A large field of view two-photon mesoscope with subcellular resolution for in vivo imaging
Figures
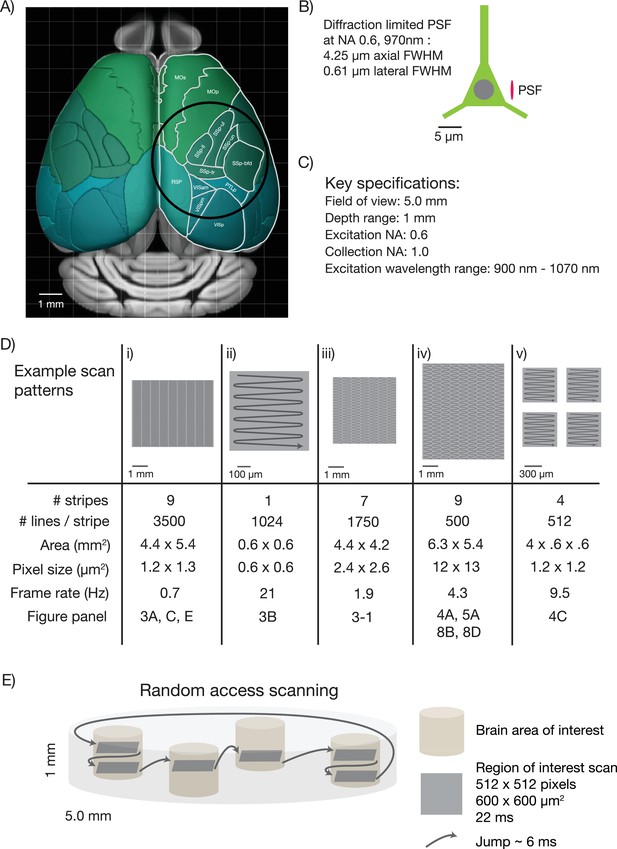
Specifications for the 2p-RAM.
(A) Schematic of the dorsal surface of the mouse cortex (from Allen Brain Atlas, Brain Explorer 2). The overlaid circle corresponds to the specified FOV (∅ 5 mm). MOp: primary somatomotor, MOs: secondary somatomotor, SSp-ul: primary somatosensory upper limb, SSp-ll: primary somatosensory lower limb, SSp-un: primary somatosensory unassigned, SSp-tr: primary somatosensory trunk, SSp-bfd: primary somatosensory barrel field, PTLp: posterior parietal association, RSP: retrosplenial, VISam: anteromedial visual, VISpm: posteromedial visual, VISp: primary visual. (B) Schematic of a neuron (approximate soma diameter, 10 μm), and a diffraction limited point spread function (PSF) corresponding to NA 0.6 and 970 nm. Subcellular resolution implies that the PSF must be contained within a single neuron. (C) Key specifications for the mesoscope. (D) Imaging parameters used in the experiments. In all cases a fast resonant scan (time per scan line, 42 µs; 24 kHz line rate) is made over patches 608 µm wide. The time per patch is 42 µs x number of lines. The entire FOV can be covered in nine stripes. A slow (0.7 Hz), high-resolution scan with 3500 lines per stripe was used to cover a large portion the FOV (i). A fast (4.3 Hz), low-resolution scan across the entire scan range has up to 500 lines per stripe (iv). More typical scans will involve sampling multiple smaller patches at high resolution and frame rates (v). See implementation for discussion of the scanning patterns. (E) Schematic showing a typical use-case. The entire imaging volume is a cylinder with a 5 mm diameter and 1 mm depth. The fast scan is made over patches in different parts of the volume.
