Nanobodies: site-specific labeling for super-resolution imaging, rapid epitope-mapping and native protein complex isolation
Figures
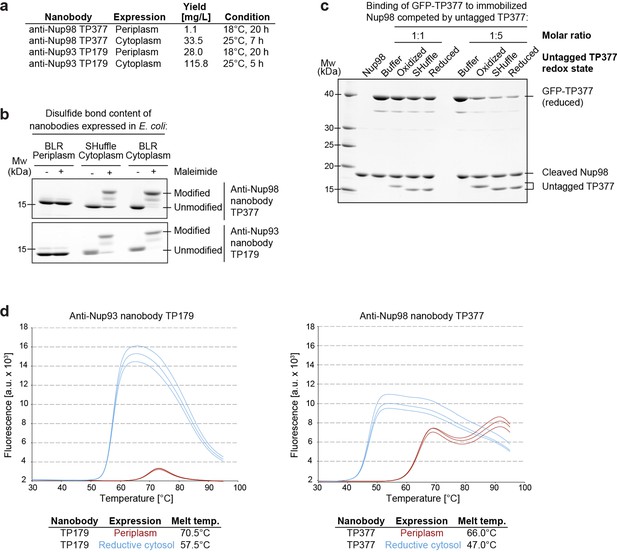
Affinity and thermostability of reduced and oxidized nanobodies.
(a) Comparison of typical yields for the anti-Nup93 nanobody TP179 and the anti-Nup98 nanobody TP377 expressed either in the Escherichia coli BLR periplasm with a C-terminal His6-tag or in the oxidative cytoplasm of E. coli SHuffle with an N-terminal His14-bdNEDD8-tag. (b) Analysis of disulfide bond content using a maleimide shift assay. Anti-Nup93 nanobody TP179 and anti-Nup98 nanobody TP377, expressed either in the oxidative periplasm of E. coli BLR, the oxidative cytosol of E. coli SHuffle or in the reductive cytoplasm of E. coli BLR, were subjected to modification with biotin-PEG23-maleimide in SDS–PAGE sample buffer (-DTT) and analyzed by non-reducing SDS–PAGE followed by Coomassie staining. (c) The redox state of the anti-Nup98 nanobody TP377 does not affect the affinity for its target. Biotinylated His14-Avi-bdSUMO-tagged Nup98716-866 was immobilized on Streptavidin agarose und used to bind the reduced GFP-tagged TP377. Binding was in the absence or presence of an equimolar amount or fivefold excess of nanobody competitor, namely untagged TP377 produced either in the oxidative periplasm, in the mildly oxidative cytoplasm of E. coli SHuffle or in the reductive cytoplasm of BLR. Bound nanobodies were then eluted by proteolytic cleavage of the bdSUMO tag of Nup98 and analyzed by SDS–PAGE followed by Coomassie staining. Note that the oxidized, disulfide bond-stabilized nanobody (produced in the periplasm) behaved like the reduced variant (produced in the E. coli BLR cytoplasm). Formation of the disulfide bond therefore does not seem to significantly contribute to the overall affinity. (d) Differential scanning fluorimetry (thermofluor, Niesen et al., 2007) analysis of nanobodies expressed in the oxidative periplasm (red) or the reductive cytosol (blue) of E. coli BLR. The anti-Nup93 and anti-Nup98 nanobodies were heated in the presence of Sypro Orange dye from 30 to 100°C and thermal unfolding curves were obtained. The melting temperature is derived from the inflection point of the curve.
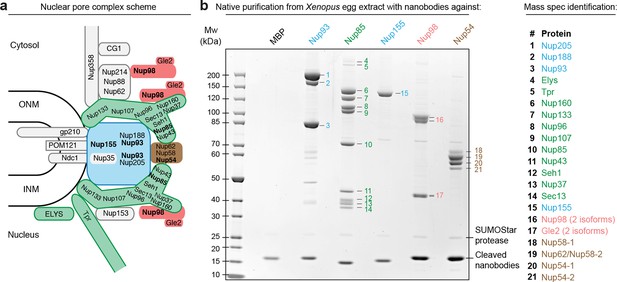
Purification and native elution of NPC subcomplexes with specific nanobodies.
