Active suppression of a leaf meristem orchestrates determinate leaf growth
Figures
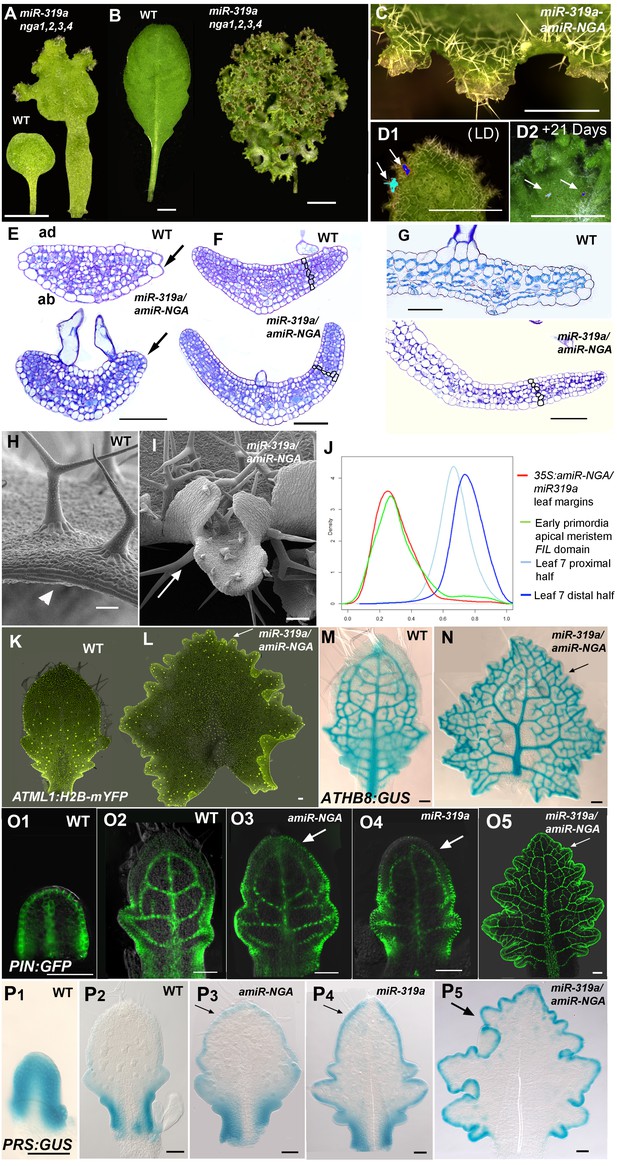
Reduction in NGA and CIN-TCP activities renders organ margins indeterminate.
(A, B) Overview of wild-type (WT) and 35S:miR319a nga1,2,3,4 (miR319a nga1,2,3,4) cotyledons (A) and leaves (B). (C-I) Close ups of leaf margins of WT and 35S:miR319a-amiR-NGA plants. (D1-D2) Third leaf of 35S:miR319a/35S:amiR-NGA plant marked with nail polish (arrows; light blue and dark blue), demonstrating the ongoing growth from the margins along 21 days. (E) Transverse sections through the distal end of developing wild-type and 35S:miR319a/35S:amiR-NGA leaves showing large versus small marginal cells (arrows). (F) Developing wild-type and 35S:miR319a/35S:amiR-NGA leaf primordia exhibit a similar six-cell-layered blade anatomy (outlined). (G) The margin of older wild-type margins have large, differentiated cells, whereas the 35S:miR319a/35S:amiR-NGA leaf margin retains the primordial blade structure. (H) The marginal cells of wild-type are elongated (arrowhead), while small isodiametric cells and initiating trichomes are found at 35S:miR319a/35S:amiR-NGA margins (I; arrow). (J) Transcriptome-based differentiation-score distributions of dissected 35S:miR319a/35S:amiR-NGA leaf margins, sorted primordia cells expressing FIL gene, and proximal or distal halves of seventh wild-type leaves (see Materials and methods for details). (K-L) Expression of ATML1:H2B-mYFP (yellow), ATHB8:GUS (M, N; blue), PIN1:PIN1-GFP (O; green), and PRS:GUS (P; blue) in developing leaves of indicated genotypes and a young wild-type leaf primordium (O1, P1). Note the distal exclusion of marker expression in slightly older wild-type leaves (O2, P2) while arrows indicate persisting expression along the distal margin with reduced NGA and CIN-TCP activities. ad, adaxial leaf side; ab, abaxial leaf side. Scale bars: A, D1-D2, 2 mm; B, 5 mm; C, 1 mm; 50 μm in other panels.
-
Figure 1—source data 1
Mean size of the leaf in wild-type and 35S:amiR-NGA plants, corresponding to the data shown in Figure 1—figure supplement 1C.
- https://doi.org/10.7554/eLife.15023.003
-
Figure 1—source data 2
Mean size of the palisade mesophyll cells in wild-type and 35S:amiR-NGA plants, corresponding to the data discussed in the legend to Figure 1—figure supplement 1D and E.
- https://doi.org/10.7554/eLife.15023.004
-
Figure 1—source data 3
CYCB1;1:GUS expression in distal wild-type and 35S:miR-NGA, correspnding to the data shown in Figure 1—figure supplement 1F–I.
- https://doi.org/10.7554/eLife.15023.005
-
Figure 1—source data 4
Effects on expression of different CIN-TCP and NGATHA family members and possible off targets in amiR-NGA and miR319a overexpressing plants- Figure 1—figure supplement 3.
- https://doi.org/10.7554/eLife.15023.006
-
Figure 1—source data 5
Differences in flowering time among wild-type, 35S:amiR-NGA, 35S:miR319a and 35S:amiR-NGA/35S:miR319a plants, corresponding to the data shown in Figure 1—figure supplement 4D.
- https://doi.org/10.7554/eLife.15023.007
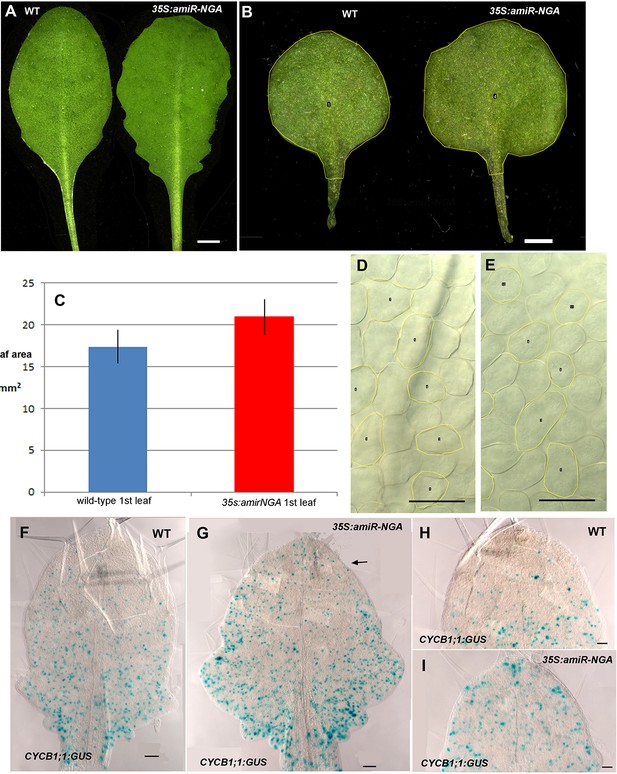
Altered growth in leaves with reduced NGATHA gene activity.
(A) Wild-type (WT) and 35S:amiR-NGA leaves showing increased serrations when activities of all four NGA genes are jointly compromised. (B) Representative first leaves of wild-type and 35S:amiR-NGA plants illustrating the larger size of the 35S:amiR-NGA leaf. The yellow outline and central numbers are annotations for leaf area analysis using ImageJ (v1.47). (C) The average leaf area of first leaves from wild-type and 35S:amiR-NGA plants (n = 20). Bars indicate mean ± SE. Leaf area was defined as depicted in B. A significant increase in leaf area was observed in 35S:amiR-NGA leaves (t-test, p=2.4 × 10–6). (D-E) Palisade mesophyll cells in wild-type, D and 35S:amiR-NGA leaves, E. The yellow outline and central numbers are annotations for leaf area analysis using ImageJ (v1.47). This analysis indicated that palisade mesophyll cells of 35S:amiR-NGA leaf size were somewhat smaller (mean cell area in wild-type = 1442 μm2, mean cell area in 35S:amiR-NGA = 1337 μm2; t-test, p=0.016, n = 80). (F–I) CYCB1;1:GUS expression (blue) in wild-type (F, H) and 35S:amiR-NGA (G, I) leaves. Strong GUS expression occurs during G2 and M phases of the cell cycle. (F) CYCB1;1:GUS expression in wild-type leaf at this stage is more abundant toward the base of the leaf and is comparatively reduced in the distal region. (G) In a similarly aged 35S:amiR-NGA leaf CYCB1;1:GUS expression is likewise more abundant toward the base of the leaf but is more frequently observed in the distal domain than in wild-type (arrow). (H) The distal region of a wild-type leaf showing low levels of CYCB1;1:GUS expression. (I) The distal region of a 35S:amiR-NGA leaf showing higher levels of CYCB1;1:GUS expression than wild-type. Expression was quantified as follows: after GUS staining and dissection, three leaves of the total length ranging from 1200 to 1500 µm were photographed for each genotype. The upper half of each image, as exemplified in F-G, was analyzed. CYCB1;1:GUS expressing area was scored as a percentage of distal leaf area. Whereas 3.2% of the distal leaf area of wild-type constituted CYCB1;1:GUS expression, 9% of the distal region of 35S:amiR-NGA leaves comprised CYCB1;1:GUS expression, suggesting that cell divisions in 35S:amiR-NGA leaves persist longer. Cell size and cell division measurements show that 35S:amiR-NGA plants with a compromised expression of all four NGA genes have larger leaves consisting of smaller cells and that prolonged cell division occurs in the distal region of the 35S:amiR-NGA leaf. Scale bars: A, 5 mm; B, 1 mm; 50 μm in the remainder.
