Viral RNA switch mediates the dynamic control of flavivirus replicase recruitment by genome cyclization
Figures
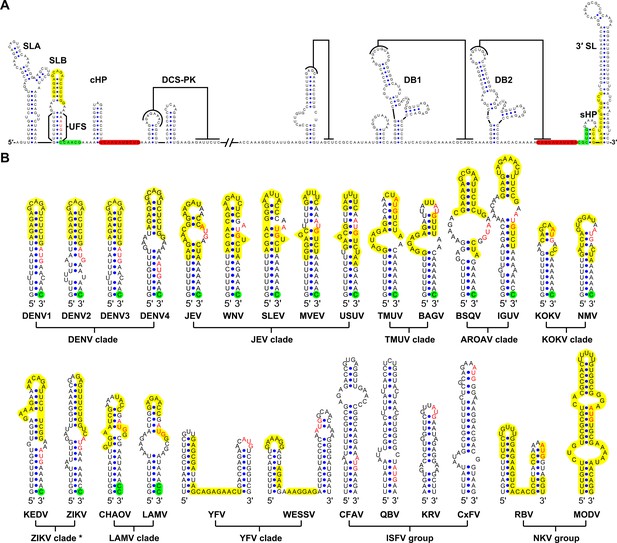
Identification and comparison of the 5′ UAR-UFS elements among flaviviruses.
(A) Terminal RNA structures of the DENV4 genome. The UAR, DAR and CS elements are highlighted in yellow, green and red, respectively. Pseudoknotted interactions are labeled. The UFS stem region is indicated by parentheses. (B) Comparison of the 5′ UAR-UFS and UFS-like elements in MBFVs, NKVs and ISFVs. *: KEDV and ZIKV were supposed to belong to the same clade herein. The MBFV 5′ UAR sequences and NKV sequences involved in genome cyclization are colored in yellow. The nucleotides that participate in the DAR interactions of MBFVs are labeled in green. Translational start codons in (A) and (B) are shown in red. Virus name abbreviations are annotated in Figure 1—figure supplement 2.
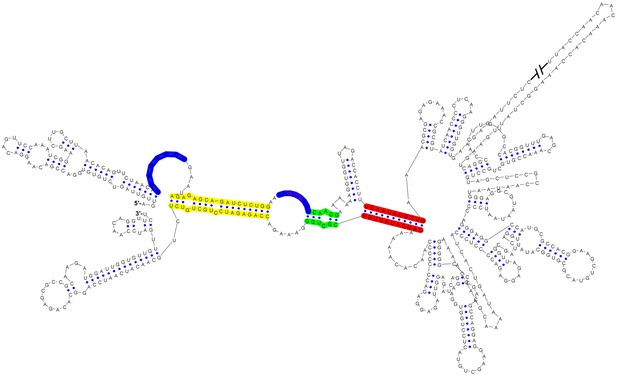
Circular conformation of the DENV4 genomic RNA.
The UAR, DAR and CS elements are shown in yellow, green and red, respectively. The sequence of the unwound UFS duplex is shown in blue.
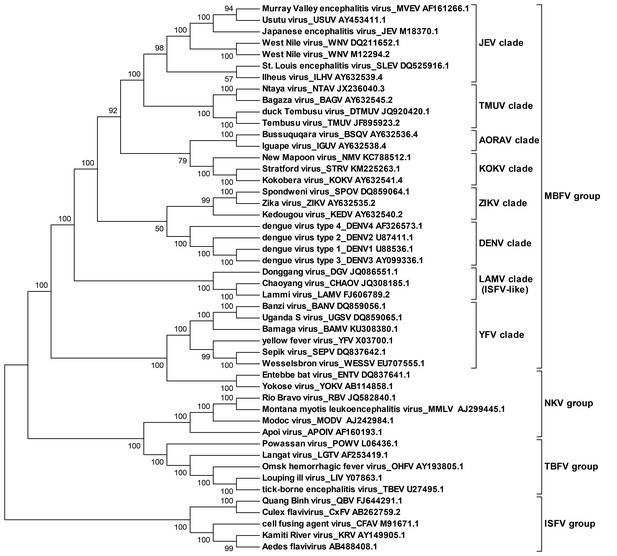
Molecular phylogenetic analysis of the flavivirus genus.
The ORF sequences of 48 flavivirus species were obtained from Genbank and aligned by the Clustal W method. The evolutionary relationships of flaviviruses were inferred by using the Maximum Likelihood method based on the Tamura-Nei model. Analyses were performed with MEGA6. Different evolutionary groups as well as various clades among the MBFV group are annotated.
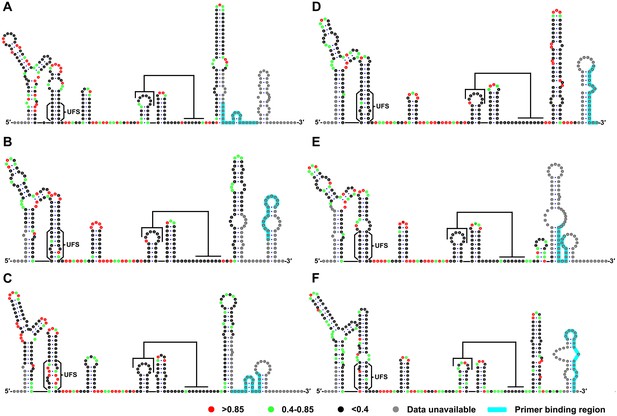
SHAPE analysis of the 5′ end RNA of representative flaviviruses.
