An experimentally validated network of nine haematopoietic transcription factors reveals mechanisms of cell state stability
Figures
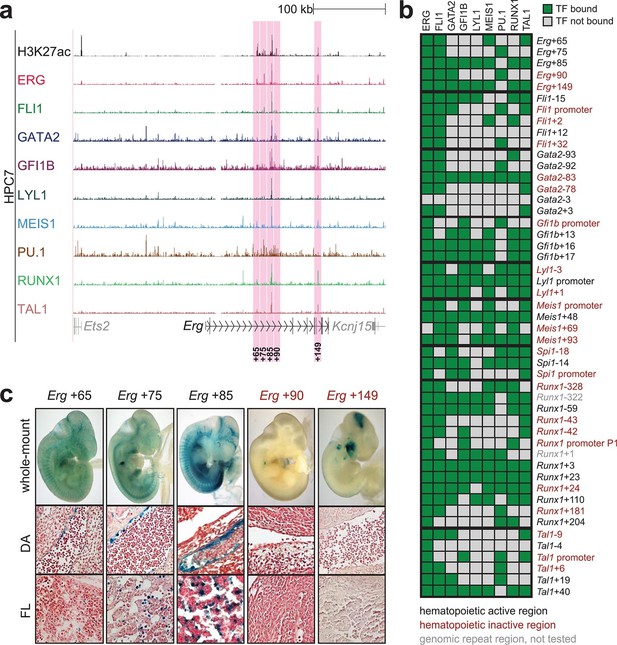
Identification of haematopoietic active cis-regulatory regions.
(a) UCSC screenshot of the Erg gene locus for ChIP-Sequencing data for nine haematopoietic TFs (ERG, FLI1, GATA2, GFI1B, LYL1, MEIS1, PU.1, RUNX1 and TAL1 [Wilson et al., 2010]) and for H3K27ac (Calero-Nieto et al., 2014) in HPC7 cells. Highlighted are all regions of the Erg gene locus that are acetylated at H3K27 and are bound by three or more TFs. Numbers indicate the distance (in kb) from the ATG start codon. (b) Summary of the identification of candidate cis-regulatory regions for all nine TFs and subsequent analysis in transgenic mouse assays. The inspection of the nine gene loci and the application of the selection criteria (≥3 TFs bound and H3K27ac) identified a total of 49 candidate cis-regulatory regions. The heatmap shows the binding pattern of the nine TFs to all candidate regulatory elements in HPC7 cells: green = bound, grey = unbound. Haematopoietic activity in E11.5 transgenic mice is indicated by the font color: black = active, red = not active. Grey indicates genomic repeat regions that were not tested in transgenic mice. Detailed experimental data corresponding to the summary heatmap can be found in Figure 1 and Figure 1—figure supplement 1–8. (c) Haematopoietic activity of the five candidate Erg cis-regulatory regions was determined in E11.5 transgenic mouse assays. Shown are X-Gal-stained whole-mount embryos and paraffin sections of the dorsal aorta (DA, ventral side on the left/top) and foetal liver (FL), sites of definitive haematopoiesis. Colour coding as in B.
-
Figure 1—source data 1
Number of PCR and LacZ positive transgenic embryos (E10.5–11.5) for each regulatory region.
- https://doi.org/10.7554/eLife.11469.004
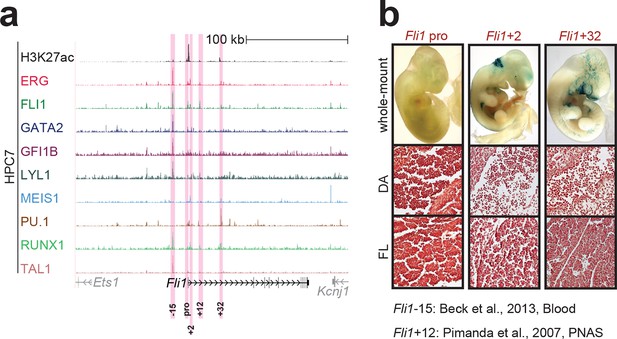
Identification of haematopoietic active cis-regulatory elements for Fli1.
(a) The candidate cis-regulatory elements were identified by ChIP-Seq analysis of the TFs ERG, FLI1, GATA2, GFI1B, LYL1, MEIS1, PU.1, RUNX1 and TAL1 as well as H3K27 acetylation in the haematopoietic stem/progenitor cell line HPC7. Highlighted in pink are the candidate cis-regulatory regions which are bound by at least three of the nine TFs and showed H3K27 acetylation. The numbering represents the direction and distance in kilobases from the start codon ATG (pro = promoter). (b) Candidate regions were assayed for haematopoietic enhancer activity in mouse transient transgenic embryos. X-Gal stained whole-mount E11.5 embryos and paraffin sections of the dorsal aorta (DA; longitudinal section, ventral side on the left/top) and foetal liver (FL) are shown for the candidate cis-regulatory regions. Transgenic mouse data are not shown for previously published regions, but relevant publications are listed.
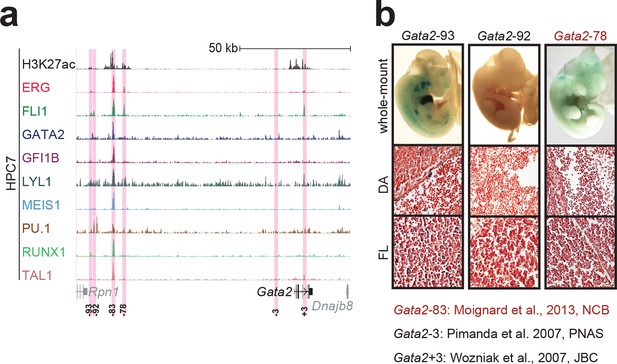
Identification of haematopoietic active cis-regulatory elements for Gata2.
(a) The candidate cis-regulatory elements were identified by ChIP-Seq analysis of the TFs ERG, FLI1, GATA2, GFI1B, LYL1, MEIS1, PU.1, RUNX1 and TAL1 as well as H3K27 acetylation in the haematopoietic stem/progenitor cell line HPC7. Highlighted in pink are the candidate cis-regulatory regions which are bound by at least three of the nine TFs and showed H3K27 acetylation. The numbering represents the direction and distance in kilobases from the start codon ATG (pro = promoter). (b) Candidate regions were assayed for haematopoietic enhancer activity in mouse transient transgenic embryos. X-Gal stained whole-mount E11.5 embryos and paraffin sections of the dorsal aorta (DA; longitudinal section, ventral side on the left/top) and foetal liver (FL) are shown for the candidate cis-regulatory regions. Transgenic mouse data are not shown for previously published regions, but relevant publications are listed.
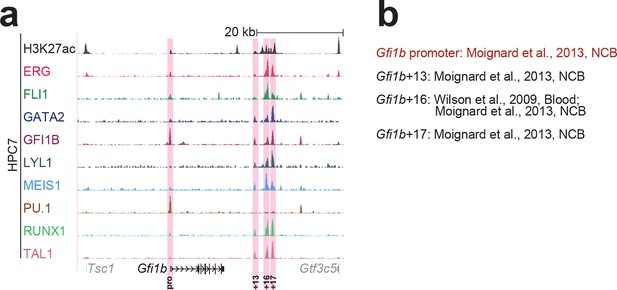
Identification of haematopoietic active cis-regulatory elements for Gfi1b.
(a) The candidate cis-regulatory elements were identified by ChIP-Seq analysis of the TFs ERG, FLI1, GATA2, GFI1B, LYL1, MEIS1, PU.1, RUNX1 and TAL1 as well as H3K27 acetylation in the haematopoietic stem/progenitor cell line HPC7. Highlighted in pink are the candidate cis-regulatory regions which are bound by at least three of the nine TFs and showed H3K27 acetylation. The numbering represents the direction and distance in kilobases from the start codon ATG (pro = promoter). (b) All candidate regions were previously published. Relevant publications are listed.
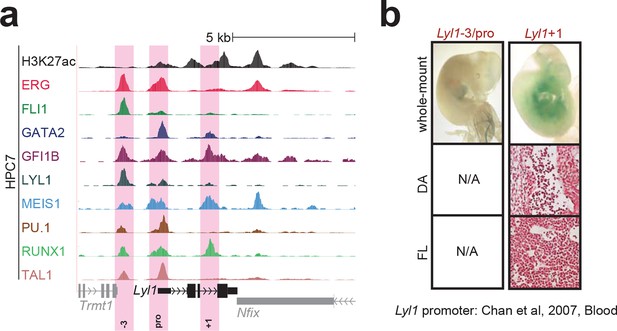
Identification of haematopoietic active cis-regulatory elements for Lyl1.
(a) The candidate cis-regulatory elements were identified by ChIP-Seq analysis of the TFs ERG, FLI1, GATA2, GFI1B, LYL1, MEIS1, PU.1, RUNX1 and TAL1 as well as H3K27 acetylation in the haematopoietic stem/progenitor cell line HPC7. Highlighted in pink are the candidate cis-regulatory regions which are bound by at least three of the nine TFs and showed H3K27 acetylation. The numbering represents the direction and distance in kilobases from the start codon ATG (pro = promoter). (b) Candidate regions were assayed for haematopoietic enhancer activity in mouse transient transgenic embryos. X-Gal stained whole-mount E11.5 embryos and paraffin sections of the dorsal aorta (DA; longitudinal section, ventral side on the left/top) and foetal liver (FL) are shown for the candidate cis-regulatory regions. Transgenic mouse data are not shown for previously published regions, but relevant publications are listed.
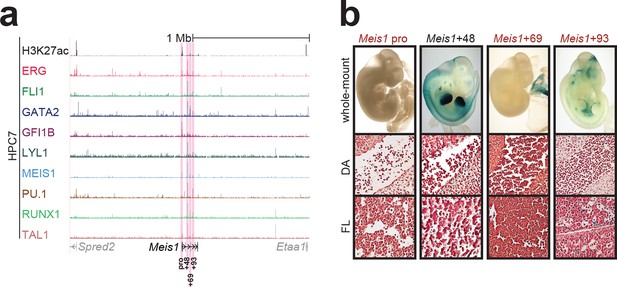
Identification of haematopoietic active cis-regulatory elements for Meis1.
(a) The candidate cis-regulatory elements were identified by ChIP-Seq analysis of the TFs ERG, FLI1, GATA2, GFI1B, LYL1, MEIS1, PU.1, RUNX1 and TAL1 as well as H3K27 acetylation in the haematopoietic stem/progenitor cell line HPC7. Highlighted in pink are the candidate cis-regulatory regions which are bound by at least three of the nine TFs and showed H3K27 acetylation. The numbering represents the direction and distance in kilobases from the start codon ATG (pro = promoter). (b) Candidate regions were assayed for haematopoietic enhancer activity in mouse transient transgenic embryos. X-Gal stained whole-mount E11.5 embryos and paraffin sections of the dorsal aorta (DA; longitudinal section, ventral side on the left/top) and foetal liver (FL) are shown for the candidate cis-regulatory regions.
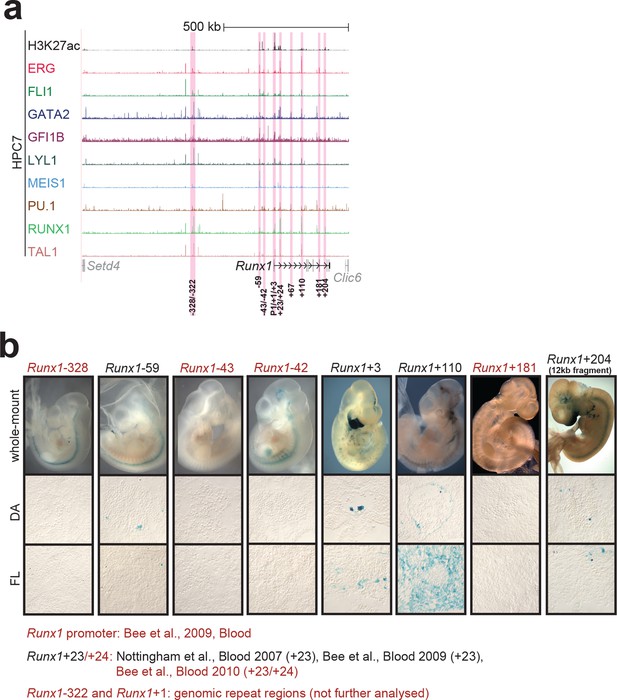
Identification of haematopoietic active cis-regulatory elements for Runx1.