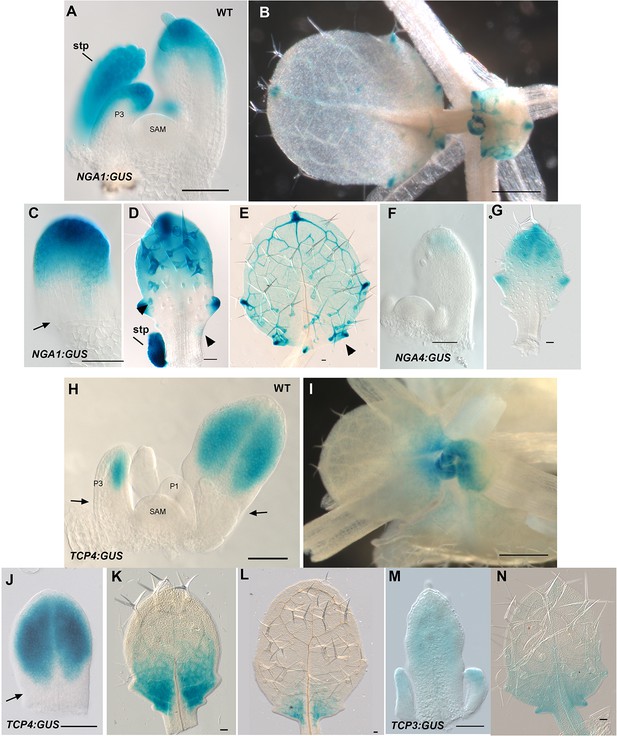
NGA1:GUS, NGA4:GUS, TCP4:GUS and TCP3:GUS expression in leaves.
(A-E) Expression pattern of NGA1:GUS (blue) in wild type. (A) NGA1:GUS expression is absent in the vegetative shoot apical meristem (SAM) and the proximal region of the older leaf primordia. The expression is initiated distally in the leaf primordium (P3). (B) Aerial view of a vegetative apex showing NGA1:GUS expression at the tips and serrations of older leaves. (C–E) Developmental series of leaves showing NGA1:GUS expression in the wild-type background. (C) An early leaf primordium shows strong NGA1:GUS expression at the leaf tip. (D) NGA1:GUS expression in an older leaf primordium. Lateral serrations express NGA1:GUS. As additional serrations form basipetally, NGA1:GUS expression in initiating serrations becomes apparent (arrowheads) at the base of the leaf. (E) A maturing wild-type leaf with additional lateral expression foci of NGA1:GUS (arrowhead). (F–G) Expression pattern of NGA4:GUS (blue) in wild type. (F) NGA4:GUS expression is absent from the SAM and is confined to the tip of the young leaves. (G) NGA4:GUS expression is confined to the tips and serrations of the older leaves. (H–L) Expression pattern of TCP4:GUS (blue) in wild type. (H) TCP4:GUS expression is absent in the vegetative shoot apical meristem (SAM), young leaf primordium (P1) and the proximal region of the older leaf primordia (arrows). TCP4:GUS expression is initiated distally in the older leaf primordia. (I) Aerial view of a vegetative apex showing TCP4:GUS expression retreating toward the base of older leaves. (J–L) TCP4:GUS expression is shown in a developmental series of leaves from young (J) to older (L). (J) A young leaf primordium showing distal TCP4:GUS expression. The proximal region lacks TCP4:GUS expression (arrow). (K) Later TCP4:GUS expression becomes excluded from the distal region and confined to the proximal region of leaf lamina. (L) In older leaves TCP4:GUS expression is further confined to the leaf base before being subsequently lost. (M–N) Expression pattern of TCP3:GUS (blue) in wild type. (M) TCP3:GUS expression is observed at the distal region of young leaves and stipules. (N) TCP3:GUS in older leaves becomes restricted to the base of the leaf. Note that the GUS reaction results in indigo blue colour. SAM, shoot apical meristem; stp, stipule. Scale bars: B, I, 1 mm; 50 μm in the remainder.
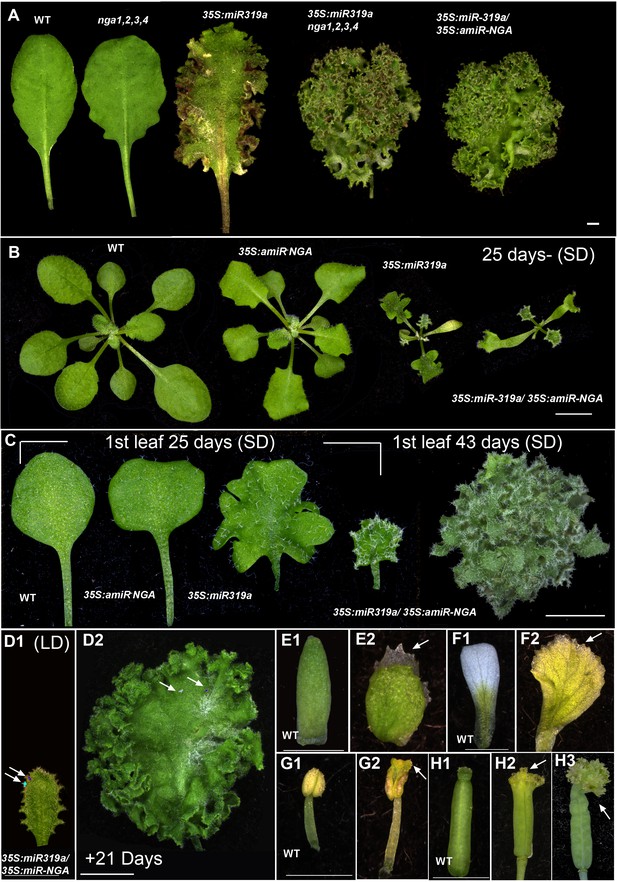

Ongoing marginal growth in leaves and floral organs with reduced CIN-TCP and NGATHA gene activities.
(A) Leaves of wild-type, nga1,2,3,4, 35S:miR319a and nga1,2,3,4 35S:miR319a and 35S:miR319a/35S:amiR-NGA leaves illustrating the synergistic effect on leaf marginal growth when both the CIN-TCP and NGATHA gene activities are reduced (for data on the effect of miR319a and amiR-NGA on the expression of targets and related gene family members see Figure 1—source data 4). (B) Representative wild-type, 35S:amiR-NGA, 35S:miR319a, and 35S:miR319a/35S:amiR-NGA plants grown for 25 days under short-day (SD) conditions. Leaf expansion is delayed, and the leaf initiation rate is decreased in 35S:miR319a/35S:amiR-NGA lines. (C) From left to right shown are the first leaves of wild-type, 35S:amiR-NGA, 35S:miR319a, and 35S:miR319a/35S:amiR-NGA plants grown for 25 days, and the rightmost is the first leaf of 35S:miR319a/35S:amiR-NGA plant grown for 43 days. The first leaves of wild-type, 35S:amiR-NGA and 35S:miR319a plants have reached their maximum sizes after 25 days of growth while the first leaf of 35S:miR319a/35S:amiR-NGA plant continues to grow from the margins reaching a large and complex shape with continued marginal growth, photographed here after 43 days. (D1–D2) Third leaf of a long-day (LD) grown 35S:miR319a/35S:amiR-NGA plant marked with nail polish (arrows; light blue and dark blue), demonstrating the growth from the margins in 21 days. (E1–H3) Floral organs from wild-type (E1, F1, G1, H1) and 35S:miR319a/35S:amiR-NGA (E2, F2, G2, H2, H3) plants, illustrating the continued growth from organ tips when CIN-TCP and NGA gene activities are compromised. Sepals (E1, E2), petals (F1, F2), stamens (G1, G2), and gynoecia (H1, H2, H3) are shown. Scale bars: A-B, D2, 5 mm; C, 2 mm; D1, E1-H3. 1 mm.
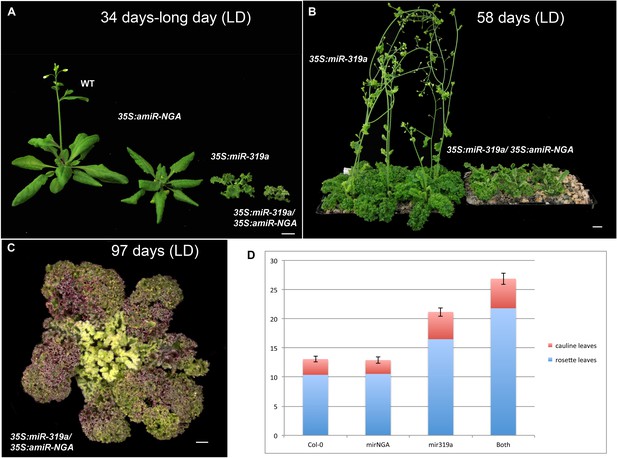
Plant growth and flowering time in plants with reduced CIN-TCP and NGATHA gene activities.
(A) Representative wild-type, 35S:amiR-NGA, 35S:miR319a and 35S:miR319a/35S:amiR-NGA plants after growing 34 days under the long-day conditions. Note that only the wild-type and 35S:amiR-NGA plants have bolted. (B) 35S:miR319a and 35S:miR319a/35S:amiR-NGA plants after growing 54 days under the long-day conditions. The 35S:miR319a/35S:amiR-NGA plants are yet to bolt. (C) 35S:miR319a/35S:amiR-NGA plant after growing 97 days under the long-day conditions. The plant is beginning to bolt. (D) Flowering time of wild-type, 35S:amiR-NGA, 35S:miR319a and 35S:miR319a/35S:amiR-NGA plants grown under the long-day conditions. Flowering time was measured as a mean number of leaves produced. While no significant difference was observed between wild-type and 35S:amiR-NGAplants (t-test, p=0.83, n = 10), differences were apparent between wild-type and 35S:miR319a (t-test, p=4.8–09, n = 10) and between 35S:miR319a and 35S:miR319a/35S:amiR-NGA plants (t-test, p=7.4-07, n = 10). Bars indicate mean ± SE. Scale bars: C, 5 mm; A–B, 1 cm.