The SHAPE reactivity results are labeled in the structure models of different flaviviruses. Highly reactive nucleotides (SHAPE reactivity > 0.85) are labeled in red, whereas nucleotides with moderate SHAPE reactivity (0.4–0.85) are labeled in green. Black-colored nucleotides indicate low or no SHAPE reactivity (<0.4). Nucleotides lacking SHAPE data are labeled in gray. The primer-binding region in the reverse transcription reaction is shown in light blue. (A) JEV, strain SA-14-14-2 (AF315119). (B) DENV1; the sequence is based on strain WestPac (U88535). (C) DENV2; the sequence is based on strain NGC (KM204118). (D) DENV3; the sequence is based on strain 80–2 (AF317645). (E) DENV4; strain 814669 (AF326573), with two artificial nonsense mutations that do not affect the 5′ end secondary structures or vRNA replication. The two mutations, located in the loop region of cHP and DCS-PK Loop 3, respectively, are labeled in black-colored font. (F) ZIKV; strain FSS13025 (KU955593). Results presented were from two biological replicates.
-
Figure 2—source data 1
Source data for Figure 2.
- https://doi.org/10.7554/eLife.17636.007
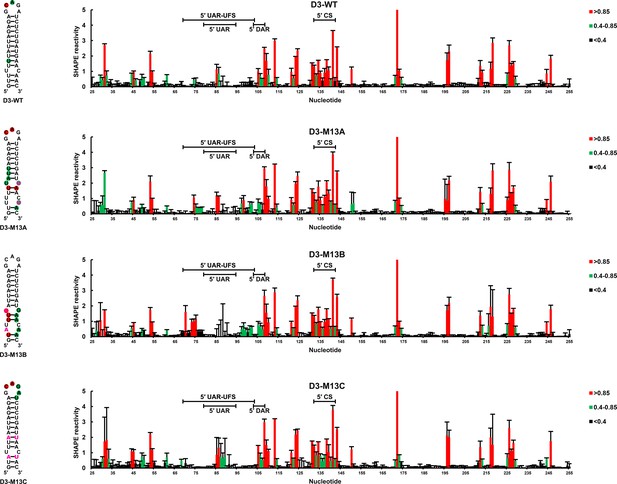
SHAPE analysis of the DENV3 UFS mutants.
SHAPE analysis of DENV3 5′-300 nt RNA molecules containing UFS mutations. the SHAPE result of WT 5′-300 nt RNA is shown in parallel. SHAPE diagrams of 5′ 25–255 nt are shown, and the regions corresponding to various 5′ elements are indicated. Negative SHAPE reactivity is set to zero. The SHAPE reactivity of the 5′ UAR-UFS region is annotated on the corresponding secondary structures. Nucleotides with SHAPE reactivity greater than 0.85 are labeled in red, moderately reactive sites (0.4–0.85) are labeled in green. Unlabeled sites have low or no SHAPE reactivity (<0.4). Two biological replicates were performed for each RNA.
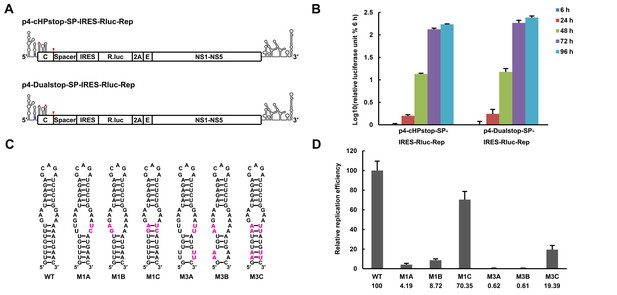
The secondary structure of the UFS is required for flavivirus vRNA replication.
(A) The organization of the DENV4 replicon constructs. In p4-cHPstop-SP-IRES-Rluc-Rep, the translation of viral nonstructural proteins is controlled by the EMCV IRES, and artificial stop codons (red inverted triangles) were introduced into the cHP loop region as well as the end of the capsid-coding region to abolish translation directed by the 5′ cap structure. In the improved version, p4-Dualstop-SP-IRES-Rluc-Rep, one additional stop codon was introduced into loop 3 of the DCS-PK element to abolish the potential translational start from a cryptic AUG in the 5′ CS element. (B) Replication characteristics of the above two replicons in BHK-21 cells. Relative luciferase units are defined as the ratio of luciferase units measured at different time points after transfection to the value measured at 6 hr post-transfection. (C) Demonstration of DENV4 UFS mutants. Mutations are shown in purple. The 'A' mutants contain mutations in the downstream strand of the UFS, and the 'B' mutants contain mutations in the upstream strand of the UFS. The 'C' mutant in each group combines the 'A' and 'B' mutations to restore the secondary structure of the UFS. Note that the secondary structures shown here are for illustration only and do not represent the actual folding of these mutants. (D) Replication efficiency of the UFS mutants. The results of one of two independent biological replicates were shown. One hundred nanograms per well of replicon RNA species were transfected into BHK-21 cells in triplicate. The results are expressed as the percentage of the relative luciferase units of a UFS mutant at 72 hr post-transfection to the value of the WT replicon (relative replication efficiency). The error bar represents the standard deviation in all figures. The mean values of relative replication efficiency are listed below the names of the replicon constructs.
-
Figure 3—source data 1
Source data for 3B, 3D and Figure 3—figure supplement 1.
- https://doi.org/10.7554/eLife.17636.010
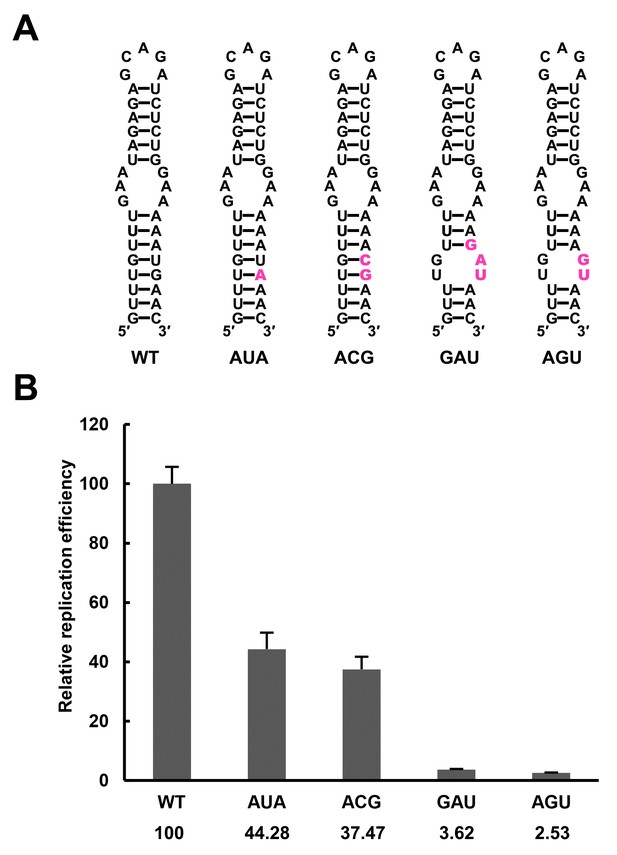
Replication of DENV4 UFS mutants targeting the AUG region.