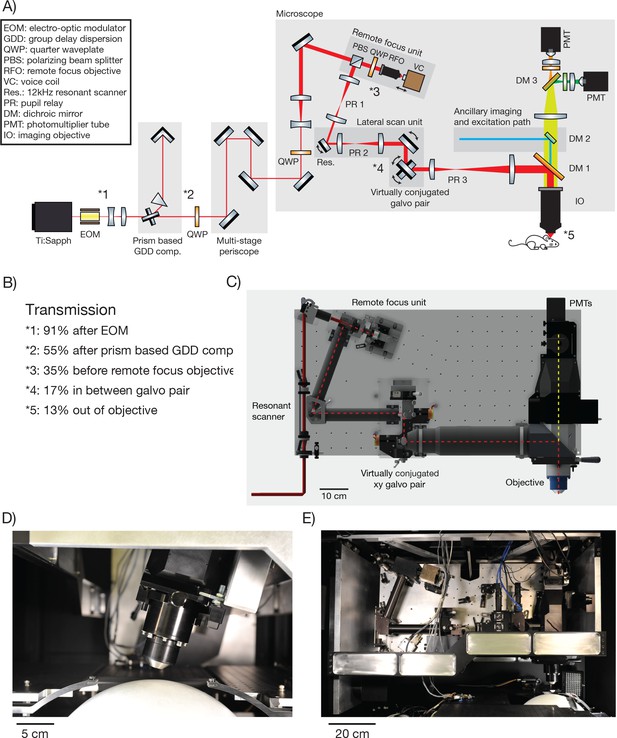
2p-RAM optics and optomechanics.
(A) Optical layout. Several routing mirrors were omitted for clarity. See text for detailed description. (B) Power transmission measurements at 970 nm. (C) CAD model of the microscope. The microscope was assembled on a vertically mounted breadboard measuring 1.0 m x 0.5 m. The objective is 63 mm wide, 120 mm long, and weighs 1.43 kg. (D) Photograph of microscope objective. (D, E, photocredit; Matt Staley). (E) Photograph of vertically mounted breadboard.
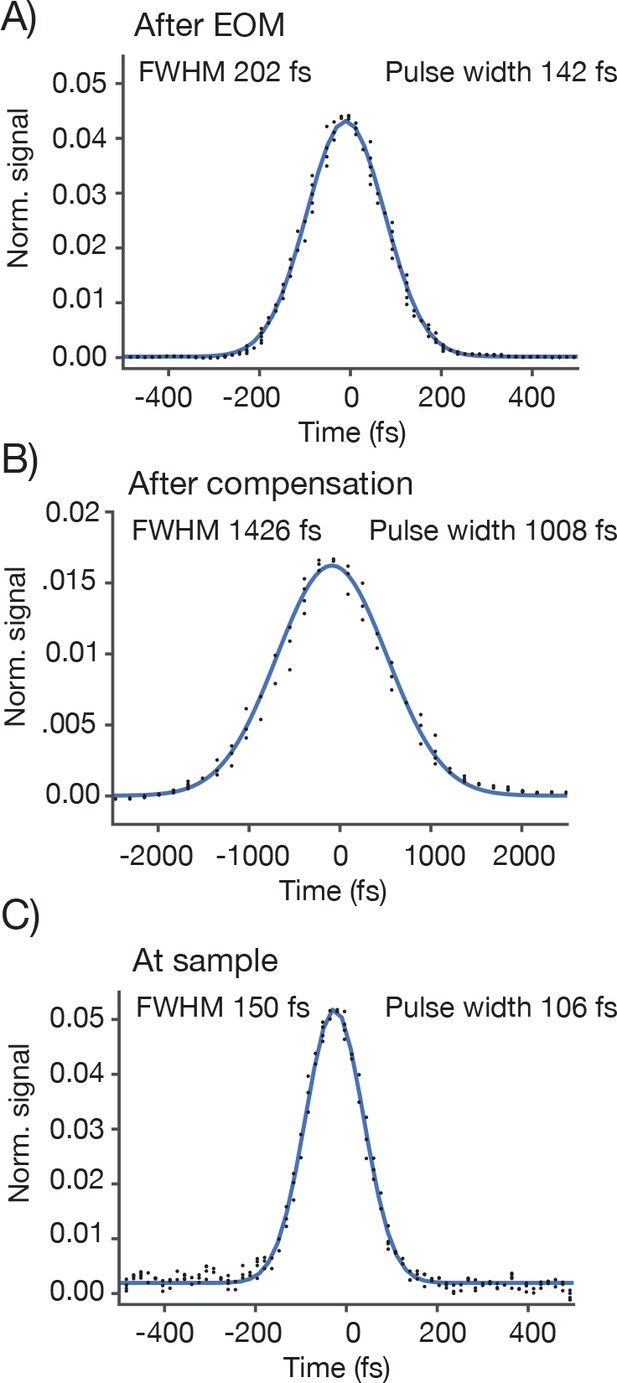
Pulse width measurements.