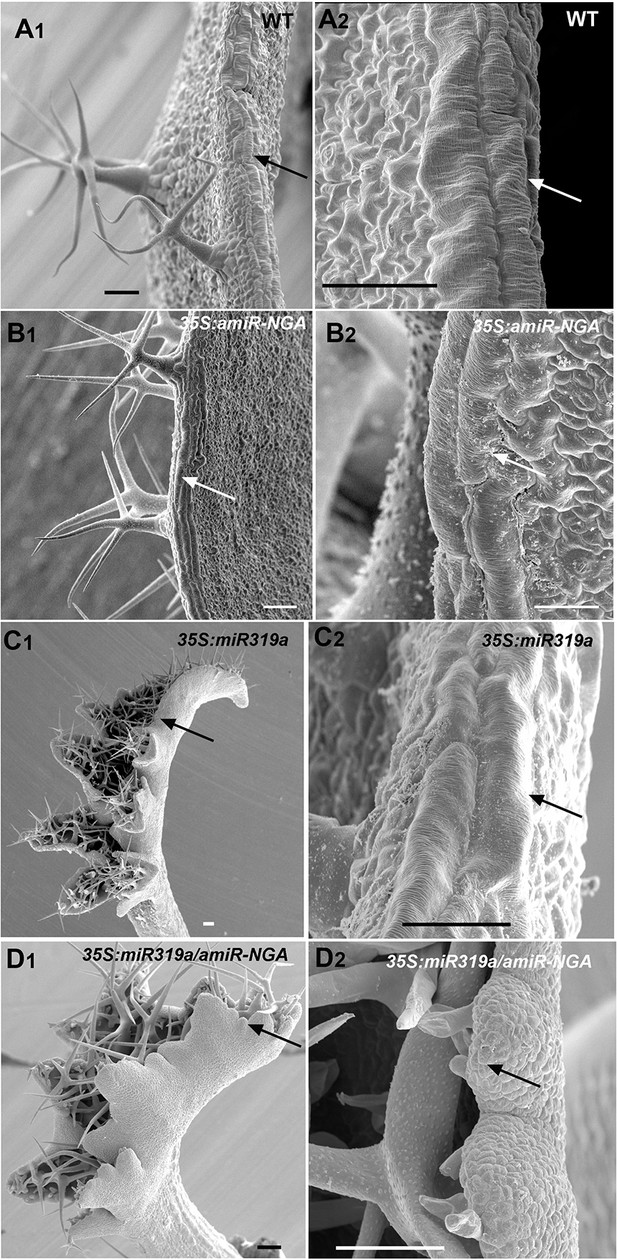
SEM of leaf margin cells with reduced CIN-TCP and NGATHA gene activities.
Wild-type, 35S:amiR-NGA, 35S:miR319a and 35S:miR319a/35S:amiR-NGA lines captured by scanning electron micrographs (SEMs) at low (1) or high (2) magnification. (A) Wild-type leaf margin at low (A1) or high (A2) magnification showing the elongated cells at the margin (arrows). (B) 35S:amiR-NGA leaf margin with elongated cells (arrows). (C) Developing 35S:miR319a leaf margin with elongated cells (arrows). (D) Developing 35S:miR319a/35S:amiR-NGA leaf margin lacking elongated cells (arrows). Scale bars: 50 μm in A to B, C, and D to D2; 25 μm in B2 and C2.
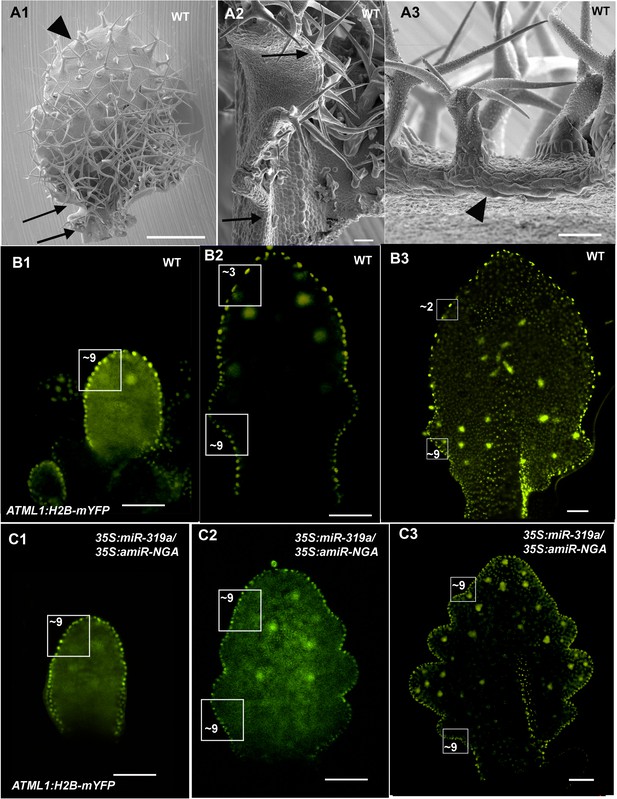
Patterns of proximal to distal leaf margin cell expansion.
(A1-A3) SEM of a differentiating wild-type leaf margin demonstrating a gradient of proximal to distal cell elongation. (A1) The adaxial surface of a wild-type leaf with regions of the left margin demarked by arrows and an arrowhead shown in detail in (A2) and (A3). (A2) Marginal region between the arrows shows a proximal to distal gradient of increasing cell length. (A3) The distal region of the leaf has prominent, elongated marginal cells (arrowhead). A developmental series of wild-type (B1-B3) and 35S:miR319a/35S:amiR-NGA (C1–C3) leaves where the margin cell nuclei are marked by ATML1:H2B-mYFP expression. Regions of 2500 μm2 are boxed and the approximate numbers of margin cell nuclei are given. (B1–B3) In older wild-type leaves the number of margin cell nuclei in the distal region is reduced relative to the proximal leaf due to cell expansion and differentiation. (C1–C3) In developing 35S:miR319a/35S:amiR-NGA leaves the number of marginal cells in the distal relative to the proximal domains does not change, reflecting lack of differentiation and expansion of the marginal cells. Scale bars: A, 1 mm; 50 μm in other panels.
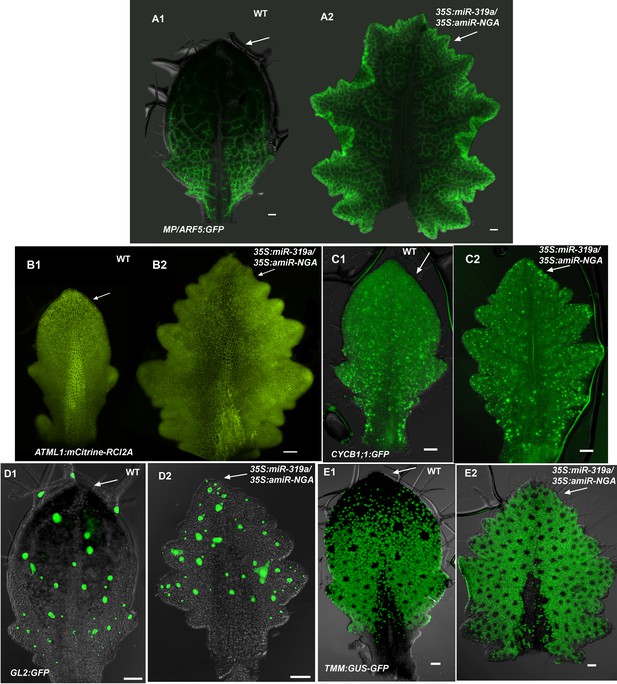
Distribution of markers in leaves with reduced CIN-TCP and NGATHA gene activities.
(A) Expression of MP/ARF5:GFP (green) occurs in provasculature and diminish as the vasculature differentiate. In wild-type differentiating leaf (A1) MP/ARF5:GFP decreases in the distal part of the leaf (arrow) but is maintained near the leaf margin and the provasculature at the leaf base. ARF5:GFP expression continues at the distal margins of older 35S:miR319a/35S:amiR-NGA leaves (A2) (arrow) associated with de novo provasculature formation and, unlike wild-type, remains active in the distal vasculature of internal maturing leaf tissue appearing more reticulate. (B) Expression of ATML1:mCitrine-RCI2A (yellow), which marks the plasma membrane of epidermal cells, outlining cells in the epidermis. In a wild-type leaf (B1), cells in the distal region of the leaf (arrow) are larger relative to those at the base. Expression of ATML1:mCitrine-RCI2A in a larger, older 35S:miR319a/35S:amiR-NGA leaf (B2) shows that the cells at the distal, marginal region remain small (arrow). (C) Expression of CYCB1;1:GFP in a developing leaf. More cells at the distal leaf margin of 35S:miR319a/35S:amiR-NGA leaf (arrow) express CYCB1;1:GFP compared to wild-type. (D) GL2:GFP expression (green) becomes reduced in the distal domain of wild-type leaves (D1) but continues to be expressed in initiating trichomes in the distal region of 35S:miR319a/35S:amiR-NGA leaves (D2, arrow). (E) TMM-GUS-GFP expression (green) is excluded from the distal wild-type leaf (E1) but continues in the distal region of 35S:miR319a/35S:amiR-NGA leaves (E2, arrow). Scale bars: 50 μm.