(A) Demonstration of mutants targeting the AUG start codon. Mutation sites are shown in purple. These mutants were designed without affecting either genome cyclization or the terminal topology of circularized genome. For detailed information, please refer to Supplementary file 3. (B) Relative replication efficiency of UFS mutants shown above at 48 hr post-transfection. Results shown were obtained from three parallel technical replicates.
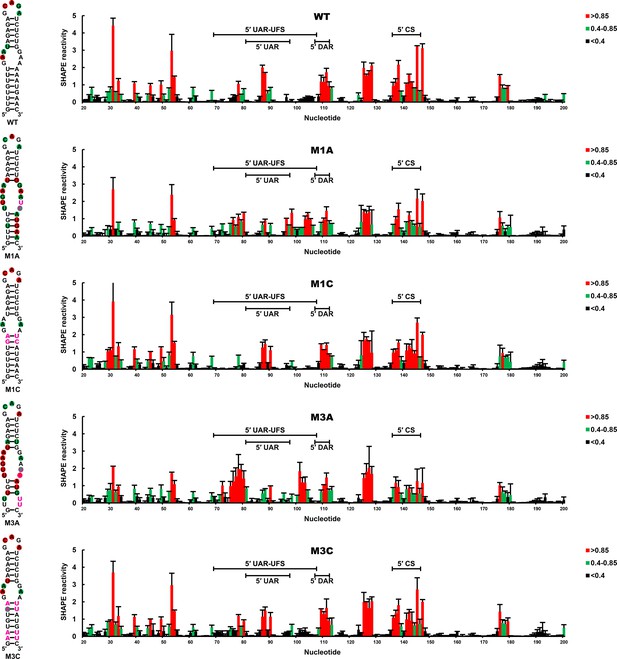
SHAPE analysis of the DENV4 UFS mutants.
SHAPE analysis was performed for DENV4 5′-300 nt RNA containing the UFS mutations M1A, M1C, M3A or M3C. The SHAPE result for WT 5′ end RNA is shown in parallel. SHAPE diagrams of 5′ 20–200 nt are shown, and the regions corresponding to various 5′ elements are indicated. Negative SHAPE reactivity is set to zero. The SHAPE reactivity of the 5′ UAR-UFS region is annotated on the corresponding secondary structures (left). Nucleotides with SHAPE reactivity greater than 0.85 are labeled in red, and moderately reactive sites (0.4–0.85) are labeled in green. Unlabeled sites have low or no SHAPE reactivity (<0.4). Two biological replicates were performed for each RNA, and the error bars represent the standard deviation.
-
Figure 4—source data 1
Source data for Figure 4.
- https://doi.org/10.7554/eLife.17636.013
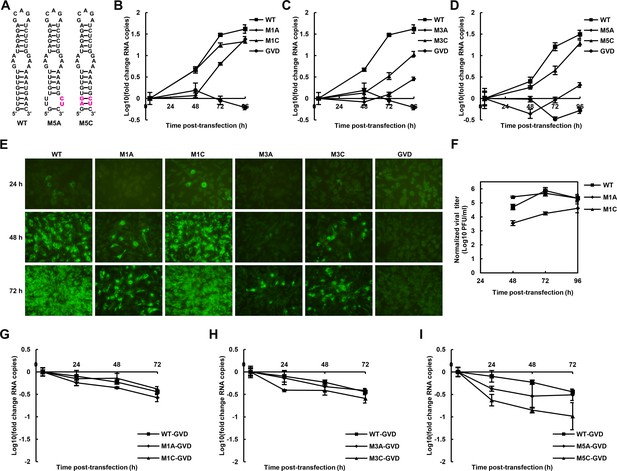
The role of the UFS in viral propagation.
(A) DENV4 UFS M5 series mutants. Mutations are shown in purple. (B–D) vRNA replication profiles of different UFS mutants in transfected BHK-21 cells. WT and NS5-inactive GVD mutant vRNA, measured in parallel, are shown as controls. (B) M1 series, (C) M3 series and (D) M5 series. (E) Indirect immunofluorescence assay of WT, GVD, and UFS-mutated vRNA-transfected BHK-21 cells. (F) Determination of the virus titers in the supernatants of the M1 series mutant-transfected BHK-21 cells. The virus titers in the supernatants of WT vRNA-transfected cells were determined in parallel. The virus titers were normalized according to input vRNA copies determined at 4 hr post-electroporation. (G–I) The RNA stability of the UFS mutants was determined by qRT-PCR and are expressed as the fold change relative to the vRNA copies at 4 hr post-transfection. (G) M1 series, (H) M3 series and (I) M5 series. The results were from one biological replicate. Experiments to determine vRNA copies and virus titers were performed in triplicate. Note that the time point '72 hr' for the GVD group in (D) contains data from only two parallel wells (technical replicates), because the result of the third sample was unreliable due to an error in the RNA isolation process.
-
Figure 5—source data 1
Source data for 5B and 5C.
