A genome-wide resource for the analysis of protein localisation in Drosophila
Figures
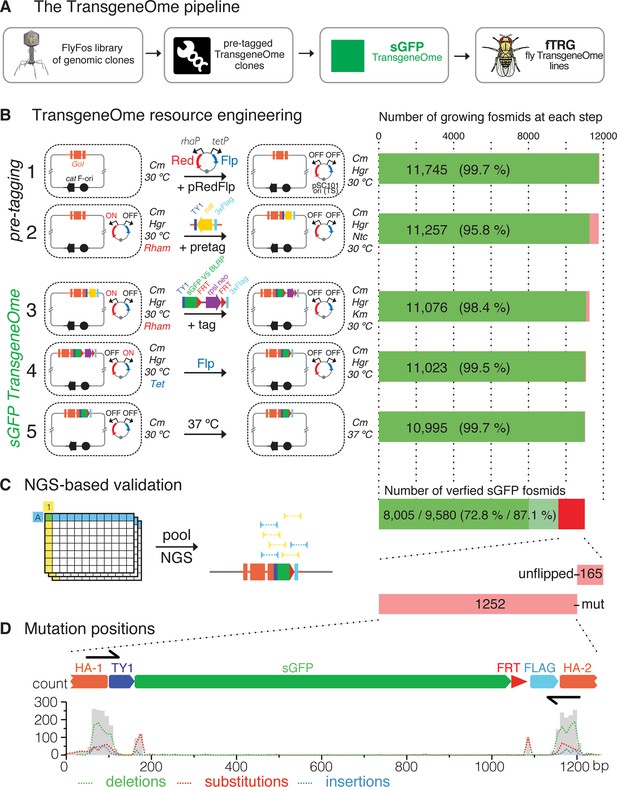
Generation of the TransgeneOme library.
(A) Overview of the tagging strategy. Liquid culture recombineering is used to insert a ‘pre-tagging’ cassette into FlyFos genomic clones in bacteria. This cassette can then be replaced by a simple, universal, recombineering reaction with any tag of choice, here, a superfolder GFP tag (sGFP) to generate the sGFP TransgeneOme clone library. These clones are transformed into flies generating transgenic FlyFos libraries that can be used for multiple in vivo applications. (B) TransgeneOme resource engineering. The steps of the recombineering pipeline are shown on the left with the success rate of each step indicated on the right (red colour denotes bacterial clones that did not grow). The E. coli cells are schematically represented with a dotted circle. With the first two steps the ‘pre-tagging’ cassette is inserted, which is replaced in the next three steps with the sGFP cassette to generate the sGFP TransgeneOme library. See text and methods section for details; abbreviations: GoI – gene of interest; cat – chloramphenicol resistance gene, F-ori – the fosmid vector replication control sequences; pRedFlp – recombineering helper plasmid with the pSC101 temperature sensitive origin of replication, which can be maintained at 30°C and is removed at the final step by temperature shift to 37°C; it carries the Red gbaA operon (Red), which drives homologous recombination in vivo under the control of the L-Rhamnose (Rham) inducible promoter (rhaP) and the Flp recombinase (Flp) under the control of tetracycline (Tet) inducible promoter (tetP); pretag – a pre-tagging cassette consisting of the Nourseothricin resistance gene (nat), flanked by the 2xTY and 3xFlag tag, which provide regions of homology to the tagging cassette (tag), consisting of the TY1 tag, superfolder GFP (sGFP), the V5 tag and the target peptide for the birA biotin ligase (blrp), the FRT-flanked selection/counter-selection operon rpsl-neo (confers streptomycin sensitivity and kanamycin resistance) and the 3xFlag tag. (C) Next-generation-sequencing (NGS)-based validation of the sGFP TransgeneOme library. Schematic of the bar coding (BC) strategy of row and column pools is shown to the left and sequencing results to the right. In this example, a clone with the coordinates A1 will receive the row A barcode (blue) and the column 1 barcode (yellow), which allow the mapping of the NGS sequence reads to the respective well. By using the mate-paired strategy the reads mapping to the tag can be assigned to a specific transgene, i.e. only reads where one mate of the pair maps to the tag and the other to the genome are used. Sequenced tags within fosmids without point mutations are shown in solid green, clones without mutation in tagging cassette but incomplete coverage in light green and clones with mutation(s) or un-flipped cassette are shown in red. (D) Statistics of the mutation distributions with deletions indicated by green, substitutions by red and insertions by blue interrupted lines. Note that most mutations reside within the recombineering primer sequences (denoted as black arrows).

Tagging cassettes.
Tags tested and used in this study. Shown is the form of the tagging cassette after insertion into the target site and flip-out of the counter selection sequences (rpsl-neo) leaving behind the FRT sequence coloured in red. While both the TY1-sGFP-FLAG and TY1-sGFP-V5-BLRP-FLAG tags can be used for localisation of the tagged protein of interest in vivo the latter offers additional affinity purification options and was the tag used for the genome-wide resource. The TY1-T2A-sGFPnls-FLAG results in cleavage at the T2A peptide and nuclear localization of the released sGFP-NLS fragment. Abbreviations: 2xTY1, 3xFlag, V5, Strep – epitope tags; BLRP – target peptide for the birA biotin ligase, allowing for tissue-specific labelling and purification (through tissue specific expression of birA); Pre (Prescission protease) and TEV (Tobacco Etch Virus endopeptidase) cleavage sites; T2A - 'self-cleaving' peptide; NLS – nuclear localization signal. The 'pre-tagging' cassette can be easily exchanged to any of the tagging cassettes through homologous recombination as it starts and ends with the same epitope tag encoding sequences.
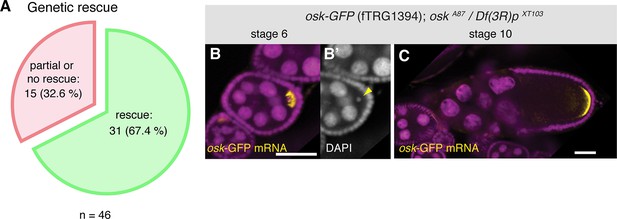
Functionality tests of the GFP-tagged fTRG lines by genetic complementation analysis.
(A) Genetic rescue statistics of null/strong mutant alleles for 46 selected fTRG lines. Note that more than two-thirds of the lines show a rescue (see Table 2). (B, C) osk-GFP mRNA (in yellow) expressed from fTRG1394 rescues egg-chamber development of an osk null allele (Jenny et al., 2006). osk-GFP mRNA enriches in the early oocyte (B, stage 6) and rescues the oogenesis arrest and the DNA condensation defect of the osk mutant (B’, yellow arrowhead). At stage 10 osk-GFP RNA enriches at the posterior pole (C) and produces sufficient protein to ensure proper embryogenesis. osk-GFP mRNA is shown in yellow, DAPI in magenta; scale bars indicate 30 µm.
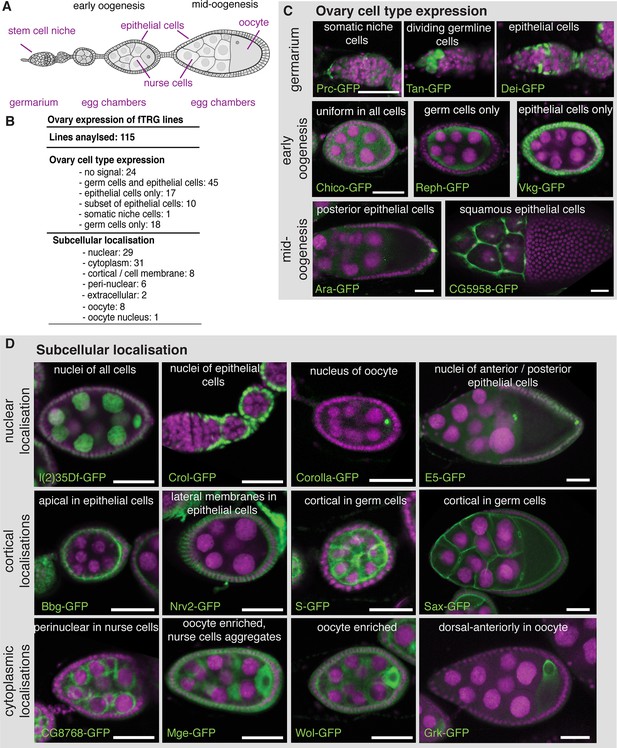
Expression of fTRG tagged proteins in ovaries.