(a) The candidate cis-regulatory elements were identified by ChIP-Seq analysis of the TFs ERG, FLI1, GATA2, GFI1B, LYL1, MEIS1, PU.1, RUNX1 and TAL1 as well as H3K27 acetylation in the haematopoietic stem/progenitor cell line HPC7. Highlighted in pink are the candidate cis-regulatory regions which are bound by at least three of the nine TFs and showed H3K27 acetylation. The numbering represents the direction and distance in kilobases from the start codon ATG (pro = promoter). (b) E10 embryos and cryosections of the DA (transverse; ventral down) and FL are shown. For the Runx1+204 region, a larger 12 kb fragment (chr16:92,620,915–92,631,936, mm9) was used for transient transgenesis, but similar results were obtained with the +204 fragment alone (data not shown). The +24 element was tested in conjunction with the +23 and did not change its tissue specificity (Bee et al., 2010). Preliminary data show that the +24 on its own does not mediate robust tissue specific expression of reporter genes. Transgenic mouse data are not shown for previously published regions, but relevant publications are listed.
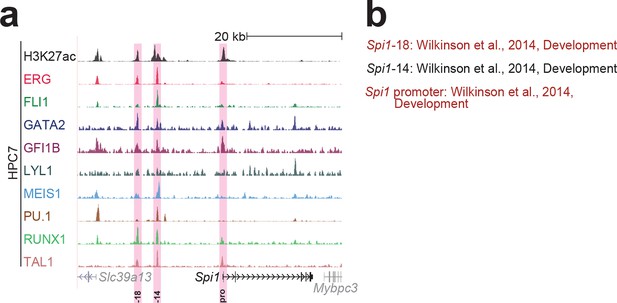
Identification of haematopoietic active cis-regulatory elements for Spi1.
(a) The candidate cis-regulatory elements were identified by ChIP-Seq analysis of the TFs ERG, FLI1, GATA2, GFI1B, LYL1, MEIS1, PU.1, RUNX1 and TAL1 as well as H3K27 acetylation in the haematopoietic stem/progenitor cell line HPC7. Highlighted in pink are the candidate cis-regulatory regions which are bound by at least three of the nine TFs and showed H3K27 acetylation. The numbering represents the direction and distance in kilobases from the start codon ATG (pro = promoter). (b) All candidate regions were previously published. Relevant publications are listed.
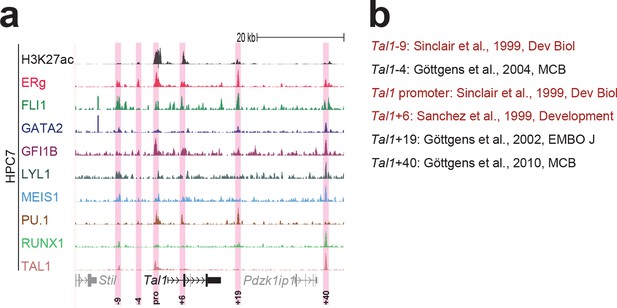
Identification of haematopoietic active cis-regulatory elements for Tal1.
(a) The candidate cis-regulatory elements were identified by ChIP-Seq analysis of the TFs ERG, FLI1, GATA2, GFI1B, LYL1, MEIS1, PU.1, RUNX1 and TAL1 as well as H3K27 acetylation in the haematopoietic stem/progenitor cell line HPC7. Highlighted in pink are the candidate cis-regulatory regions which are bound by at least three of the nine TFs and showed H3K27 acetylation. The numbering is based on the distance (in kb) to promoter 1a. (b) All candidate regions were previously published. Relevant publications are listed.
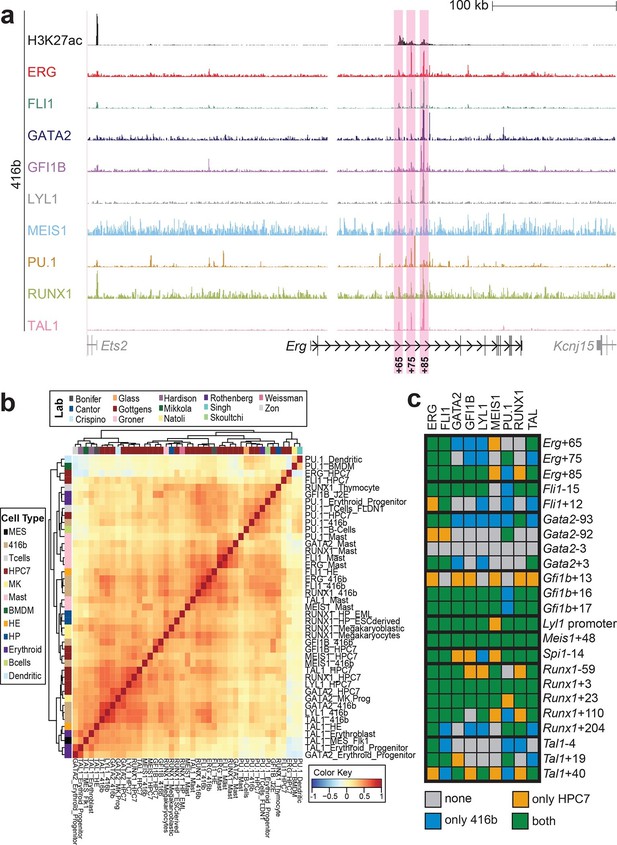
Comparison of TF binding pattern at haematopoietic active cis-regulatory regions in two haematopoietic progenitor cell lines, HPC7 and 416b.
(a) UCSC screenshot of the Erg gene locus for ChIP-Sequencing data for nine haematopoietic TFs (ERG, FLI1, GATA2, GFI1B, LYL1, MEIS1, PU.1, RUNX1 and TAL1) and for H3K27ac in 416b cells. Highlighted are those haematopoietic active Erg cis-regulatory regions that were identified based on acetylation of H3K27 and TF binding in HPC7 cells followed by transgenic mouse assays. Numbers indicate the distance (in kb) from the ATG start codon. (b) Hierarchical clustering of the binding profiles for HPC7, 416b and other published datasets. The heatmap shows the pairwise correlation coefficient of peak coverage data between the pairs of samples in the row and column. The order of the samples is identical in columns and rows. Details about samples listed can be found in Figure 2—source data 1. (c) Pair-wise analysis of binding of the nine TFs to haematopoietic active cis-regulatory regions of the nine TFs in HPC7 versus 416b cells. Green = bound in both cells types, blue = only bound in 416b cells, orange = only bound in HPC7 cells, grey = not bound in either cell type.
-
Figure 2—source data 1
List of ChIP-Seq samples included in the heatmap in Figure 2b.
- https://doi.org/10.7554/eLife.11469.014
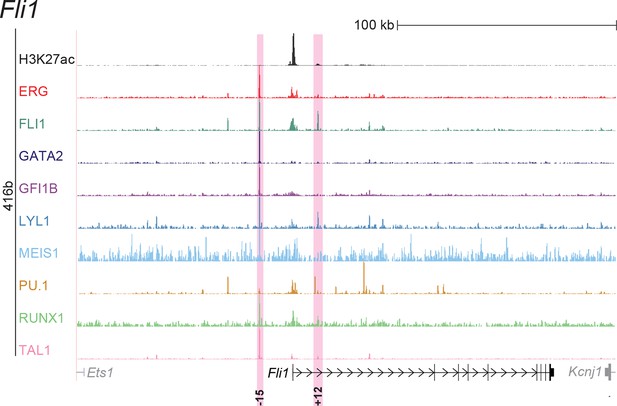
UCSC screenshot for the Fli1 gene locus demonstrating binding patterns for nine key haematopoietic TFs and H3K27ac in 416b cells.
Highlighted in pink are cis-regulatory regions that were identified based on the selection criteria (≥3 TFs bound and H3K27ac) in HPC7 cells and were shown to possess haematopoietic activity. The numbering represents the distance (in kb) from the start codon ATG.
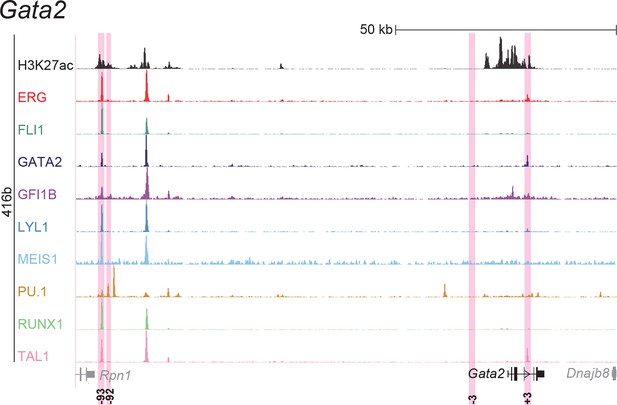
UCSC screenshot for the Gata2 gene locus demonstrating binding patterns for nine key haematopoietic TFs and H3K27ac in 416b cells.
Highlighted in pink are cis-regulatory regions that were identified based on the selection criteria (≥3 TFs bound and H3K27ac) in HPC7 cells and were shown to possess haematopoietic activity. The numbering represents the distance (in kb) from the start codon ATG.
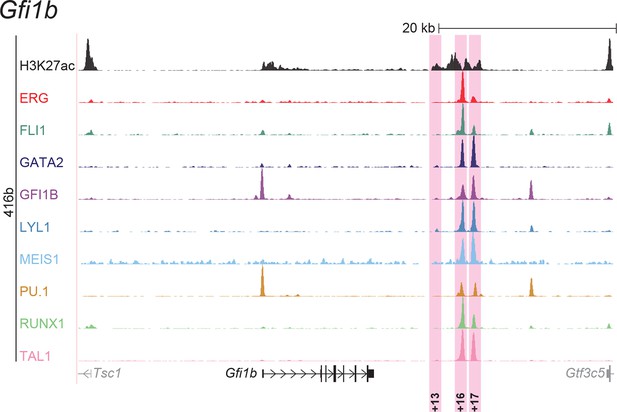
UCSC screenshot for the Gfi1b gene locus demonstrating binding patterns for nine key haematopoietic TFs and H3K27ac in 416b cells.
Highlighted in pink are cis-regulatory regions that were identified based on the selection criteria (≥3 TFs bound and H3K27ac) in HPC7 cells and were shown to possess haematopoietic activity. The numbering represents the distance (in kb) from the start codon ATG.
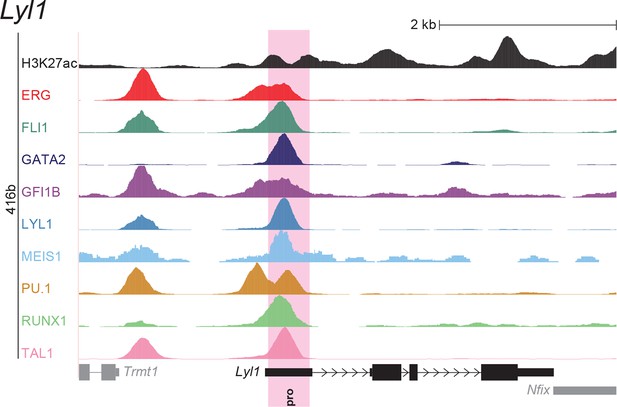
UCSC screenshot for the Lyl1 gene locus demonstrating binding patterns for nine key haematopoietic TFs and H3K27ac in 416b cells.
Highlighted in pink is the promoter ('pro') that was identified based on the selection criteria (≥3 TFs bound and H3K27ac) in HPC7 cells and was shown to possess haematopoietic activity. The promoter is labelled with 'pro'.