- https://doi.org/10.7554/eLife.17636.015
-
Figure 5—source data 2
Source data for 5D
- https://doi.org/10.7554/eLife.17636.016
-
Figure 5—source data 3
Source data for 5F
- https://doi.org/10.7554/eLife.17636.017
-
Figure 5—source data 4
Source data for 5G, 5H and 5I.
- https://doi.org/10.7554/eLife.17636.018
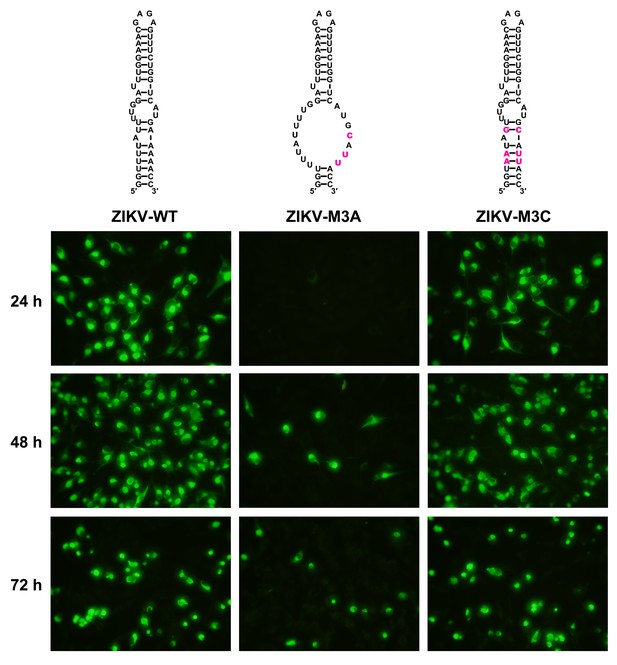
Replication of ZIKV UFS mutants in BHK-21 cells.
Indirect immunofluorescence assay of WT and UFS-mutated ZIKV vRNA-transfected BHK-21 cells were performed. The secondary structures of mutants were demonstrated and mutation sites are shown in purple.
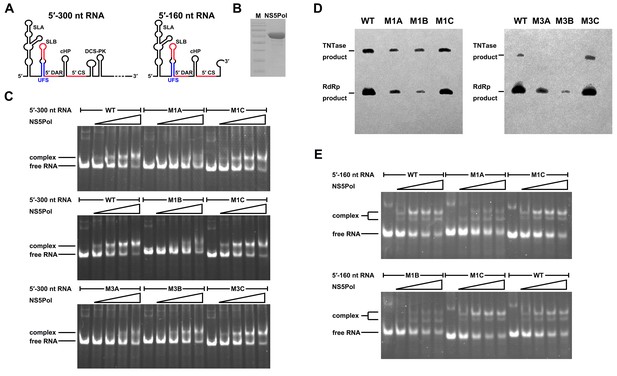
UFS is crucial for RdRp recruitment and de novo RNA synthesis.
(A) Simplified diagrams of DENV4 virus 5′ end RNA constructs used for RdRp binding and/or in vitro RdRp activity assays. The region corresponding to the UFS element is shown in blue, and the red regions represent the genome cyclization elements. (B) SDS-PAGE of purified recombinant NS5Pol of DENV4. (C) The binding of DENV4 NS5Pol to 5′-300 nt RNA molecules containing UFS mutations was analyzed by EMSA. No NS5Pol was present in the left first lane of each group. The NS5Pol concentrations in the reactions were approximately 2.0, 3.5, 5.0 and 7.5 μM (from left to right). (D) In vitro RdRp activity assay using 5′-160 nt RNA as a template. The reactions were incubated at 30°C for 20 min. Detection was based on the incorporation of biotin-11-CTP into the products. Left, the results of the M1 series; right, the results of the M3 series. Two bands were detectable in the blot: the upper template-sized band was generated by the terminal addition of biotin-11-CTP to the input template due to the terminal nucleotide transferase activity of NS5Pol in the presence of Mn2+, and the lower band represented the dsRNA product of de novo RNA synthesis, which was confirmed by comparison of the electrophoretic patterns of the RdRp products and annealed dsRNA molecules (Figure 6—figure supplement 2). (E) EMSA was performed for the 5′-160 nt RNA molecules containing M1 series mutations. The NS5Pol concentrations in the reactions were approximately 3.6, 7.2, 10.7 and 14.3 μM (left to right).
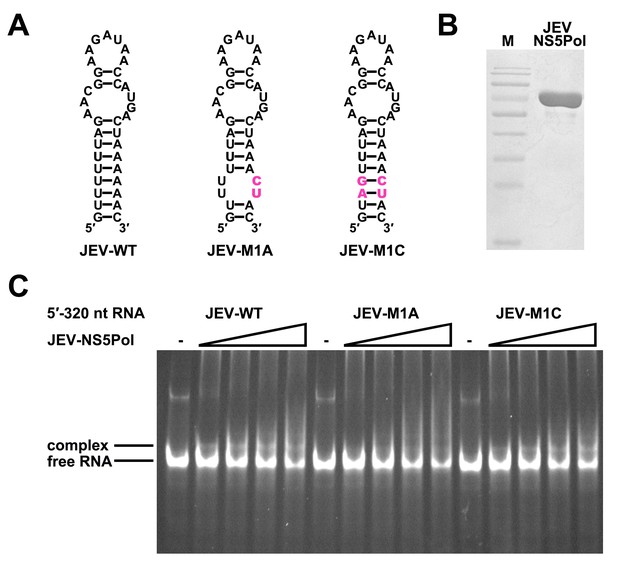
The UFS is required for efficient NS5 binding to 5′ RNA in JEV.
(A) JEV UFS mutants. Mutations are shown in purple. (B) SDS-PAGE of purified recombinant JEV NS5Pol. (C) The binding of JEV NS5Pol to 5′-320 nt RNA molecules containing UFS mutations was analyzed by EMSA. No NS5Pol was present in the left first lane of each group. The NS5Pol concentrations in the reactions were approximately 3.3, 5.0, 6.6 and 7.5 μM (from left to right).