(a) Schematic representation of the subcomplex organization and relative localization of Nups within an asymmetric unit of the eightfold rotational symmetric vertebrate NPC (ONM/INM = outer and inner nuclear membrane). The nuclear and cytoplasmic rings of the structural NPC scaffold are mainly composed of the Nup107-Nup160 Y-shaped complex (green). The central inner ring of the scaffold is composed of the Nup93 subcomplex (blue). The scaffold is bound to the nuclear envelope via transmembrane Nups and further anchors FG-repeat nucleoporins (e.g. Nup98 [red] and the Nup62•Nup58•Nup54 complex [brown]) within the central channel, where they form the permeability barrier. Nups against which nanobodies were raised are highlighted in bold. (b) Native purification of major NPC scaffold subcomplexes and FG-repeat nucleoporins from Xenopus egg extract. Biotinylated His14-Avi-(GlySer)9-SUMOStar-(GlySer)9-tagged nanobodies were immobilized on magnetic Streptavidin beads and then incubated with Xenopus egg extract. After washing, nanobodies were gently eluted along with their bound target complexes by SUMOStar protease cleavage. One tenth of the eluates were analyzed by SDS–PAGE and Coomassie staining. All labeled bands were identified via mass spectrometry. The color code represents the subcomplex organization of the NPC as illustrated in (a). A nanobody raised against Escherichia coli Maltose-binding protein (MBP) served as a negative control. NPC, nuclear pore complex.
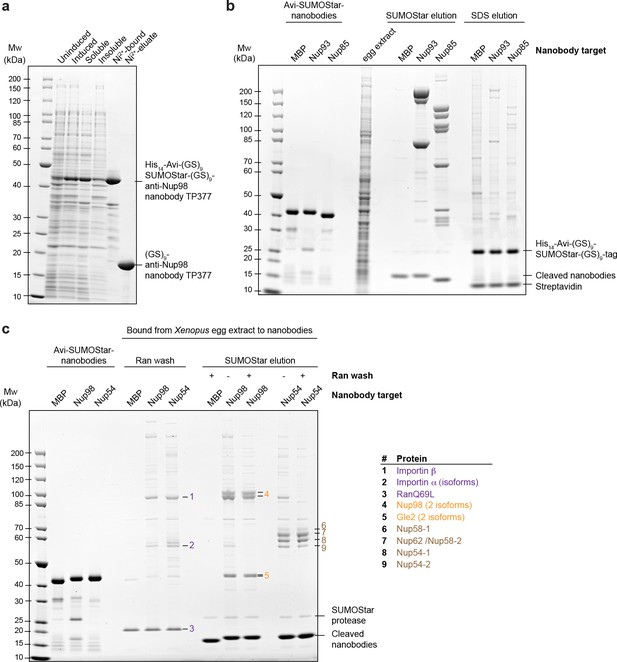
Optimization of native protein complex purification using nanobodies.
(a) SDS–PAGE and Coomassie staining showing the expression of the anti-Nup98 nanobody TP377 carrying a protease-cleavable affinity tag (His14-Avi-(GlySer)9-SUMOStar-(GlySer)9) in the Escherichia coli cytoplasm and its one-step purification using Ni2+ chelate affinity chromatography and imidazole elution. The Avi-tag mediates binding to Streptavidin after biotinylation by the biotin ligase BirA (Beckett et al., 1999; Schatz, 1993). (b) Analysis of natively purified and remaining bead-bound material. Anti-Nup93 and anti-Nup85 nanobodies were used to purify their respective target complexes from crude Xenopus egg extract. After native elution with SUMOStar protease, the beads were heated in SDS–PAGE sample buffer containing 400 mM urea for 10 min at 97°C. Note that protease cleavage released the cognate complexes very efficiently and that the remaining bead-bound material essentially represents just the non-specific background, cleaved tags and leaked streptavidin. (c) Effect of a RanQ69L•GTP wash on FG-repeat Nup purification. FG repeat-bound nuclear transport receptor•cargo complexes were efficiently removed by washing the beads for 10 min at 4°C with 100 µl 1 µM RanQ69L5-180•GTP before elution with SUMOStar protease.