(A) Schematic overview of oogenesis stages and cell types. (B) Summary of the identified expression patterns; see also Supplementary file 3. (C) Selected examples for cell type specific fTRG expression patterns at germarium, early- and mid-oogenesis stages visualised by anti-GFP antibody staining. (D) Selected examples of subcellular localisation patterns, highlighting nuclear, cortical and cytoplasmic patterns at different oogenesis stages. GFP is show in green, DAPI in magenta; scale bars indicate 30 µm.
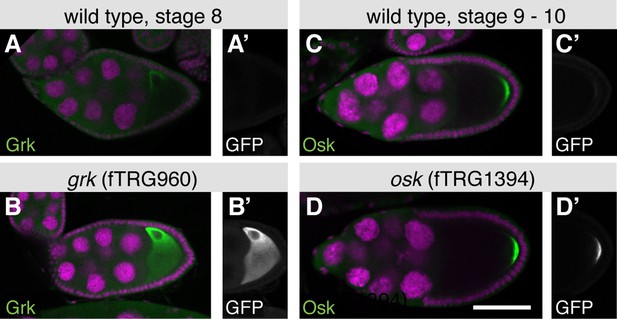
Co-localisation of fTRG derived tagged proteins with endogenous proteins during oogenesis.
(A, B) Endogenous untagged Grk protein detected with an anti-Grk antibody localises similarly in wild-type stage 8 oocytes (A), as in fTRG960 oocytes expressing Grk-GFP detected by an anti-GFP antibody (B). Note that both Grk and GFP antibody patterns are indistinguishable (compare B and B’). (C, D) Osk protein detected by an anti-Osk antibody in wild type stage 9–10 oocytes (C) compared to Osk antibody labelled protein in an fTRG1394 oocyte expressing Osk-GFP in addition to endogenous Osk (D). Note the co-localisation of anti-Osk and anti-GFP antibodies (D and D’). Scale bars indicate 30 µm.
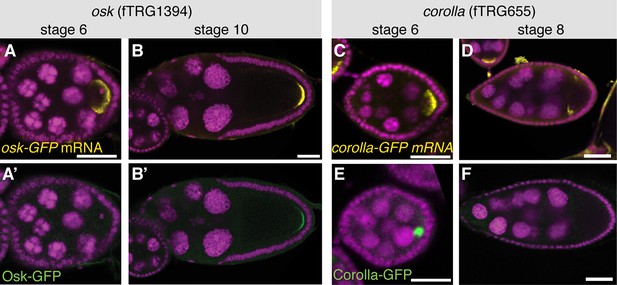
Posttranscriptional regulation of protein expression during oogenesis.
(A, B) osk-GFP mRNA visualised by an anti-GFP labelled RNA probe (yellow, DAPI in magenta) at stage 6 and stage 10 of oogenesis. (A’, B’) Osk-GFP protein visualised by anti-GFP antibody (green, DAPI in magenta) at stage 6 and stage 10. Note that Osk-GFP protein is not detectable at stage 6. (C, D) corolla-GFP mRNA (yellow, DAPI in magenta) at stage 6 and stage 8. (E, F) Corolla-GFP protein (green, DAPI in magenta) at stage 6 and stage 8. Note that Corolla-GFP protein is only detectable at stage 6 but not stage 8. Scale bars indicate 30 µm.
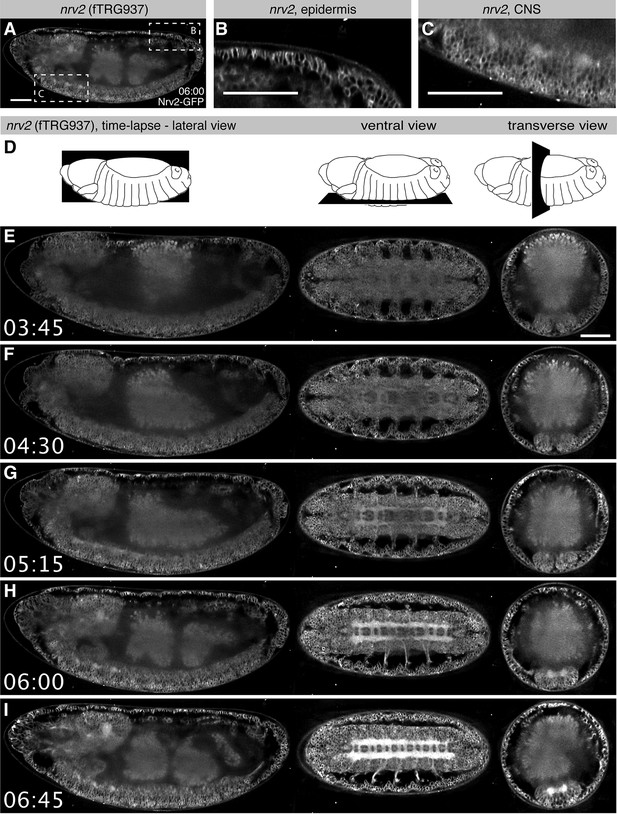
Live in toto imaging during embryogenesis with SPIM.
(A-C) Nrv2-GFP protein is enriched in cell membrane of the epidermis and the CNS of late stage 16 embryos, as shown by a lateral section (A) and high magnifications of the posterior epidermis (B) and the ventral CNS (C). (D) Schemes of the lateral, ventral and transverse optical section views through the embryo shown in (E-I). (E-I) Still image from a Nrv2-GFP time-lapse Video with lateral section views on the left, ventral sections in the middle and transverse sections on the right. Note that Nrv2-GFP is first expressed in the developing epidermal epithelial cells (E, F) and then becomes enriched in the CNS (G-I, see Video 1). Scale bars indicate 50 µm.
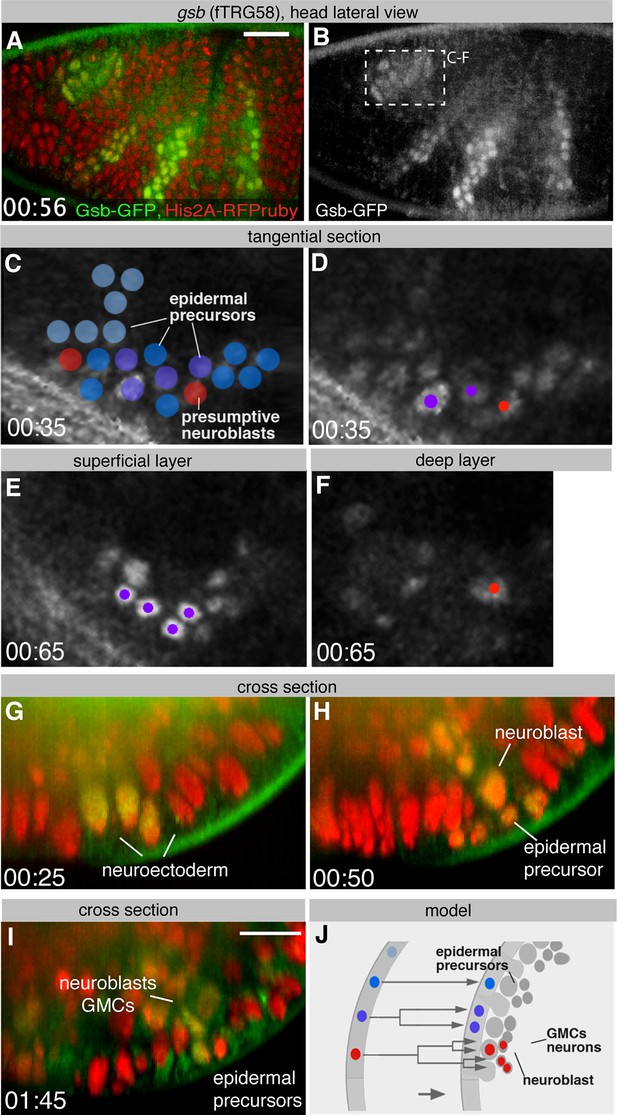
Live Gsb-GFP imaging during embryogenesis with SPIM.
