The autophagy gene Atg16l1 differentially regulates Treg and TH2 cells to control intestinal inflammation
Figures
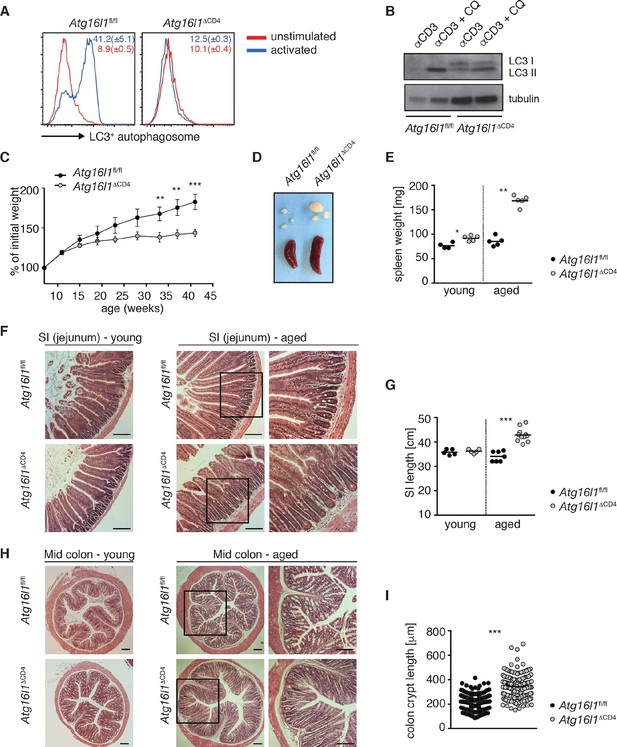
Aged Atg16l1ΔCD4 mice develop intestinal inflammation.
(A) FACS analysis of LC3+ autophagosome formation in CD4+ T cells from cLP of Atg16l1ΔCD4 and Atg16l1fl/fl mice after overnight activation with or without α-CD3 (5 μg/ml) and α-CD28 (1 μg/ml). (B) Western blot analysis of LC3 lipidation in naïve splenic CD4+ T cells isolated from Atg16l1ΔCD4 mice and Atg16l1fl/fl mice after 3hr activation with α-CD3 (5 μg/ml) and α-CD28 (1 μg/ml) with or without chloroquine (CQ, inhibitor of lysosomal degradation, 50 μM). (C) Weight curves of Atg16l1ΔCD4 and Atg16l1fl/fl littermates. (D) Representative images of spleens and mesenteric lymph nodes (mLN) from aged Atg16l1ΔCD4 and Atg16l1fl/fl littermates and (E) spleen weights of young and aged Atg16l1ΔCD4 and Atg16l1fl/fl littermates. (F,H) Representative photomicrographs of haemotoxilin and eosin (H&E) stained sections of (F) jejunum and (H) mid-colon from young and aged Atg16l1ΔCD4 and Atg16l1fl/fl littermates, scale bar 150 μm. (G,I) Quantification of (G) SI lengths and (I) mid-colon crypt lengths in aged Atg16l1ΔCD4 and Atg16l1fl/fl littermates. Data are representative of at least three independent experiments (A-E, F, H) or combined from two (G) or three (I) independent experiments, with at least 3 mice per group. Data shown as mean ± s.e.m (A,C). Each dot represents an individual mouse and horizontal bars denote means (E,G). In (I) each dot represents an individual crypt measurement and horizontal bars denote means. Statistical significance was determined using two-way analysis of variance (ANOVA) with Bonferroni’s correction for multiple comparisons (C) or the Mann–Whitney test (E,G,I), **p<0.01; ***p<0.001. SI LP– small intestine lamina propria, cLP – colonic lamina propria. Young mice: 8–12 weeks old, aged mice >5 months old.
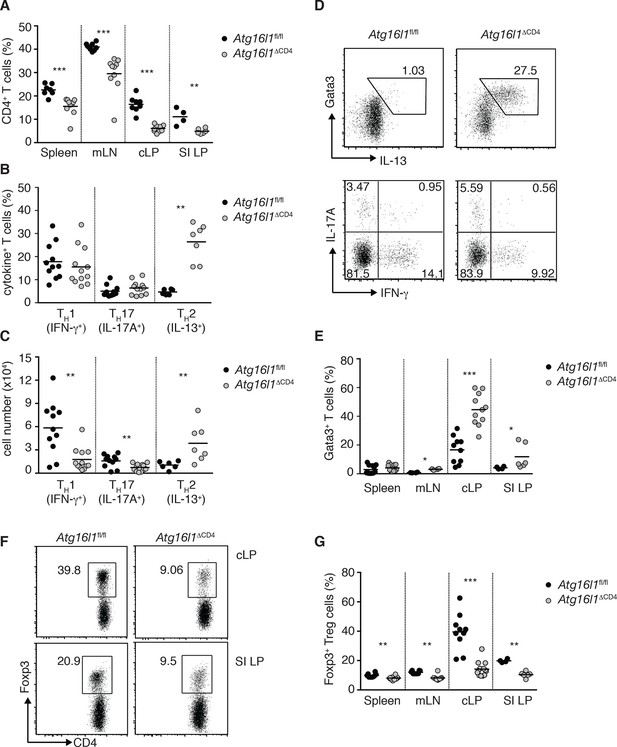
Atg16l1ΔCD4 mice exhibit reciprocal dysregulation of intestinal TH2 and Treg cells before the onset of intestinal inflammation.
(A) Frequencies of CD4+ T cells as a proportion of live cells in young Atg16l1ΔCD4 and Atg16l1fl/fl littermates. (B) Frequencies and (C) total numbers of IFN-γ+ TH1, IL-17A+ TH17 and IL-13+ TH2 cells isolated from cLP of young Atg16l1ΔCD4 and Atg16l1fl/fl littermates (gated on CD4+ T cells). (D) Representative FACS plots of Gata3 and IL-13 (top) or IFN-γ and IL-17A (bottom) expression by cLP CD4+ T cells isolated from young Atg16l1ΔCD4 and Atg16l1fl/fl littermates (gated on CD4+ TCRβ+ Foxp3- live cells). (E) Frequencies of Gata3+ CD4+ T cells in young Atg16l1ΔCD4 and Atg16l1fl/fl littermates (gated on CD4+ TCRβ+ Foxp3- cells). (F) Representative FACS plots and (G) frequencies of Foxp3+ Treg cells in young Atg16l1ΔCD4 and Atg16l1fl/fl littermates (gated on CD4+ TCRβ+ cells). Data are combined from three or more independent experiments with at least two mice per group (A,B, D, E, G) or are representative of four independent experiments with at least four mice per group (D, F). Each dot represents an individual mouse and horizontal bars denote means. Numbers indicate percentage of cells in gates or quadrants. Statistical significance was determined using the Mann–Whitney test, *p<0.05; **p<0.01; ***p<0.001. SI LP– small intestine lamina propria, cLP – colonic lamina propria. Young mice: 8–12 weeks old.