We measured the pulse width (full-width-at-half-max) using an autocorrelator. The autocorrelator was placed after the first telescope and EOM. For a Gaussian laser pulse, the width of the autocorrelation is 1.41 times the pulse width of the individual laser pulse. (A) Autocorrelation at the autocorrelator without GDD compensation. (B) Autocorrelation at the autocorrelator with GDD compensation. The pulse is now very broad due to the pre-chirp. (C) The autocorrelation at the specimen. As a sample we used a uniform fluorescent solution (Sigma F6377, 100 µM, 1 mm thick) and recorded fluorescence using the PMT.
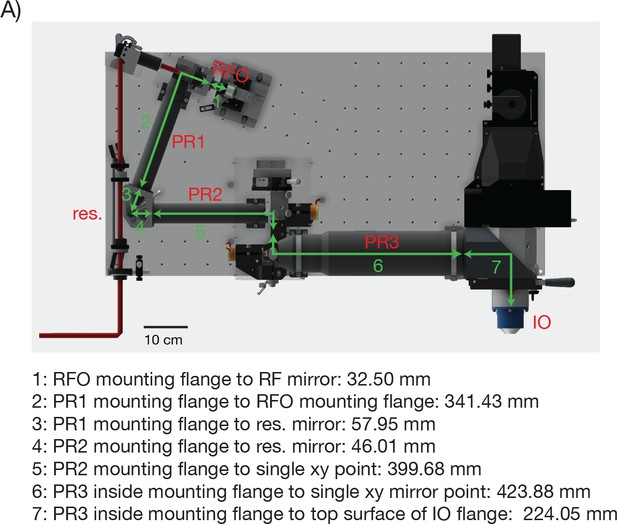
Distances between the critical 2p-RAM components.
(A) CAD model of the microscope with critical distances for assembling the 2p-RAM. PR: pupil relay, RFO: remote focus objective, IO: immersion objective, res.: resonant scan mirror.
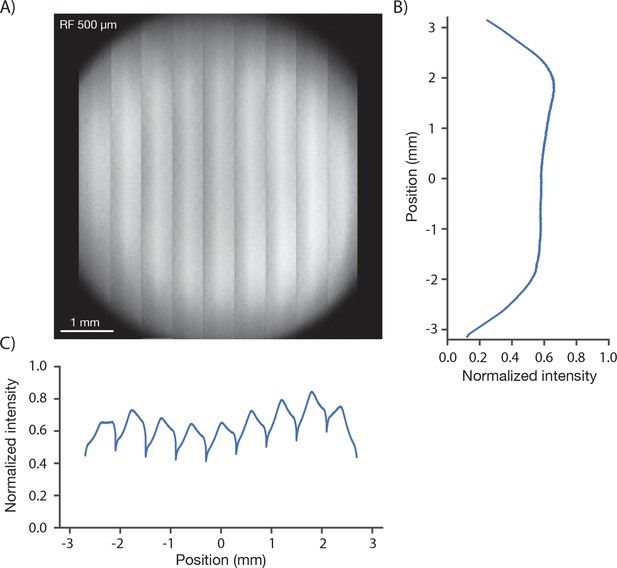
Image brightness.
(A) Image of a uniform fluorescent sample (fluorescein solution, Sigma F6377, 100 µM, 1 mm thick under 450 µm of coverglass, average of 30 frames). The image was acquired in 9 stripes, each approximately 600 µm wide (see Figure 1D, iv for details). The intensity variations in the image are caused by changes both in excitation and collection efficiency across the field. (B) Y profile (along slow scan axis) through slice at the central RF position normalized to the peak signal, which was located off the main two axes. (C) X profile (along fast scan axis) profile through central slice. The dips in signal at the edge of the stripes are caused by aberrations at large resonant scan angles.
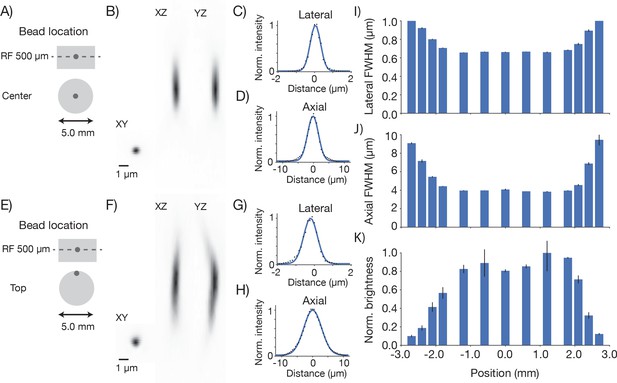
Point spread functions.