(A, B) Lateral view of a live stage 10 embryo expressing Gsb-GFP (green in A) and Histone2A-mRFPruby (red in A); anterior is to the left (see Video 2). (C-F) Confocal sections of gsb-positive deuterocerebral proneural domain (recorded with a regular confocal microscope at a similar position as boxed in B). Fate of cells is symbolized by different colours (blue: epidermal precursor undergoing no further mitosis; purple: epidermal precursor undergoing one mitosis; red: neuroblasts). (C) and (D) show an optical section through the neurectoderm of stage 10 embryo prior to neuroblast delamination; (E) and (F) display the same region 30 min later, after neuroblasts have delaminated (stage 11), with a superficial optical section of surface ectoderm (E), and deep section of the neuroblast layer (F). (G-I) Optical cross sections (rotated by 90 degrees) of a similar embryo as in (A) expressing GFP-tagged Gsb (green) and Histone-2A-mRFPruby (red) at stage 10 (G), early 11 (H) and late 11 (I) showing neuroblasts delaminating from Gsb-GFP domain. (J) Schematic cross section of stage 10 (left) and stage 11 (right) ectoderm illustrating fate of cells forming part of Gsb-positive pro-neural domain. Scale bars indicate 25 µm (A, B) and 10 µm (G-I).
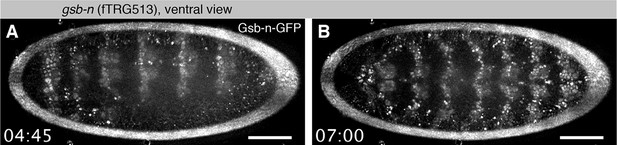
Live Gsb-n-GFP imaging during embryogenesis with SPIM.
(A, B). Ventral view of Gsb-n-GFP expression of a stage 12 embryo during germ-band retraction (A) and stage 14 during head involution (B). Note that Gsb-n-GFP remains expressed in neuronal precursors during stage 14 (Video 3). Scale bars indicate 50 µm.
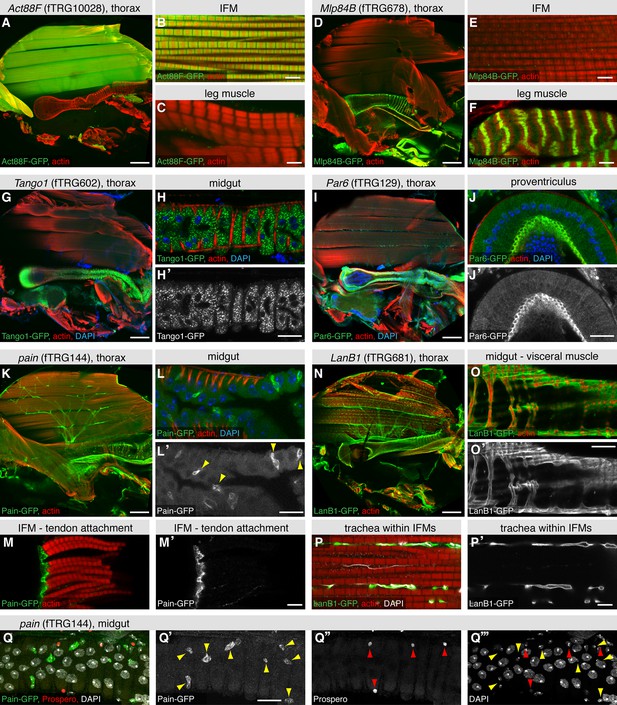
Expression of fTRG tagged proteins in tissues of the adult thorax.
Antibody stainings of the adult thorax with anti-GFP antibody (green) and phalloidin (red). (A-F) Act88-GFP expression is specific to the IFMs, where it labels the thin filaments (B), whereas Mlp84B specifically labels the Z-discs of leg muscles (F). (G-J) Tango1-GFP concentrates in a vesicle-like pattern in the gut epithelium (H, H'), whereas Par6-GFP is highly expressed in trachea (I) and the gut epithelium, where it concentrates at the apical membrane, as shown for a cross-section of the proventriculus (J, J'), nuclei are labelled with DAPI (blue). (K-M) Pain-GFP expression in the flight muscle motor neurons (K), small cells within the midgut epithelium (L, L') and tendon cells (M, M'). (N-P) LanB1-GFP labels the extracellular matrix surrounding the IFMs, the motor neurons and the trachea (N), as well as the visceral muscles (O). Even the finest trachea marked by UV auto-fluorescence (white) (P) are surrounded by LanB1-GFP (P’). (Q) Pain-GFP positive cells in the midgut do not overlap with Prospero positive nuclei of enterocytes (in red) and contain small nuclei, as visualised by DAPI co-stain in white (Q-Q’’’). Scale bars indicate 100 µm (A, D, I, K, N), 20 µm (H,J, L, O, Q) and 5 µm (B,C,E,F,M,P).
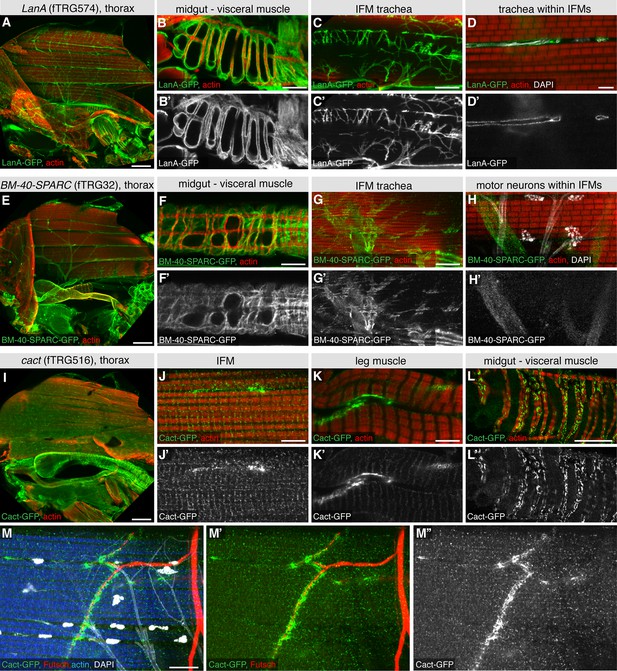
Extracellular matrix and synaptic markers of the adult thorax.
(A-H) Antibody stainings of the adult thorax with anti-GFP antibody (green or white in the single colour images) and phalloidin (red). LanA-GFP and BM-40-SPARC-GFP labels the ECM around motor neurons, visceral muscle and trachea. Note that thin trachea within the IFMs (marked by UV auto-fluorescence in white) are surrounded by LanA-GFP but not BM-40-SPARC-GFP (D, H). (I-M) Cact-GFP (green) shows a distinct localisation in IFMs, leg and visceral muscle reminiscent of a neuromuscular junction pattern. Note the partial co-localisation with the motor neuron marker Futsch (in red, M, M’), whereas no co-localisation with trachea in IFMs (in white, M). Scale bars indicate 100 µm (A, E, I), 20 µm (B, C, F, G, L), 10 µm (M) and 5 µm (D, H, J, K).