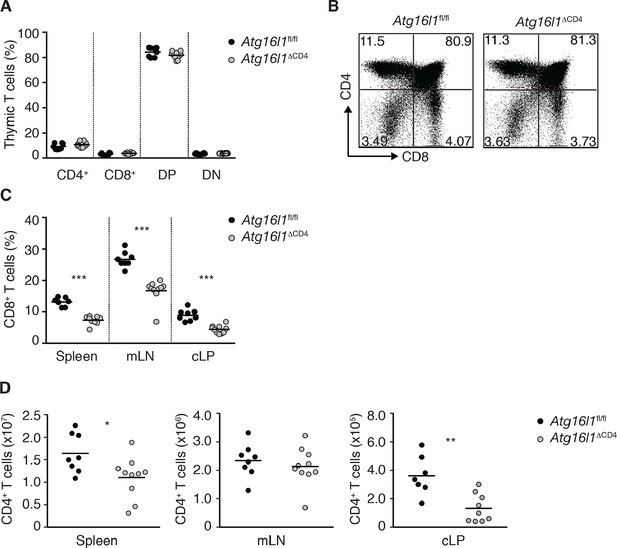
Characterization of immune cell compartments in young Atg16l1ΔCD4mice.
(A) Frequencies and (B) representative FACS plots of single positive CD4+, single positive CD8+, double positive (DP) CD4+ CD8+ and double negative (DN) CD4- CD8- thymocytes in young Atg16l1ΔCD4 and Atg16l1fl/fl littermates. (C) Frequencies of CD8+ T cells in young Atg16l1ΔCD4 and Atg16l1fl/fl littermates. (D) Total numbers of CD4+ T cells in spleen, mLN, and cLP of young Atg16l1ΔCD4 and Atg16l1fl/fl littermates. Data are combined from (A,C,D) or representative of (B) two or three independent experiments with at least 4 mice per group. Each dot represents an individual mouse and horizontal bars denote means. Numbers indicate percentage of cells in quadrants. Statistical significance was determined using the Mann–Whitney test, *p<0.05; **p<0.01; ***p<0.001. cLP – colonic lamina propria, mLN - mesenteric lymph nodes. Young mice: 8–12 weeks old.
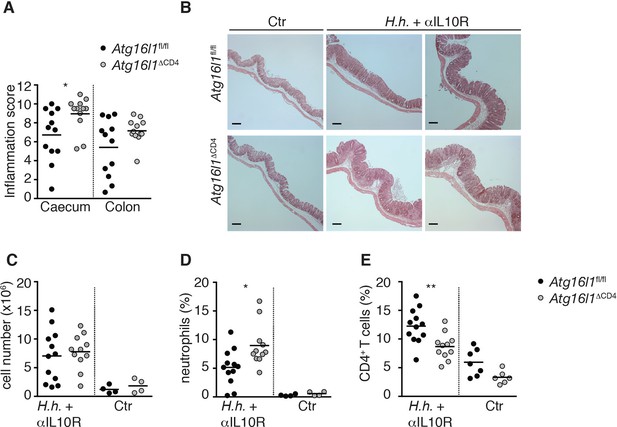
Atg16l1ΔCD4 mice have increased susceptibility to T-cell-mediated experimental IBD.
Cohorts of young Atg16l1ΔCD4 and Atg16l1fl/fl littermates were infected with Helicobacter hepaticus by oral gavage (three feeds of 1x108 CFU) and treated with anti-IL-10R mAb (1mg/mouse i.p. given weekly; H.h + αIL10R) or left untreated (Ctr). Two weeks post-infection mice were sacrificed for analyses. (A) Caecum and colon histopathology scores in H.h + αIL10R-treated Atg16l1ΔCD4 and Atg16l1fl/fl littermates. (B) Representative photomicrographs of H&E stained caecum of untreated Ctr (left panels) or H.h + αIL10R-treated (middle and right panels) Atg16l1ΔCD4 and Atg16l1fl/fl littermates, scale bar 150 μm. (C) Total lamina propria leukocyte numbers and (D,E) frequencies of (D) neutrophils (Gr1hi CD11b+) and (E) CD4+ T cells in cLP isolated from untreated Ctr or H.h + αIL10R-treated Atg16l1ΔCD4 and Atg16l1fl/fl littermates. Data are combined from two or three independent experiments with at least two mice per group. Each dot represents an individual mouse and horizontal bars denote means. Statistical significance was determined using the Mann–Whitney test, *p<0.05; **p<0.01.
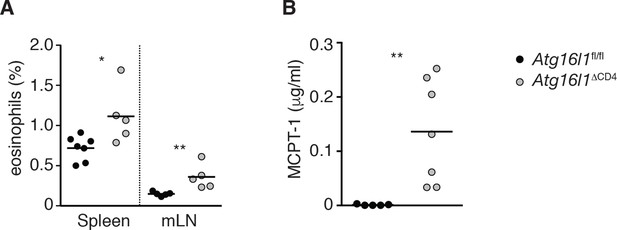
Elevated type 2 innate responses in Atg16l1ΔCD4 mice.
(A) Frequencies of eosinophils (Ly6Clow Ly6Glow CD11b+ F4/80-) in spleen and mLN of young Atg16l1ΔCD4 and Atg16l1fl/fl littermates. (B) Serum MCPT-1 levels in young Atg16l1ΔCD4 and Atg16l1fl/fl littermates were measured by ELISA. Data are combined from (A) or representative of (B) two or three independent experiments with at least four mice per group. Each dot represents an individual mouse and horizontal bars denote means. Statistical significance was determined using the Mann–Whitney test, *p<0.05; **p<0.01. mLN - mesenteric lymph nodes. Young mice: 8–12 weeks old.
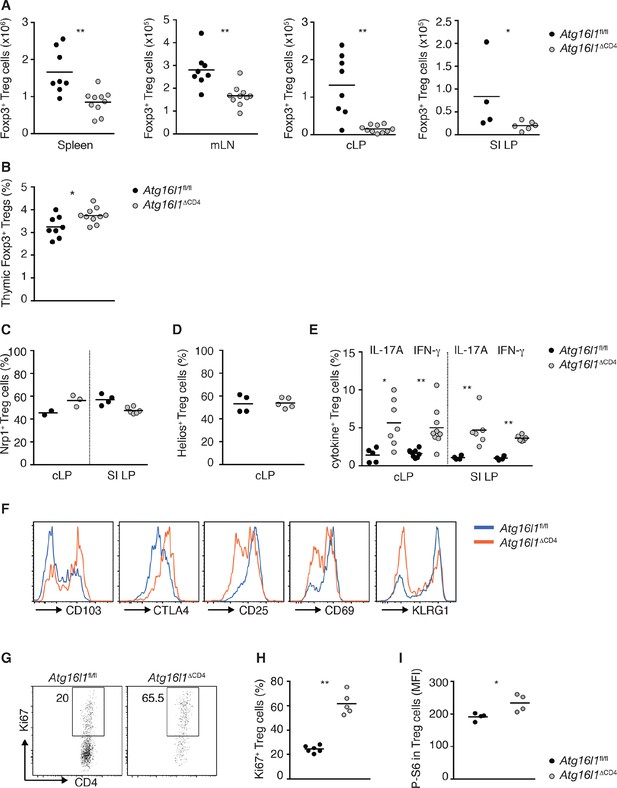
Characterization of Atg16l1-deficient Treg cells.