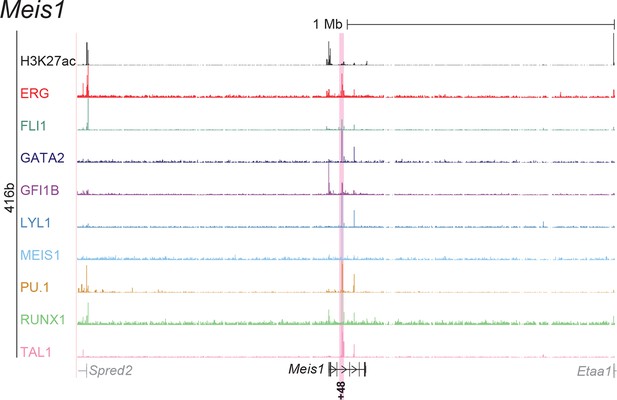
UCSC screenshot for the Meis1 gene locus demonstrating binding patterns for nine key haematopoietic TFs and H3K27ac in 416b cells.
Highlighted in pink is the cis-regulatory region that was identified based on the selection criteria (≥3 TFs bound and H3K27ac) in HPC7 cells and was shown to possess haematopoietic activity. The numbering represents the distance (in kb) from the start codon ATG.
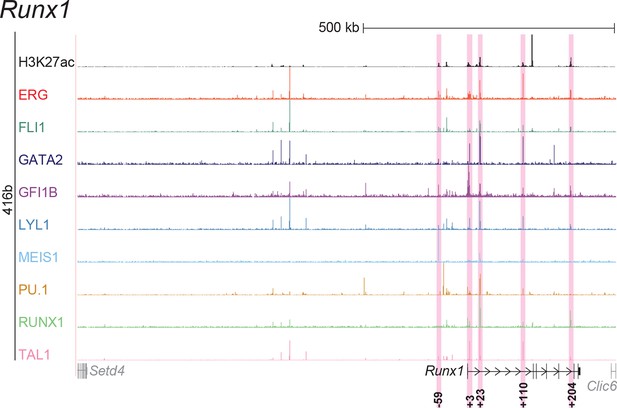
UCSC screenshot for the Runx1 gene locus demonstrating binding patterns for nine key haematopoietic TFs and H3K27ac in 416b cells.
Highlighted in pink are cis-regulatory regions that were identified based on the selection criteria (≥3 TFs bound and H3K27ac) in HPC7 cells and were subsequently shown to possess haematopoietic activity. The numbering represents the distance (in kb) from the start codon ATG.
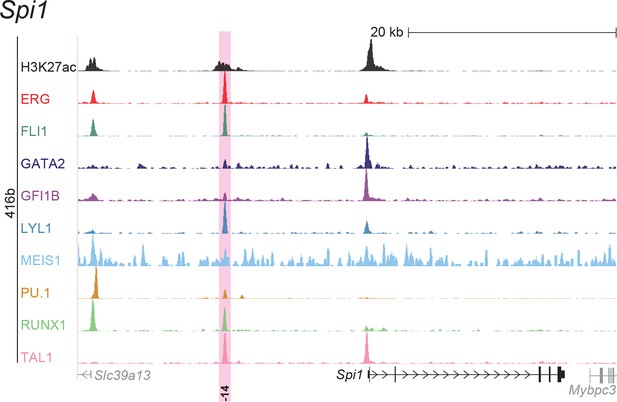
UCSC screenshot for the Spi1 gene locus demonstrating binding patterns for nine key haematopoietic TFs and H3K27ac in 416b cells.
Highlighted in pink is the cis-regulatory region that was identified based on the selection criteria (≥3 TFs bound and H3K27ac) in HPC7 cells and was shown to possess haematopoietic activity. The numbering represents the distance (in kb) from the start codon ATG.
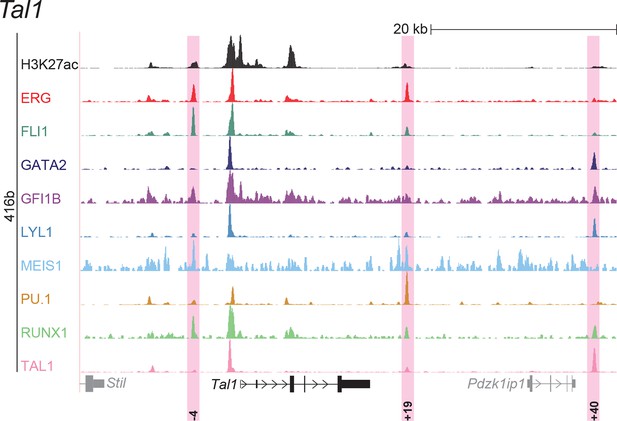
UCSC screenshot for the Tal1 gene locus demonstrating binding patterns for nine key haematopoietic TFs and H3K27ac in 416b cells.
Highlighted in pink are cis-regulatory regions that were identified based on the selection criteria (≥3 TFs bound and H3K27ac) in HPC7 cells and were shown to possess haematopoietic activity. The numbering represents the distance (in kb) from promoter 1a.
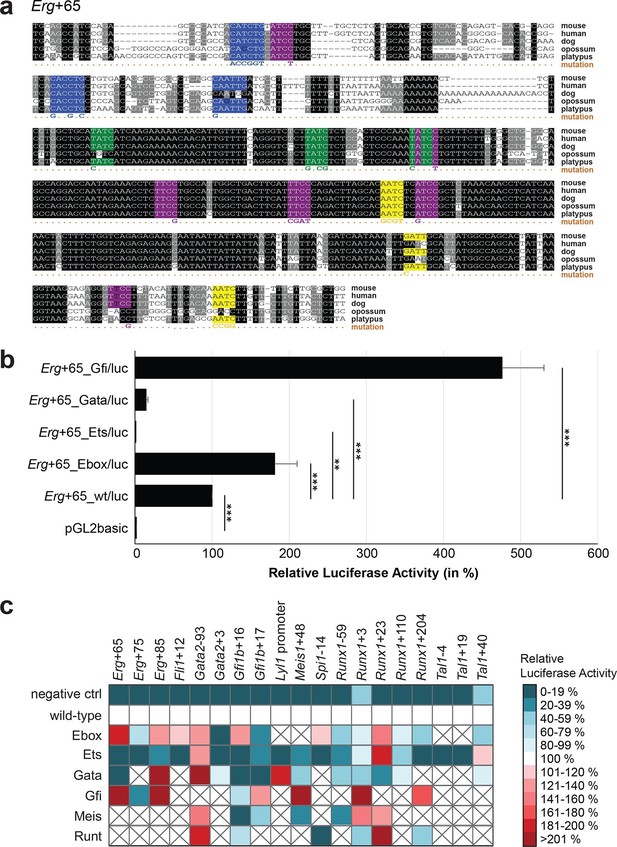
TFBS mutagenesis reveals enhancer-dependent effects of TF binding on gene expression.
(a) Multiple species alignment of mouse (mm9), human (hg19), dog (canFam2), opossum (monDom5) and platypus (ornAna1) sequences for the Erg+65 region. Nucleotides highlighted in black are conserved between all species analysed, nucleotides highlighted in grey are conserved between four of five species. Transcription factor binding sites (TFBS) are highlighted in: blue = Ebox, purple = Ets, green = Gata, yellow = Gfi, red = Meis. The nucleotides that were changed to mutate the TFBSs are indicated below the alignment. All binding sites of one motif family (e.g. all Ebox motifs) were mutated simultaneously. (b) Luciferase assay for the Erg+65 wild-type and mutant enhancer in stably transfected 416b cells. Each bar represents the averages of at least three independent experiments with three to four replicates within each experiment. The results are shown relative to the wild-type enhancer activity, which is set to 100%. Error bars represent the standard error of the mean (SEM). Stars indicate significance: **=p-value <0.01, ***=p-value <0.001. p-values were calculated using t-tests, followed by the Fisher’s method. (c) Summary of luciferase assay results for all 19 high-confidence haematopoietic active regulatory regions. Relative luciferase activity is illustrated in shades of blue (down-regulation) and red (up-regulation). Crossed-out grey boxes indicate that there is no motif for the TF and/or the TF does not bind to the region. Detailed results and corresponding alignments with highlighted TFBSs and their mutations can be found in Figure 3—figure supplements 1–18.
-
Figure 3—source data 1
List of TF binding sites and the TFs that bind to them.
- https://doi.org/10.7554/eLife.11469.024
-
Figure 3—source data 2
List of co-ordinates and primer sequences for the regulatory regions analysed in this study.
- https://doi.org/10.7554/eLife.11469.025
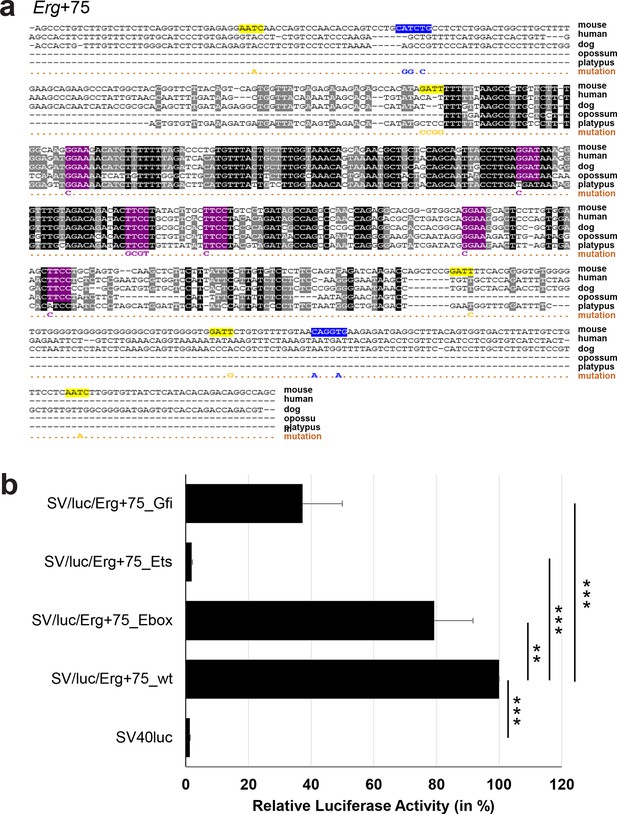
Multiple species alignment and luciferase assay results for Erg+75.
(a) Multiple species alignment (MSA) with the following species: mouse (mm9), human (hg19), dog (canFam2), opossum (monDom5) and platypus (ornAna1). Nucleotides highlighted in black are conserved between all species analysed, nucleotides highlighted in grey are conserved between four of five species. Transcription factor binding sites (TFBS) are highlighted in: blue = Ebox, purple = Ets, yellow = Gfi. The nucleotides that were changed to mutate the TFBSs are indicated below the MSA. All conserved binding sites of one motif family (e.g. all Ebox motifs) were mutated simultaneously. Where TF binding was observed in ChIP-Seq experiments in 416b cells, but the TFBS was not conserved, the motifs present in the mouse sequence only were mutated. (b) For the luciferase reporter assays in stably transfected 416b cells, the averages of at least three independent experiments with three to four replicates within each experiment are shown. Error bars represent the standard error of the mean (SEM). Stars indicate significance: **=p-value <0.01, ***=p-value <0.001. p-values were generated using t-tests, followed by the Fisher’s method and if necessary Stouffer’s z trend.
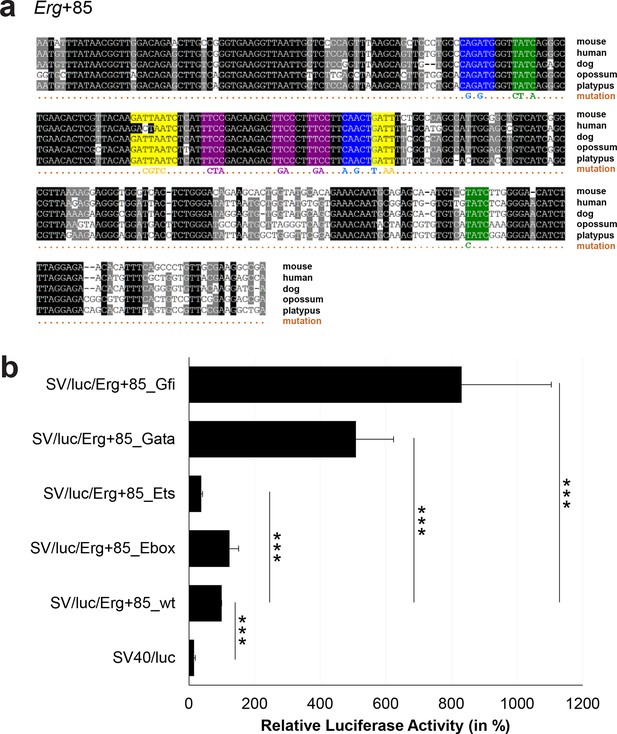
Multiple species alignment and luciferase assay results for Erg+85.