Structural analysis of natively purified Nup93 complexes.
(a) Anti-Nup93 nanobody TP179-purified Nup93•Nup188 and Nup93•Nup205 complexes were subjected to the GraFix procedure (Kastner et al., 2008) and negative staining for analysis by electron microscopy. (b) Gallery of 12 selected class averages of Nup93•Nup188 and Nup93•Nup205 particles.
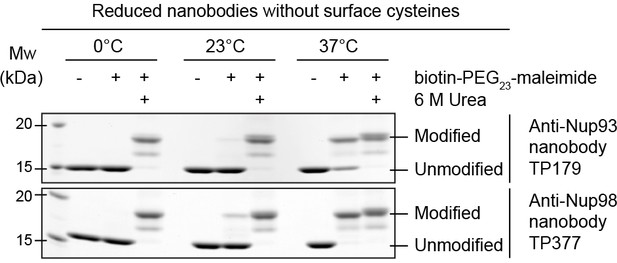
Maleimide modification of the internal cysteines of reduced nanobodies upon thermal unfolding.
Indicated nanobodies, expressed in the reductive cytoplasm of Escherichia coli BLR, were incubated at the indicated temperatures in the presence or absence of a two-fold molar excess of biotin-PEG23-maleimide (1.45 kDa) in buffer. The addition of 6 M urea served as a positive control for maleimide modification of the internal cysteines upon chemical unfolding.
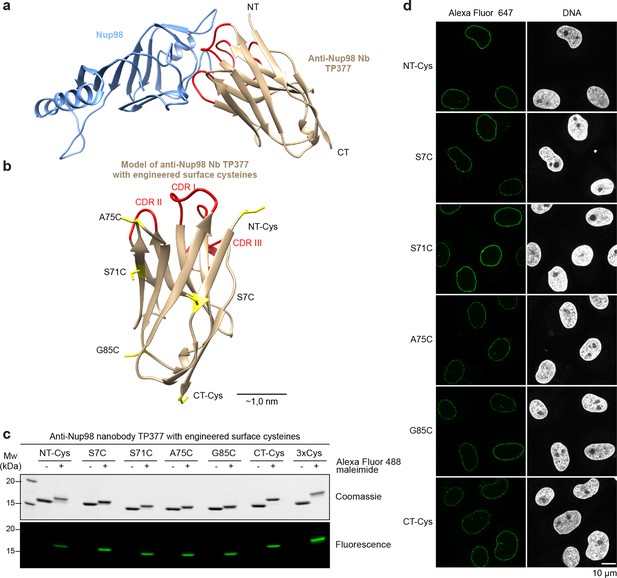
Site-specific fluorescent labeling of nanobodies.
(a) Crystal structure of the Nup98 NPC anchor domain (Nup98716-866, blue) in complex with the anti-Nup98 nanobody TP377 (beige). The three antigen-binding loops (CDR I-III) of TP377 are colored red. (NT = N-terminus, CT = C-terminus) (b) Tested positions of engineered cysteines (yellow) illustrated for nanobody TP377. Antigen-binding loops are shown in red. (c) Quantitative labeling of TP377 with Alexa Fluor 488 maleimide. TP377 with cysteines at the indicated positions can be quantitatively labeled with Alexa Fluor 488 maleimide. Labeling introduces a size shift in SDS–PAGE. Detection was either by Coomassie staining or by in-gel fluorescence. (3xCys = NT-Cys + S7C + S71C) (d) Digitonin-permeabilized Xenopus XL177 cells were incubated with 10 nM TP377 carrying a single Alexa Fluor 647 molecule at the indicated position. Cells were then washed, fixed, and counterstained with DAPI (DNA). A characteristic nuclear rim stain indicates labeling of NPCs. Note that labeling of TP377 very close to its antigen-binding loops did not perturb binding.