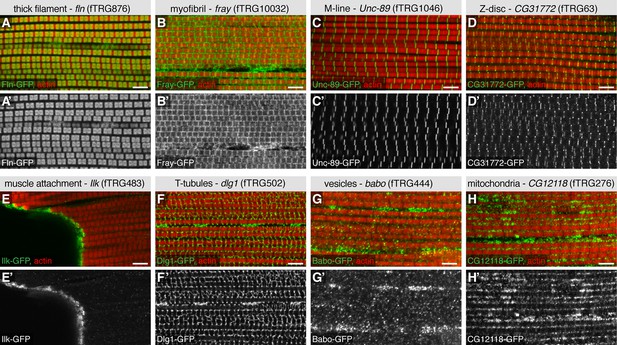
Subcellular expression patterns in adult flight muscles.
Antibody stainings of the adult thorax with anti-GFP antibody (green or white in the single colour images) and phalloidin (red). (A-D) Localisation to specific myofibrillar sub-regions; Fln-GFP marks the thick filaments (A, A’), Fray-GFP surrounds the myofibrils with an enrichment at M-lines and Z-discs (B, B’), Unc-89-GFP marks only M-lines (C, C’) and CG31772-GFP only Z-discs (D, D’). (E-H) Ilk-GFP strongly concentrates at the muscle-tendon attachment sites (E, E’), Dlg1-GFP labels the T-tubular membranes (F, F’), Babo-GFP shows a dotty, vesicular pattern (G, G’) and CG12118-GFP displays a mitochondrial pattern (H, H’). Scale bars indicate 5 µm.

CG12118-GFP localises to mitochondria.
Co-staining of adult IFMs from fTRG276 (CG12118-GFP) with anti-GFP antibody (green) and the mitochondrial marker anti-complex V α (ATPase Vα, in red). Note the strong overlap of both signals. Scale bar indicate 5 µm.
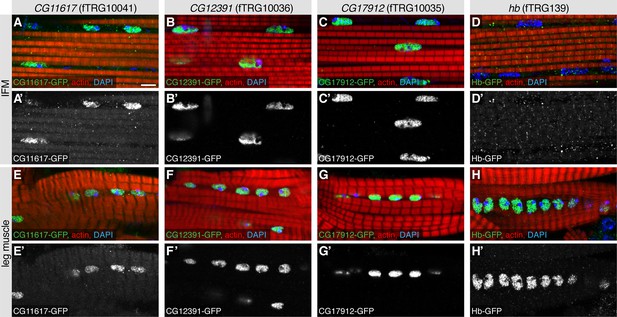
Nuclear localisations in adult flight muscles.
Antibody stainings of the adult thorax with anti-GFP antibody (green or white in the single colour images), phalloidin (red) and DAPI (blue). (A-H) CG11617-GFP (A, E), CG12391-GFP (B, F) and CG17912 (C, G) are localised to the nuclei of IFMs and leg muscles, whereas Hb-GFP is only found in leg muscle nuclei (H) and not detectable in IFM nuclei (D). Scale bars indicate 5 µm.
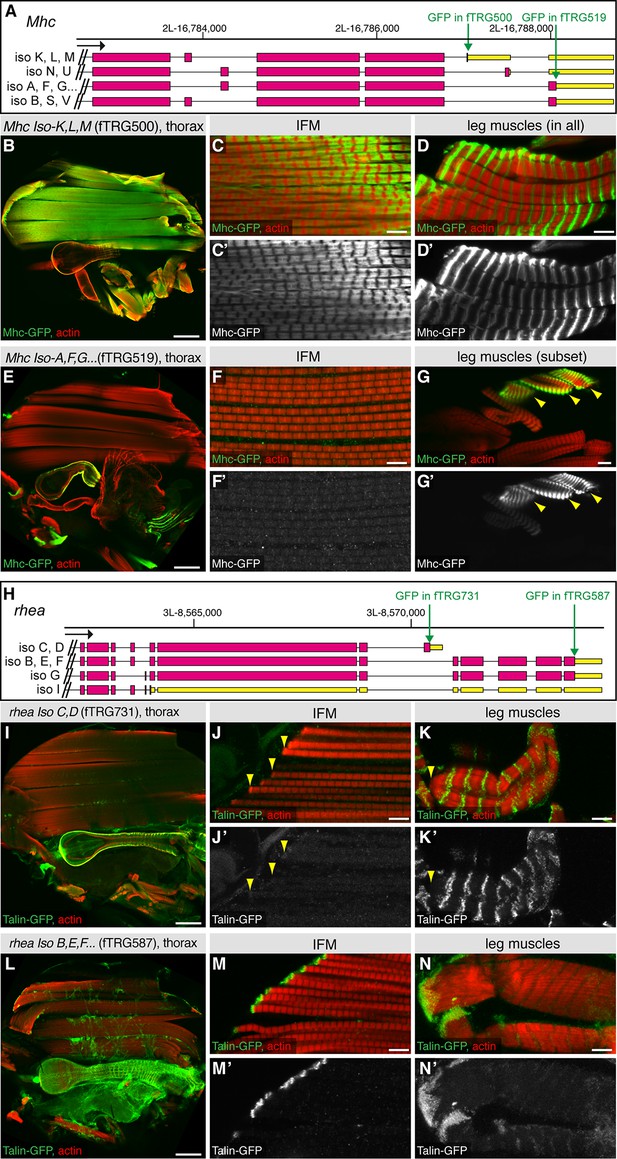
Alternative splicing into alternative C-termini.
Antibody stainings of the adult thorax with anti-GFP antibody (green or white in the single colour images) and phalloidin (red). (A, H) Gene models of the 3’ end of Mhc (A) and rhea (H) listing the predicted isoforms; coding exons are shown in pink, 3’-UTRs in yellow boxes. The positions of the GFP tag insertions are marked by green arrows. (B-G) The shorter Mhc-GFP isoforms (Iso K, L, M) are expressed in IFMs and all leg muscles (B-D), whereas the slightly longer Mhc-GFP isoforms (Iso A, F, G etc.) are not detectable in IFMs but present in visceral muscles and a subset of leg muscles (E-G). (I-N) The shorter Talin-GFP isoforms (rhea Iso C, D) are not detectable at muscle-tendon attachment sites in IFMs (J, arrowheads) and leg muscles (K, arrowhead), however do localise to costamers of leg muscles (K). However, the long Talin-GFP isoforms (rhea Iso B, E, F, G) do localise to muscle-tendon attachment sites in IFMs (M) and leg muscles (N). Scale bars indicate 100 µm (B, E, I, L), 10µm (G) and 5 µm (C, D, F, J, K, M, N).