Point spread functions were measured at 13 different positions in the field at the middle depth of the imaging volume. The sample consisted of fluorescent microbeads (diameter, 0.5 µm, embedded in 1 mm thick 1.5% agar, under 450 µm of coverglass) imaged at 970 nm. The excitation intensity was 8.75 mW, below excitation saturation of the beads. At each position, a 61 µm x 61 µm x 35 µm volume (average of 30 acquisitions) was acquired with 8.4 x 8.4 x 2 pixels per µm. Bead that were separated by at least 2 µm x 2 µm x 12 µm from the edges of the volume and other beads were analyzed. 2–13 PSFs per volume were measured. (A) Example measurement at the center of the FOV. (B) Mean intensity projections through the XY, XZ, and YZ slices of the bead. (C) Gaussian fit through the lateral slice through the bead image. (D) Same as (C), axial slice. (E–H) Same as (A–D) for a bead at the edge of the FOV (2.4 mm from the center on the top). (I) Average lateral full-width-at-half-max (FWHM) of the central XY slice of a bead, averaged across the X and Y directions. (J) Average axial FWHM. (K) Average normalized brightness of the beads. Error bars, standard deviation.
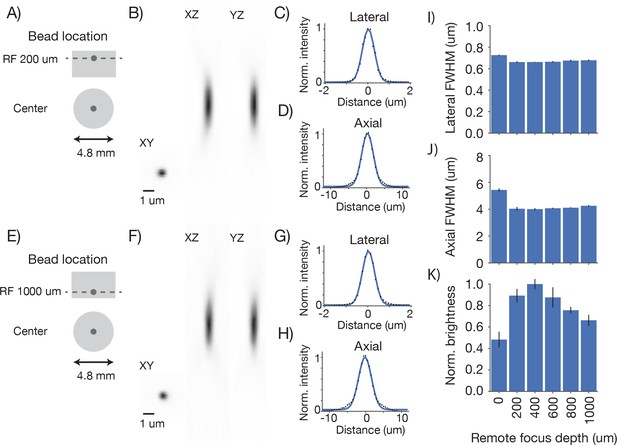
Point spread functions across imaging depth Point spread functions were measured at five different depths at the center of the FOV.
Same sample and imaging conditions as Figure 4. (A) Example measurement at the center of the FOV close to the cover glass at the top of the sample. (B) Mean intensity projections through the XY, XZ, and YZ slices of the bead. (C) Gaussian fit through the lateral slice through the bead image. (D) Same as (C), axial slice. (E–H) Same as (A–D) for a bead at the bottom of the imaging volume (RF set to 1 mm deep). (I) Average lateral full-width-at-half-max (FWHM) of the central XY slice of a bead, averaged across the X and Y directions. (J) Average axial FWHM. (K) Average normalized brightness of the beads. Error bars, standard deviation.
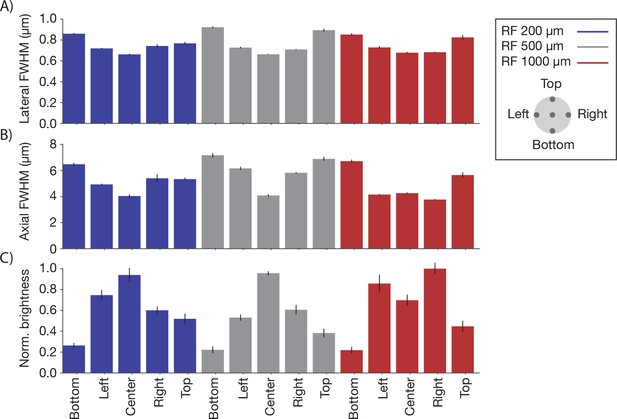
Point spread functions across the imaging volume Point spread functions were measured at 15 different locations across the imaging volume.
Three planes (depths; 0 µm, blue; 300 µm, grey; 800 µm, red) were sampled using the RF system. In each plane, five regions of the FOV were sampled, including the center and four extrema (2.4 mm from the center) in all quadrants (see legend). Same sample and imaging conditions as Figure 4. (A) Average lateral full-width-at-half-max (FWHM) of the central XY slice of a bead, averaged across the X and Y directions. (B) Average axial FWHM. (C) Average normalized brightness of the beads. Error bars, standard deviation.