(a) Multiple species alignment (MSA) with the following species: mouse (mm9), human (hg19), dog (canFam2), opossum (monDom5) and platypus (ornAna1). Nucleotides highlighted in black are conserved between all species analysed, nucleotides highlighted in grey are conserved between four of five species. Transcription factor binding sites (TFBS) are highlighted in: blue = Ebox, purple = Ets, green = Gata, yellow = Gfi. The nucleotides that were changed to mutate the TFBSs are indicated below the MSA. All conserved binding sites of one motif family (e.g. all Ebox motifs) were mutated simultaneously. (b) For the luciferase reporter assays in stably transfected 416b cells, the averages of at least three independent experiments with three to four replicates within each experiment are shown. Error bars represent the standard error of the mean (SEM). Stars indicate significance: ***=p-value <0.001. p-values were generated using t-tests, followed by the Fisher’s method and if necessary Stouffer’s z trend.
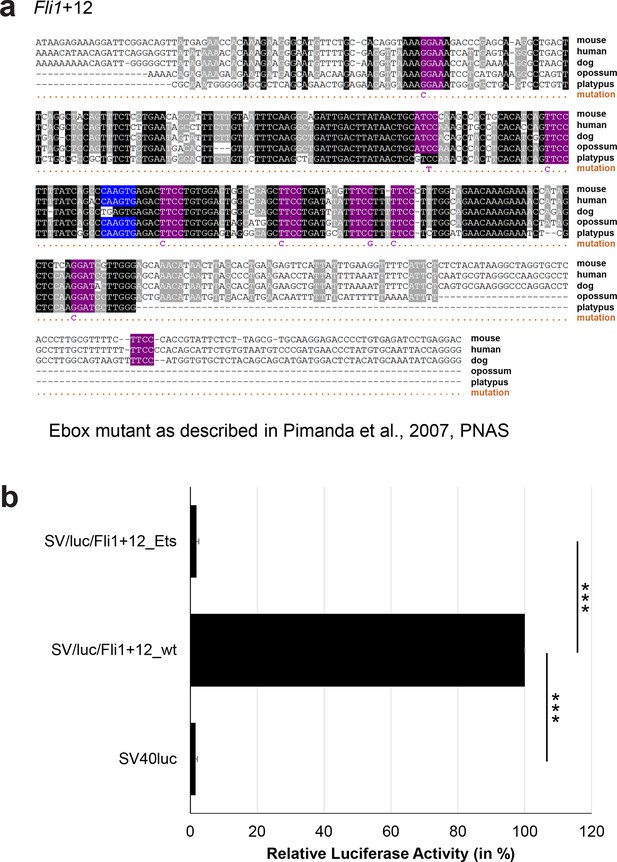
Multiple species alignment and luciferase assay results for Fli1+12.
(a) Multiple species alignment (MSA) with the following species: mouse (mm9), human (hg19), dog (canFam2), opossum (monDom5) and platypus (ornAna1). Nucleotides highlighted in black are conserved between all species analysed, nucleotides highlighted in grey are conserved between four of five species. Transcription factor binding sites (TFBS) are highlighted in: blue = Ebox, purple = Ets. The nucleotides that were changed to mutate the TFBSs are indicated below the MSA. All conserved binding sites of one motif family (e.g. all Etsmotifs) were mutated simultaneously. (b) For the luciferase reporter assays in stably transfected 416b cells, the averages of at least three independent experiments with three to four replicates within each experiment are shown. Error bars represent the standard error of the mean (SEM). Stars indicate significance: ***=p-value <0.001. p-values were generated using t-tests, followed by the Fisher’s method and if necessary Stouffer’s z trend.
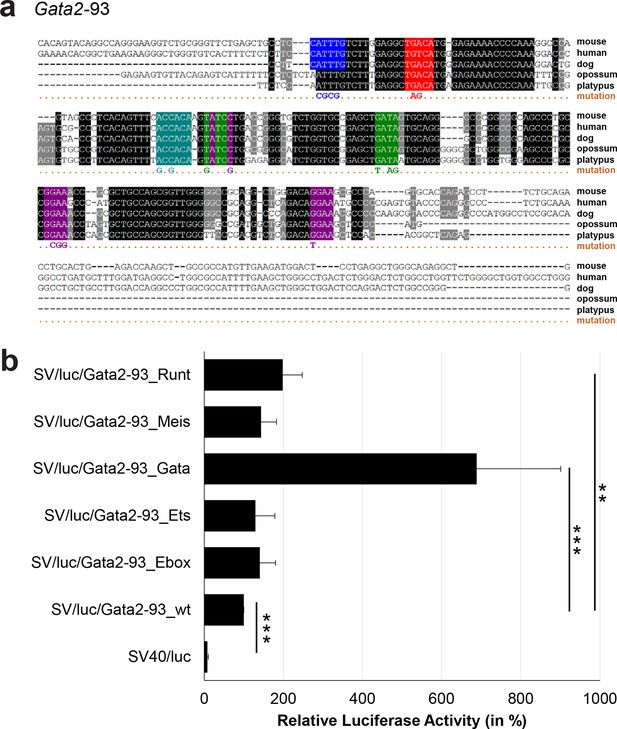
Multiple species alignment and luciferase assay results for Gata2-93.
(a) Multiple species alignment (MSA) with the following species: mouse (mm9), human (hg19), dog (canFam2), opossum (monDom5) and platypus (ornAna1). Nucleotides highlighted in black are conserved between all species analysed, nucleotides highlighted in grey are conserved between four of five species. Transcription factor binding sites (TFBS) are highlighted in: blue = Ebox, purple = Ets, green = Gata, red = Meis, turquoise = Runt. The nucleotides that were changed to mutate the TFBSs are indicated below the MSA. All conserved binding sites of one motif family (e.g. all Ets motifs) were mutated simultaneously. (b) For the luciferase reporter assays in stably transfected 416b cells, the averages of at least three independent experiments with three to four replicates within each experiment are shown. Error bars represent the standard error of the mean (SEM). Stars indicate significance: **=p-value <0.01, ***=p-value <0.001. p-values were generated using t-tests, followed by the Fisher’s method and if necessary Stouffer’s z trend.
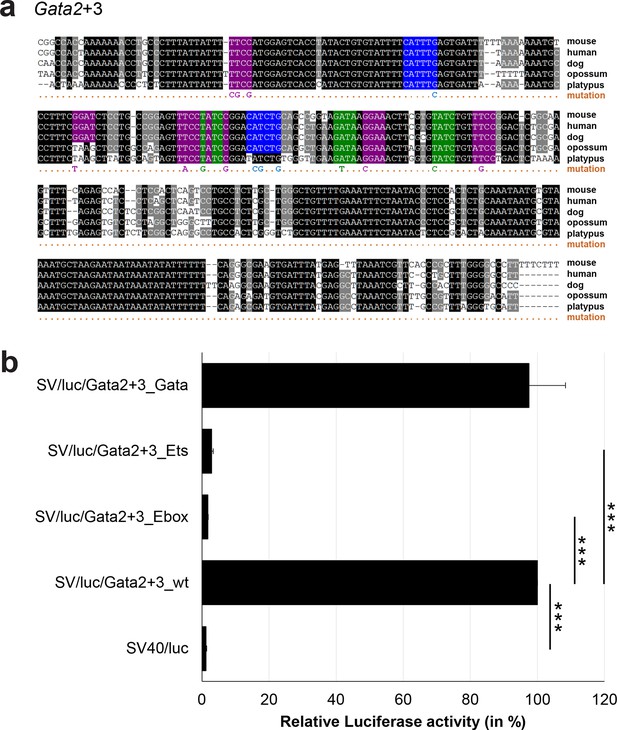
Multiple species alignment and luciferase assay results for Gata2+3.
(a) Multiple species alignment (MSA) with the following species: mouse (mm9), human (hg19), dog (canFam2), opossum (monDom5) and platypus (ornAna1). Nucleotides highlighted in black are conserved between all species analysed, nucleotides highlighted in grey are conserved between four of five species. Transcription factor binding sites (TFBS) are highlighted in: blue = Ebox, purple = Ets, green = Gata. The nucleotides that were changed to mutate the TFBSs are indicated below the MSA. All conserved binding sites of one motif family (e.g. all Ebox motifs) were mutated simultaneously. (b) For the luciferase reporter assays in stably transfected 416b cells, the averages of at least three independent experiments with three to four replicates within each experiment are shown. Error bars represent the standard error of the mean (SEM). Stars indicate significance: ***=p-value <0.001. p-values were generated using t-tests, followed by the Fisher’s method and if necessary Stouffer’s z trend.
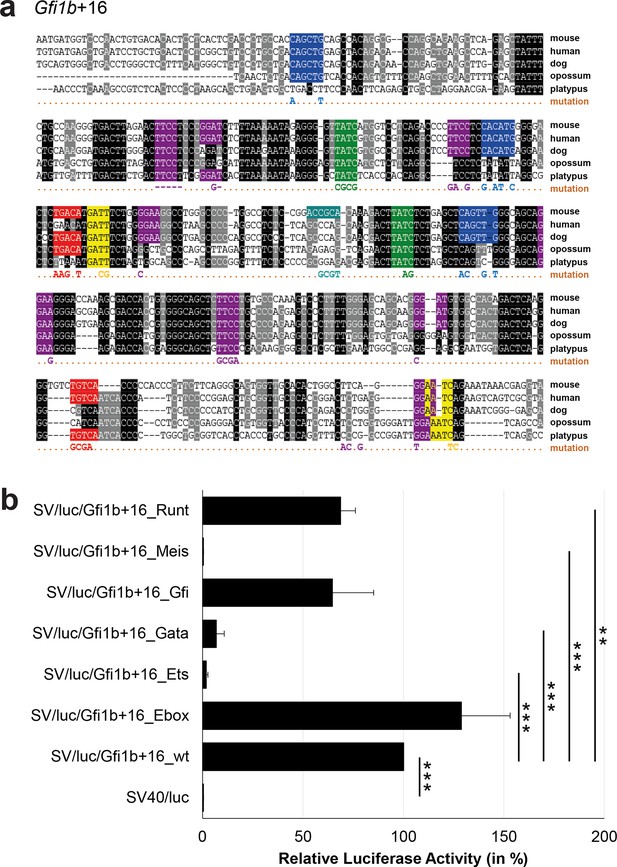
Multiple species alignment and luciferase assay results for Gfi1b+16.
(a) Multiple species alignment (MSA) with the following species: mouse (mm9), human (hg19), dog (canFam2), opossum (monDom5) and platypus (ornAna1). Nucleotides highlighted in black are conserved between all species analysed, nucleotides highlighted in grey are conserved between four of five species. Transcription factor binding sites (TFBS) are highlighted in: blue = Ebox, purple = Ets, green = Gata, yellow = Gfi, red = Meis, turquoise = Runt. The nucleotides that were changed to mutate the TFBSs are indicated below the MSA. All conserved binding sites of one motif family (e.g. all Ebox motifs) were mutated simultaneously. Where TF binding was observed in ChIP-Seq experiments in 416b cells, but the TFBS was not conserved, the motifs present in the mouse sequence only were mutated. (b) For the luciferase reporter assays in stably transfected 416b cells, the averages of at least three independent experiments with three to four replicates within each experiment are shown. Error bars represent the standard error of the mean (SEM). Stars indicate significance: **=p-value <0.01, ***=p-value <0.001. p-values were generated using t-tests, followed by the Fisher’s method and if necessary Stouffer’s z trend.