(A) Foxp3+ Treg cell numbers in spleen, mLN, cLP and SI LP of young Atg16l1ΔCD4 and Atg16l1fl/fl littermates. (B) Frequencies of Foxp3+ Treg in the thymus of young Atg16l1ΔCD4 and Atg16l1fl/fl littermates (gated on single positive CD4+ TCRβ+ cells). (C) Frequencies of Neuropilin-1+ (Nrp1+) Foxp3+ Treg cells in the cLP and SI LP of young Atg16l1ΔCD4 and Atg16l1fl/fl littermates (gated on CD4+ TCRβ+ Foxp3+ T cells). (D) Frequencies of Helios+ Foxp3+ Treg cells in the cLP of young Atg16l1ΔCD4 and Atg16l1fl/fl littermates (gated on CD4+ TCRβ+ Foxp3+ T cells). (E) Frequencies of IL-17A+ or IFN-γ+ of Foxp3+ Treg cells in the cLP and SI LP of young Atg16l1ΔCD4 and Atg16l1fl/fl littermates (gated on Foxp3+ CD4+ TCRβ+ T cells). (F) Expression of CD103, CTLA4, CD25, CD69 and KLRG1 by cLP Foxp3+ Treg cells from young Atg16l1ΔCD4 and Atg16l1fl/fl littermates (gated on Foxp3+ CD4+ TCRβ+ T cells). (G) Representative FACS plots and (H) frequencies of Ki67+ Foxp3+ Treg cells in cLP of young Atg16l1ΔCD4 and Atg16l1fl/fl littermates (gated on Foxp3+ CD4+ TCRβ+ T cells). (I) Mean fluorescence intensity (MFI) of phospo-S6 (P-S6) in Foxp3+ Treg cells in cLP of young Atg16l1ΔCD4 and Atg16l1fl/fl littermates. Data are combined from two independent experiments with at least four mice per group (A,B,E), are representative from two independent experiments with at least four mice per group (C,F-H), or are from one experiment (D,I). Each dot represents an individual mouse and horizontal bars denote means. Numbers indicate percentage of cells in gates. Statistical significance was determined using the Mann–Whitney test, *p<0.05; **p<0.01. mLN - mesenteric lymph nodes, SI LP– small intestine lamina propria, cLP – colonic lamina propria. Young mice: 8–12 weeks old.
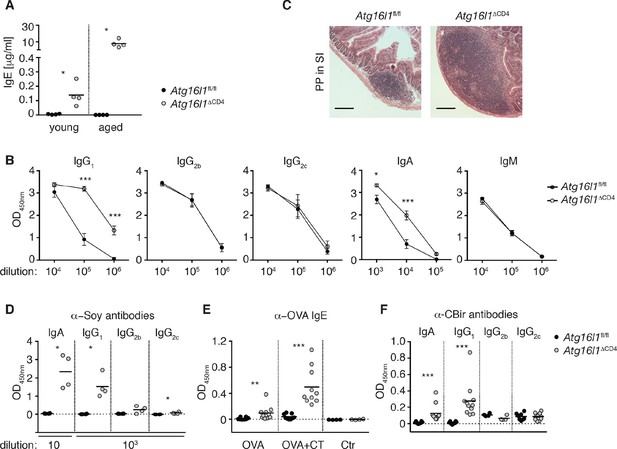
Atg16l1ΔCD4 mice develop elevated TH2-associated antibodies against intestinal luminal antigens.
(A) Serum IgE concentrations in cohorts of young and aged Atg16l1ΔCD4 and Atg16l1fl/fl littermates were measured by ELISA. (B) Serum antibody IgG1, IgG2b, IgG2c, IgA and IgM isotype levels in aged Atg16l1ΔCD4 and Atg16l1fl/fl littermates were measured by ELISA. (C) Representative photomicrographs of H&E stained sections of Peyer’s patch (PP) in the SI (jejunum) of aged Atg16l1ΔCD4 and Atg16l1fl/fl littermates, scale bar 150 μm. (D) Serum levels of Soy-specific IgA, IgG1, IgG2b, IgG2c antibodies in aged Atg16l1ΔCD4 and Atg16l1fl/fl littermates were measured by ELISA. (E) Young Atg16l1ΔCD4 and Atg16l1fl/fl littermates were fed with ovalbumin (OVA) alone or with cholera toxin (CT) as described in methods and levels of OVA-specific serum IgE were measured 8 weeks after first challenge by ELISA. (F) Levels of CBir1-specific IgA, IgG1, IgG2b and IgG2c antibodies in serum of aged Atg16l1ΔCD4 and Atg16l1fl/fl littermates were measured by ELISA, serum was diluted 50x. Data are representative from at least two independent experiments with at least three mice per group (A-D) or combined from two (E) or three (F) independent experiments with at least three mice per group. Each dot represents an individual mouse and horizontal bars denote means (A,D,E,F). Serum isotype levels are shown as mean ± s.e.m (B). Statistical significance was determined using the Mann–Whitney test (A,D-F) or two-way analysis of variance (ANOVA) with Bonferroni’s correction for multiple comparisons (B), *p<0.05; **p<0.01; ***p<0.001. SI – small intestine. Young mice: 8–12 weeks old, aged mice > 5 months old.
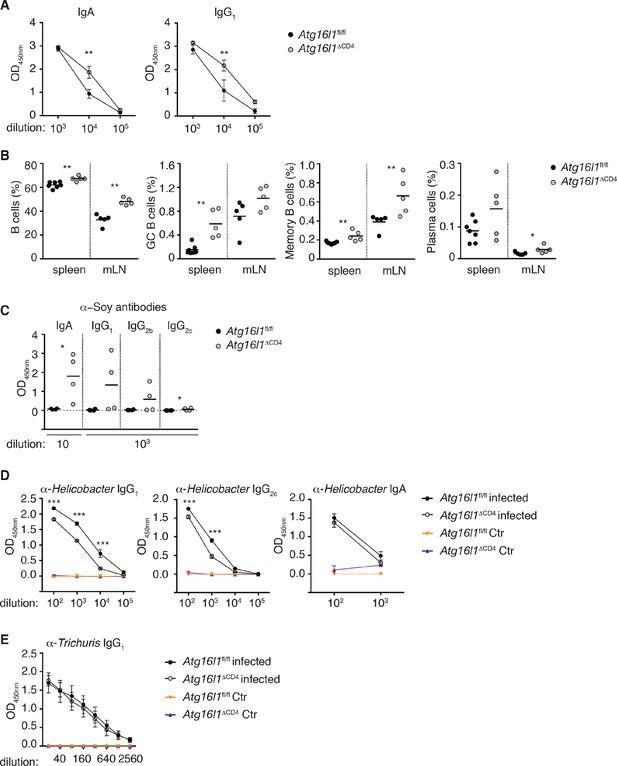
Dysregulated humoral responses in young Atg16l1ΔCD4 mice.