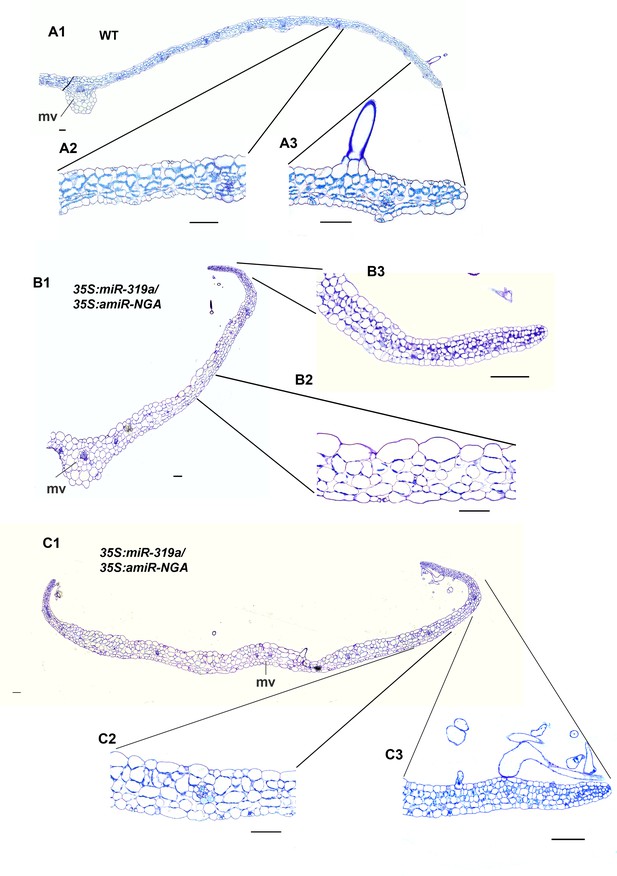
Transverse sections of leaves with reduced NGATHA and CIN-TCP activities.
(A1–C3) Transverse sections through wild-type (A1–A3), and two 35S:miR319a/35S:amiR-NGA leaves (B1-B3, C1-C3) with higher magnification images of marginal (A2, B2, C2) and internal (A3, B3, C3) regions of each leaf. In contrast to the differentiated margins of wild-type leaves (A3), the leaf margins of 35S:miR319a/35S:amiR-NGA plants (B3, C3) have the similar five/six-cell-layered blade of small, densely cytoplasmic cells typical of early wild-type leaf primordia (see Figure 1F–G). The wild-type leaf is of regular thickness (A1, A2) whereas 35S:miR319a/35S:amiR-NGA leaves are thicker, less regular, and appear to be composed of larger cells (B1-B2 and C1-C2). mv, midvein. Scale bars: 50 μm.
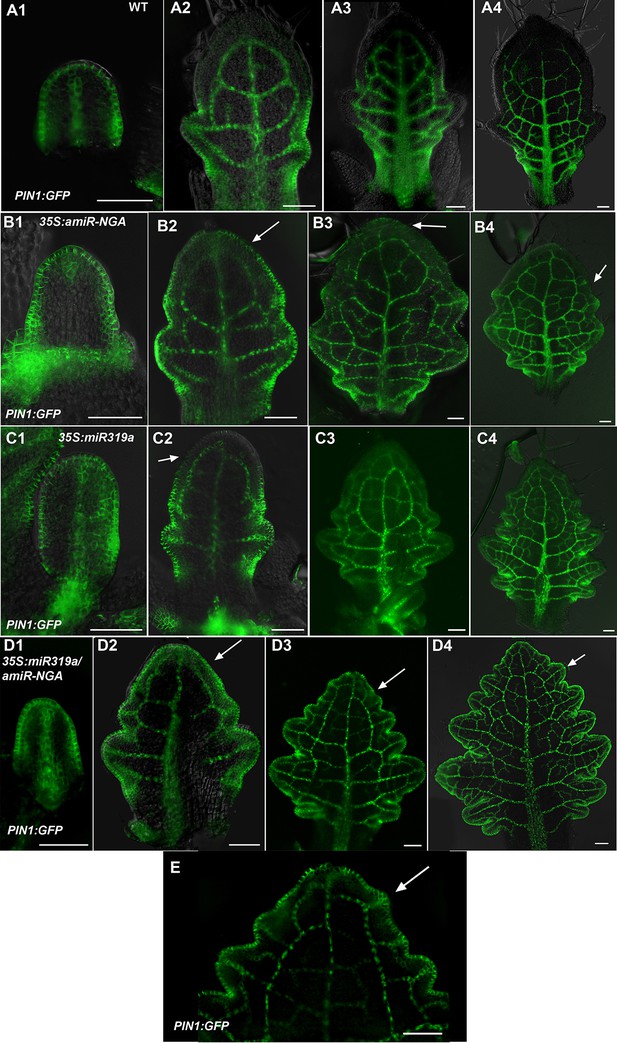
Changes in PIN1-GFP expression when CIN-TCPs and NGATHA gene activities are reduced.
(A1-A4) PIN1:PIN1-GFP expression (green) in developing wild-type leaves. (A1) In the early leaf primordium PIN1:PIN1-GFP expression is seen around the entire margin and in the primary provascualar strand. (A2-A4) In older primordia PIN1:PIN1-GFP expression is gradually excluded from the distal margin but is maintained at the proximal margins, leaf sinuses and provasculature. (B1–B4) PIN1:PIN1-GFP expression in developing 35S:amiR-NGA leaves. (B1) In early 35S:amiR-NGA leaf primordia PIN1:PIN1-GFP expression occurs around the margin as in wild-type. (B2–B4) PIN1:PIN1-GFP expression is maintained for a longer period at the distal margins of 35S:amiR-NGA leaf primordia than in wild-type (arrows). (C1-C4) PIN1:PIN1-GFP expression in 35S:miR319a leaves. (C1) In early 35S:miR319a leaves PIN1:PIN1-GFP is expressed around the entire margin. (C2–C4) In older leaves PIN1:PIN1-GFP is maintained for longer at the distal margin than in wild-type (C2; arrow) but subsequently becomes excluded from the distal margin (C3, C4). (D1–D4) PIN1:PIN1-GFP expression in 35S:miR319a/35S:amiR-NGA leaves. (D1) PIN1:PIN1-GFP is expressed around the margin of early 35S:miR319a/35S:amiR-NGA leaf primordia. (D2–D4) PIN1:PIN1-GFP expression continues at the margins of older 35S:miR319a/35S:amiR-NGA leaves (arrows). (E) The distal region of an older PIN1:PIN1-GFP 35S:miR319a/35S:amiR-NGA leaf showing presumptive auxin canalization marked by PIN1:PIN1-GFP expression along the margin (arrow). Scale bars: 50 μm.
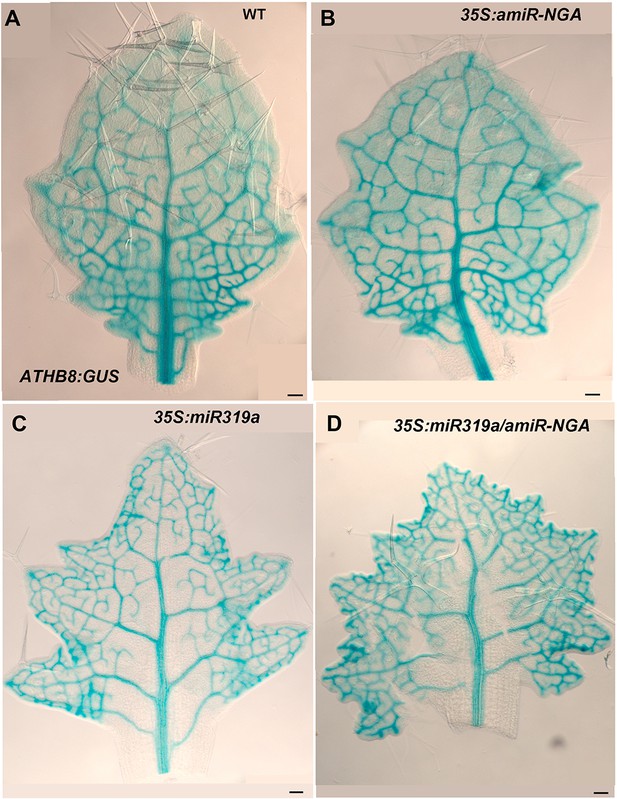
ATHB8:GUS expression in leaves with reduced CIN-TCPs and NGATHA gene activities.
ATHB8:GUS expression (blue) marks the developing provasculature and is illustrated here in similarly sized, developing leaves. (A) ATHB8:GUS expression in a developing wild-type leaf. Vasculature expression is diminished and GUS expression lacks from the margin in the distal leaf while GUS expression is strong and extends to the margin at the base of the leaf. (B) ATHB8:GUS expression in 35S:amiR-NGA leaf is largely similar to wild-type, but the expression is maintained longer distally. (C) ATHB8:GUS expression in a 35S:miR319a leaf identifies reticulated marginal expression maintained in the distal leaf. (D) ATHB8:GUS expression in a 35S:miR319a/35S:amiR-NGA leaf showing strong, reticulate expression extending to all margins. Scale bars: 50 μm.
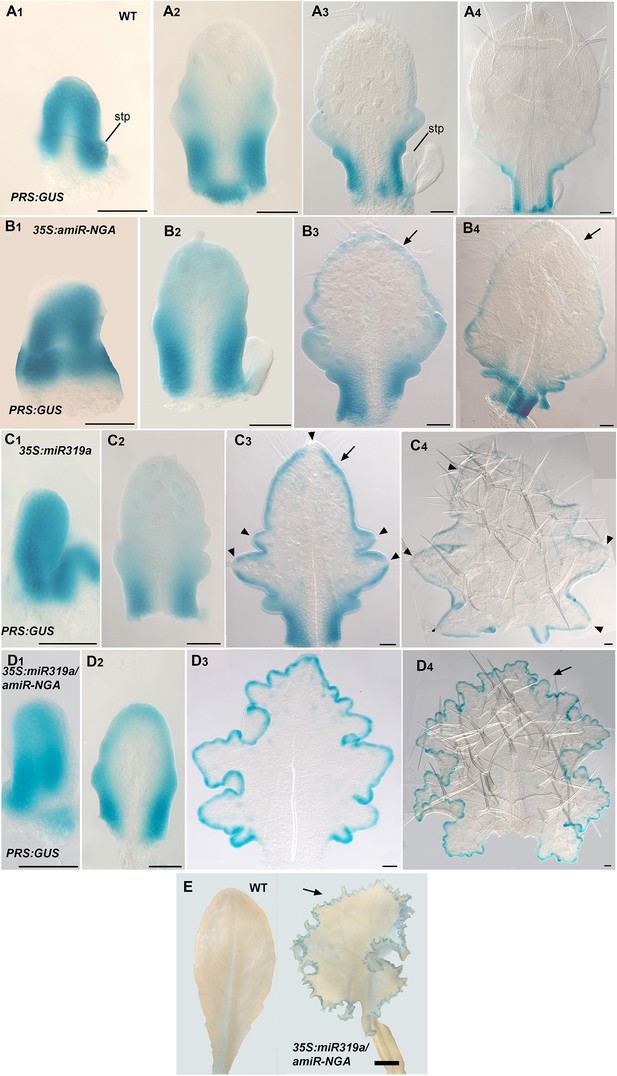
PRS:GUS expression is maintained longer at the leaf margins when CIN-TCP and NGATHA gene activities are reduced.