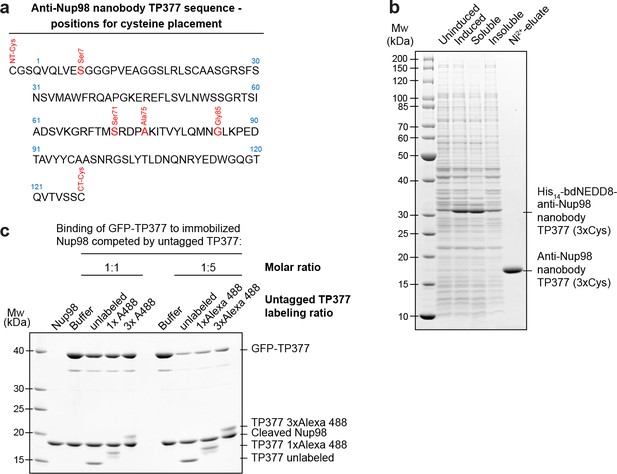
Expression and relative affinity of anti-Nup98 nanobody TP377 with engineered surface cysteines.
(a) Amino acid sequence of anti-Nup98 nanobody TP377 illustrating the positions chosen for mutation to cysteine (red). (b) SDS–PAGE and Coomassie staining showing the expression and purification of TP377 with three engineered cysteines (NT-Cys, Ser7Cys, CT-Cys) in the Escherichia coli cytoplasm. Single-step purification was performed using Ni2+ chelate affinity chromatography and cleavage using the bdNEDP1 protease. (c) Relative affinity of TP377 with different labeling ratio to Nup98716-866. GFP-tagged TP377 was incubated with biotinylated His14-Avi-bdSUMO-tagged Nup98716-866 immobilized on Streptavidin agarose. For competition, unlabeled, 1x or 3x Alexa Fluor 488-labeled TP377 was added in equimolar amount or fivefold molar excess relative to GFP-TP377. Bound nanobodies were eluted by bdSEN1P cleavage and analyzed by SDS–PAGE followed by Coomassie staining.
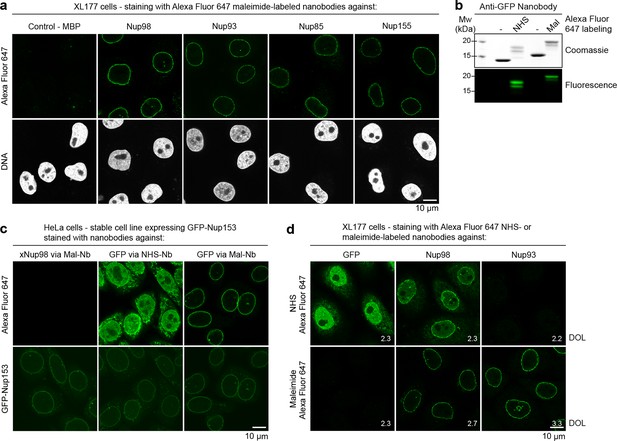
Immunofluorescence with site-specifically labeled anti-Nup nanobodies.