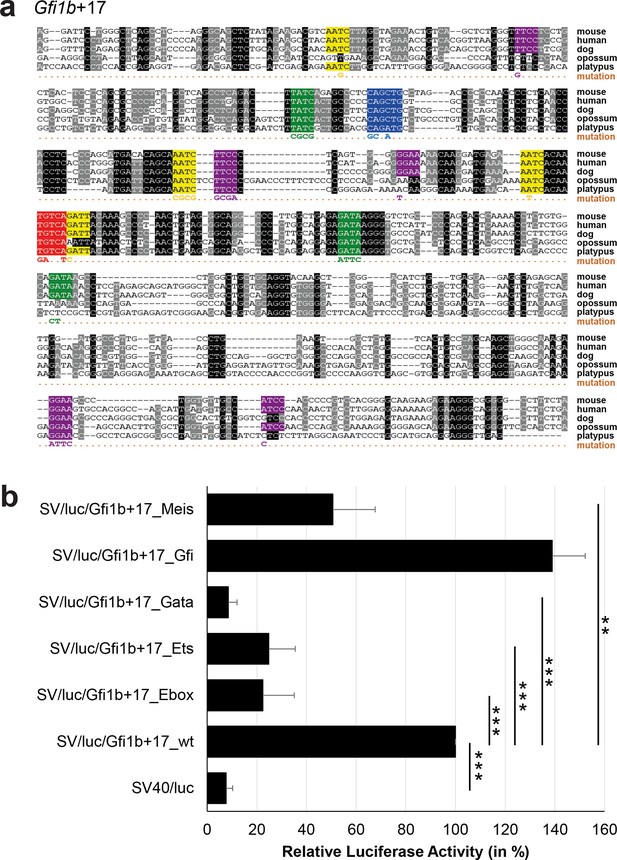
Multiple species alignment and luciferase assay results for Gfi1b+17.
(a) Multiple species alignment (MSA) with the following species: mouse (mm9), human (hg19), dog (canFam2), opossum (monDom5) and platypus (ornAna1). Nucleotides highlighted in black are conserved between all species analysed, nucleotides highlighted in grey are conserved between four of five species. Transcription factor binding sites (TFBS) are highlighted in: blue = Ebox, purple = Ets, green = Gata, yellow = Gfi, red = Meis. The nucleotides that were changed to mutate the TFBSs are indicated below the MSA. All conserved binding sites of one motif family (e.g. all Ets motifs) were mutated simultaneously. (b) For the luciferase reporter assays in stably transfected 416b cells, the averages of at least three independent experiments with three to four replicates within each experiment are shown. Error bars represent the standard error of the mean (SEM). Stars indicate significance: **=p-value <0.01, ***=p-value <0.001. p-values were generated using t-tests, followed by the Fisher’s method and if necessary Stouffer’s z trend.
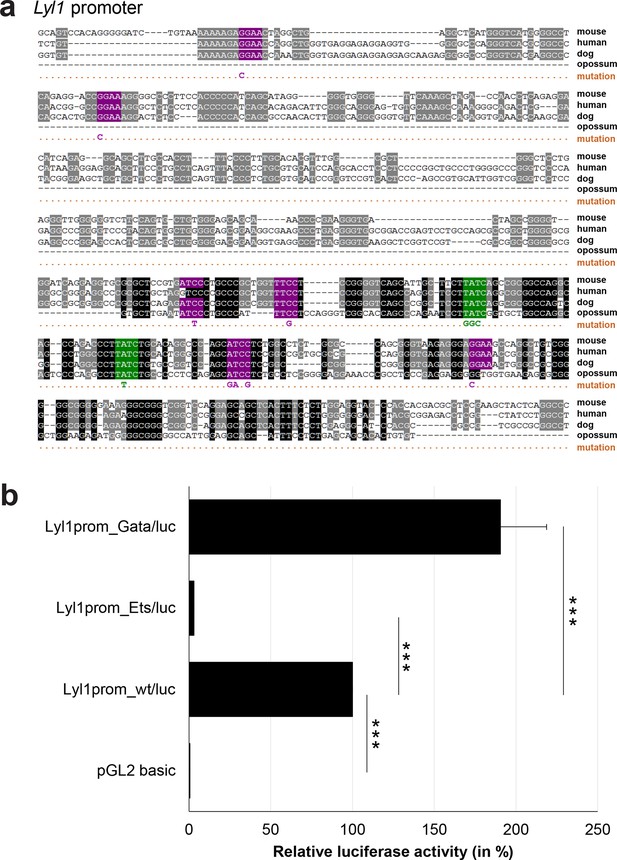

Multiple species alignment and luciferase assay results for Lyl1 promoter.
(a) Multiple species alignment (MSA) with the following species: mouse (mm9), human (hg19), dog (canFam2) and opossum (monDom5). Nucleotides highlighted in black are conserved between all species analysed, nucleotides highlighted in grey are conserved between three of four species. Transcription factor binding sites (TFBS) are highlighted in: purple = Ets, green = Gata. The nucleotides that were changed to mutate the TFBSs are indicated below the MSA. All conserved binding sites of one motif family (e.g. all Ets motifs) were mutated simultaneously. (b) For the luciferase reporter assays in stably transfected 416b cells, the averages of at least three independent experiments with three to four replicates within each experiment are shown. Error bars represent the standard error of the mean (SEM). Stars indicate significance: ***=p-value <0.001. p-values were generated using t-tests, followed by the Fisher’s method and if necessary Stouffer’s z trend.
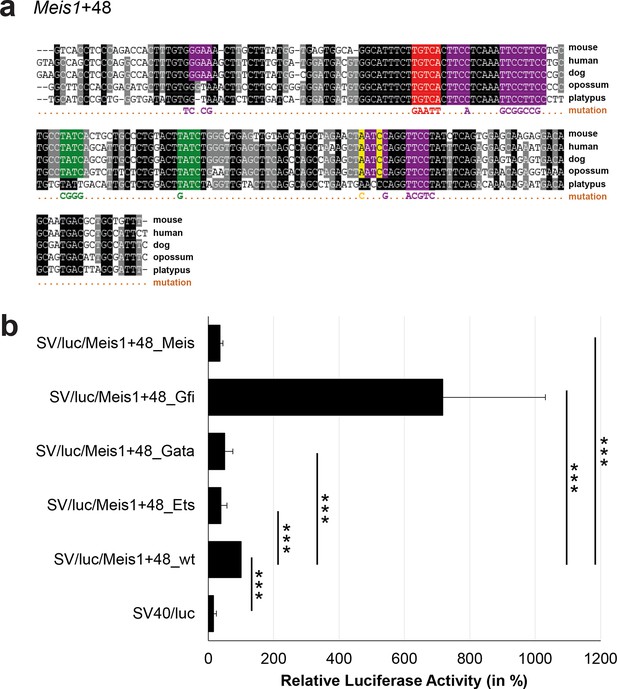
Multiple species alignment and luciferase assay results for Meis1+48.
(a) Multiple species alignment (MSA) with the following species: mouse (mm9), human (hg19), dog (canFam2), opossum (monDom5) and platypus (ornAna1). Nucleotides highlighted in black are conserved between all species analysed, nucleotides highlighted in grey are conserved between four of five species. Transcription factor binding sites (TFBS) are highlighted in: purple = Ets, green = Gata, yellow = Gfi, red = Meis. The nucleotides that were changed to mutate the TFBSs are indicated below the MSA. All conserved binding sites of one motif family (e.g. all Ets motifs) were mutated simultaneously. (b) For the luciferase reporter assays in stably transfected 416b cells, the averages of at least three independent experiments with three to four replicates within each experiment are shown. Error bars represent the standard error of the mean (SEM). Stars indicate significance: ***=p-value <0.001. p-values were generated using t-tests, followed by the Fisher’s method and if necessary Stouffer’s z trend.
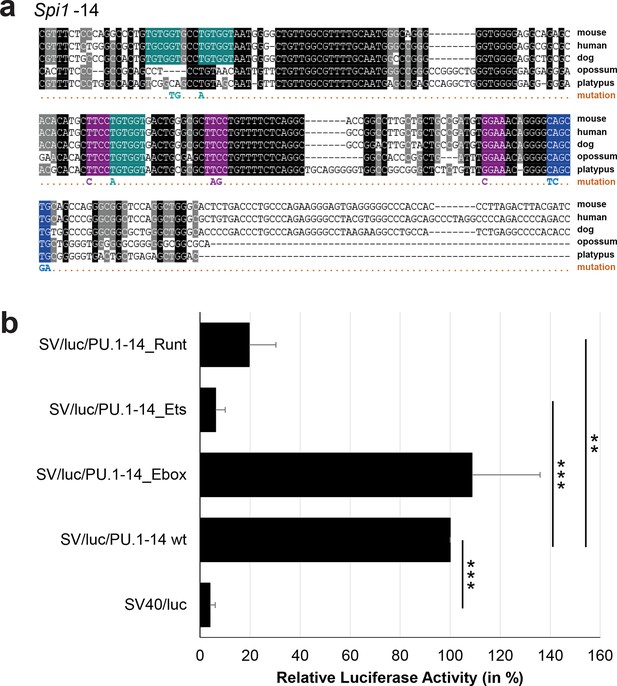
Multiple species alignment and luciferase assay results for Spi1-14.
(a) Multiple species alignment (MSA) with the following species: mouse (mm9), human (hg19), dog (canFam2), opossum (monDom5) and platypus (ornAna1). Nucleotides highlighted in black are conserved between all species analysed, nucleotides highlighted in grey are conserved between four of five species. Transcription factor binding sites (TFBS) are highlighted in: blue = Ebox, purple = Ets, turquoise = Runt. The nucleotides that were changed to mutate the TFBSs are indicated below the MSA. All conserved binding sites of one motif family (e.g. all Ets motifs) were mutated simultaneously. (b) For the luciferase reporter assays in stably transfected 416b cells, the averages of at least three independent experiments with three to four replicates within each experiment are shown. Error bars represent the standard error of the mean (SEM). Stars indicate significance: **=p-value <0.01, ***=p-value <0.001. p-values were generated using t-tests, followed by the Fisher’s method and if necessary Stouffer’s z trend.
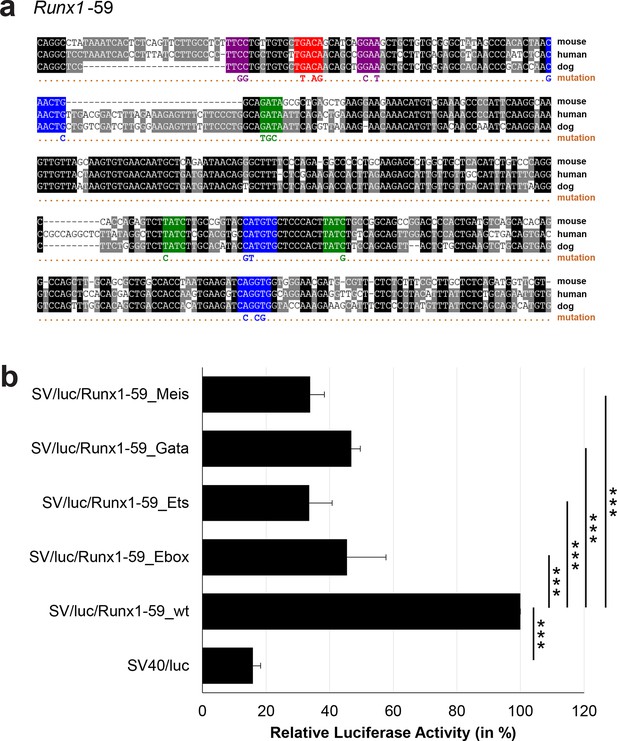
Multiple species alignment and luciferase assay results for Runx1-59.
(a) Multiple species alignment (MSA) with the following species: mouse (mm9), human (hg19) and dog (canFam2). Nucleotides highlighted in black are conserved between all species analysed, nucleotides highlighted in grey are conserved between two of three species. Transcription factor binding sites (TFBS) are highlighted in: blue = Ebox, purple = Ets, green = Gata, red = Meis. The nucleotides that were changed to mutate the TFBSs are indicated below the MSA. All conserved binding sites of one motif family (e.g. all Ebox motifs) were mutated simultaneously. (b) For the luciferase reporter assays in stably transfected 416b cells, the averages of at least three independent experiments with three to four replicates within each experiment are shown. Error bars represent the standard error of the mean (SEM). Stars indicate significance: ***=p-value <0.001. p-values were generated using t-tests, followed by the Fisher’s method and if necessary Stouffer’s z trend.