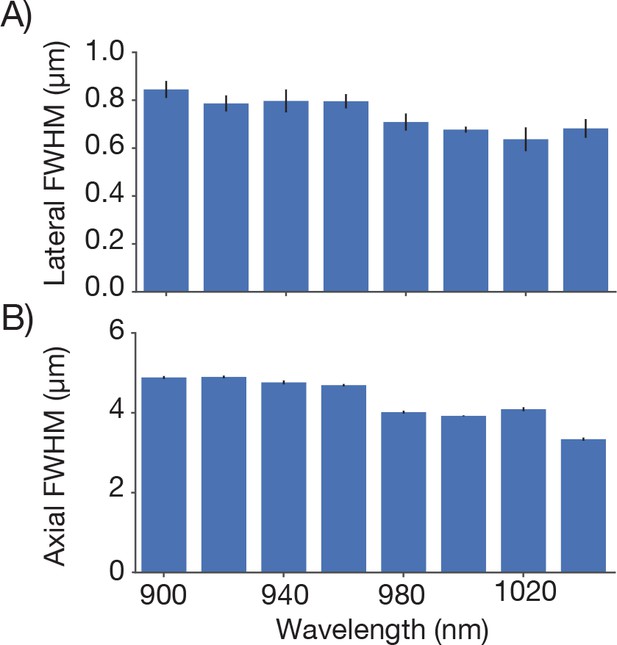
Point spread functions at different excitation wavelengths.
Point spread functions were measured at the center of the FOV for different excitation wavelengths. For each wavelength dispersion compensation was adjusted to give maximum signal from a fluorescein bath and same average power was used (conditions otherwise as in Figure 4). (A) Average lateral full-width-at-half-max (FWHM). (B) Average axial FWHM.
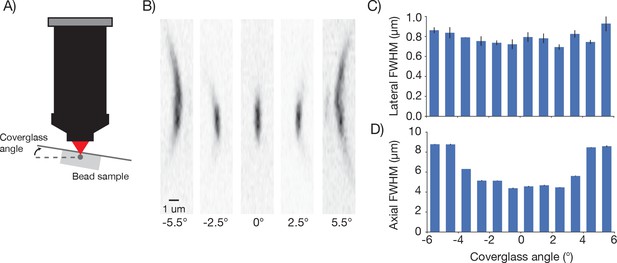
Point spread functions as a function of coverglass angle.
(A) Schematic illustrating how PSF size was measured at different coverglass angles. (B) Mean intensity projections through the XZ slices of a bead at different coverglass angles. (C) Average lateral full-width-at-half-max (FWHM). (D) Average axial FWHM.
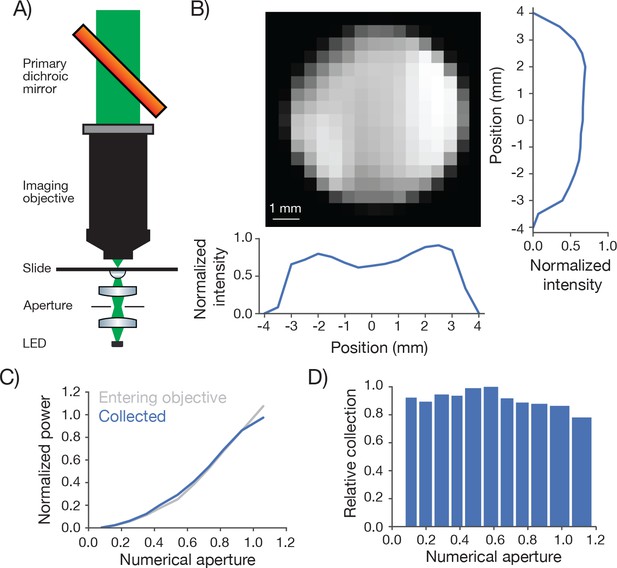
Measurement of collection efficiency.