(A) Serum antibody IgA and IgG1 isotype levels in young Atg16l1ΔCD4 and Atg16l1fl/fl littermates were measured by ELISA. (B) Frequencies of B cells (B220+), germinal center B cells (GC: B220+ GL7+ CD95+), memory B cells (B220+ Gl7- IgM- IgG+) and plasma cells (CD138+) in the spleen and mLN of young Atg16l1ΔCD4 and Atg16l1fl/fl littermates (gated on live CD45+ cells). (C) Serum levels of Soy-specific IgA, IgG1, IgG2b, IgG2c antibodies in young Atg16l1ΔCD4 and Atg16l1fl/fl littermates were measured by ELISA. (D) Cohorts of young Atg16l1ΔCD4 and Atg16l1fl/fl littermates were infected with Helicobacter hepaticus by oral gavage (three feeds of 1x108 CFU) and levels of serum Helicobacter-specific IgG1, IgG2c and IgA antibodies were determined three weeks later by ELISA. (E) Cohorts of young Atg16l1ΔCD4 and Atg16l1fl/fl littermates were orally infected with Trichuris muris (200 eggs) and levels of T. muris-specific IgG1 in serum were determined 34 days later by ELISA. Data are from one experiment with at least three mice per group (A-D) or representative from two independent experiments with five mice per group (E). Each dot represents an individual mouse and horizontal bars denote means (B,C) or data represent mean ± s.e.m (A,D,E). Statistical significance was determined using the Mann–Whitney test (A,C) or two-way analysis of variance (ANOVA) with Bonferroni’s correction for multiple comparisons (B,D,E): statistical significance was determined between infected groups (b,c), *p<0.05; **p<0.01. mLN - mesenteric lymph nodes. Young mice: 8–12 weeks old.
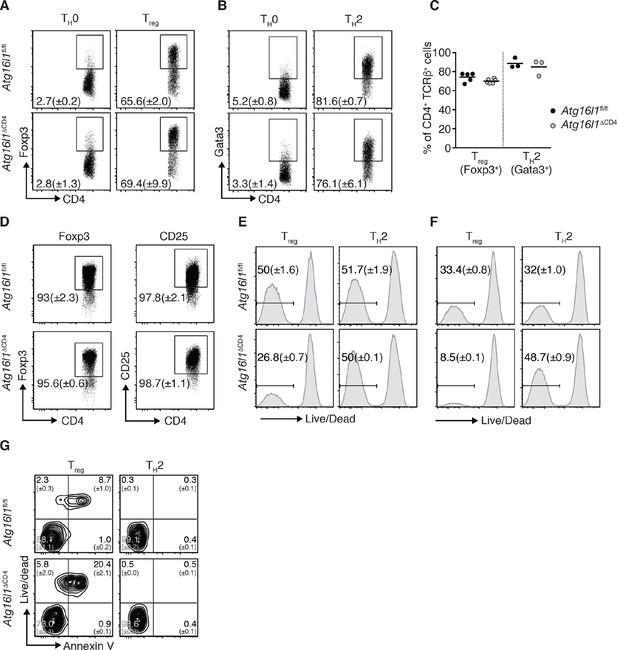
Atg16l1 promotes survival of Treg cells and limits TH2 cell survival.
(A,B) Atg16l1ΔCD4 or Atg16l1fl/fl naïve CD4+ T cells were cultured in TH0, Treg, or TH2 polarizing conditions for 48 hr and analyzed by FACS. Representative FACS plots show (A) Foxp3 and (B) Gata3 expression (gated on CD4+ TCRβ+ T cells). (C) Frequencies of Treg cells (Foxp3+) and TH2 cells (Gata3+) arising from Atg16l1ΔCD4 or Atg16l1fl/fl naïve CD4+ T cells cultured in Treg or TH2 polarizing conditions for 5 days. (D) Atg16l1ΔCD4 or Atg16l1fl/fl Treg cells were cultured with anti-CD3 (3 μg/ml) and anti-CD28 (1 μg/ml) for 48 hr, then maintained in the presence of IL-4 and IL-13 for a further 5 days before FACS analysis of Foxp3 and CD25 expression of live CD4+ T cells. (E,F) Naïve Atg16l1ΔCD4 or Atg16l1fl/fl CD4+ T cells were cultured with (E) 1 μg/ml or (F) 5 μg/ml anti-CD3 plus anti-CD28 (1 μg/ml) for 48 hr in Treg or TH2 polarizing conditions, then maintained in polarizing conditions for a further 5 days before FACS analysis of cell survival. Histograms show gates and frequencies of live CD4+ T cells. (G) Representative FACS plots of viability dye and Annexin V staining of Treg cells and TH2 cells from the cLP of young Atg16l1ΔCD4 and Atg16l1fl/fl littermates, gated on CD4+ TCRβ+ Foxp3+ (left panel), or CD4+ TCRβ+ Gata3+ (right panel). Data are representative from two (D,G) or three independent experiments (A,B,E,F), or are combined from three independent experiments (C). Each dot represents an individual cell culture (C) or data are shown as mean ± s.e.m (A,B,D-F). Numbers indicate percentage of cells in quadrants (G). cLP – colonic lamina propria. Young mice: 8–12 weeks old.
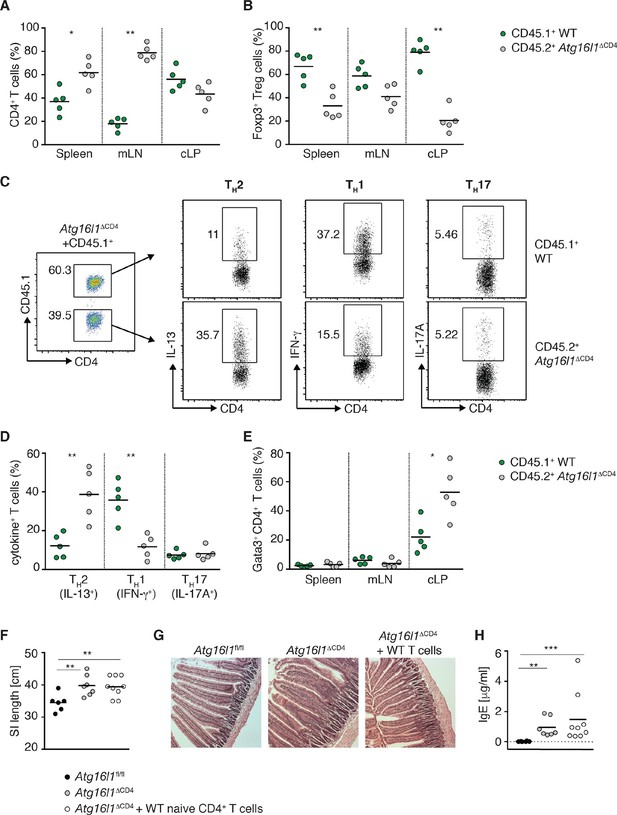
Autophagy contributes to the elevated TH2 responses in Atg16l1ΔCD4 mice in a cell-intrinsic manner.