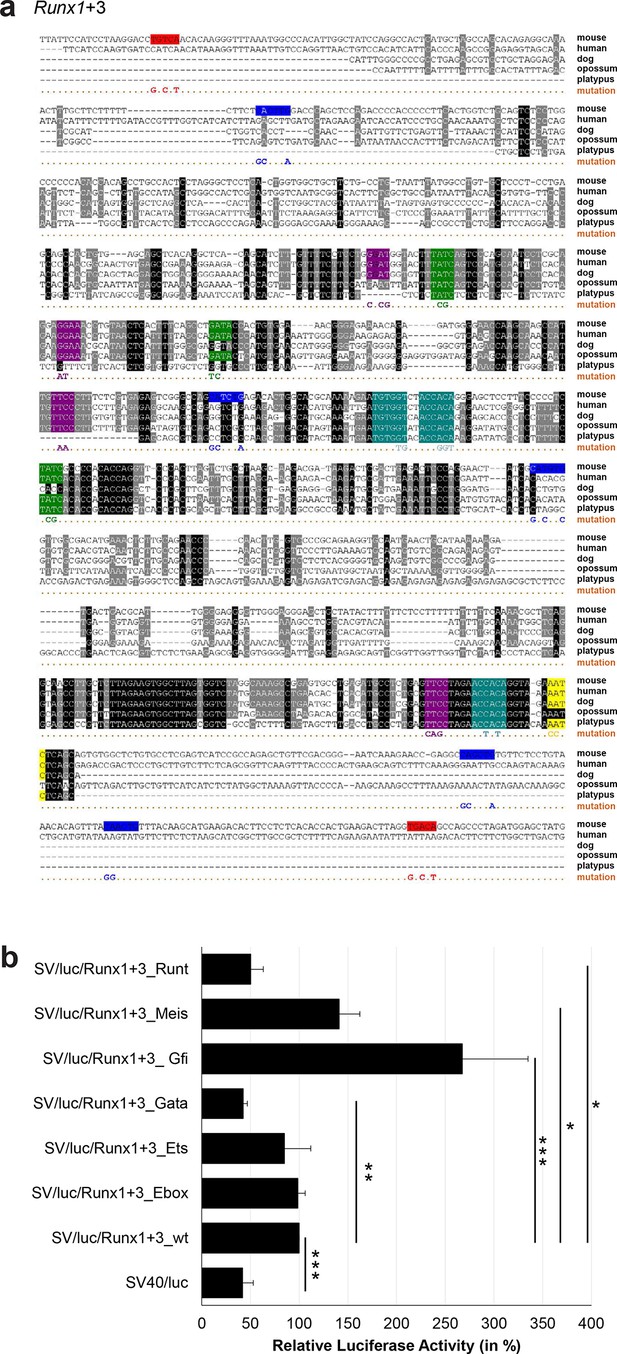
Multiple species alignment and luciferase assay results for Runx1+3.
(a) Multiple species alignment (MSA) with the following species: mouse (mm9), human (hg19), dog (canFam2), opossum (monDom5) and platypus (ornAna1). Nucleotides highlighted in black are conserved between all species analysed, nucleotides highlighted in grey are conserved between four of five species. Transcription factor binding sites (TFBS) are highlighted in: blue = Ebox, purple = Ets, green = Gata, yellow = Gfi, red = Meis, turquoise = Runt. The nucleotides that were changed to mutate the TFBSs are indicated below the MSA. All conserved binding sites of one motif family (e.g. all Ets motifs) were mutated simultaneously. Where TF binding was observed in ChIP-Seq experiments in 416b cells, but the TFBS was not conserved, the motifs present in the mouse sequence only were mutated. (b) For the luciferase reporter assays in stably transfected 416b cells, the averages of at least three independent experiments with three to four replicates within each experiment are shown. Error bars represent the standard error of the mean (SEM). Stars indicate significance: *=p-value <0.05, **=p-value <0.01, ***=p-value <0.001. p-values were generated using t-tests, followed by the Fisher’s method and if necessary Stouffer’s z trend.
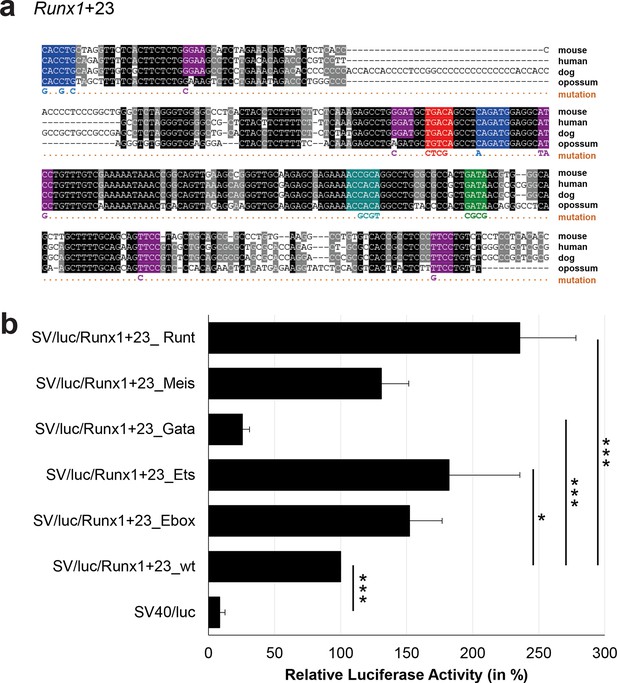
Multiple species alignment and luciferase assay results for Runx1+23.
(a) Multiple species alignment (MSA) with the following species: mouse (mm9), human (hg19), dog (canFam2) and opossum (monDom5). Nucleotides highlighted in black are conserved between all species analysed, nucleotides highlighted in grey are conserved between three to four species. Transcription factor binding sites (TFBS) are highlighted in: blue = Ebox, purple = Ets, green = Gata, red = Meis, turquoise = Runt. The nucleotides that were changed to mutate the TFBSs are indicated below the MSA. All conserved binding sites of one motif family (e.g. all Ebox motifs) were mutated simultaneously. (b) For the luciferase reporter assays in stably transfected 416b cells, the averages of at least three independent experiments with three to four replicates within each experiment are shown. Error bars represent the standard error of the mean (SEM). Stars indicate significance: *=p-value <0.05, ***=p-value <0.001. p-values were generated using t-tests, followed by the Fisher’s method and if necessary Stouffer’s z trend.
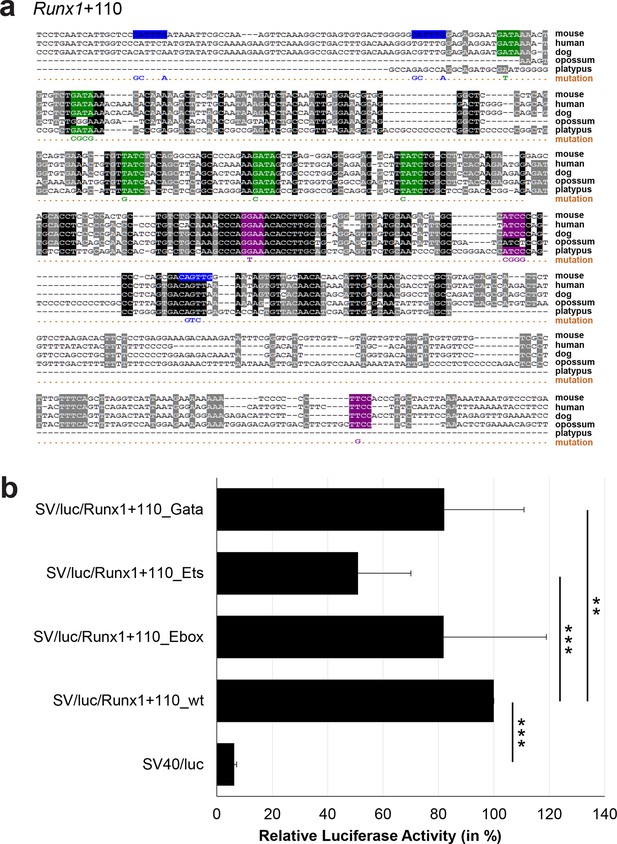

Multiple species alignment and luciferase assay results for Runx1+110.
(a) Multiple species alignment (MSA) with the following species: mouse (mm9), human (hg19), dog (canFam2), opossum (monDom5) and platypus (ornAna1). Nucleotides highlighted in black are conserved between all species analysed, nucleotides highlighted in grey are conserved between four of five species. Transcription factor binding sites (TFBS) are highlighted in: blue = Ebox, purple = Ets, green = Gata. The nucleotides that were changed to mutate the TFBSs are indicated below the MSA. All conserved binding sites of one motif family (e.g. all Ets motifs) were mutated simultaneously. Where TF binding was observed in ChIP-Seq experiments in 416b cells, but the TFBS was not conserved, the motifs present in the mouse sequence only were mutated. (b) For the luciferase reporter assays in stably transfected 416b cells, the averages of at least three independent experiments with three to four replicates within each experiment are shown. Error bars represent the standard error of the mean (SEM). Stars indicate significance: **=p-value <0.01, ***=p-value <0.001. p-values were generated using t-tests, followed by the Fisher’s method and if necessary Stouffer’s z trend.
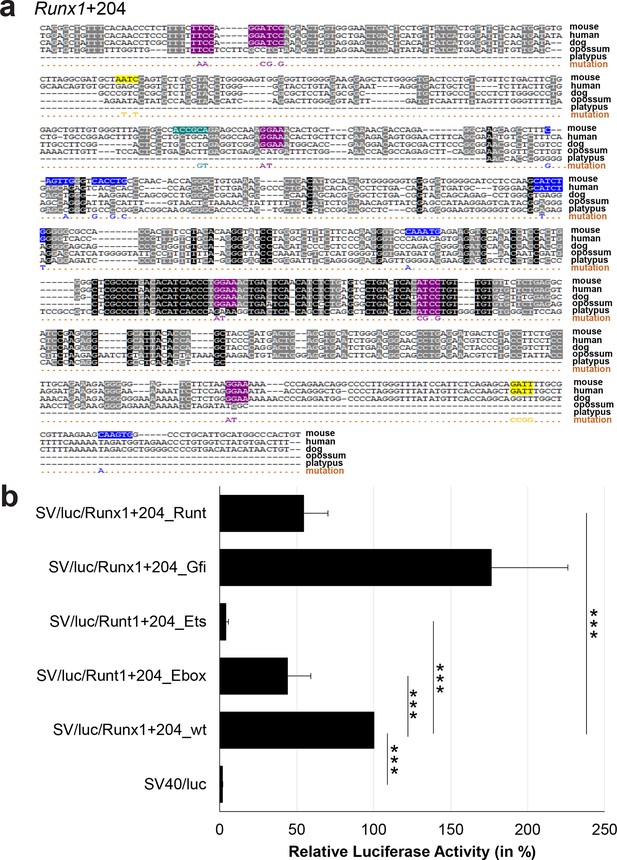
Multiple species alignment and luciferase assay results for Runx1+204.
(a) Multiple species alignment (MSA) with the following species: mouse (mm9), human (hg19), dog (canFam2), opossum (monDom5) and platypus (ornAna1). Nucleotides highlighted in black are conserved between all species analysed, nucleotides highlighted in grey are conserved between four of five species. Transcription factor binding sites (TFBS) are highlighted in: blue = Ebox, purple = Ets, yellow = Gfi, turquoise = Runt. The nucleotides that were changed to mutate the TFBSs are indicated below the MSA. All conserved binding sites of one motif family (e.g. all Ets motifs) were mutated simultaneously. Where TF binding was observed in ChIP-Seq experiments in 416b cells, but the TFBS was not conserved, the motifs present in the mouse sequence only were mutated. (b) For the luciferase reporter assays in stably transfected 416b cells, the averages of at least three independent experiments with three to four replicates within each experiment are shown. Error bars represent the standard error of the mean (SEM). Stars indicate significance: ***=p-value <0.001. p-values were generated using t-tests, followed by the Fisher’s method and if necessary Stouffer’s z trend.