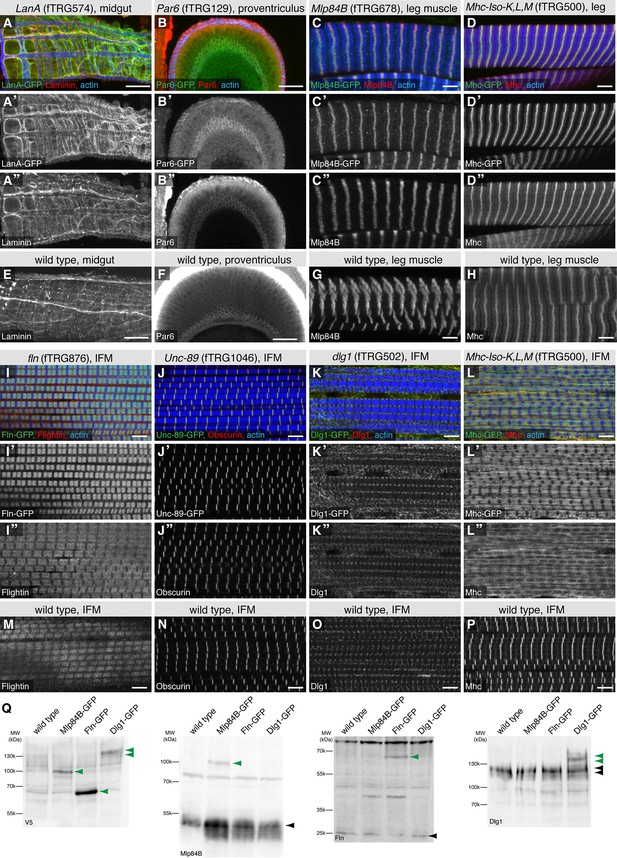

Co-localisation of fTRG tagged proteins with endogenous proteins.
Antibody stainings of the adult thorax from fTRG or wild-type lines with anti-GFP antibody (green or white in the single colour images) and antibodies against various fly proteins (red). (A-D) Co-localisation of LanA-GFP with anti-Laminin antibody stain around the midgut (A), of Par6-GFP with anti-Par6 at the apical side of the proventriculus epithelium (B) and of Mlp84B-GFP as well as Mhc-GFP with anti-Mlp84B and anti-Mhc antibody stain in leg muscles, respectively (C, D). (E-H) Adult thoraces from wild-type flies show very similar patterns with the respective antibodies. (I-L) Adult IFMs showing the co-localisation of Fln-GFP with anti-Fln antibody staining (I), Unc-89/Obscurin-GFP with anti-Obscurin antibody staining (J), Dlg1-GFP with anti-Dlg1 antibody staining (K) and Mhc-GFP with anti Mhc antibody staining (L). (M-P) The same antibodies result in very similar patterns in wild-type IFMs apart from the a sharp versus a diffuse Mhc pattern comparing wild-type to Mhc-GFP flies (L, P). (Q) Western blots loaded with total protein extract from wild-type, Mlp84B-GFP, Fln-GFP and Dlg1-GFP adult males probed with anti-V5 (included in the GFP tag) anti-Mlp84B, anti-Fln and anti-Dlg1 antibodies. Note the about 40 kDa size shift of the tagged proteins in the respective lanes (marked with green arrow heads) versus the untagged protein band (black arrow heads).
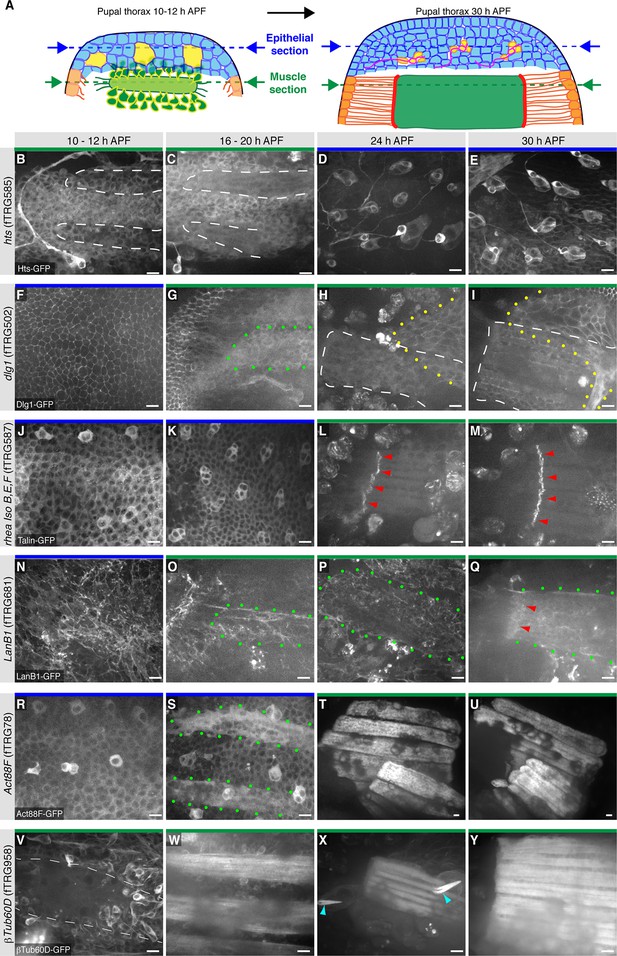
Live imaging of fTRG tagged proteins in living pupal thorax.
(A) Schematic drawing of a 10–12 hr (left) and a 30h pupal thorax (right). The developing epidermis is shown in blue, with the SOP precursors in yellow (developing neurons in red), the differentiating tendons are shown in orange, the myoblasts and muscle fibers in green, and the muscle-tendon junction in red. The schematic positions of the optical sections through epithelium and muscles are indicated with blue and green dotted lines, respectively. (B-Y) Live imaging of pupal thoraces at the indicated stages acquired with a spinning disc confocal (except S and T, which were acquired with a two-photon microscope). Blue bars above the image indicate epithelial sections and green bars indicate muscle sections (as explained in A). Hts-GFP is expressed in fusing myoblasts (B, C) and strongly in developing SOPs (D, E). Dlg1-GFP labels the epithelial junctions (F), internal muscle structures (green dots, G) and an unidentified additional developing epithelium (yellow dots, H, I). Talin-GFP is higher expressed in developing SOPs (J, K) and strongly localised to the muscle-tendon junction from 24 hr APF (red arrowheads, L, M). LanB1-GFP localises to the basal side of the developing epithelium (N) and surrounds the forming muscle fibers (green dots, O-Q) with a slight concentration at the muscle-tendon junction at 30 hr APF (red arrowheads, Q). Act88F-GFP weakly labels the developing epithelium, with a slight concentration in the SOPs until 20 hr APF (R, S) and very strongly marks the IFMs from 24 hr onwards (T, U). βTub60D-GFPis expressed in the fusing myoblasts (V, W) and also labels the microtubule bundles in the developing muscle fibers (X, Y) and hair cells of the developing sensory organs (light blue arrow heads in X). Scale bars indicate 10 µm.
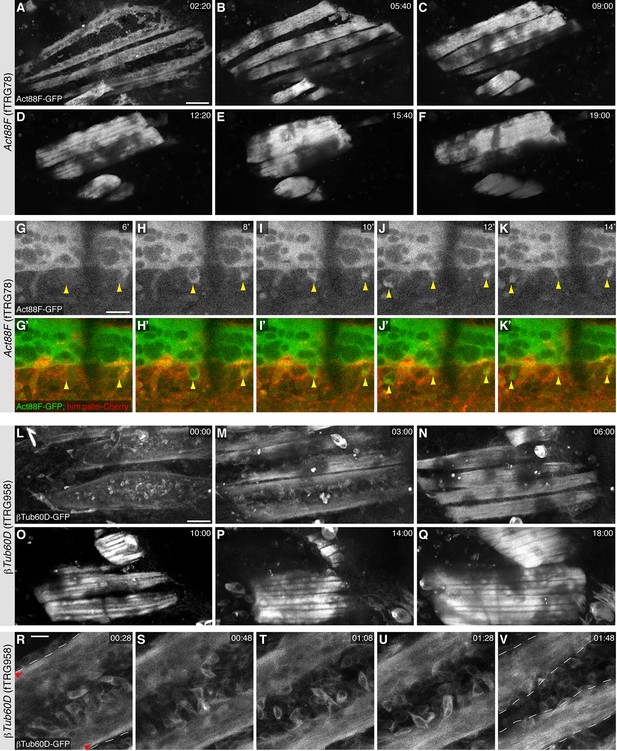
Live imaging during pupal development.