(A) Apparatus for measuring signal collection across the FOV and determining the NA of the collection optics. The light source consisted of a LED (530 nm) and adjacent pinhole, with a miniature optical relay that projected the pinhole onto the top of a microscope slide. The projected spot had a 460 µm diameter. An aperture in the relay could be adjusted to change the NA of the light source from 0 to 1. (B) Normalized signal intensity measured with the PMT (not shown in A) as a function of position of the light source. The objective was moved in a 0.5 mm grid relative to the light source. Right, Y slice through the center of the image. Left, X slice through the center of the image. The peak signal was located off the main two axes. (C) Signal intensity measured by the PMT as a function of the numerical aperture of the light source (blue) overlaid on the power entering the objective (gray). As the NA of the light source increases the power entering the objective scales approximately as the square of the NA, as expected. The power entering the objective was measured with a high NA accepting power meter (Thor; S170C). Measurement was taken with the 460 µm spot in the center of the field. (D) Peak normalized ratio of the intensity measured by the PMT to the power entering the objective as a function of the NA of the light source (axis sampled non-uniformly). The signal intensity measured through the collection optics scales with the power entering the objective until it drops off around NA 1.0, significantly larger than the 0.6 excitation NA of the objective.
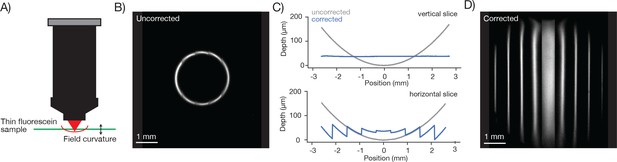
Characterization and correction of field curvature.
(A) Field curvature was measured using a thin fluorescein sample (11.2 µm, sandwiched between a microscope slide and 450 µm of coverglass, imaging conditions as in Figure 3A). At a particular RF position the image of the sample was a fluorescent ring. We measured the diameter of the ring as a function of RF position. (B) Image of the sample without any correction for field curvature. The sample appears as a thin ring. (C) Measured field curvature (gray: before field curvature correction, blue: after field curvature correction). (D) The RF mirror was programmed to compensate for the field curvature. The compensation is done on a line-by-line basis using the average position of the resonant mirror during the line as the point to correct. This compensation is able to correct the field curvature along each stripe well (top), but is unable to correct the field curvature within a resonant scan as the resonant mirror moves too fast (bottom).
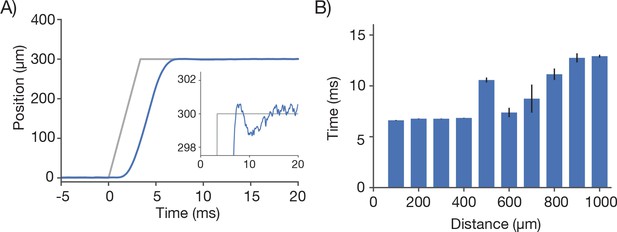
Characterization of remote focus mirror positioner (voice coil).
(A) Motion profile (blue) during of the RF mirror during a 300 µm move of the focal position. The command signal (grey) contained a 3.3 ms ramp to prevent oscillations of the actuator. Inset corresponds to a zoom during the settle period. (B) The step and settle times (to <2.5 µm) as a function of step size.
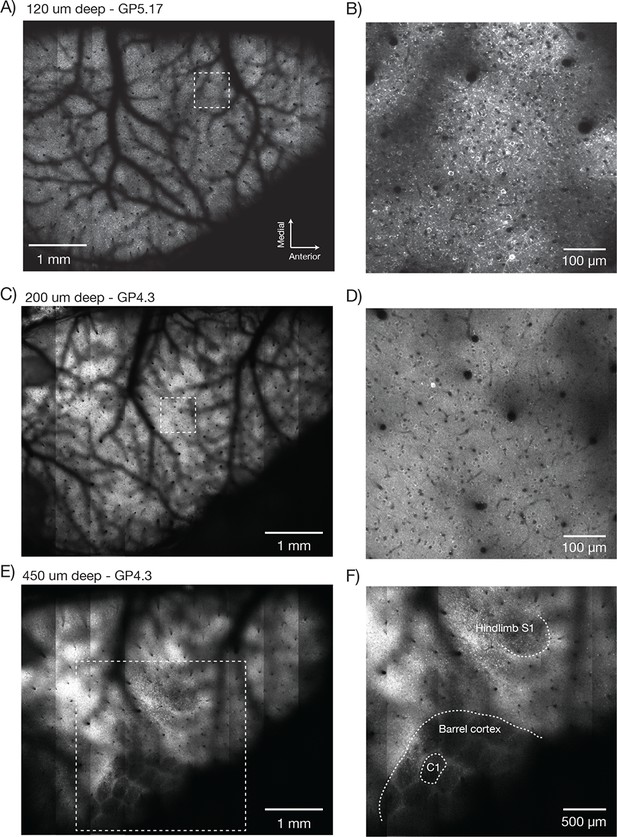
Imaging fluorescent proteins in anesthetized transgenic mice.