(A1–D4) PRS:GUS expression (blue) at various developmental stages from early (left) to late (right) in each row. (A1–A4) PRS:GUS expression in wild-type leaves. (A1) PRS:GUS is expressed marginally in a young leaf primordium. (A2) In an older wild-type primordium PRS:GUS expression becomes restricted to the proximal leaf. (A3) PRS:GUS expression is excluded from the distal region but is maintained at the proximal region including the incipient petiole. (A4) PRS:GUS expression is only present at the base of the leaf and incipient petiole. (B1-B4) PRS:GUS expression in 35S:amiR-NGA leaves. (B1) An early 35S:amiR-NGA leaf primordium showing PRS:GUS expression around the entire margin. (B2) In an older 35S:amiR-NGA leaf primordium PRS:GUS expression is strongest at the base and is also present distally. (B3, B4) In an older leaf primordium PRS:GUS expression is strongest at the leaf base and incipient petiole and is also maintained at the distal margin (arrows). (C1–C4) PRS:GUS expression in 35S:miR319a leaves. (C1) A young 35S:miR319a leaf primordium with PRS:GUS expression around the entire margin. (C2) In an older 35S:miR319a leaf primordium PRS:GUS expression is observed around the margin and is strongest at the base of the leaf. (C3, C4) Older 35S:miR319a leaf primordia showing characteristic deep serrations. PRS:GUS expression is observed around the margin (arrow; C3) but is absent from the tip of the leaf and serration tips (arrowheads). (C4) PRS:GUS expression is still maintained at low levels. Arrowheads indicate the lack of PRS:GUS expression at serration tips. (D1–D4) PRS:GUS expression in 35S:miR319a/35S:amiR-NGA leaves. (D1) An early 35S:miR319a/35S:amiR-NGA leaf primordium with PRS:GUS expression around the entire margin. (D2) A young 35S:miR319a/35S:amiR-NGA leaf primordium with PRS:GUS expression at the margins. (D3, D4) In older 35S:miR319a/35S:amiR-NGA leaves with elaborate leaf margins, PRS:GUS expression is maintained seamlessly at the margins (arrow; D4). All leaves are arranged with the adaxial surface facing up. stp, stipule. Scale bars: E, 2 mm; all other panels, 50 μm.
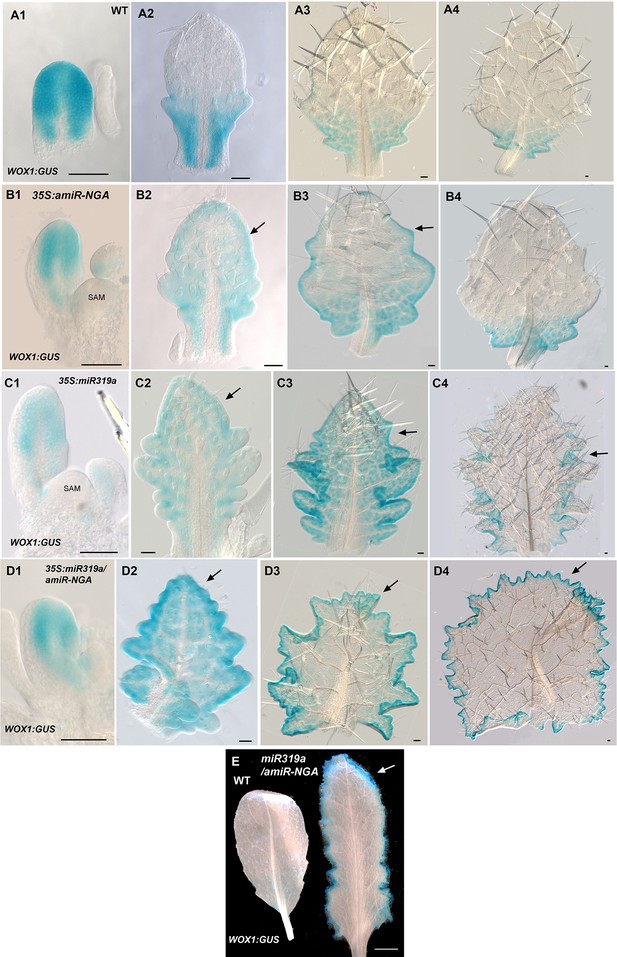
WOX1:GUS expression is maintained longer at the leaf margins when CIN-TCP and NGATHA gene activities are reduced.
(A1–D4) WOX1:GUS expression (blue) at various developmental stages from early (left) to late (right) in each row. (A1–A4) WOX1:GUS expression in wild-type leaves. (A1) WOX1:GUS is expressed around the margin of an early leaf primordium. (A2) In an older leaf primordium WOX1:GUS expression becomes restricted to the proximal leaf including the incipient petiole. (A3) WOX1:GUS expression is absent from the distal region but is maintained at the basal region. (A4) In an older leaf residual WOX1:GUS expression occurs only at the leaf base. (B1–B4) WOX1:GUS expression in 35S:amiR-NGA leaves. (B1) An early 35S:amiR-NGA leaf primordium with WOX1:GUS expression at the margin. (B2) An older 35S:amiR-NGA leaf primordium with WOX1:GUS expression maintained throughout the leaf (arrow). (B3) An older 35S:amiR-NGA leaf primordium where WOX1:GUS expression is strongest at the base and is maintained at the distal margin (arrow). (B4) An older leaf primordium with WOX1:GUS expression restricted to the leaf base (arrow). (C1–C4) WOX1:GUS expression in 35S:miR319a leaves. (C1) Young 35S:miR319a leaf primordia with WOX1:GUS expression at the margin. (C2) An older 35S:miR319a leaf primordium with WOX1:GUS expression maintained distally (arrow). (C3) An older 35S:miR319a leaf with WOX1:GUS expression observed throughout the leaf including the distal domain (arrow). (C4) In an older 35S:miR319a leaf primordium WOX1:GUS expression is reduced at the leaf tip and serrations but is still present in the sinuses (arrow). (D1–D4) WOX1:GUS expression in 35S:miR319a/35S:amiR-NGA leaves. (D1) An early 35S:miR319a/35S:amiR-NGA leaf primordium with WOX1:GUS expression at the margins. (D2) A young 35S:miR319a/35S:amiR-NGA leaf primordium shows WOX1:GUS expression throughout the leaf with stronger expression around the margin. (D3) An older 35S:miR319a/35S:amiR-NGA leaf with elaborated leaf margins. WOX1:GUS is continuously expressed at the margins including the distal margin (arrow). (D4) In an older 35S:miR319a/35S:amiR-NGA leaf with WOX1:GUS expression at all margins including the distal margin (arrow). (E) WOX1:GUS expression in older wild-type and 35S:miR319a/amiR-NGA leaves showing strong and continued expression at the margins of the 35S:miR319a/amiR-NGA leaf (arrow). Note the absence of GUS expression in the wild-type leaf. All leaves are arranged with the adaxial surface facing up. SAM, shoot apical meristem. Scale bars: A1–D4, 50 μm; E, 5 mm.
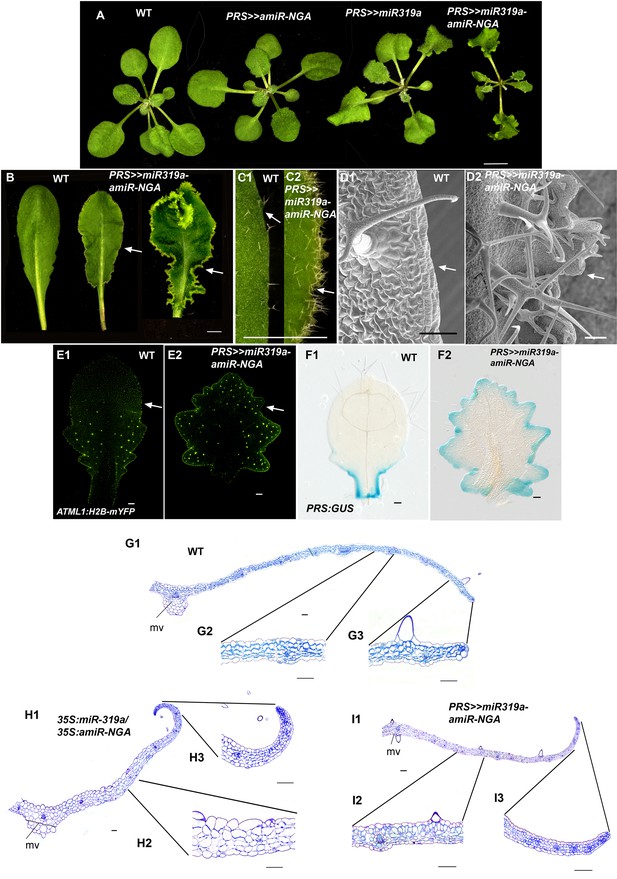
Reduced NGATHA and CIN-TCP gene activities in the PRESSED FLOWER (PRS) domain alters leaf marginal growth.