(A-F) Stills from live two-photon imaging of an intact 14 hr APF pupa expressing Act88F-GFP strongly labelling the developing IFMs (see Video 4). (G-K) Stills from a two-colour spinning disc Video expressing Act88F-GFP (green) and him-GAL4, UAS-palm-Cherry (red) labelling the myoblasts (see Video 5). Note the sudden green label of single myoblasts after fusion had occurred (yellow arrowheads, see Video 5). (L-Q) Stills from two-photon Video of an intact 14 hr APF pupa expressing βTub-60D-GFP in fusing myoblasts and developing myofibers (See Video 6). (R-V) Stills from a high resolution two-photon Video of an intact 16 hr APF pupa expressing βTub-60D-GFP. Single myoblast during fusion can be resolved (See Video 7). Strong microtubule bundles (red arrow heads) are visible close to the edges of the splitting myotube (white dashed lines, R); splitting is complete in (V). Scale bars indicate 50 µm (A-F, L-Q) and 10 µm (G-K) and (R-V).
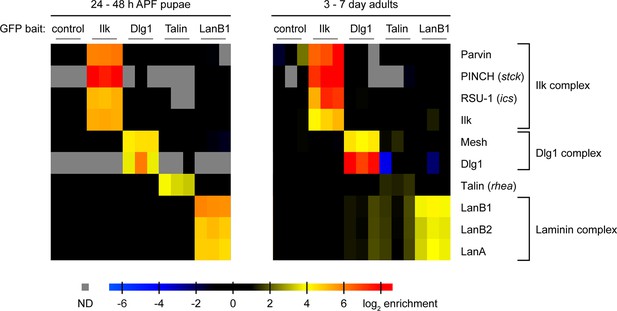
Proteomics with fTRG bait proteins.
GFP-tagged Ilk, Dlg1, Talin, and LanB1, respectively, were affinity-enriched from protein extracts generated from whole pupae (left) or adult flies (right) using anti-GFP immunoprecipitation. A wild-type fly strain not expressing any GFP-tagged protein served as control. Proteins were quantified using mass spectrometry and the MaxLFQ label-free quantification algorithm in MaxQuant. Selected proteins are visualized by their enrichment factors in individual samples over the control (or simulated noise level, if not detected in the control). Specific interaction partners are characterised by the similarity of the quantitative profiles and co-enrichment with the respective bait proteins
Videos
Multi-view SPIM Video of a stage 12 Nrv2-GFP expressing embryo.
A stack was acquired every 15 min, lateral, dorsal, ventral and transverse views of the same time points are displayed. From stage 11 onwards Nrv2-GFP is present ubiquitously in the plasma membrane. Later, its expression increases in the CNS, particularly in the neuropil and the motor neurons. Video plays with 7 frames per second. Time is given in hh:mm. Scale bar indicates 50 µm.
Lateral head section from a SPIM Video of a stage 10 Gsb-GFP (green, white in the top Video), Histone-2A-mRFPruby (red) embryo.
A stack was acquired every 7 min. The segmentally re-iterated stripe-like gsb expression domain in the head neuroectoderm is visible. Later, gsb is expressed in ganglion mother cells and nerve cells that are the progeny of gsb expressing neuroblasts. Video plays with 7 frames per second. Time is given in hh:mm. Scale bar indicates 50 µm.
Ventral view of a SPIM Video of a stage 6 Gsb-n-GFP embryo.
A stack was acquired every 15 min. Gsb-n-GFP is only detectable at the end of germ-band extension. During germ-band retraction, it is expressed in characteristic L-shaped expression domains in the hemi-segments of the trunk. In the late stage embryo, Gsb-n-GFP is present in the neurons of the shortening ventral nerve cord. Video plays with 7 frames per second. Time is given in hh:mm. Scale bar indicates 50 µm.
Z-projection of a two-photon Video of an about 14 hr APF pupa expressing Act88F-GFP.
A stack was acquired every 20 min for 19 hr. Expression of Act88F-GFP increases in the indirect flight muscles dramatically, thus contrast was reduced several times in course of the Video to avoid over-exposure. Video plays with 5 frames per second. Time is given in hh:mm.
Single plane of a spinning disc confocal Video of an about 14 hr APF old pupa expressing Act88F-GFP (green) in the flight muscle myotubes and him-GAL4; UAS-palm-Cherry in the myoblasts.
An image stack was acquired every two minutes. Note the newly fused myoblasts acquired the GFP label within a single time interval (highlighted by green arrows). Video plays with 5 frames per second. Time is given in minutes.
Z-projection of a two-photon Video of an about 14 hr APF pupa expressing βTub60D-GFP.
A stack was acquired every 20 min for 25 hr. Note the high expression of βTub60D-GFP in fusing myoblasts and the thick microtubules bundles in the developing flight muscles. Hair cells of the developing sensory organs also show strong expression, however, move out of the Z-stack over time. Video plays with 5 frames per second. Time is given in hh:mm.
Single plane of a two-photon Video of an about 16 hr APF old pupa expressing βTub60D-GFP in myoblasts and the forming flight muscle myotubes.
An image stack was acquired every two minutes for more than 3 hr. Note that single myoblasts can be followed during fusion. Most myoblasts fuse at the center of the myotube, which gradually splits into two myotubes. Video plays with 5 frames per second. Time is given in hh:mm.
Tables
TransgeneOme constructs and fTRG lines - overview of TransgeneOme constructs generated and verified by sequencing for the different pilot sets and the genome-wide set, including the respective numbers of the transgenic fTRG lines generated.
| Tagged constructs and transgenic lines | |||
|---|---|---|---|
| constructs | verified constructs | transgenic lines | |
| ‘pre-tagging’ - TransgeneOme | 11257 | -- | -- |
| TY1-sGFP-V5-BLRP-FLAG | 799 | ||
| - TransgeneOme | 10995 | 9580 | |
| - pilot set | 1328 | 1328 | |
| TY1-T2A-sGFPnls-FLAG | |||
| - pilot set | 273 | 273 | 30 |
| TY1-sGFP-FLAG | |||
| - pilot set | 644 | 483 | 51 |
| unique constructs | 23169 | 10711 | 880 |
| unique genes | 11257 | 9993 | 826 |
Genetic rescue of mutants with the fTRG lines Table listing fTRG lines and respective genetic alleles as well as rescue assays that were used to assess the functionality of the fTRG lines. Note that about two-thirds of the lines rescue the mutant phenotypes.