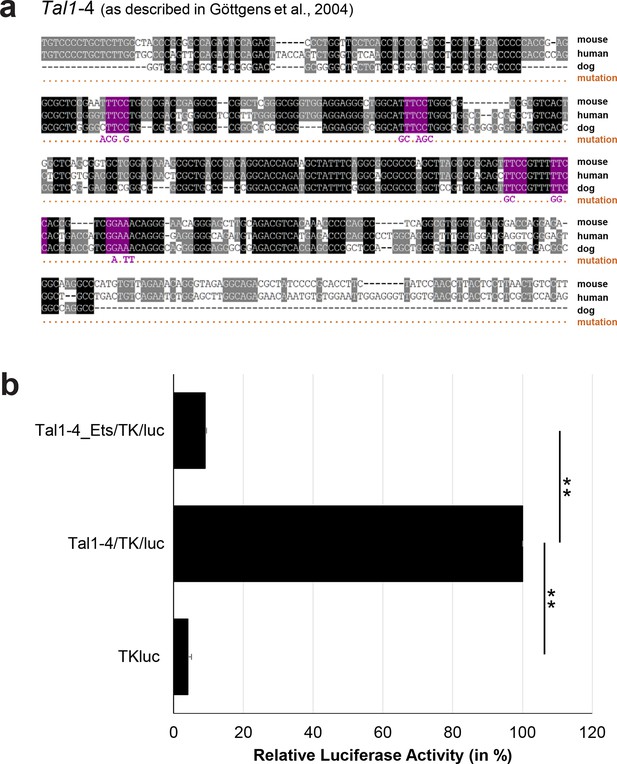
Multiple species alignment and luciferase assay results for Tal1-4.
(a) Multiple species alignment (MSA) with the following species: mouse (mm9), human (hg19) and dog (canFam2). Nucleotides highlighted in black are conserved between all species analysed, nucleotides highlighted in grey are conserved between two of three species. Transcription factor binding sites (TFBS) are highlighted in: purple = Ets. The nucleotides that were changed to mutate the TFBSs are indicated below the MSA. All conserved binding sites of the Ets motif family were mutated simultaneously. (b) For the luciferase reporter assays in stably transfected 416b cells, the averages of at least three independent experiments with three to four replicates within each experiment are shown. Error bars represent the standard error of the mean (SEM). Stars indicate significance: **=p-value <0.01. p-values were generated using t-tests, followed by the Fisher’s method and if necessary Stouffer’s z trend.
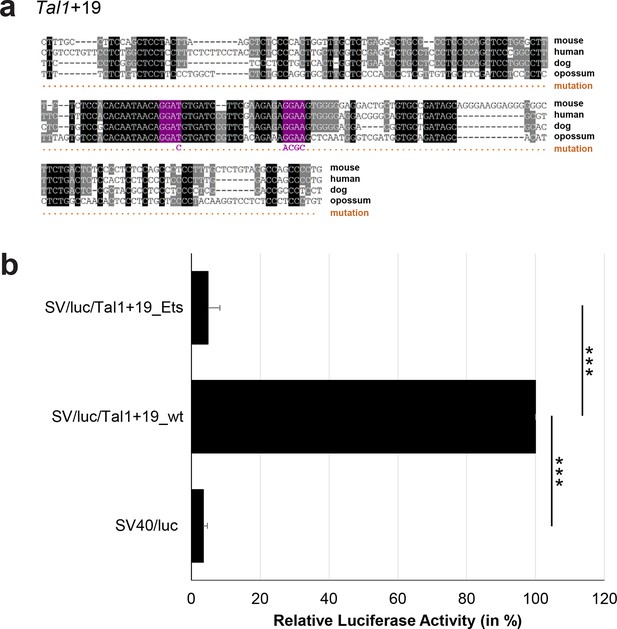
Multiple species alignment and luciferase assay results for Tal1+19.
(a) Multiple species alignment (MSA) with the following species: mouse (mm9), human (hg19), dog (canFam2) and opossum (monDom5). Nucleotides highlighted in black are conserved between all species analysed, nucleotides highlighted in grey are conserved between three of four species. Transcription factor binding sites (TFBS) are highlighted in: purple = Ets. The nucleotides that were changed to mutate the TFBSs are indicated below the MSA. All conserved binding sites of the Ets motif family were mutated simultaneously. (b) For the luciferase reporter assays in stably transfected 416b cells, the averages of at least three independent experiments with three to four replicates within each experiment are shown. Error bars represent the standard error of the mean (SEM). Stars indicate significance: ***=p-value <0.001. p-values were generated using t-tests, followed by the Fisher’s method and if necessary Stouffer’s z trend.
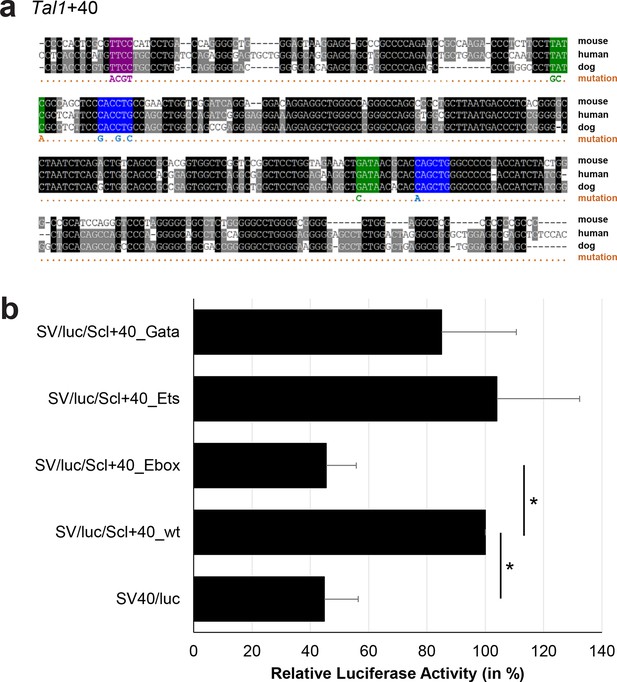
Multiple species alignment and luciferase assay results for Tal1+40.
(a) Multiple species alignment (MSA) with the following species: mouse (mm9), human (hg19) and dog (canFam2). Nucleotides highlighted in black are conserved between all species analysed, nucleotides highlighted in grey are conserved between two of three species. Transcription factor binding sites (TFBS) are highlighted in: blue = Ebox, purple = Ets, green = Gata. The nucleotides that were changed to mutate the TFBSs are indicated below the MSA. All conserved binding sites of one motif family (e.g. all Ebox motifs) were mutated simultaneously. (b) For the luciferase reporter assays in stably transfected 416b cells, the averages of at least three independent experiments with three to four replicates within each experiment are shown. Error bars represent the standard error of the mean (SEM). Stars indicate significance: *=p-value <0.05. p-values were generated using t-tests, followed by the Fisher’s method and if necessary Stouffer’s z trend.
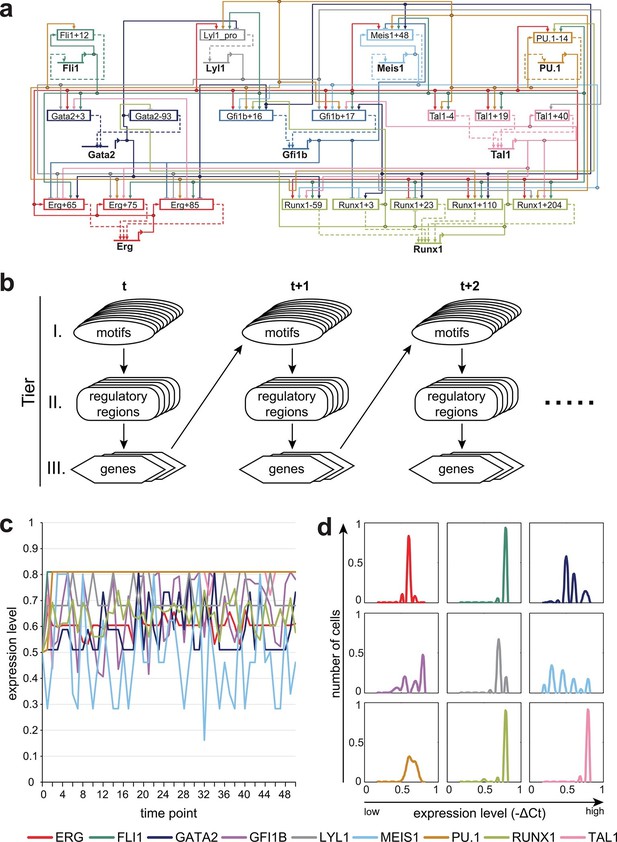
A three-tier dynamic Bayesian network (DBN) incorporating transcriptional regulatory information can recapitulate the HSPC expression state.
(a) Representation of the complete network diagram generated using the Biotapestry software (Longabaugh et al., 2005). (b) Schematic diagram describing the DBN which contains three tiers: I. TF binding motifs within regulatory regions, II. cis-regulatory regions influencing the expression levels of the various TFs, and III. genes encoding the TFs. The output of tier III, namely the expression levels of the TF, feed back into the TF binding at the various motifs of tier I. The model therefore is comprised of successive time slices (t). (c) Simulation of a single cell over time. The expression levels of all 9 TFs are the same at the beginning (0.5). The simulation rapidly stabilizes with characteristic TF expression levels. (d) Simulation of a cell population by running the model 1000 times. The scale of the x-axis is linear. Each simulation was run as described in (c).
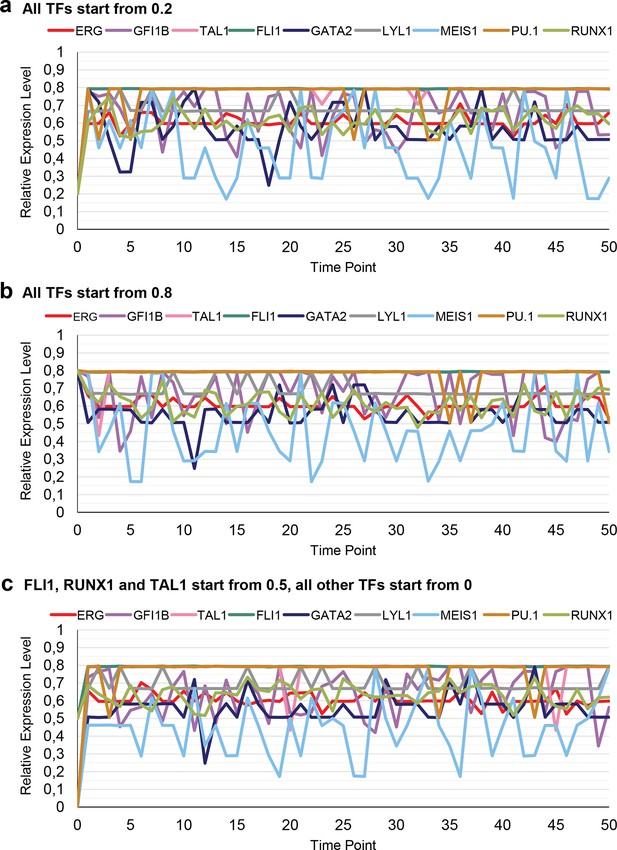
Simulation of a single cell over time with different expression levels at the beginning.
The simulation rapidly stabilizes with characteristic TF expression levels irrespective of the starting conditions. (a) The expression levels of all 9 TFs are 0.2 at the start of the simulation. (b) The expression levels of all 9 TFs are 0.8 at the start of the simulation. (c) The expression levels for FLI1, RUNX1 and TAL1 are set to be 0.5 at the beginning, with all other TFs not being expressed (value of 0).