Young Atg16l1ΔCD4 mice (CD45.2+) were adoptively transferred with 4-5x106 naïve WT CD4+ T cells (CD45.1+) and analyzed 3 months later. (A) Frequencies of WT (CD45.1+) and Atg16l1-deficient (CD45.2+) CD4+ T cells in the spleen, mLN and cLP. (B) Frequencies of WT (CD45.1+) and Atg16l1-deficient (CD45.2+) Foxp3+ Treg cells in the spleen, mLN and cLP (gated on CD4+ TCRβ+ T cells). (C) Representative FACS plots showing gating of WT (CD45.1+) and Atg16l1-deficient (CD45.1-) CD4+ T cells and expression of IL-13 (TH2), IFN-γ (TH1) and IL-17A (TH17) in the cLP (gated on CD4+ TCRβ+ Foxp3- T cells). (D) Frequencies of WT (CD45.1+) and Atg16l1-deficient (CD45.2+) TH2 (IL-13+), TH1 (IFN- γ+) and TH17 (IL-17A+) cells among CD4+ TCRβ+ Foxp3- T cells in the cLP. (E) Frequencies of WT (CD45.1+) and Atg16l1-deficient (CD45.2+) Gata3+ CD4+ T cells in the spleen, mLN and cLP (gated on CD4+ TCRβ+ Foxp3- T cells). (F) SI lengths and (G) representative photomicrographs of jejunum of control untreated Atg16l1fl/fl or Atg16l1ΔCD4 littermates and reconstituted Atg16l1ΔCD4 mice, scale bar 150 μm. (H) Serum IgE concentrations in control untreated Atg16l1fl/fl or Atg16l1ΔCD4 littermates and adoptively transferred Atg16l1ΔCD4 mice were measured by ELISA. Data are representative of two independent experiments with at least four mice per group (A-E,G) or combined from two independent experiments (F,H). Each dot represents cells coming from the donor or the hosts within the individual transferred mouse (A,B,D,E) or each dot represents an individual mouse (F,H), horizontal bars denote mean. Numbers indicate percentage of cells in gates. Statistical significance was determined using the Mann–Whitney test, *p<0.05; **p<0.01. mLN - mesenteric lymph nodes, cLP – colonic lamina propria. Young mice: 10–12 weeks old.
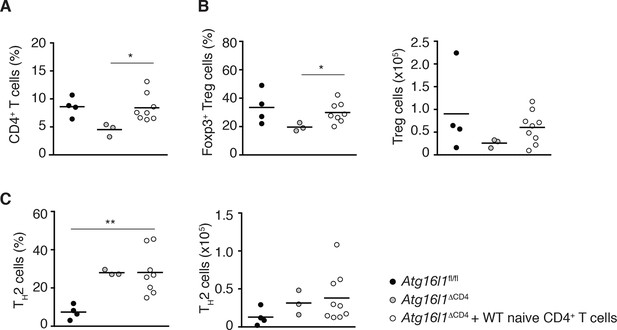
Reconstitution of intestinal CD4+ T cell compartments in adoptively transferred Atg16l1ΔCD4 mice.
Young Atg16l1ΔCD4 mice (CD45.2+) were adoptively transferred with 4-5x106 naïve WT CD4+ T cells (CD45.1+) i.v. and cLP populations analyzed 3 months later. (A) Frequencies of total CD4+ T cells. (B) Frequencies and total numbers of Foxp3+ Treg cells (gated on CD4+ TCRβ+ T cells). (C) Frequencies and total numbers of TH2 (IL-13+) cells (gated on CD4+ TCRβ+ Foxp3- T cells). Data are combined from two independent experiments with at least four mice per group. Each dot represents an individual mouse, horizontal bars denote mean. Statistical significance was determined using the Mann–Whitney test, *p<0.05; **p<0.01. cLP – colonic lamina propria. Young mice: 10–12 weeks old.
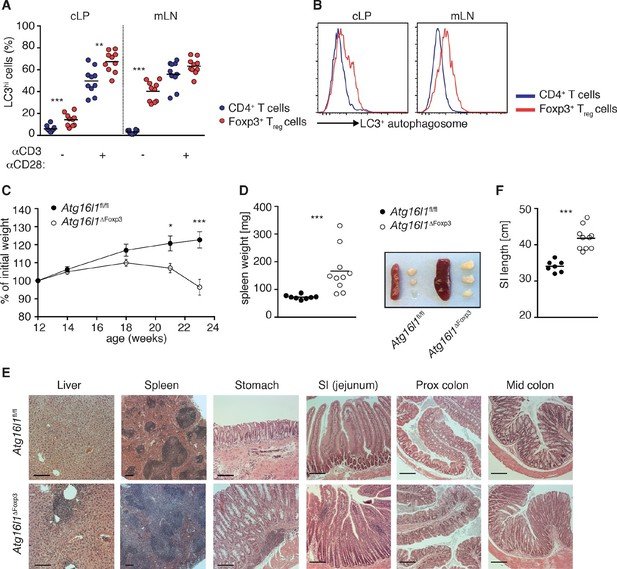
Aged Atg16l1ΔFoxp3 mice develop spontaneous multi-organ inflammation.
(A) LC3+ autophagosome formation by Foxp3- CD4+ T cells and Foxp3+ Treg cells from cLP and mLN of WT mice in unstimulated cells or after overnight activation with α-CD3 (5 μg/ml) and α-CD28 (1 μg/ml). (B) Representative LC3 staining of unstimulated cells (gated on Foxp3+ CD4+ TCRβ+ Treg cells or Foxp3- CD4+ TCRβ+ T cells). (C) Weight curves and (D) spleen weights and representative images of spleen and mLN of aged Atg16l1ΔFoxp3 and Atg16l1fl/fl littermates. (E) Representative photomicrographs of H&E stained sections of liver, spleen, stomach, SI (jejunum), proximal colon and mid-colon of aged Atg16l1ΔFoxp3 and Atg16l1fl/fl littermates, scale bar 150 μm. (F) Quantification of SI length. Data are combined from two to four independent experiments with two to five mice per group (A,D,F) or are representative of two to three independent experiments with two to five mice per group (B,C,E). Each dot represents an individual mouse and horizontal bars denote means (A,D,F). Data shown as mean ± s.e.m (C). Statistical significance was determined using two-way analysis of variance (ANOVA) with Bonferroni’s correction for multiple comparisons (C) or using the Mann–Whitney test (A,D,F), *p<0.05; **p<0.01; ***p<0.001. mLN - mesenteric lymph nodes, SI – small intestine lamina propria, cLP – colonic lamina propria. Aged mice >5 months old.