(A) Low magnification image (one section, 120 µm below the dura). The mouse expressed GCaMP6f under the thy-1 promoter (GP 5.17 line). Scanning parameters as in Figure 1Di. Field curvature correction was enabled. Average of 30 scans; power 120 mW. (B) Higher magnification image (dashed box in A). Scanning parameters as in Figure 1Dii. (C) Low magnification image (one section, 200 µm below the dura). The mouse expressed GCaMP6s under the thy-1 promoter (GP 4.3 line). Scanning parameters as in Fig. 1Di. Field curvature correction was enabled. Average of 30 scans; power 120 mW. Data corresponds to Video 1. (D) Higher magnification image (dashed box in C). Single neurons with nuclear excluded GCaMP are clearly visible. (E) Low magnification image at a deeper focal plane (one section, 450 µm below the dura). Same mouse and scanning parameters as C); power 175 mW. (F) Higher magnification image (dashed box in E) showing major somatosensory areas in layer 4. These areas appear as regions with reduced fluorescence since GCaMP6s expression is minimal in layer 4 stellate cells.
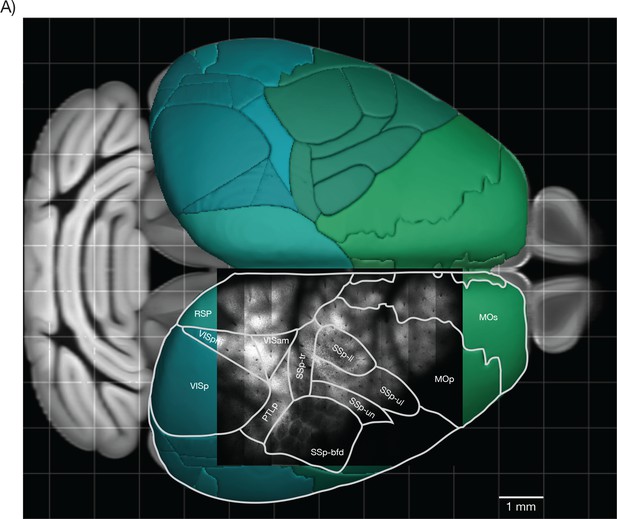
Overlay of a typical field of view with the reference atlas.
(A) Overlay of the image from Figure 7E with the mouse cortex (from Allen Brain Atlas, Brain Explorer 2). MOp: primary somatomotor, MOs: secondary somatomotor, SSp-ul: primary somatosensory upper limb, SSp-ll: primary somatosensory lower limb, SSp-un: primary somatosensory unassigned, SSp-tr: primary somatosensory trunk, SSp-bfd: primary somatosensory barrel field, PTLp: posterior parietal association, RSP: retrosplenial, VISam: anteromedial visual, VISpm: posteromedial visual, VISp: primary visual.
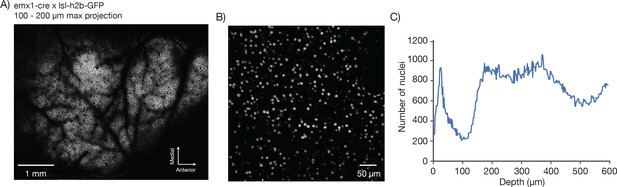

Counting nuclei.
(A) Max projection, 100 µm to 200 µm deep, imaged in a mouse expressing GFP in the nuclei of excitatory neurons (emx1-cre x lsl-h2b-GFP). The image was locally contrast enhanced with a contrast-limited adaptive histogram equalization (CLAHE, scikit-image). (B) Blow-up of a single section showing automatically detected nuclei in red. Nuclei were detected in 3D using a local peak finding algorithm (peak_local_max, scikit-image) after median filtering. (C) Number of detected nuclei per 2 µm thick slice, smoothed with a length 3 median filter.