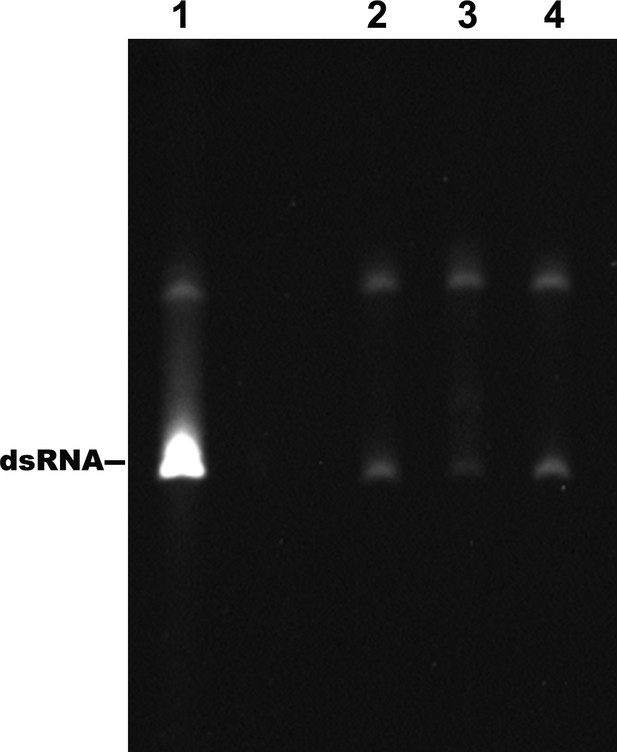
Identification of the products of the RdRp reactions.
Lane 1: dsRNA prepared by the annealing of the positive and negative strand of DENV4 5′-160 nt RNA. Lane 2–4: products of RdRp assays using different 5′-160 nt RNAs as templates. Lane 2: WT, Lane 3: M2A, Lane 4: M2C.
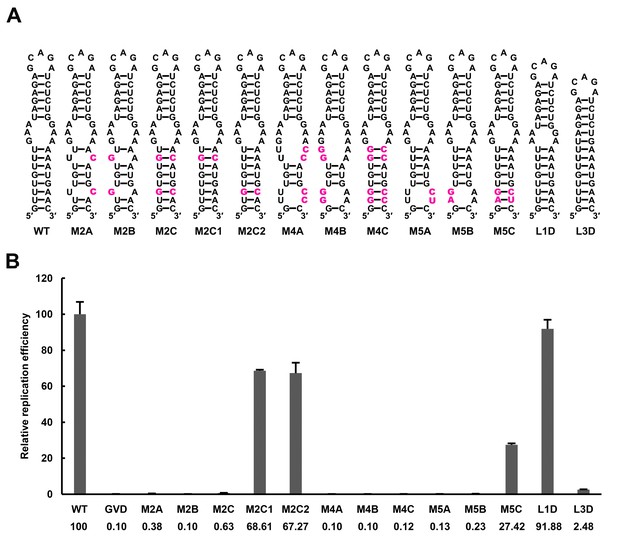
Effect of UFS stability on vRNA replication.
(A) Demonstration of UFS mutants. The mutations are indicated in purple. The M2C mutant contained two UA-to-GC base pair substitutions, whereas the M4C mutant contained four substitutions in the UFS secondary structure. It should be noted that the secondary structures are meant for illustration purposes only, and the structures displayed for UFS-disrupted mutants are not the most thermodynamically favorable ones. (B) Relative replication efficiency of UFS mutants shown in (A) at 72 hr post-transfection. The results of one of two independent biological replicates were shown. Fifty nanograms per well of replicon RNA were transfected into 96-well plates. Experiments were performed in triplicate.
-
Figure 7—source data 1
Source data for Figure 7 and Figure 7—figure supplement 1.
- https://doi.org/10.7554/eLife.17636.024
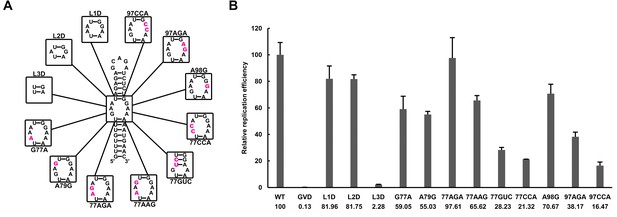
Effects of mutations in the SLB-UFS internal loop on vRNA replication.
(A) Demonstration of mutants targeting the internal loop between SLB and UFS. Mutation sites are shown in purple. (B) Relative replication efficiency of UFS mutants shown above at 72 hr post-transfection. Results shown were obtained from two parallel technical replicates.
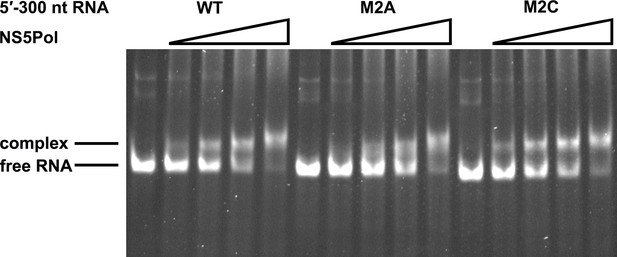
NS5Pol binding assay of the DENV4 5′-300 nt M2A and M2C mutants.
The binding of NS5Pol to 5′-300 nt WT, M2A and M2C RNA was analyzed by EMSA assay. The left first lane in each group contained no NS5Pol. The NS5Pol concentrations in the reactions were approximately 2.0, 3.5, 5.0 and 7.5 μM (from left to right).
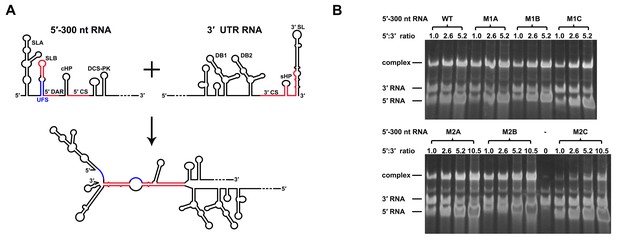
Increasing the stability of the UFS hinders vRNA cyclization.