All lines are obtained using transactivation (PRS:LhG4). (A–F2) Plants and leaves are placed with the adaxial face up. (A) Wild-type, PRS>>amiR-NGA, PRS>>miR-319a and PRS>>miR-319a-amiR-NGA plants showing the increased leaf marginal growth when activity of the NGATHAs, CIN-TCPs and both NGATHAs and CIN-TCPs are reduced in the PRS expression domain. (B) Wild-type and PRS>>miR-319a-amiR-NGA leaves showing indeterminate leaf margins (arrows). (C1–C2) Higher magnification image of the margins of wild-type (C1) and PRS>>miR-319a-amiR-NGA (C2) leaves showing active margins (arrow). (D1–D2) SEM of wild-type (D1) and PRS>>miR-319a-amiR-NGA leaves (D2) illustrating the elongate cells of wild-type (arrow) relative to the jagged margins with small cells and initiating trichomes of PRS>>miR-319a-amiR-NGA leaves (arrow). (E1–E2) Wild-type (E1) and PRS>>miR-319a-amiR-NGA leaves (E2) with the epidermal nuclei are marked by ATML1:H2B-mYFP expression (yellow). Sparse ATML1:H2B-mYFP expression is observed in the distal part of the wild-type leaf (arrow). By contrast a high density of expression is apparent around the distal margins of the PRS>>miR-319a-amiR-NGA leaf and the nuclei of developing trichomes which have undergone endoreduplication (arrow). (F1–F2) PRS:GUS expression (blue) in a differentiating wild-type (F1) and PRS>>miR-319a-amiR-NGA leaf (F2). Whereas PRS:GUS expression is confined to the proximal margins of the wild-type leaf, expression occurs around the entire PRS>>miR-319a-amiR-NGA leaf margin. (G1–13) Transverse sections of wild-type (G1–G3), 35S:miR319a/35S:amiR-NGA (H1–H3) and PRS>>miR-319a-amiR-NGA (I1–I3) leaves. The margins of 35S:miR319a/35S:amiR-NGA (H3) and PRS>>miR-319a-amiR-NGA (I3) leaves have the similar five/six-cell-layered blade of small cells that characterizes the development of early wild-type leaf primordia (see Figure 1F–G) in contrast to the differentiated margins of wild-type (G3). Internally, the structure of the PRS>>miR-319a-amiR-NGA (12) more closely resembles that of wild-type (G2) than 35S:miR319a/35S:amiR-NGA (H2), where the blade is thicker and the cells appear enlarged. mv, midvein. Scale bars: A, B, 5 mm; C1–C2, 2 mm; 50 μm in other panels.
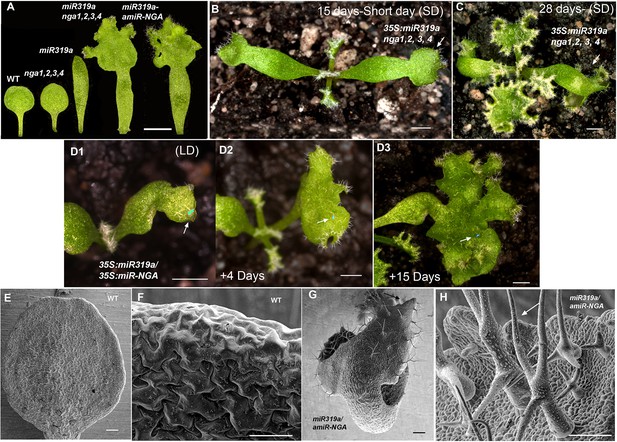

Ongoing marginal growth in cotyledons with reduced CIN-TCP and NGATHA gene activities.
(A) From left to right shown are the cotyledons of wild-type, nga1,2,3,4, 35S:miR319a, nga1,2,3,4 35S:miR319a and 35S:miR319a-amiR-NGA plants, showing differences in cotyledon shape and growth. (B) 35S:miR319a nga1,2,3,4 seedling growing for 15 days under the short-day conditions, showing extended growth at the cotyledon tips (arrow) and first leaves. (C) The same 35S:miR319a nga1,2,3,4 seedling as in (B) after 28 days of growth, illustrating the continued growth from the cotyledon tips (arrow) and rosette leaves. (D1–D3) Cotyledon of the same 35S:miR319a/35S:amiR-NGA plant marked with nail polish (light blue; arrows), illustrating marginal growth of the cotyledon over 15 days under the long-day conditions. (E–H) Scanning electron micrographs of the adaxial surface of wild-type (E, F) and 35S:miR319a/35S:amiR-NGA (G, H) cotyledons. The distal margin of cotyledons is shown at higher magnification in F, H. The distal region of 35S:miR319a/35S:amiR-NGA cotyledons continues to grow with leaf-specific characteristics, such as stellate trichomes (arrow). Large epidermal cells are seen in wild-type (F) whereas small cells and initiating trichomes are observed in 35S:miR319a/35S:amiR-NGA (H, arrow). Scale bars: A, 2 mm; B–D3, 1 mm; E–H, 50 μm.
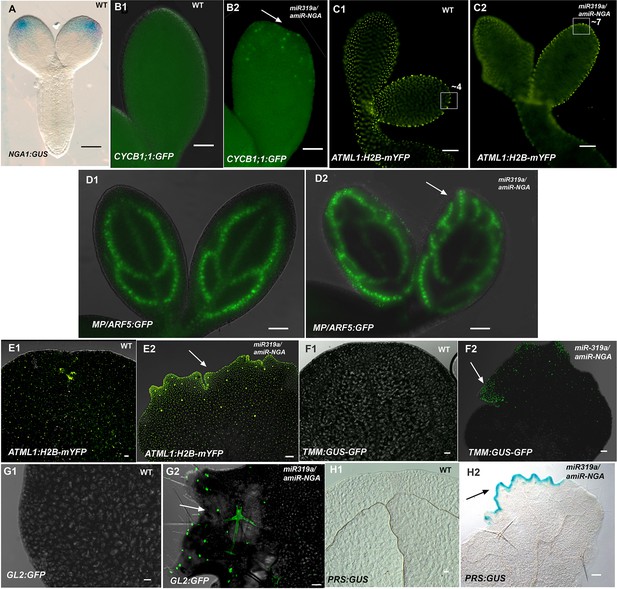
Morphological and marker analyses in cotyledons with reduced CIN-TCP and NGATHA gene activities.
(A) NGA1:GUS (blue) is expressed at the cotyledon tips of embryo eight days after pollination. (B1–D2) Marker expression in embryos nine days after pollination. (B1, B2) CYCB1;1:GFP expression (green) in embryo cotyledons. Little or no CYCB1;1:GFP expression was detected in wild-type (B1) while extensive expression was observed in the cotyledon tips (arrow) in 35S:miR319a/35S:amiR-NGA (B2). (C1, C2) Expression of ATML1-H2B-mYFP (yellow), which marks the nuclei of the epidermal cells. (C1) Expression at the margins of developing wild-type cotyledons indicates regular, enlarging cells. (C2) At the margins of the developing 35S:miR319a/35S:amiR-NGA cotyledons the cells appear smaller and more numerous. Regions of 2500 μm2 are boxed and the approximate numbers of margin-cell-nuclei are given. (D1, D2) MP/ARF5:GFP (green) expression in the developing wild-type cotyledons. (D1) MP/ARF5:GFP expression is strongest in the younger, peripheral, provascular stands and is diminished in the older central strand of wild-type. (D2) MP/ARF5:GFP expression in the developing 35S:miR319a/35S:amiR-NGA cotyledons reveals extensive expression in presumptive de novo provasculature extending from the margins (arrow).(E1–H2) Expression of markers in the cotyledons of wild-type and 35S:miR319a/35S:amiR-NGA plants after germination. (E1, E2) Adaxial cotyledon surface of plants expressing ATML1-H2B-mYFP (yellow), which marks the nuclei of the epidermal cells. (E1) Dispersed YFP signal suggests well-spaced nuclei, reflecting the presence of large, differentiated cells in wild-type cotyledons. (E2) In a 35S:miR319a/35S:amiR-NGA cotyledon, the nuclear YFP expression is observed in high density at and near the margins, suggesting the presence of numerous small cells. Large YFP dots are associated with endoreduplicating trichome cells (arrow) not observed in wild-type. (F1) Adaxial surface and margin of a wild-type cotyledon expressing the leaf meristemoid regulator TMM:GUS-GFP (green). No GFP signal is observed. (F2) Adaxial surface and margin of a 35S:miR319a/35S:amiR-NGA cotyledon with TMM:GUS-GFP expression apparent in cells associated with the margin (arrow). (G1) Adaxial surface and margin of a wild-type cotyledon expressing the trichome marker GL2:GFP. No GFP signal is observed. (G2) Adaxial surface and margin of a 35S:miR319a/35S:amiR-NGA cotyledon with GL2:GFP expression (green) associated with the initiating and developing trichomes (arrow). (H1, H2) Cleared cotyledon tips of plants expressing PRS:GUS (blue), a reporter construct for the PRS gene. (H1) No GUS signal was observed in wild-type. (H2) Strong PRS:GUS expression (blue) is seen at the margin (arrow) in 35S:miR319a/35S:amiR-NGA plants. Scale bars: 50 μm.
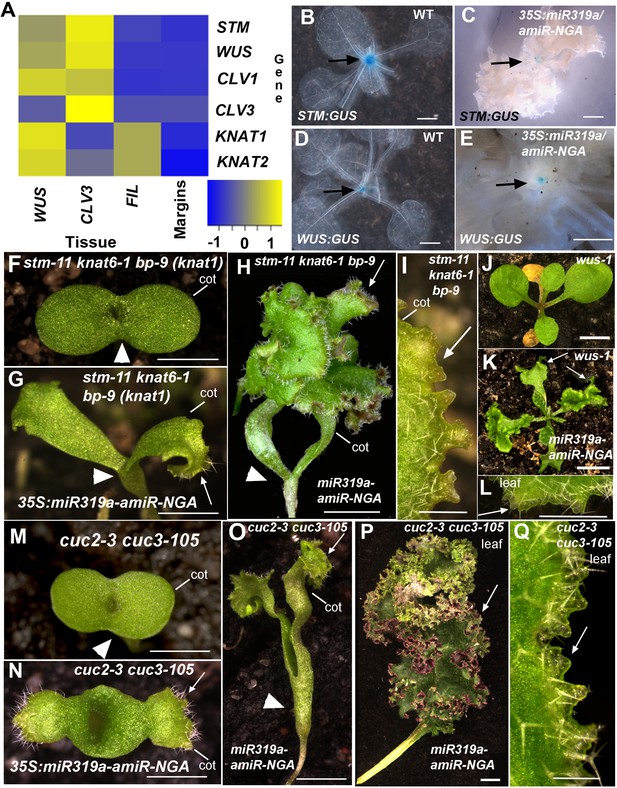

Regulators of SAM maintenance and leaf margin elaboration are dispensable for maintenance of the marginal leaf meristem.