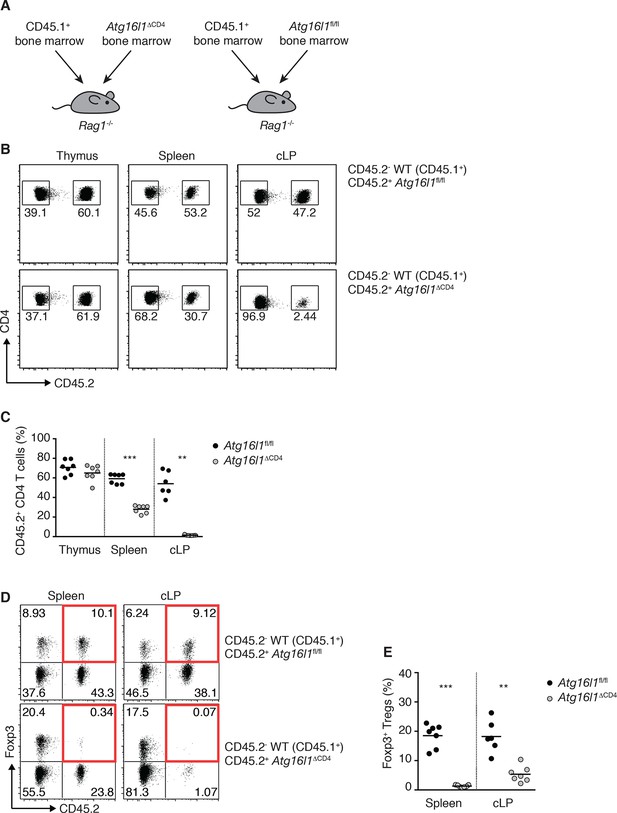
Impaired reconstitution of mixed bone marrow chimeras by Atg16l1-deficient T cells.
(A) Experimental design for the generation of mixed bone marrow (BM) chimeras. BM cells isolated from WT (CD45.1+) and Atg16l1fl/fl or Atg16l1ΔCD4 (CD45.2+) mice were injected at a 1:1 ratio into lethally irradiated (1100 Rad) Rag1-/- recipients (total of 1x107 cells per mouse). (B) Representative FACS plots showing frequencies of Atg16l1ΔCD4 or Atg16l1fl/fl (CD45.2+) and WT (CD45.2-) CD4+ T cells in the thymus, spleen and cLP of mixed BM chimeras (gated on CD4+ TCRβ+ T cells). (C) Frequencies of Atg16l1ΔCD4 or Atg16l1fl/fl (CD45.2+) CD4+ T cells in mixed BM chimeras (shown as percentage of total CD4+ TCRβ+ T cells). (D) Representative FACS plots and (E) frequencies of Foxp3+ Treg cells derived from Atg16l1ΔCD4 or Atg16l1fl/fl (CD45.2+) cells in mixed BM chimeras (gated on CD4+ TCRβ+ T cells). Highlighted top right quadrants indicate Treg cells derived from Atg16l1fl/fl orAtg16l1ΔCD4 BM. Data are representative from two independent experiments with at least seven mice per group. Each dot represents an individual mouse and horizontal bars denote means. Numbers indicate percentage of cells in gates. Statistical significance was determined using the Mann–Whitney test, **p<0.01; ***p<0.001.
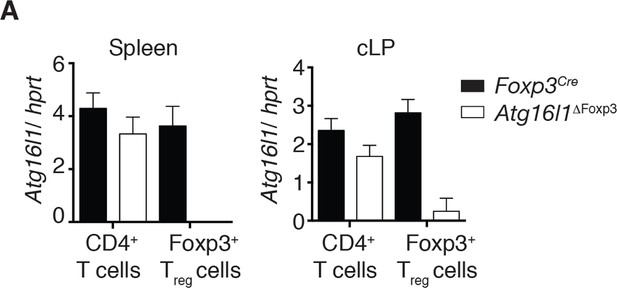
Analysis of Atg16l1 expression in Atg16l1ΔFoxp3 mice.
(A) qPCR analysis of Atg16l1 exon 3 levels in sorted CD4+ Foxp3- T cells and Foxp3+ Treg cells from spleen and cLP of Atg16l1ΔFoxp3 and Foxp3Cre mice. Data are representative from two independent experiments with 5 mice per group. Atg16l1 exon 3 levels are shown as mean ± s.e.m of three technical replicates, normalised to expression of hprt. cLP – colonic lamina propria.
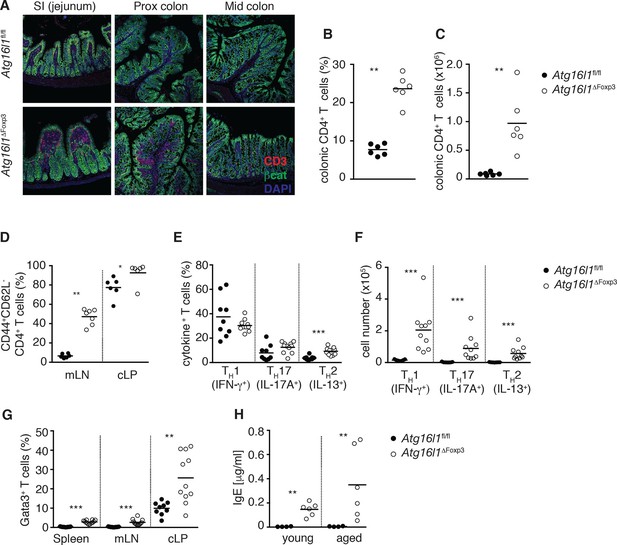
Atg16l1ΔFoxp3 mice cannot control pro-inflammatory TH effector responses.
(A) Representative immunofluorescence images of small intestine and proximal and mid colon of aged Atg16l1ΔFoxp3 and Atg16l1fl/fl littermates stained for CD3 (red), β-catenin (green) and DAPI (blue). (B) Frequencies and (C) total numbers of cLP CD4+ TCRβ+ T cells in aged Atg16l1ΔFoxp3 and Atg16l1fl/fl littermates. (D) Frequencies of effector (CD44+CD62L-) CD4+ T cells in the mLN and cLP of aged Atg16l1ΔFoxp3 and Atg16l1fl/fl littermates (gated on CD4+ TCRβ+ Foxp3- T cells). (E) Frequencies and (F) total numbers of TH1 (IFN-γ+), TH17 (IL-17A+), TH2 (IL-13+) T cells in the cLP of aged Atg16l1ΔFoxp3 and Atg16l1fl/fl littermates (gated on CD4+ TCRβ+ Foxp3- T cells). (G) Frequencies of Gata3+ CD4+ T cells in aged Atg16l1ΔFoxp3 and Atg16l1fl/fl littermates (gated on CD4+ TCRβ+ Foxp3- T cells). (H) Serum IgE concentrations in Atg16l1ΔFoxp3 and Atg16l1fl/fl littermates were measured by ELISA. Data are combined from two to four independent experiments with two to five mice per group (B-H) or are representative of two independent experiments with two to five mice per group (A). Each dot represents an individual mouse and horizontal bars denote means. Statistical significance was determined using the Mann–Whitney test *p<0.05; **p<0.01; ***p<0.001. mLN - mesenteric lymph nodes, cLP – colonic lamina propria. Young mice: 8–12 weeks old, aged mice >5 months old.