| Gene | Chromosome | fTRG line | Tag | Rescue? | Rescue assay | Alleles, deficiencies used in trans for rescue assay | Reference |
|---|---|---|---|---|---|---|---|
| amos | 2nd | fTRG_218 | 2xTY1-sGFP-V5-preTEV-BLRP-3xFLAG | yes | antenna size and bristle number rescued to normal | amos[3] | |
| anterior open (aop, Yan) | 2nd | fTRG_142 | 2xTY1-sGFP-V5-preTEV-BLRP-3xFLAG | yes | embryonic lethality rescued to viable adults | aop[1] (BL-3101); aop[Yan1] (BL-8780) | |
| aubergine (aub) | 2nd | fTRG_581 | 2xTY1-sGFP-V5-preTEV-BLRP-3XFLAG | yes | female sterility entirely rescued | aub[HN2] (BL-8517); Df(2L)BSC145 (BL-9505) | |
| baboon (babo) | 2nd | fTRG_444 | 2xTY1-sGFP-V5-preTEV-BLRP-3xFLAG | yes | lethality rescued to viable adults | babo[32] (BL-5399); babo[k16912] (BL-11207) | |
| bag of marbles (bam) | 3rd | fTRG_3 | 2xTY1-sGFP-V5-preTEV-BLRP-3xFLAG | yes | female sterility entirely rescued | bam[delta86]; Df(3R)exel9020 | Christian Bökel, pers. comm. |
| cactus (cact) | 2nd | fTRG_516 | 2xTY1-sGFP-V5-preTEV-BLRP-3xFLAG | yes | lethality and female sterility rescued | cact[1]; cact[4] | |
| CG32121 | 3rd | fTRG_92 | 2xTY1-sGFP-V5-preTEV-BLRP-3xFLAG | yes | flightlessness rescued | CG32121 [XZ1] (X. Zhang and F.S., unpublished); Df(3L)ED4502 (BL-8097) | |
| CG6509 (dlg5) | 2nd | fTRG_10251 | 2xTY1-sGFP-3xFLAG | yes | lethality rescued to viable adults | CG6509 [KG006748] (BL13692); Df(2L)BSC244 (BL-9718) | |
| discs large 1 (dlg1) | X | fTRG_502 | 2xTY1-sGFP-V5-preTEV-BLRP-3xFLAG | yes | male lethality rescued to viable adults | Dlg1[5] (BL-36280) | |
| dorsal (dl) | 2nd | fTRG_29 | 2xTY1-sGFP-V5-preTEV-BLRP-3xFLAG | yes | bristle number rescued to normal | dl[1]; dl[4] | |
| ebi | 2nd | fTRG_10141 | 2xTY1-sGFP-3xFLAG | yes | lethality rescued to viable adults | ebi[CCS-8] (BL-8397); ebi[E90] (BL-30720) | |
| escargot (esg) | 2nd | fTRG_10170 | 2xTY1-sGFP-3xFLAG | no | lethality not rescued | esg[35Ce-1] (BL-3900); esg[35Ce-3] (BL-30475) | |
| eyes absent (eya) | 2nd | fTRG_492 | 2xTY1-sGFP-V5-preTEV-BLRP-3XFLAG | no | lethality not rescued | eya[C0233]; eya[C0275] | |
| fat (ft) | 2nd | fTRG_10233 | 2xTY1-T2A-nlsGFP-3xFLAG | yes | lethality rescued to viable adults | ft[G-rv] (BL-1894); ft[8] (BL-5406) | |
| 48 related 2 (Fer2) | 3rd | fTRG_334 | 2xTY1-sGFP-V5-preTEV-BLRP-3XFLAG | yes | defective climbing rescued to wild type | Fer2[e03248] | Dib et al., 2014 |
| fizzy (fzy) | 2nd | fTRG_10250 | 2xTY1-T2A-nlsGFP-3xFLAG | no | lethality not rescued | fzy[1] (BL-2492); fzy[3] (BL-25143) | |
| flightless I (fliI) | X | fTRG_467 | 2xTY1-sGFP-V5-preTEV-BLRP-3xFLAG | yes | lethality or flightlessness rescued | fli[14] (BL-7481); fli[3] (BL-4730) | |
| Hand | 2nd | fTRG_10163 | 2xTY1-sGFP-3xFLAG | yes | semi-lethality rescued to viable adults | Hand[173] | |
| hippo (hpo) | 2nd | fTRG_10130 | 2xTY1-sGFP-3xFLAG | yes | larval lethality rescued to viable adults | hpo[KS240] (BL-25085); hpo[KC202] (BL-25090) | |
| HLH54F | 2nd | fTRG_153 | 2xTY1-sGFP-V5-preTEV-BLRP-3xFLAG | yes | lethality rescued to viable adults | bHLH54F[598]; Df(2R) Exel7150 (BL-7891) | |
| Integrin linked kinase (Ilk) | 3rd | fTRG_483 | 2xTY1-sGFP-V5-preTEV-BLRP-3xFLAG | yes | embryonic lethality rescued to viable adults (wing blisters) | Ilk[1] (BL-4861); Df(3L)BSC733 (BL-26831) | |
| Kinesin heavy chain (Khc) | 2nd | fTRG_10243 | 2xTY1-T2A-nlsGFP-3xFLAG | yes | lethality rescued to viable adults | Khc[8] (BL-1607); Khc[1ts] (BL-31994) | |
| LanB1 | 2nd | fTRG_681 | 2xTY1-sGFP-V5-preTEV-BLRP-3XFLAG | yes | lethality rescued to viable adults | LanB1 [KG03456] (BL-13957); Df(2L) Exel7032 (BL-7806) | |
| multiple ankyrin repeats single KH domain (mask) | 3rd | fTRG_486 | 2xTY1-sGFP-V5-preTEV-BLRP-3xFLAG | yes | lethality rescued to viable adults | mask[10.22]/ Df(3R)BSC317 | Barry Thompson, pers. comm |
| midline (mid) | 2nd | fTRG_490 | 2xTY1-sGFP-V5-preTEV-BLRP-3xFLAG | no | lethality not rescued | mid[B1295]; mid[C2372] | |
| numb | 2nd | fTRG_25 | 2xTY1-sGFP-V5-preTEV-BLRP-3xFLAG | yes | lethality rescued to viable adults | numb[1] (BL-4096); Df(2L)30A-C (BL-3702) | |
| odd skipped (odd) | 2nd | fTRG_47 | 2xTY1-sGFP-V5-preTEV-BLRP-3xFLAG | no | lethality not rescued | odd[5] (BL-5345); Df(2L) Exel7018 (BL-7789) | |
| optomotor-blind-related-gene-1 (org-1) | X | fTRG_485 | 2xTY1-sGFP-V5-preTEV-BLRP-3xFLAG | no | male lethality not rescued | org-1[OJ487] | |
| oskar (osk) | 3rd | fTRG_1394 | 2XTY1-SGFP-V5-preTEV-BLRP-3XFLAG | yes | female sterility entirely rescued | osk[A87]/ Df(3R)p-XT103 | |
| Pabp2 | 2nd | fTRG_565 | 2xTY1-sGFP-V5-preTEV-BLRP-3XFLAG | no | lethality not rescued | Pabp2[01] (BL-9838); Pabp2[55] (BL-38390) | |
| patched (ptc) | 2nd | fTRG_82 | 2xTY1-sGFP-V5-preTEV-BLRP-3xFLAG | yes | lethality rescued to viable adults | ptc[9] (BL-3377); ptc[16] (BL-35500) | |
| retina abarrent in pattern (rap) | X | fTRG_1253 | 2xTY1-sGFP-V5-preTEV-BLRP-3XFLAG | yes | lethality rescued to viable adults | rap[ie28] | Yuu Kimata, pers. comm. |
| rhea (Talin) | 3rd | fTRG_587 | 2xTY1-sGFP-V5-preTEV-BLRP-3XFLAG | yes | embryonic lethality rescued to viable adults | rhea[1]; rhea[79] | |
| RhoGEF2 | 2nd | fTRG_591 | 2xTY1-sGFP-V5-preTEV-BLRP-3XFLAG | yes | embryonic lethality rescued to viable adults | RhoGEF2 [04291] | Jörg Großhans, pers. comm. |
| roundabout (robo) | 2nd | fTRG_567 | 2xTY1-sGFP-V5-preTEV-BLRP-3XFLAG | no | lethality not rescued | robo[1] (BL-8755); robo[2] (BL-8756) | |
| saxophone (sax) | 2nd | fTRG_10070 | 2xTY1-sGFP-3xFLAG | yes | lethality rescued to viable adults | sax[4] (BL-5404); sax[5] (BL-8785) | |
| scribbler (sbb) | 2nd | fTRG_443 | 2xTY1-sGFP-V5-preTEV-BLRP-3xFLAG | no | lethality not rescued | sbb[04440] (BL-11376); Df(2R)BSC334 (BL-24358) | |
| Sin3A | 2nd | fTRG_596 | 2xTY1-sGFP-V5-preTEV-BLRP-3XFLAG | no | lethality not rescued | Sin3A[08269] (BL-12350); Sin3A [B0948] | |
| smoothened (smo) | 2nd | fTRG_599 | 2xTY1-sGFP-V5-preTEV-BLRP-3XFLAG | yes | lethality rescued to viable adults | smo[3] (BL-3277); smo[119B6] (BL-24772) | |
| snail (sna) | 2nd | fTRG_71 | 2xTY1-sGFP-V5-preTEV-BLRP-3xFLAG | no | lethality not rescued | sna[18] (BL-2311); sna[1] (BL-25127) | |
| spalt major (salm) | 2nd | fTRG_165 | 2xTY1-sGFP-V5-preTEV-BLRP-3xFLAG | no | lethality not rescued | salm[1] (BL-3274); Df(2L)32FP-5 (BL-29717) | |
| Target of rapamycin (Tor) | 2nd | fTRG_713 | 2xTY1-sGFP-V5-preTEV-BLRP-3XFLAG | no | lethality not rescued | Tor[deltaP] (BL-7014); Df(2L) Exel7055 (BL-7823) | |
| traffic jam (tj) | 2nd | fTRG_163 | 2xTY1-sGFP-V5-preTEV-BLRP-3xFLAG | no | sterility not rescued | tj[PL3] (BL-4987); Df(2L)Exel8041 (BL-7849) | |
| viking (vkg) | 2nd | fTRG_595 | 2xTY1-sGFP-V5-preTEV-BLRP-3XFLAG | no | lethality not rescued | vkg[01209] (BL-11003); Df(2L) Exel7022 (BL-7794) | |
| Unc-89/ Obscurin | 2nd | fTRG_1046 | 2xTY1-sGFP-V5-preTEV-BLRP-3XFLAG | yes | flightlessness rescued | Unc-89[EY15484] | |
| yorkie (yki) | 2nd | fTRG_875 | 2xTY1-sGFP-V5-preTEV-BLRP-3XFLAG | yes | lethality rescued to viable adults | yki[B5] | Barry Thompson, pers. comm. |
in totoSPIM imaging of fTRG lines in the embryo Table listing the fTRG lines that were imaged in the embryo using Zeiss Lightsheet Z.1 from multiple angles over time. nrv2, gsb and gsb-n are discussed in the text. For the remaining lines, we list broad categorisation of the expression detected by SPIM imaging.