(A) Schematic diagram of experimental design. As 5′-300 nt RNA is incubated with 3′ UTR RNA, a 5′-3′ RNA bimolecular complex is formed due to the interactions that promote genome cyclization. Various RNA structural elements in the DENV4 genomic ends are shown. The regions involved in terminal interactions are colored in red, and the UFS element is labeled in blue. (B) RNA binding assay of the UFS mutants. 3′ UTR RNA (20 ng/μl) was incubated with different amounts (16, 40, 80 and 160 ng/μl) of various 5′-300 nt RNA mutants in 20 μl reactions. The formation of RNA complexes was then analyzed by native TBE PAGE gels. A gel image of the M1 series mutants is shown on the top, and the results for the M2 series mutants are shown below. The molar ratios between 5′ and 3′ RNA are indicated above the gel lanes.
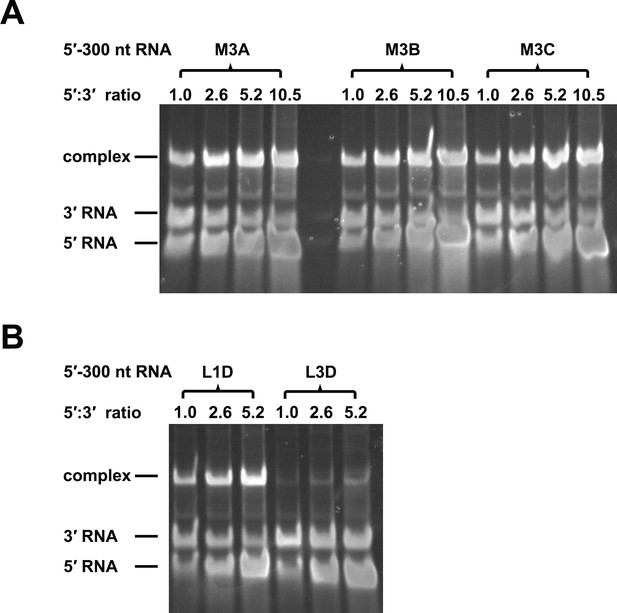
RNA binding assay of the UFS L1D, L3D and M3 series mutants.
3′ UTR RNA (20 ng/μl) was incubated with different amount (16, 40, 80 and 160 ng/μl) of the corresponding 5′-300 nt RNA mutants in 20 μl reactions. The formation of RNA complexes was then analyzed by native TBE PAGE gels. The molar ratios between 5′ and 3′ RNA are indicated above the gel lanes.
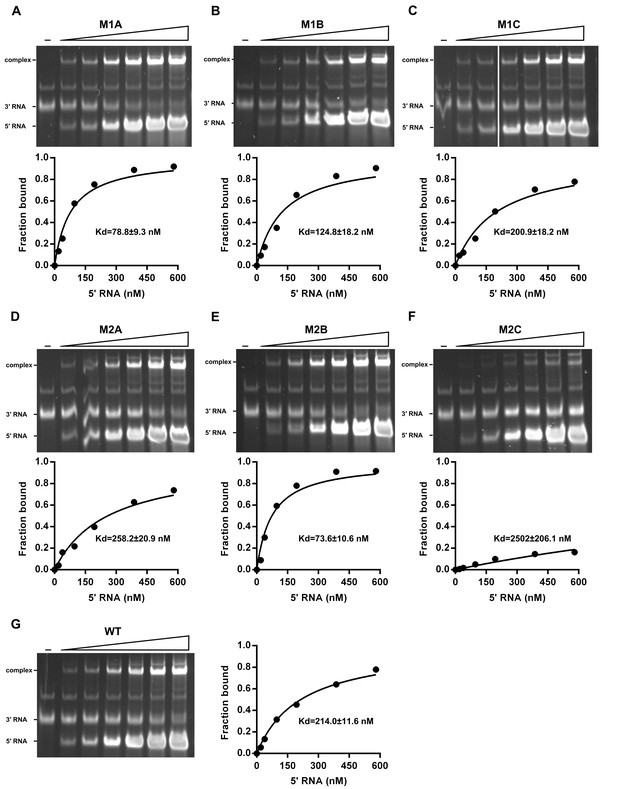
The calculation of binding affinity of 3′ UTR for the 5′-300 nt RNAs.
Reactions containing a series of concentrations of different 5′-300 nt RNAs and fixed concentration (approximately 36.9 nM) of 3′ UTR RNA were resolved in native PAGE gels. The gels were analyzed by ImageJ software and the percentages of bound 3′ UTR RNA were calculated. We assumed that the band intensity has a positive linear correlation with the molecular weights of the corresponding molecules, and that was taken into account in the calculation. The plotted data was fitted with % Bound=[X]/(Kd+[X]) using GraphPad Prism 6 (GraphPad Software, La Jolla, California, USA), where % Bound is the percentage of bound 3′ UTR RNA, and [X] is the concentration of 5′-300 nt RNA. By this method, the Kd was estimated and showed along with the binding curve.
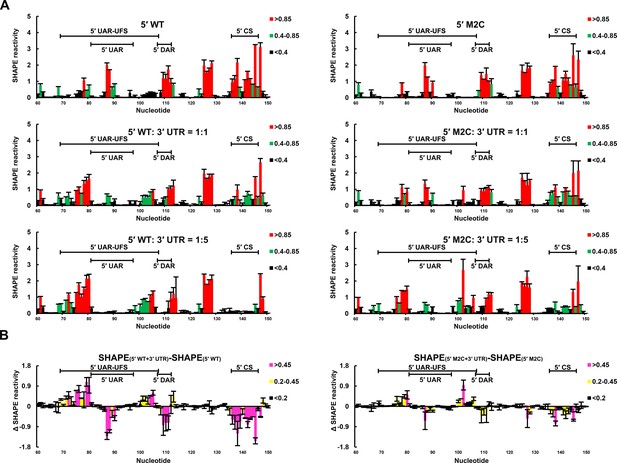
SHAPE analysis of the interactions between DENV4 genomic ends.