(A) Relative mRNA expression levels of meristem regulators in cells collected from apices and in indeterminate leaf margins. Levels of three class 1 KNOX genes, STM, KNAT1 and KNAT2, as well as WUS and CLV1/3 are shown. Levels were determined in sorted cells expressing WUS or CLV3 (meristem-expressed) or FIL (expressed in developing organs) (see Materials and methods for details) and compared with those in 35S:miR319a/35S:amiR-NGA indeterminate leaf margins (labeled as Margins). Heatmap color represents the row Z-score. (B, C) STM:GUS and (D, E) WUS:GUS expression (blue) is confined to the vegetative shoot meristem (arrows) both in wild-type and 35S:miR319a/35S:amiR-NGA plants. (F–I) stm-11 knat6-1 bp-9, (J–L) wus-1, and (M-Q) cuc2-3 cuc3-105 seedlings have a disrupted apical meristem. Arrowheads denote fused cotyledons. In the presence of the 35S:miR319a-amiR-NGA transgene, cotyledons (G, H, I, K, N, O) and leaves (K, L, P, Q) grow indeterminately (arrows) in these mutants. Close-ups of the indeterminate margin are shown in I, L, Q. Leaves in P and Q are produced from a SAM that grows through fused cuc2-3 cuc3-105 cotyledons such as those depicted in O. cot, cotyledon. Scale bars: B-E, J, K, 5 mm; F-H, M-O, 2 mm; I, L, Q, 0.5 mm; P, 1 cm.
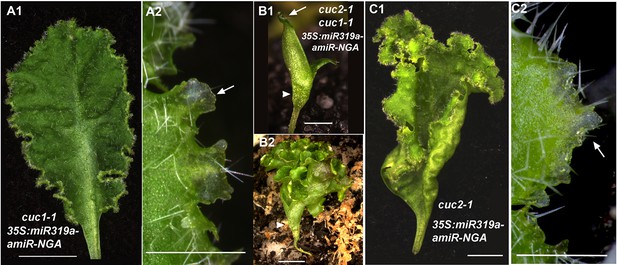

CUC activities are dispensable for indeterminate leaf margin growth.
(A1, A2) A cuc1-1 35S:miR319a-amiR-NGA leaf with indeterminate margins. (A2) Close up of cuc1-1 35S:miR319a-amiR-NGA margins shows translucent, meristematic features of marginal cells (arrow). (B1, B2) cuc2-1 cuc1-1 35S:miR319a-amiR-NGA seedling with fused cotyledons and no shoot apical meristem. Cotyledon margin shows on-going marginal growth. (B2) Older cuc2-1 cuc1-1 35S:miR319a-amiR-NGA seedling illustrating the elaborated margin due to continued marginal growth. (C1) The margins of cuc2-1 35S:miR319a-amiR-NGA leaves maintain indeterminate growth. (C2) Close-up of the indeterminate margin of the cuc2-1 35S:miR319a-amiR-NGA leaf. Initiating trichomes and translucent tissue (arrow) are apparent. Scale bars: A1, B2–C1, 5 mm; A2, B1, C2, 1 mm.
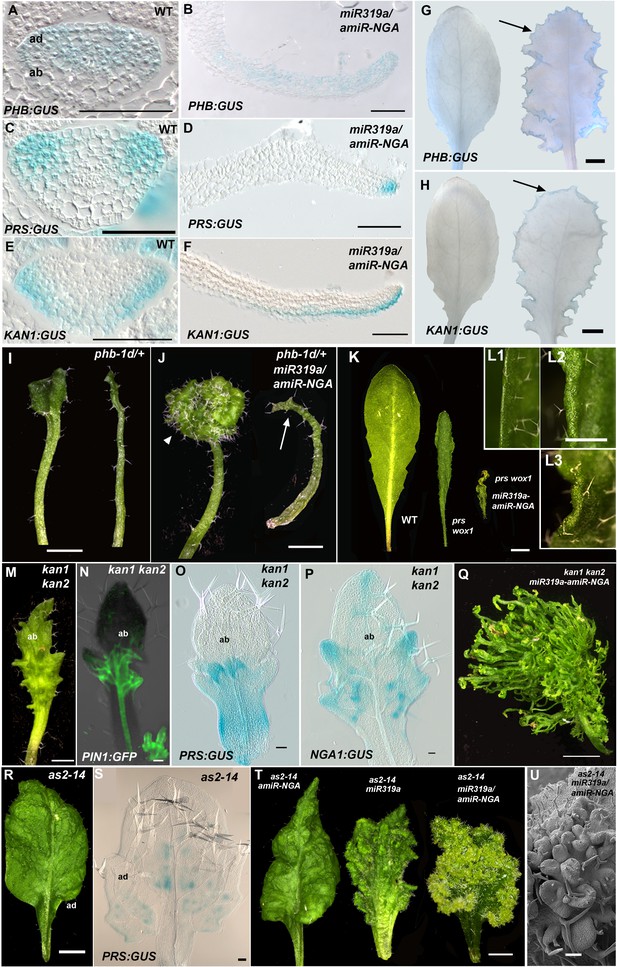
Marginal meristem activity requires juxtaposition of adaxial/abaxial polarity factors.
(A–F) Adaxial PHB:GUS (A, B), central PRS:GUS (C, D) and abaxial KAN1:GUS (E, F) expression domains (blue) in transverse sections with abaxial sides facing upward. Young wild-type leaf primordia (A, C, E) and older 35S:miR319a/35S:amiR-NGA leaf margins (B, D, F) are shown. (G–H) PHB:GUS (G) and KAN1:GUS (H) in whole wild-type (left) and 35S:miR319a/35S:amiR-NGA (right) leaves. Arrows denote continued expression. (I) Completely (right) and partially (left) radialized leaves of phb-1d/+ mutants. (J) phb-1d /+ 35S:miR319a/35S:amiR-NGA leaves. The distal lamina exhibits continual marginal growth (arrowhead) whereas a radialized leaf lacks such growth (arrow). (K) Wild-type, prs wox1, and prs wox1 35S:miR319a-amiR-NGA leaves at equivalent age. (L) Close-ups of differentiated leaf margins of wild-type (L1), prs wox1 (L2) and prs wox1 35S:miR319a-amiR-NGA (L3; compare with the indeterminate leaf margin in Figure 1C). (M–P) kan1 kan2 leaves showing abaxial outgrowths (M), and ectopic expression of PIN1:PIN1-GFP (N), PRS:GUS (O) and NGA1:GUS (P) associated with abaxial outgrowths. (Q) A kan1 kan2 35S:miR319a-amiR-NGA leaf showing proliferative tissue outgrowth. (R, S) as2-14 leaves showing ectopic, adaxial PRS:GUS expression (S). (T) From left to right shown are as2-14 35S:amiR-NGA, as2-14 35S:miR319a, as2-14 35S:miR319a/35S:amiR-NGA leaves with increasing adaxial outgrowths. (U) Close-up of the adaxial surface of as2-14 35S:miR319a/35S:amiR-NGA leaf. ad/ab, adaxial and abaxial leaf sides. Scale bars: G–J, M, 2 mm; L, 1 mm; K, Q, R, T, 5 mm; 50 μm in other panels.
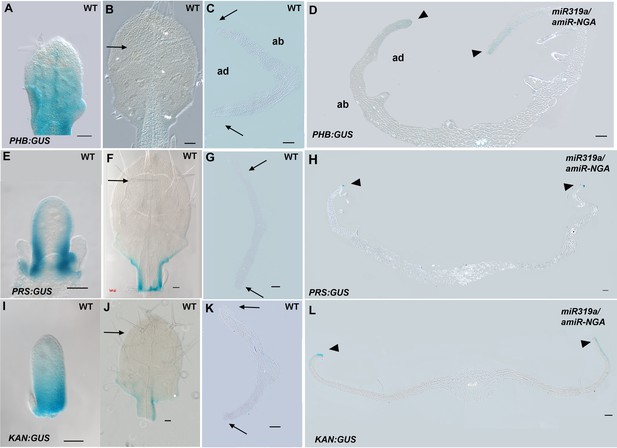
Expression of adaxial/abaxial polarity and central WOX genes is maintained at the leaf margins when CIN-TCP and NGATHA gene activities are reduced.
(A-D) PHB:GUS expression (blue) in wild-type (A–C) and 35S:miR319a/35S:amiR-NGA (D) leaves. (A) In young wild-type leaves adaxial PHB:GUS expression diminishes in the distal region. (B) In older leaves PHB:GUS expression is not apparent (arrow) except for faint expression in the base of the leaf. (C) PHB:GUS expression in a leaf of equivalent age to the one presented in (B) in transverse section. No expression is detected at the margins (arrows) or elsewhere in the leaf. (D) In older, larger 35S:miR319a/35S:amiR-NGA leaves PHB:GUS expression is detected adaxially at the margins (arrowheads; see Figure 3B for detail). (E-H) PRS:GUS expression (blue) in wild-type (E–G) and 35S:miR319a/35S:amiR-NGA (H) leaves. (E) In early wild-type leaves PRS:GUS expression starts to diminish at the distal leaf margin. (F) In older leaves PRS:GUS expression is absent from the distal region (arrow). (G) Transverse section of a leaf of equivalent age to that in (F). The margin (arrows) shows no evidence of PRS:GUS expression. (H) Transverse section of an older, larger 35S:miR319a/35S:amiR-NGA leaf shows ongoing marginal PRS:GUS expression (arrowheads; see Figure 3D for detail). (I–L) KAN1:GUS expression (blue) in wild-type (I–K) and 35S:miR319a/35S:amiR-NGA (L) leaves. (I) As young leaf primordia grow, KAN1:GUS expression diminishes distally. (J) In older leaves KAN1:GUS expression cannot be detected in the upper part of the leaf (arrow). (K) Transverse section of a leaf of equivalent age to that in (K) shows no evidence of KAN1:GUS expression at the margin (arrows). (L) Transverse section of an older, larger 35S:miR319a/35S:amiR-NGA leaf shows abaxial KAN1:GUS expression at the margins (arrowheads; see Figure 3F for detail). ad-adaxial. ab-abaxial. Scale bars: A–L, 50 μm.