| fTRG number | FBgn_id | Gene symbol | Signal | Embryonic expression | Movie | Beads |
|---|---|---|---|---|---|---|
| 58 | FBgn0001148 | gsb | strong | tissue-specific expression | Yes | Yes |
| 71 | FBgn0003448 | snail | weak | tissue-specific expression | Yes | Yes |
| 88 | FBgn0025360 | Optix | medium | tissue-specific expression | Yes | Yes |
| 94 | FBgn0010433 | ato | weak | tissue-specific expression | Yes | Yes |
| 137 | FBgn0259685 | crb | medium | tissue-specific expression | Yes | Yes |
| 155 | FBgn0029123 | SoxN | strong | tissue-specific expression | Yes | Yes |
| 349 | FBgn0024294 | spn43Aa | strong | late expression, deposited in the cuticle | Yes | Yes |
| 513 | FBgn0001147 | gsb-n | medium | tissue-specific expression | Yes | Yes |
| 937 | FBgn0015777 | nrv2 | strong | ubiquitous expression, membrane signal | Yes | Yes |
Summary of adult muscle fTRG expression patterns 54 detected adult muscle localisation patterns (flight muscle, leg muscle and visceral muscle) from Supplementary file 4 are summarised. fTRG line number is listed in brackets.
| Thick filament | Thin filament / myofibril | M-line | Z-disc | Muscle attachment site | T-tubules / sarcolemma | Dotty pattern / vesicles (?) | Mitochondria | Nucleus | Neuro-muscular junction |
|---|---|---|---|---|---|---|---|---|---|
| Fln (876, IFM) | Act88F (78, 10028, IFM) | Prm (475, IFM) | CG31772 (63) | Ilk (483) | Dlg1(502) | Babo (444) | CG12118 (276) | Adar (570) | Cact (516) |
| Mf (Iso-A,G, N, 501) | Fray (125, 10032) | Unc-89 (1046) | Kettin (Sls-Isoform, 569) | Talin (rhea, Iso-B, E, F, G, 587) | Sax (10070) | CG5885 (10017, leg m.) | Blimp-1 (10149) | Veli (10125) | |
| Mhc (Iso-K, L, M, 500) | Hsp83 (10010) | Lmpt (584, I-band, leg m.) | β-PS Integrin (mys, 932) | CLIP-190 (156) | CG11617 (10041) | ||||
| Mhc (Iso-A, F, 519, leg m. subset & visceral. m.) | TpnC25D (1257, leg m. & visceral m.) | Mask (486, IFM) | Dlg5 (CG6509, 10251, IFM) | CG12391 (10036) | |||||
| Prm (475, leg m. & visceral m.) | TpnI (wupA, 925, leg m. & visceral m.) | Mlp60A (709, leg m. & visceral m.) | Hts (585) | CG17912 (10035) | |||||
| Mlp84B (678, leg m. & visceral m.) | Mask (486, leg m.) | CG32121 (92) | |||||||
| Talin (Iso-C, D, 731, leg m. & visceral m.) | Pyd3 (53) | Dorsal (29, leg m.) | |||||||
| Rho1 (31) | E2F2 (115) | ||||||||
| Sc2 (79, 10039) | Gro (21) | ||||||||
| Tango11 (699) | Hand (10163, visceral m.) | ||||||||
| Tsc1 (59) | Hb (139, leg m.) | ||||||||
| Vps35 (CG5625, 57) | Mnt (34) | ||||||||
| P53 (84) | |||||||||
| Salm (165) | |||||||||
| Vri (182) |
Additional files
-
Supplementary file 1
TransgeneOme constructs.
Construct and clone names, tag locations, as well the sequencing validation data are listed for all TransgeneOme constructs generated. Sheet 1 lists the sGFP TransgeneOme (TY1-sGFP-V5-BLRP-FLAG tag, NGS sequenced), sheet 2 the TY1-sGFP-FLAG pilot set clones (junctions Sanger sequenced, only exact matches are counted as verified), sheet 3 the TY1-T2A-sGFPnls-FLAG pilot set clones (entire tag Sanger sequenced) and sheet 4 the TY1-sGFP-V5-BLRP-FLAG pilot set clones (entire tag Sanger sequenced). Sheet 5 summarises all verified tagged genes in these sets.
- https://doi.org/10.7554/eLife.12068.033
-
Supplementary file 2
fly TransgeneOme lines (fTRG) lines.
Table listing all 880 transgenic FlyFos (fTRG) lines, with fTRG numbers, construct and clones names, as well as nature of the tag and the used landing site. The second sheet compares the genes tagged by the fTRG lines to the available GFP gene trap lines. 765 genes are only found in the TransgeneOme resource.
- https://doi.org/10.7554/eLife.12068.034
-
Supplementary file 3
fTRG expression in ovaries.
Table listing the expression patterns for 115 fTRG lines in ovaries. Expression was detected in 94 lines by anti-GFP antibody stainings. Cell type specific expression and subcellular localisations were monitored for these lines.
- https://doi.org/10.7554/eLife.12068.035
-
Supplementary file 4
fTRG expression in the adult thorax.
Table listing the expression pattern for 121 fTRG lines in adult thoraces. Expression was detected in 101 lines by anti-GFP antibody stainings. Cell type specific expression and subcellular localisations were monitored for these lines.
- https://doi.org/10.7554/eLife.12068.036
-
Supplementary file 5
Proteomics quantification.
Quantitative mass spectrometry values of all detected protein obtained with the MaxQuant software suite for all the GFP-enrichment experiments are listed.
- https://doi.org/10.7554/eLife.12068.037