(A) SHAPE analysis was performed for DENV4 5′-300 nt WT and M2C RNA in the presence of various amounts of 3′ UTR RNA. For convenience, only the results for nucleotides 60–150, which include critical elements for genome cyclization, are shown. The regions corresponding to various 5′ elements are indicated. Negative SHAPE reactivity is set to zero for clear demonstration. (B) The changes in the SHAPE reactivity of the 5′ 60–150 region were obtained by subtracting the SHAPE reactivity when 3′ UTR RNA was absent from the corresponding values when 3′ UTR RNA was present in a 1:1 ratio with 5′-300 nt RNA. The results were from two biological replicates, and the error bars represent the standard deviation.
-
Figure 9—source data 1
Source data for Figure 9.
- https://doi.org/10.7554/eLife.17636.031
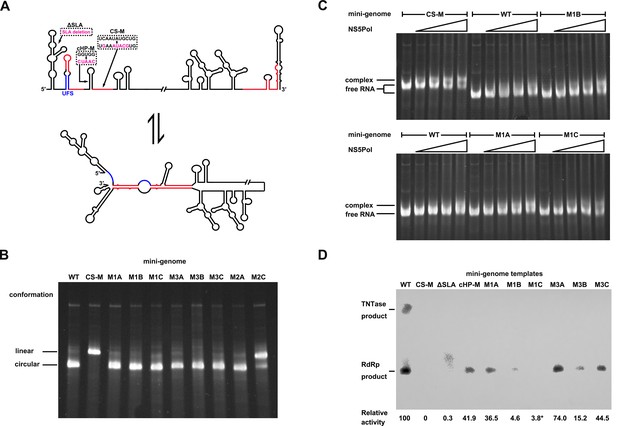
Genome cyclization disables the function of the UFS in NS5 recruitment.
(A) Schematic diagram showing the conformational changes of the DENV4 genome. Two major conformations of vRNA, the linear and circular form, exist in equilibrium. The regions involved in terminal interactions are colored in red, and the UFS element is labeled in blue. The locations and sequences of the ΔSLA, cHP-M and CS-M mutants used in this section are indicated. (B) The conformational equilibrium of mini-genome RNA containing different mutations was analyzed using native PAGE. The bands of the linear and circular conformations were identified using the CS-M mutant as a control. The faint and much larger bands were likely to be formed by misfolded RNA molecules. (C) The binding of NS5Pol to different mini-genome RNA molecules was analyzed by EMSA. Mini-genomes containing the M1 series mutations in the UFS were assayed, and the WT and CS-M mini-genomes served as control reactions. No NS5Pol was present in the left first lane of each group. The NS5Pol concentrations in the reactions were approximately 2.0, 3.5, 5.0 and 7.5 μM (from left to right). (D) In vitro RdRp activity assay using various mini-genome RNA molecules as templates. The reactions were performed at 30°C for 40 min. Note that the signal of the ΔSLA lane did not correspond to the dsRNA product generated by de novo initiation. The blot was analyzed by ImageJ software and RdRp activity was expressed as percentage of WT. Note that the template activity of M1C-mini was calculated by analyzing a longer exposed version of the same blot shown.
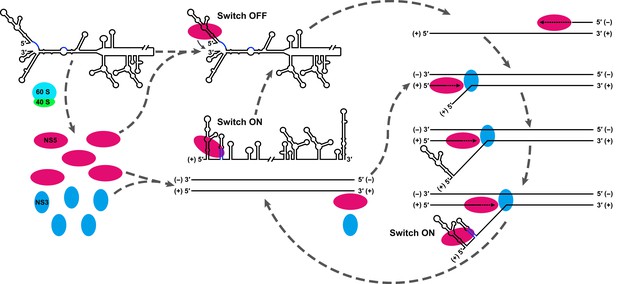
Model explaining the functional mechanism of the UFS switch.
A proposed mechanistic model of flavivirus vRNA replication. (i) After the viral genome is released into the cytoplasm, first, translation occurs on the circularized genome to generate a sufficient level of viral proteins for downstream vRNA synthesis. (ii) Accumulation of viral replication proteins (only NS5 and NS3 are shown) to high concentrations results in the formation of an RNA-viral replicase complex and initiates (−) RNA synthesis to generate the double-stranded replication form (RF). (iii) Then, the synthesis of nascent (+) RNA using RF as the template is initiated by NS5/viral replicase. During the process, 5′ local structures including UFS are formed in the displaced, parental (+) strand RNA. A free NS5/viral replicase is recruited to bind the 5′ end of the displaced (+) RNA. (iv) As soon as the nascent (+) strand RNA synthesis completes, the released (+) ssRNA genome is circularized by terminal interactions, and the next round of (−) strand synthesis is ready to start.
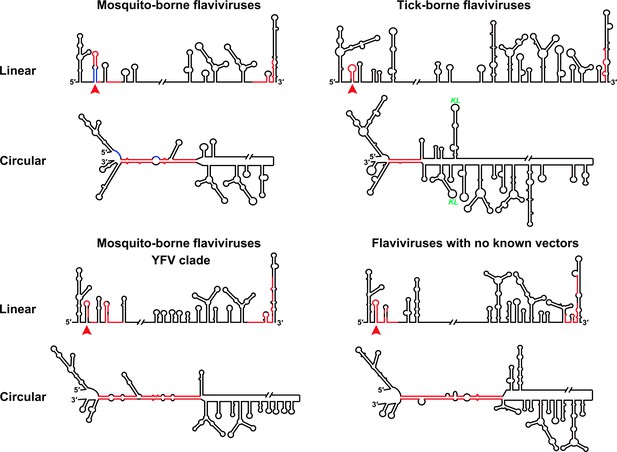
Genome conformation models of major groups of flaviviruses.