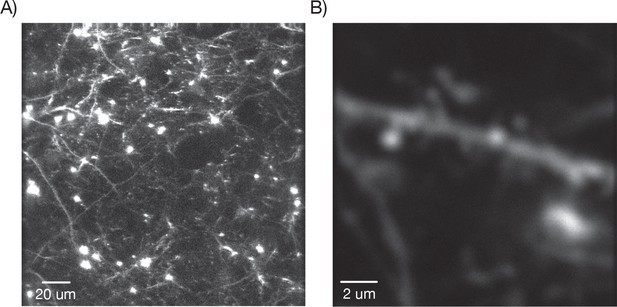
Dendritic images.
(A) One slice through a mouse expressing YFP in a subset of neurons. (B) Zoom in over a dendrite with clearly visible spines.
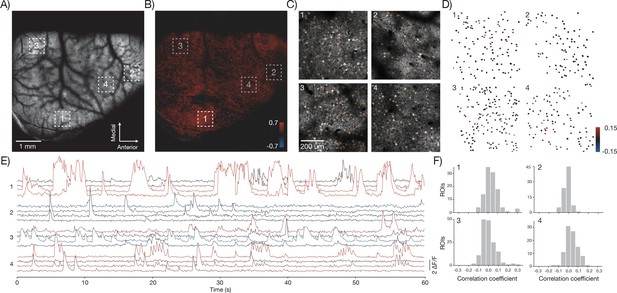
In vivo imaging of neural activity.
(A) Low magnification image from a mouse expressing GCaMP6f under the thy-1 promoter (GP 5.17 line) (parameters described in Figure 1Div; 120 mW average laser power). Sampling rate, 4.3 Hz. (B) Cross-correlation map of the mean activity in region 1 with the activity in all pixels after smoothing with a 5x5 pixel boxcar filter. A highly correlated spot of activity is visible around region 1. (C) Four fields of view (corresponding to boxes in A) that were acquired at high resolution and frame rate (scan parameters describe in Figure 1Dv; power, 120 mW; sampling rate, 9.6 Hz). Regions of interest (orange) were manually drawn around individual neurons. (D) Each neuron from the four fields of view colored according to its correlation value with the average activity in region 1. The average activity in region 1 was computed by taking the maximum values across all the Δ F / F traces in that region. (E) Δ F / F traces for 16 neurons extracted from the four separate regions (four per region) colored according to their correlation coefficient shown in D. Sampling rate, 9.6 Hz. Data corresponds to Video 3. (F) Histograms of the correlation coefficients for the neurons in D in the four separate regions.
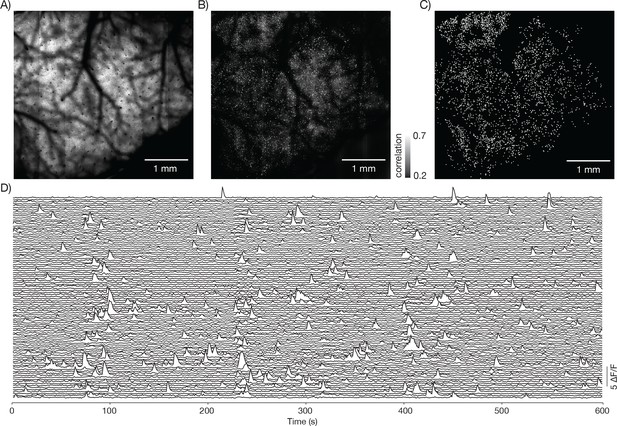
Neural activity in a large field of view.
(A) Image from a mouse expressing GCaMP6s under the thy-1 promoter (GP 4.3 line) (parameters described in Figure 1Diii, FOV 4.4 mm x 4.2 mm; 120 mW average laser power; sampling rate 1.9 Hz).
Image corresponds to Video 3. (B) Local correlation image, showing the result of cross-correlating each pixel with a 4-pixel neighborhood around it. (C) 3179 regions of interest were manually drawn around active neurons, using the local correlation image as a reference. (D) Δ F / F traces for 100 neurons. The neurons were selected based on having high skewness during that time period.
Videos
Multi-resolution information in a 2p-RAM image stack.
Same data as in Figure 7C.
Functional imaging of a large field of view.
Same data as in Figure 8—figure supplement 1. Movie data was smoothed and downsampled by a factor of two in time.
Low resolution imaging at low speed, followed by multi-area imaging at higher speeds.
Same data as in Figure 8A,C. Low resolution movie data smoothed and downsampled by a factor of three in time. High resolution movie data median filtered in space with filter size two, and smoothed and down sampled by a factor of five in time.