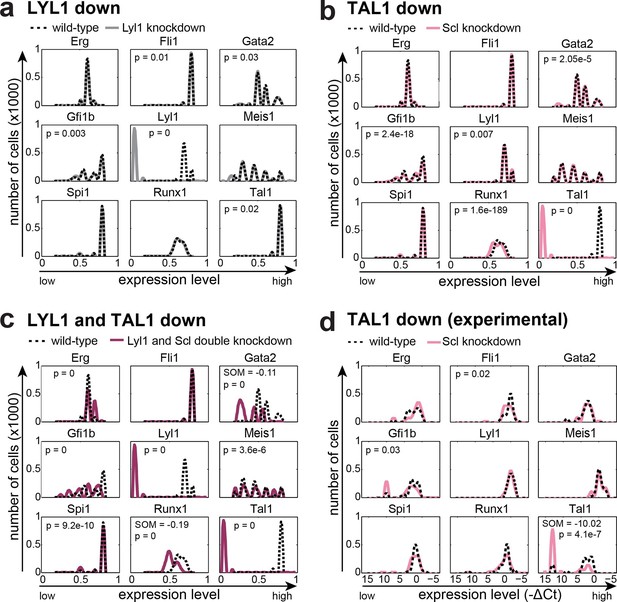
The DBN recapitulates the consequences of TAL1 and LYL1 single and double perturbations as seen in vivo and in vitro.
Computational prediction of gene expression patterns for the nine TFs of interest after perturbation of TAL1 (a), LYL1 (b) or both (c). Deletion of TAL1 or LYL1 on their own has no major consequences on the expression levels of the other eight TFs of the gene regulatory network, but simultaneous deletion of both TAL1 and LYL1 caused changes in expression of several genes, mainly a decrease in Gata2 and Runx1. This major disruption of the core GRN for blood stem/progenitor cells is therefore consistent with TAL1/LYL1 double knockout HSCs showing a much more severe phenotype than the respective single knock-outs. One thousand simulations were run for each perturbation to determine the TFs expression levels in a 'cell population' by selecting expression levels at random time points after reaching its initial steady state. Expression levels of 0 resemble no expression, whereas expression levels of 1 stand for highest expression level that is possible in this system. The scale of the x-axes is linear. (d) Gene expression levels measured in single 416b cells transfected with siRNA constructs against Tal1 or a control. The density plots of gene expression levels after perturbation of TAL1 indicate the relative number of cells (y-axes) at each expression level (x-axes). The scale of the x-axes is linear. The values indicate the results of the Wilcoxon rank-sum test: alterations to the expression profiles are indicated by the p-value (statistical significance: p <0.001 for computational data and p <0.05 for experimental data); substantial shifts in median expression level are indicated by the shift of median (SOM) (SOM >0.1 for computational data and >1 for experimental data). For details, see Figure 5—figure supplement 1; for full expression data, see Figure 5—source data 1 .
-
Figure 5—source data 1
Raw and normalised data for the single cell gene expression experiments presented in this study.
1) TAL1 down-regulation (related to Figure 5 d), 2) PU.1 down-regulation (related to Figure 6 a), 3) GFI1B up-regulation (related to Figure 6b) and 4) AML-ETO9a perturbation (related to Figure 6 c)
- https://doi.org/10.7554/eLife.11469.047
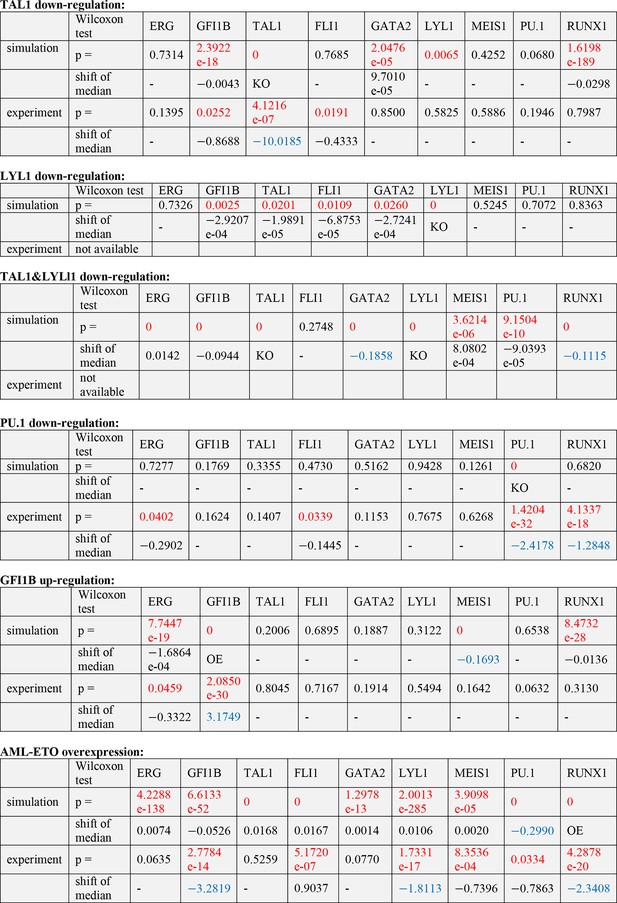
Significance tests for the computational and experimental data after TF perturbations.
To determine statistical significance the Wilcoxon rank-sum test was used. Alterations to the expression profiles are indicated by the p-value; with statistically significance defined as follows: p <0.001 for computational data and p <0.05 for experimental data. Significance of a substantial shift in median expression levels are as follows: shift of median >0.1 for computational data and >1 for experimental data (because of different scales). If the number for the shift of median is negative, the median of the perturbation data is smaller than that of the wild-type control; if the number is positive, the median of the perturbation is larger than that of the control. For simplicity, all significant changes are highlighted in red (p-value) and blue (shift of median).
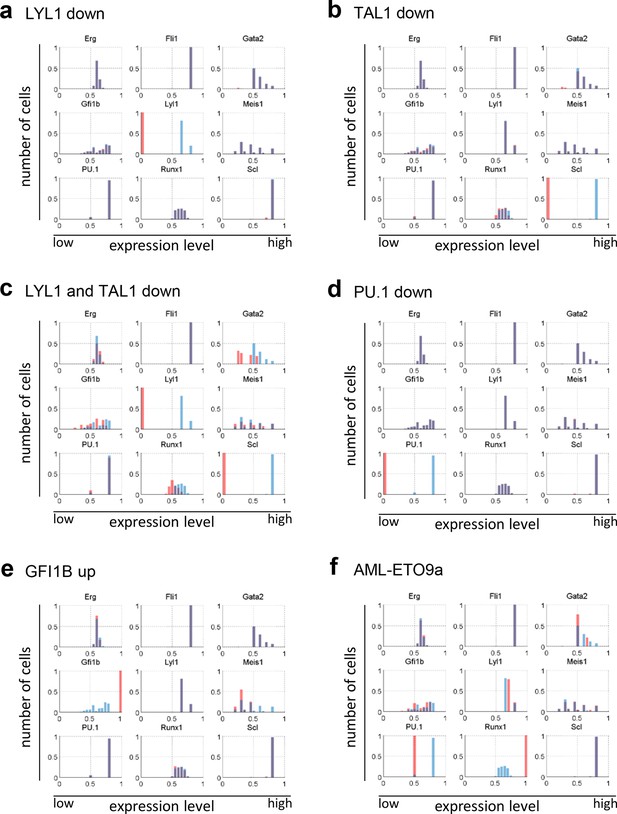
Histogram plots showing the gene expression distributions of all nine genes of the network for the perturbations presented in this study.
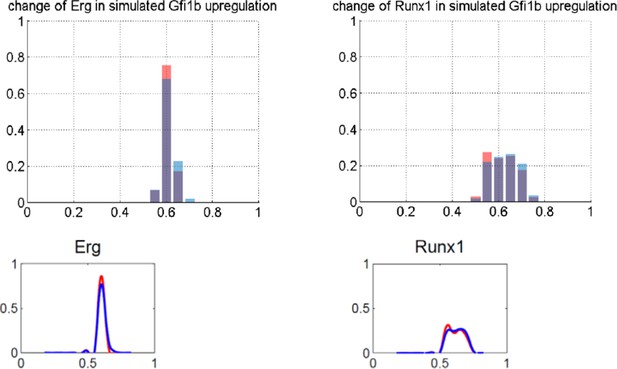
(a) LYL1 down-regulation; (b) TAL1/SCL down-regulation; (c) LYL1 and TAL1/SCL down-regulation; (d) PU.1 down-regulation; (e) GFI1B up-regulation; and (f) AML-ETO9a simulation.
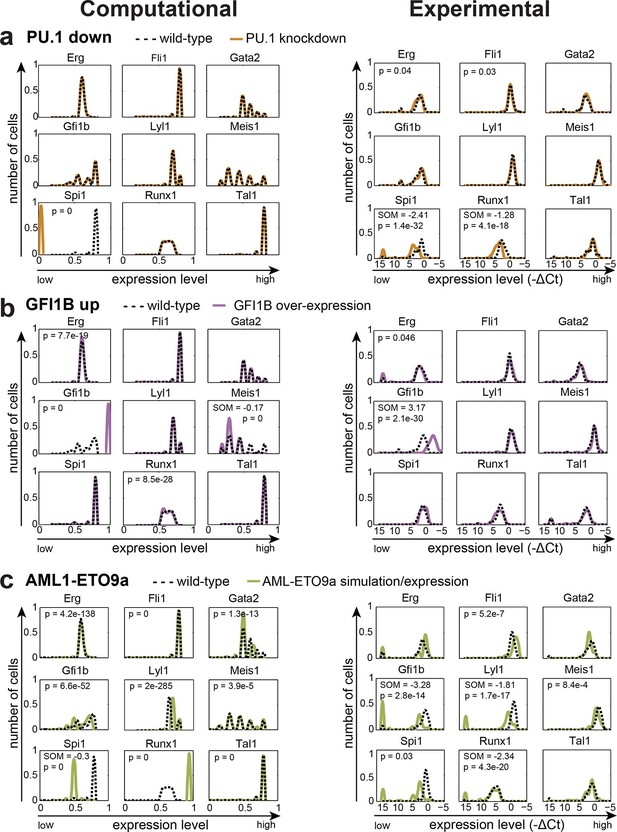
The DBN captures the transcriptional consequences of network perturbations.
Left panel: Computational prediction of gene expression after perturbation of specific TFs. 1000 simulations were run for each perturbation to determine expression levels in a 'cell population' (expression at 0 resembles no expression, whereas expression of 1 represents the highest possible expression level). The scale of the x-axes is linear. Right panel: Density plots of gene expression levels in single 416b cells after perturbation of specific TFs indicating the relative number of cells at each expression level. The scale of the x-axes is linear. The values indicate the results of the Wilcoxon rank-sum test: alterations to the expression profiles are indicated by the p-value (statistical significance: p<0.001 for computational data and p<0.05 for experimental data); substantial shifts in median expression level are indicated by the shift of median (SOM) (SOM >0.1 for computational data and >1 for experimental data). For details, see Figure 5—figure supplement 1. (a) PU.1 down-regulation: (Left) Computational prediction of gene expression after PU.1 knockdown (Spi1 was set to 0 after reaching its initial steady state). (Right) Gene expression levels measured in single 416b cells transduced with shRNA constructs against shluc (wild-type) or shPU.1 (PU.1 knockdown). (b) GFI1B overexpression: (Left) Computational prediction of gene expression after overexpression of GFI1B (Gfi1b was set to 1 after reaching its initial steady state). (Right) Gene expression levels in single 416b cells transduced with a GFI1B-expressing vector compared to an empty vector control (wild-type). (c) Consequences of the AML-ETO9a oncogene: (Left) Computational prediction of gene expression patterns after introducing the dominant-negative effect of the AML-ETO9a oncogene (Runx1 was fixed at the maximum value of 1 after reaching its initial steady state and in addition all Runt binding sites were set to have a repressive effect). (Right) Gene expression levels measured in single 416b cells transduced with an AML-ETO9a expressing vector fused to mCherry. mCherry positive cells were compared to mCherry negative cells (wild-type).
-
Figure 6—source data 1
Summary of all computational simulations for perturbations of one or two TFs.
The results for a total of 162 simulations are shown. The data can be accessed using the embedded hyperlinks. The y-axes show the number of cells and the x-axes the relative expression level. Blue curves represent wild-type data and red curves represent perturbation data.
- https://doi.org/10.7554/eLife.11469.051