogeny tree was obtained using Clustal W2 and visualized using iToL (Letunic and Bork, 2011). Numbers represent simulated branch lengths.
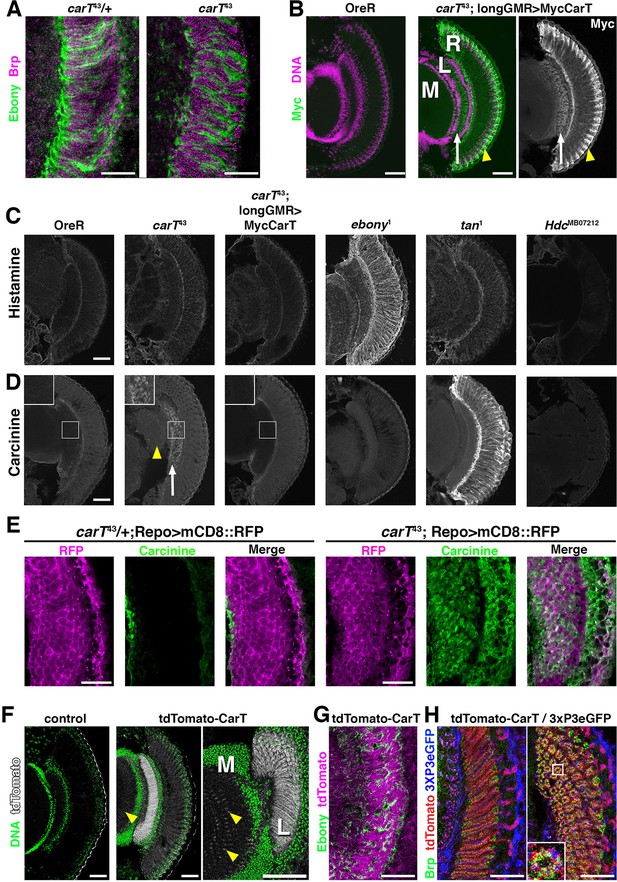
Loss of CarT in photoreceptors increases laminal carcinine.
(A) Confocal sections of photoreceptor axonal endings within the lamina stained for the synapse marker Bruchpilot (Brp) and the glia-specific Ebony. (B) Micrographs of cryo-sections from control (OreR) and flies expressing the UAS-Myc-CarT transgene driven by longGMR-Gal4 in the carT43 background. Retina (R), lamina (L), and medulla (M) neuropiles are stained for DNA (magenta) and Myc-tagged CarT (green). Arrow points to Myc staining in photoreceptor axonal endings and arrowhead to distal retina. (C,D) Micrographs of cryo-sections from control (OreR) and mutant flies affecting the histamine–carcinine cycle: carT43, carT43;GMR-Gal4/UAS-Myc-CarT, ebony1, tan1, and HdcMB07212 stained for histamine (C) or carcinine (D). Arrow and arrowhead in D point to carcinine accumulations in the lamina and medulla, respectively. (E) Micrographs from control (carT43/+ ) or carT43 flies expressing a UAS-mCD8::RFP transgene driven by repo-Gal4 and stained for carcinine. (F) Micrographs showing tdTomato fluorescence of the in-frame tdTomato-CarT allele compared with wild type control. DNA staining is shown in green. Arrowheads point to tdTomato signal at R7 and R8 photoreceptor terminals within the medulla. (G) Confocal section of laminal region of a tdTomato-CarT fly stained for glia-specific Ebony. (H) Confocal sections of flies heterozygous for tdTomato-CarT (red) and photoreceptor-specific 3xPax3-eGFP (blue) stained for Brp (green). Sections are parallel to and across photoreceptor axons, respectively. Scale bars are 20 µm in A, E, G and H and 50 µm in B–D and F.
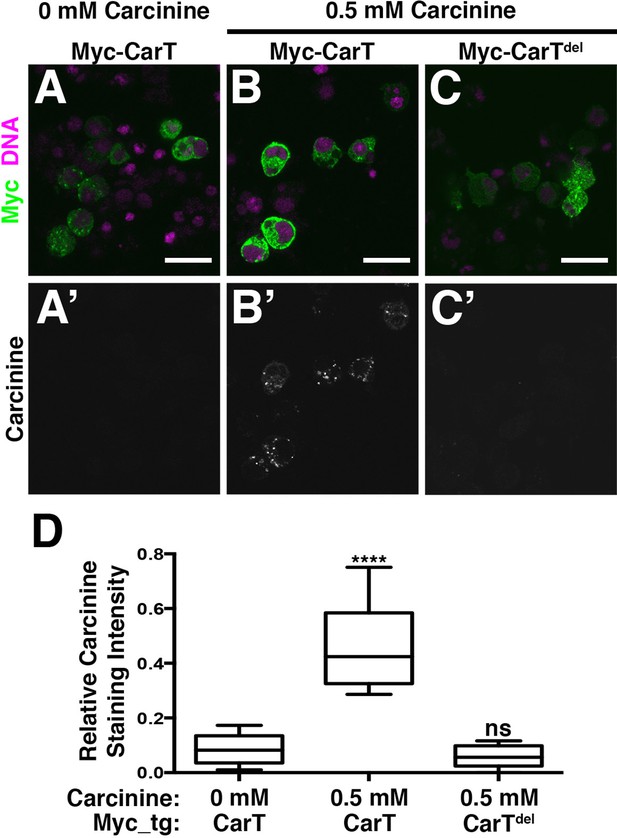
CarT is a carcinine transporter.
(A) Micrographs of S2 cells transfected with the Myc-CarT transgene (A,A’ and B, B’) or the Myc-CarTdel transgene deleted for 10 internal transmembrane domains (C,C’) cultured in media lacking carcinine (A,A’) or supplemented with 0.5 mM carcinine (B,B’ and C,C’) and stained for DNA and the Myc epitope (A–C) or carcinine (A’–C’). (D) Quantification of carcinine signals normalized to signals for the Myc epitope. ****p < 0.001.