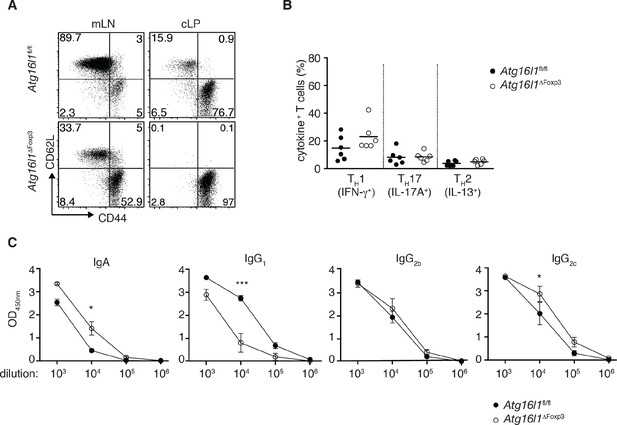
Additional characterization of Atg16l1ΔFoxp3 mice.
(A) Representative FACS plots of CD44 and CD62L expression by CD4+ T cells from cLP of aged Atg16l1ΔFoxp3 and Atg16l1fl/fl littermates (gated on CD4+ TCRβ+ Foxp3- T cells). (B) Frequencies of TH1 (IFN-γ+), TH17 (IL-17A+), TH2 (IL-13+) T cells in the cLP of young Atg16l1ΔFoxp3 and Atg16l1fl/fl littermates (gated on CD4+ TCRβ+ Foxp3- T cells). (C) Serum antibody IgA, IgG1, IgG2b, IgG2c isotype levels in aged Atg16l1ΔFoxp3 and Atg16l1fl/fl littermates. Data are combined from three independent experiments with two to five mice per group (B) or are representative from two to three independent experiments with two to five mice per group (A,C). Each dot represents an individual mouse and horizontal bars denote means. Numbers indicate percentage of cells in gates. Data shown as mean ± s.e.m (C). Statistical significance was determined using the Mann–Whitney test, *p<0.05; **p<0.01; ***p<0.001. mLN - mesenteric lymph nodes, cLP – colonic lamina propria. Young mice: 8–12 weeks old, aged mice >5 months old.
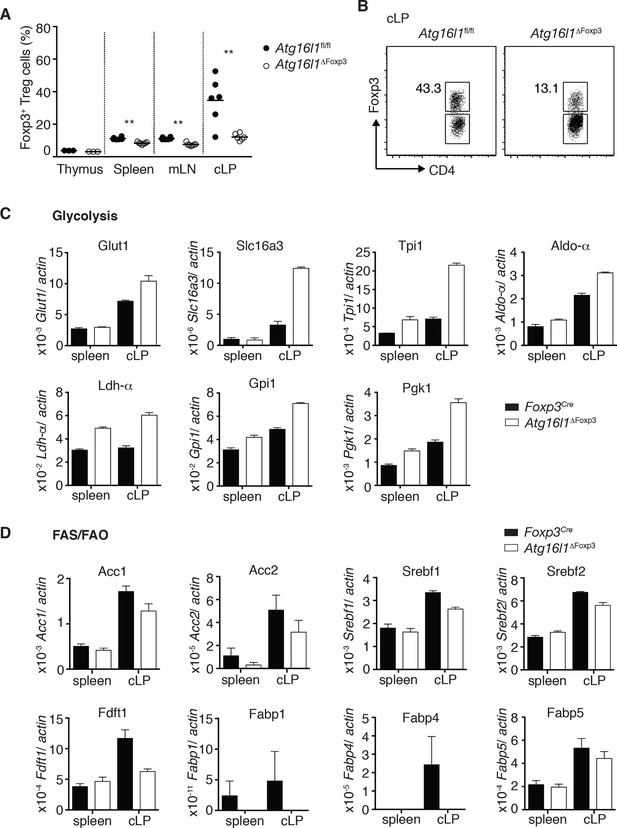
Cell-intrinsic autophagy is required for metabolic adaptation and survival of intestinal Foxp3+ Treg cells.
(A) Foxp3+ Treg cell frequencies among CD4+ TCRβ+ T cells in Atg16l1ΔFoxp3 and Atg16l1fl/fl littermates and (B) representative FACS plots of Foxp3 expression in cLP CD4+ T cells from young Atg16l1ΔFoxp3 and Atg16l1fl/fl littermates (gated on CD4+ TCRβ+ T cells). (C) qPCR analysis of glycolytic gene levels in sorted Foxp3+ Treg cells from spleen and cLP of young Atg16l1ΔFoxp3 and Foxp3Cre mice (sorted for CD4+ TCRβ+ YFP+). (D) qPCR analysis of FAS and FAO gene levels in Foxp3+ Treg cells from the spleen and cLP of young Atg16l1ΔFoxp3 and Foxp3Cre mice (sorted for CD4+ TCRβ+ YFP+). FAS: fatty acid synthesis, FAO: fatty acid oxidation, Glut1: glucose transporter 1, Slc16ac: solute carrier family 16 member 3 (lactic acid and pyruvate transporter), Tpi1: triosephosphate isomerase 1, Aldo–α: aldolase α, Ldh-α: lactate dehydrogenase α, Gpi1: Glucose phosphate isomerase 1, Pgk1: Phosphoglycerate kinase 1, Acc1: acetyl-CoA carboxylase 1, Acc2: acetyl-CoA carboxylase 2, Srebf1: sterol regulatory element binding transcription factor 1, Srebf2: sterol regulatory element binding transcription factor 2, Fdft1: farnesyl-diphosphate farnesyltransferase 1, Fabp: Fatty acid-binding protein. Data are representative from two (C,D) or three independent experiments (A,B). Each dot represents individual mouse (A) or data are shown as mean ± s.e.m (C,D). Gene expression levels are shown as mean ± s.e.m of three technical replicates (C,D). Numbers indicate percentage of cells in gates (B). cLP – colonic lamina propria. Young mice: 8–12 weeks old.
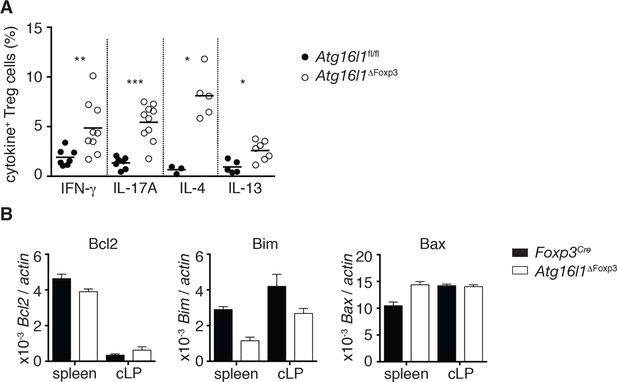
Atg16l1-deficient colonic Treg cells exhibit increased cytokine secretion.
(A) Frequencies of IFN-γ+, IL-17A+, IL-4+, IL-13+ Foxp3+ Treg cells in the cLP of aged Atg16l1ΔFoxp3 and Atg16l1fl/fl littermates (gated on Foxp3+ CD4+ TCRβ+ T cells). (B) qPCR analysis of Bcl2, Bim and Bax levels in Foxp3+ Treg cells from young cLP of Atg16l1ΔFoxp3 and Foxp3Cre mice (sorted for CD4+ TCRβ+ YFP+). Data are combined from three independent experiments with two to five mice per group (A) or are representative from two independent experiments (B). Each dot represents an individual mouse and horizontal bars denote means (A). Gene expression levels are shown as mean ± s.e.m of three technical replicates (B). cLP – colonic lamina propria. Young mice: 8–12 weeks old.