Adaxial/abaxial polarity factors are necessary for and spatially define the marginal meristem, which is suppressed by CIN-TCP and NGATHA gene activities.
(A) phb-1d/+ and phb-1d /+ 35S:miR319a-amiR-NGA plants. phb-1d/+ plants show varied leaf phenotypes, and most dramatically affected leaves are completely adaxialized and radialized lacking lamina as can occur in phb-1d/phb-1d homozygotes where all lateral organs are radialized and small (left inset). In phb-1d /+ 35 S:miR319a-amiR-NGA leaves where lamina is present, marginal indeterminate growth and abaxial lamina proliferation occur (arrow; right inset). (B) Wild-type seedling showing flat cotyledons. (C) kanadi1-2 kanadi2-1 (kan1 kan2) seedling illustrating upwardly curved cotyledons with abaxial ridges. (D) kan1 kan2 35S:miR319a-amiR-NGA seedling. Note the pronounced abaxial outgrowths from the cotyledon tips (arrow). (E) Representative leaves of kan1 kan2, kan1 kan2 35S:amiR-NGA, kan1 kan2 35S:miR319a and kan1 kan2 35S:miR319a-amiR-NGA from left to right showing the progressive increase in abaxial outgrowth proliferation. (F) as2-14 (left) and as2-14 35S:miR319a/35S:amiR-NGA (right) plants of equivalent age. The cotyledons of the as2-14 35S:miR319a/35S:amiR-NGA plants have extensive adaxial outgrowths (arrow). (G) Representative leaves of as2-14 and as2-14 in combination with the transgene 35S:amiR-NGA, 35S:miR319a and 35S:miR319a/35S:amiR-NGA. as2-14 mutant leaves have a slightly uneven adaxial surface. The surface curvature phenotype is enhanced in as2-14 35S:amiR-NGA leaves. Adaxial outgrowths develop on as2-14 35S:miR319a leaves. In as2-14 35S:miR319a/35S:amiR-NGA leaves tissue proliferates from the adaxial surface and grows indeterminately. (H) Close up of the adaxial surface of as2-14 35S:miR319a/35S:amiR-NGA leaf showing proliferating lamina tissue. Scale bars: A, E, F, G, 5 mm; inset in A, B–D, H, 1 mm.
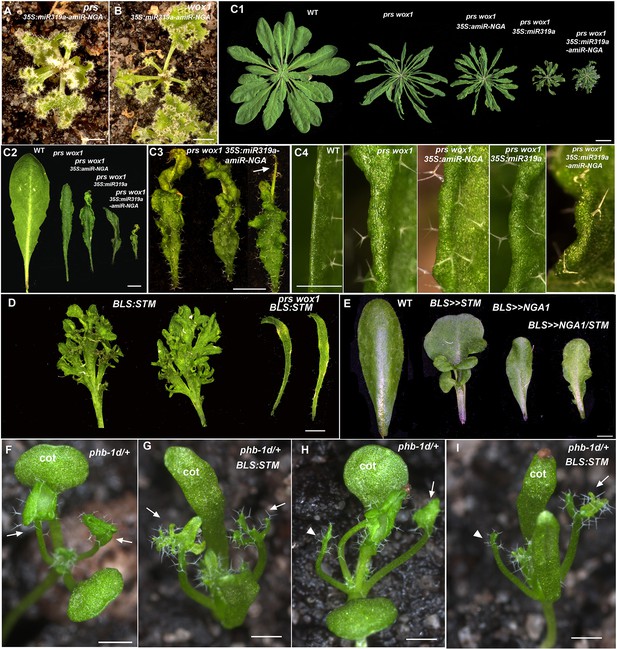

Changes in PRS/WOX1, NGATHA, or polarity factor activities affects leaf growth and morphologies.
(A, B) prs 35S:miR319a-amiR-NGA (A) and wox1 35S:miR319a-amiR-NGA (B) plants showing indeterminate leaf margins. (C1–C4) Morphology of leaves and leaf margins in prs wox1 double mutant plants with compromised NGA/CIN-TCP activities. (C1) Representative wild-type, prs wox1, prs wox1 35S:amiR-NGA, prs wox1 35S:miR319a and prs wox1 35S:miR319a-amiR-NGA plants. prs wox1 plants have narrow leaves which are increasingly curled when combined with miR319a-amiR-NGA expression. No evidence of indeterminate growth is observed at margins of the prs wox1 35S:miR319a-amiR-NGA plant. (C2) Representative leaves of wild-type, prs wox1, prs wox1 35S:amiR-NGA, prs wox1 35S:miR319a and prs wox1 35S:miR319a-amiR-NGA plants. prs wox1 is not completely epistatic to 35S:miR319a-amiR-NGA as 35S:miR319a-amiR-NGA prs wox1 leaves are of smaller size, darker green and epinastic. However, continued leaf marginal growth is absent in prs wox1 35S:miR319a-amiR-NGA leaves. (C3) Close up of representative prs wox1 35S:miR319a-amiR-NGA leaves showing the curling phenotype and the lack of continued marginal growth. Some leaves are very thin at their distal region (arrow). (C4) The morphology of margins of wild-type, prs wox1, prs wox1 35S:amiR-NGA, prs wox1 35S:miR319a and prs wox1 35S:miR319a-amiR-NGA leaves. All leaf margins in the prs wox1 background appear similar lacking ongoing proliferation. (D) The prs wox1 background suppresses the dissected leaf phenotype caused by ectopic expression of a class 1 KNOX gene, SHOOT MERISTEMLESS (STM). Shown are two representative leaves detached from BLS:STM and prs wox1 BLS:STM plants at an equivalent age. Leaves of BLS:STM plants are highly dissected. In contrast, leaves are small and simple in prs wox1 BLS:STM plants. Plants shown here are in the Columbia background. (E) Leaf-specific expression of NGA1 (BLS>>NGA1) suppresses the leaf complexity phenotype caused by ectopic STM expression (BLS>>STM). Concomitant expression of NGA1 and STM by the BLS:LhG4 driver in the transactivation system (BLS>>NGA1/ STM) results in simple leaves. Plants used in this analysis are in the Ler background (Shani et al., 2009). (F-I) The radialized leaf phenotype of phb-1d/+ suppresses the dissected leaf phenotype caused by ectopic expression of STM. (F) A phb-1d/+ seedling where the first two leaves have lamina (arrows). (G) A phb-1d/+ BLS:STM seedling where the first leaves exhibit elaboration (arrows). (H) A phb-1d/+ seedling where one of the first two leaves has lamina (arrow) while the other is radialized (arrowhead). (I) A phb-1d/+ BLS:STM seedling where one of the first two leaves exhibits elaboration (arrow) while the other is radialized (arrowhead). cot, cotyledon. All plants and leaves are shown with the adaxial side facing up. Scale bars: A, B, C3, D, E, 2 mm; C1, 1 cm; C2, 5 mm; C4, F–I, 1 mm.
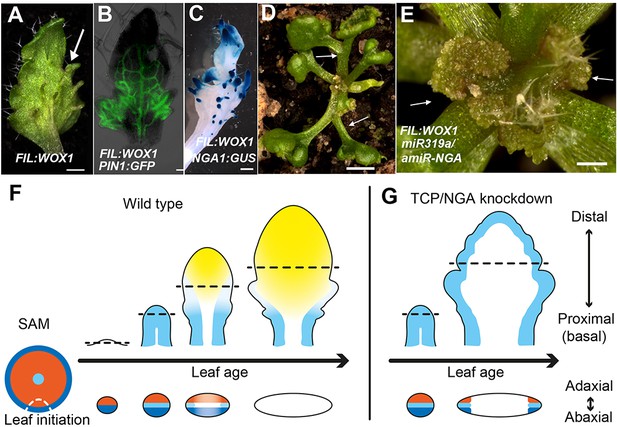
Dynamic restriction of the leaf meristem.
(A–C) FIL:WOX1 leaf with developing abaxial outgrowths (arrow in A). These outgrowths show prolonged PIN1:PIN1-GFP (B) and NGA1:GUS (C) expression. (D, E) In FIL:WOX1 35S:miR319a/35S:amiR-NGA first leaves show occasional bifurcation (arrows in D) and later emerging leaves are highly proliferative (arrows in E) in the distal domain. (F, G) Scheme depicting developing wild-type (F) and CIN-TCP/NGA compromised (G) leaves shown from a proximo-distal perspective (above horizontal arrows) and abaxial-adaxial perspective captured at the dashed-line (below horizontal arrows). Leaves are physically and evolutionarily derived from shoot apical meristems (SAM; left most cartoon in F). The SAM is radially patterned with external (blue), internal (red) and central WOX (aqua) domains. In wild-type leaf primordia (F) the pre-pattern at the SAM (white dash line) is converted into juxtaposed abaxial and adaxial domains directing WOX activation in an intervening domain. Feedback between these three domains stabilizes the leaf meristem, promotes lamina growth and maintains pluripotency (Nakata and Okada, 2012) before meristem activity is restricted to the proximal marginal domains by CIN-TCP/NGA activities (yellow), permitting prolonged growth only at the proximal region of the leaf. In leaves where CIN-TCP/NGA activities are reduced (G), meristem activity is maintained at all margins in a pattern reminiscent of initiating leaf primordia. Scale bars: A, C, D, 2 mm; E, 1 mm; 50 μm in B.
Additional files
-
Supplementary file 1
Primers used for PCR-mediated cloning.
- https://doi.org/10.7554/eLife.15023.030