(a) Xenopus XL177 cells were digitonin-permeabilized and stained with anti-Nup nanobodies carrying a single N-terminal Alexa Fluor 647 maleimide dye before fixation and DAPI staining. A characteristic nuclear rim stain indicates labeling of NPCs. A nanobody raised against Escherichia coli Maltose-binding protein (MBP) served as a negative control. (b) Labeling of the anti-GFP nanobody Enhancer with Alexa Fluor 647 NHS ester at lysines or at three engineered cysteines using Alexa Fluor 647 maleimide. Labeling introduces a size shift in SDS–PAGE. Detection was either by Coomassie staining or by in-gel fluorescence. (c) Staining of HeLa cells stably expressing GFP-tagged Nup153 with the anti-GFP nanobody labeled via NHS ester or maleimide Alexa Fluor 647. The nanobody TP377, raised against Xenopus (x)Nup98, does not cross-react with human Nup98 and served as a negative control. The NHS-labeled GFP nanobody produced strong background-staining, while its maleimide-labeled version yielded bright nuclear rim stains. (d) Staining of XL177 cells with nanobodies labeled with Alexa Fluor 647 either at their internal lysine residues (NHS ester dye) or via engineered cysteines (maleimide dye). Note that the widely used anti-GFP nanobody Enhancer produces significant background staining when labeled via lysines but not when using engineered cysteines and a maleimide dye. All nanobodies were used at a concentration of 10 nM and all images were obtained under identical settings. DOL, degree of labeling.
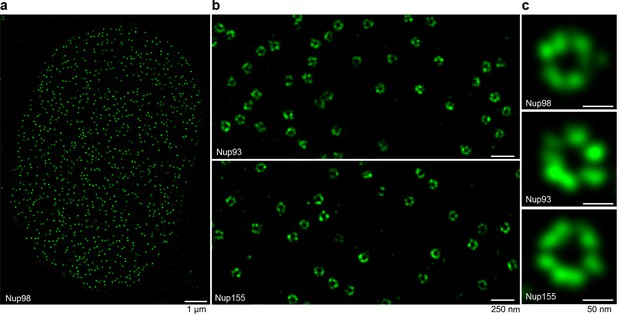
STORM imaging of nuclear pore complexes stained with site-specifically labeled anti-Nup nanobodies.
(a) STORM image of an entire XL177 cell nucleus stained with anti-Nup98 nanobody TP377 carrying a single N-terminal Alexa Fluor 647 maleimide. (b) Close-up view of XL177 cell nuclear envelope regions stained with anti-Nup93 nanobody TP179 (upper panel) or an anti-Nup155 nanobody (lower panel) containing multiple NPCs. (c) STORM images of individual NPCs stained with indicated anti-Nup nanobodies.
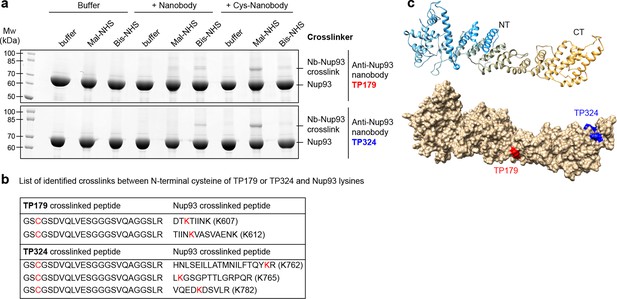
Rapid epitope mapping via crosslinking mass spectrometry.
(a) Crosslinking of two different anti-Nup93 nanobodies (TP179 and TP324) to Nup93 using amine-to-amine ('Bis-NHS'; BS3; 11.4 Å linker length) or thiol-to-amine ('Mal-NHS'; BMPS; 5.9 Å linker length) crosslinking reagents. The combination of the very short Mal-NHS crosslinker with an engineered cysteine close to the antigen-binding loops provided for both nanobodies by far the highest yield of crosslinked nanobody•Nup93 adduct. (b) List of identified crosslinked peptides involving Nup93 lysines and Cys-TP179 or Cys-TP324. The crosslinked amino acids are highlighted in red (see also Figure 8—figure supplement 1). (c) Crosslinked lysines of Nup93 to the N-terminal cysteine on anti-Nup93 nanobodies TP179 (red) or TP324 (blue) are depicted on a structural model of Nup93168-end generated by I-TASSER (Zhang, 2008). Based on the orthologous yeast crystal structures (Jeudy and Schwartz, 2007; Schrader et al., 2008), Nup93 is predicted to form a similar J-shaped structure (color gradient: NT = N-terminus in blue to CT = C-terminus in orange). Whereas TP179 binds to the central portion, TP324 binds to the C-terminus of Nup93.