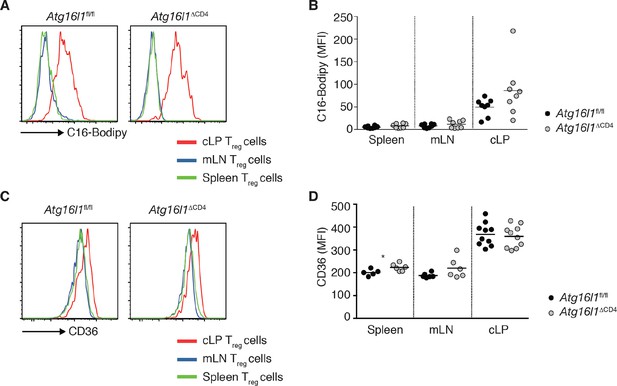
Increased lipid uptake by intestinal Treg cells.
(A,B) Atg16l1fl/fl and Atg16l1ΔCD4 littermates were injected i.p. with 50 μg of fluorescent 16-carbon fatty acid analog BODIPY C-16 and culled 1 hr later and tissue was collected for analysis by flow cytometry. (A) Representative FACS plots and (B) quantification of C16-Bodipy uptake by Foxp3+ Treg cells in the spleen, mLN and cLP (gated on Foxp3+ CD4+ TCRβ+ T cells). (C) Representative FACS plots and (D) quantification of CD36 expression by Foxp3+ Treg cells in the spleen, mLN and cLP of Atg16l1fl/fl and Atg16l1ΔCD4 littermates (gated on Foxp3+ CD4+ TCRβ+ T cells). Data are combined from (B,D) or are representative of (A,C) two independent experiments with 3–5 mice per group. Each dot represents an individual mouse and horizontal bars denote means. Statistical significance was determined using the Mann–Whitney test, *p<0.05. mLN - mesenteric lymph nodes, cLP – colonic lamina propria. Young mice: 8–12 weeks old.
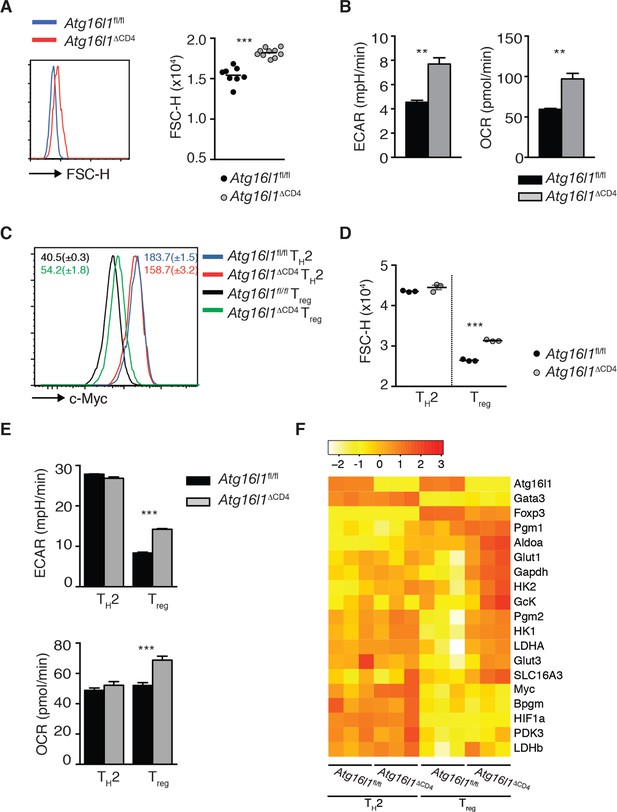
TH2 cells exhibit an enhanced glycolytic metabolic profile that is independent of autophagy.
(A) Representative FACS plot and quantification of the cell size (FSC-H) of naïve (CD44- CD62L+) CD4+ T cells from the spleen of Atg16l1ΔCD4 or Atg16l1fl/fl littermates. (B) Basal level of oxygen consumption rate (OCR) and extracellular acidification rate (ECAR) in naïve (CD44- CD62L+) unstimulated CD4+ T cells isolated from the spleen of Atg16l1ΔCD4 or Atg16l1fl/fl littermates measured using the Seahorse metabolic flux analyzer. (C,D) Expression of c-Myc (C) and cell size (D) was analyzed by FACS in Atg16l1ΔCD4 or Atg16l1fl/fl CD4+ T cells that were cultured in TH2 or Treg-polarizing conditions for 3 days and rested for one day in the presence of polarizing cytokines. (E) Basal level of OCR and ECAR were measured by Seahorse metabolic flux analyzer in Atg16l1ΔCD4 or Atg16l1fl/fl CD4+ T cells cultured in TH2 or Treg- polarizing conditions for 3 days and rested for 2 days in the presence of polarizing cytokines. (F) qPCR analysis of glycolytic gene levels in Atg16l1ΔCD4 or Atg16l1fl/fl CD4+ T cells cultured in TH2 or Treg polarizing conditions for 3 days and rested for 1 day in the presence of polarizing cytokines. Data are combined from two independent experiments (A), or are representative of two independent experiments (B-D), or are from one experiment (E,F). Each dot represents an individual mouse (A) or individual cell culture (D). ECAR and OCAR data represent mean ± s.e.m values of T cell populations that were assayed in triplicates or quadruplicates (B,E). Gene expression data of triplicate cultures represent normalized expression values for each gene that were scaled to a mean of 0 and a standard deviation of 1 (F). Statistical significance was determined using the Mann–Whitney test (A) or unpaired Student’s t –test (B,D,E), **p<0.01; **p<0.001.
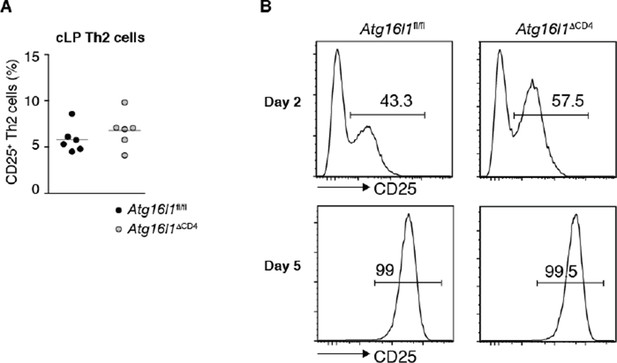
Autophagy deficiency does not influence expression of IL-2 receptor on TH2 cells (A) Expression of IL-2Ra (CD25) by TH2cells in the cLP of young Atg16l1ΔCD4 and Atg16l1fl/fl littermates (gated on CD4+ TCRβ+ Foxp3- Gata3+ T cells), data are representative of two independent experiments.
(B) Expression of CD25 was measured in naïve Atg16l1ΔCD4 or Atg16l1fl/fl CD4+ T cells cultured with anti-CD3 (5μg/ml) and anti-CD28 (1μg/ml) in TH2 polarizing conditions for 2 or 5 days, data are from one experiment.