Sequences involved in terminal interactions are shown in red. The 5′ RNA structures that are immediately downstream of SLA elements and consistently involved in genome cyclization are indicated by red arrows. The typical UFS element in MBFVs is shown in blue. The demonstrated structures are based on DENV4 for the MBFVs; tick-borne encephalitis virus (TBEV) for TBFVs; Wesselsbron virus (WESSV) for the YFV clade of the MBFVs and RBV for the NKVs. The kissing loop interaction identified recently in the tick-borne group (Tsetsarkin et al., 2016) is labeled by green letters (KL).
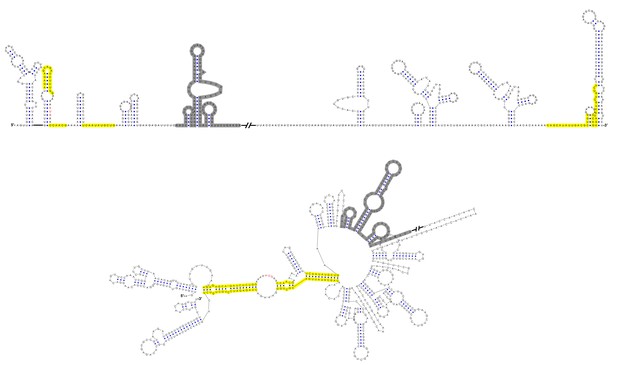
Terminal genomic RNA structures of the MBFV group.
The demonstrated sequence was based on DENV4. Sequences involved in genome cyclization are highlighted in yellow. The gray regions are not included in the demonstration of Figure 12. The mfold software and VARNA application were utilized to generate the structural models.
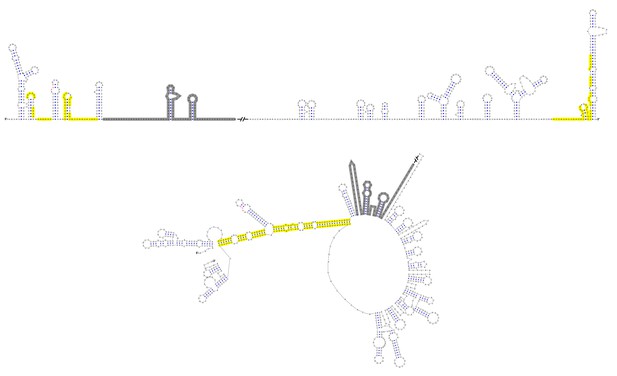
Terminal genomic RNA structures of the YFV clade in the MBFV group.
The demonstrated sequence was based on WESSV. Sequences involved in genome cyclization are highlighted in yellow. The gray regions are not included in the demonstration of Figure 12. The mfold software and VARNA application were utilized to generate the structural models.
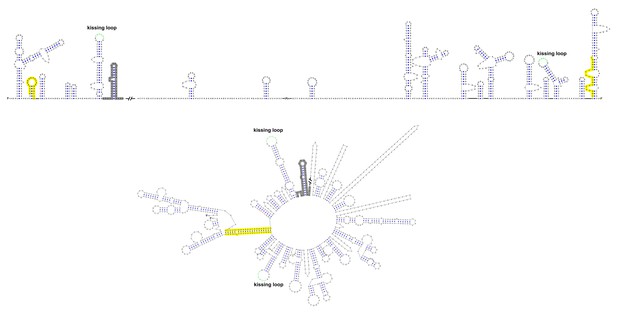
Terminal genomic RNA structures of the TBFV group.
The demonstrated sequence was based on TBEV. Sequences involved in genome cyclization are highlighted in yellow. The gray regions are not included in the demonstration of Figure 12. The mfold software and VARNA application were utilized to generate the structural models.
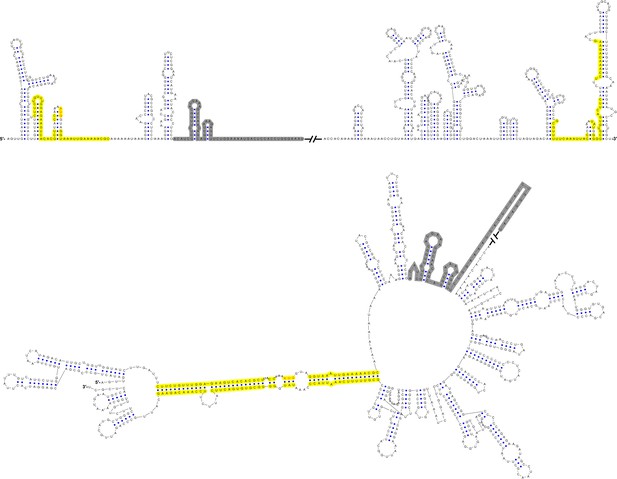
Terminal genomic RNA structures of the NKV group.
The demonstrated sequence was based on RBV. Sequences involved in genome cyclization are highlighted in yellow. The gray regions are not included in the demonstration of Figure 12. The mfold software and VARNA application were utilized to generate the structural models.
Additional files
-
Supplementary file 1
Mfold prediction for the terminal RNA structures of different flaviviruses.
- https://doi.org/10.7554/eLife.17636.039
-
Supplementary file 2
Mfold prediction for the 5′ end RNA structures of yokose virus clade of NKV group.
- https://doi.org/10.7554/eLife.17636.040
-
Supplementary file 3
Mfold prediction for analysis of the influence of DENV4 UFS mutations on the overall genome terminal RNA structures.
- https://doi.org/10.7554/eLife.17636.041
-
Supplementary file 4
Structure models of DENV4 5′ WT, M1A, M1C, M3A and M3C RNA generated by RNAstructure software using SHAPE constraints.
- https://doi.org/10.7554/eLife.17636.042