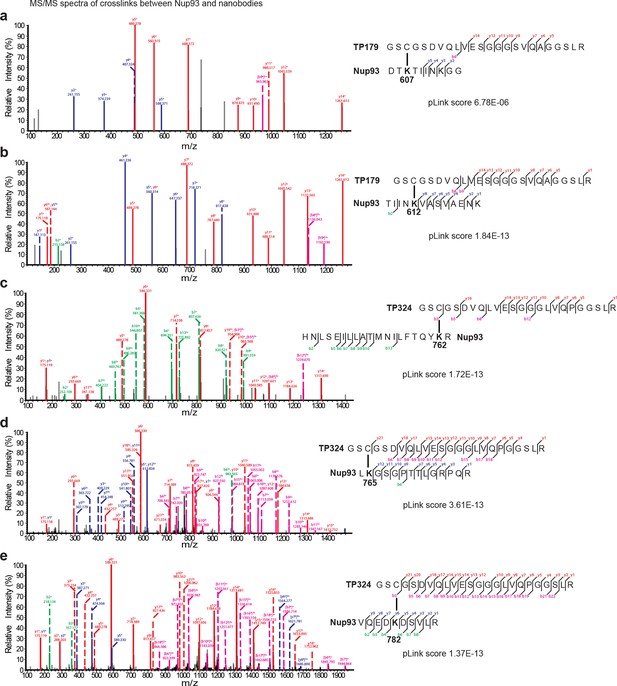
Representative MS/MS spectra of the crosslinked peptides derived from Nup93•nanobody complexes.
The spectra with the best pLink score are shown for the crosslinks between: (a) TP179 Cys3 - Nup93 Lys607, (b) TP179 Cys3 - Nup93 Lys612, (c) TP324 Cys3 - Nup93 Lys762, (d) TP324 Cys3 - Nup93 Lys765, and (e) TP324 Cys3 - Nup93 Lys782. The peaks of the b and y ions are labeled with their charge stages and m/z values. The b and y ions of the longer peptide in a crosslink pair are highlighted in magenta and red, respectively, and the b and y ions of the shorter peptide are highlighted in green and blue, respectively. Fragment ions with superscript 'x' represent those fragment ions with the other peptide crosslinked.
Tables
Crystallographic data collection and refinement statistics
| Nup98•Nb TP377 complexa | |
|---|---|
| Data collection | |
| Space group | P41 |
| Cell dimensions | |
| a, b, c (Å) | 66.59, 66.59, 87.90 |
| α, β, γ (°) | 90.00, 90.00, 90.00 |
| Resolution (Å) | 47.00-1.90 (1.95-1.90)b |
| Rsym or Rmerge | 0.128 (>1)b |
| I / σI | 27.7 (2.6)b |
| Completeness (%) | 99.5 (98.7)b |
| Redundancy | 27.4 (27.4)b |
| Refinement | |
| Resolution (Å) | 47.00-1.90 |
| No. reflections | |
| Measured | 823105 |
| Unique | 30218 |
| Rwork/Rfree | 0.167 / 0.196 |
| No. atoms | |
| Protein | 2176 |
| Water | 145 |
| Wilson B-factor (Å2) | 27.4 |
| R.m.s. deviations | |
| Bond lengths (Å) | 0.010 |
| Bond angles (°) | 1.08 |
| Ramachandran statistics (%) | |
| Favored | 98.0 |
| Allowed | 2.0 |
| Outliers | - |
-
aA single crystal was used for data collection.
-
bValues in parentheses are for highest resolution shell.





