Live-cell single-molecule tracking reveals co-recognition of H3K27me3 and DNA targets polycomb Cbx7-PRC1 to chromatin
Figures
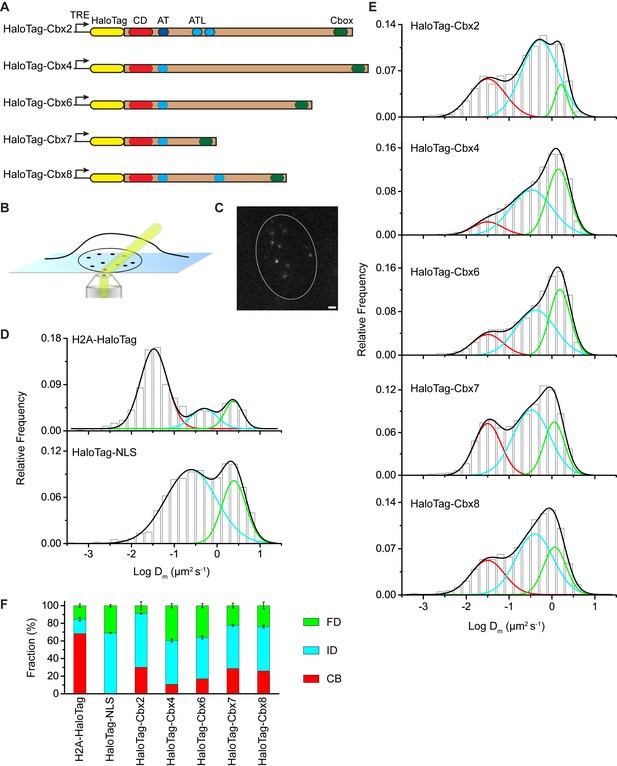
The Cbx family members exhibit distinct dynamics in living mES cells.
(A) The sequences encoding the five Cbx proteins were fused with HaloTag to generate the HaloTag-Cbx fusions that were stably expressed in wild-type (PGK12.1) mES cells. The expression level of HaloTag-Cbx fusions was controlled by Tet-responsive element (TRE). HaloTag is shown in yellow, CD (chromodomain) in red, AT (AT-hook) motif in light blue; ATL (AT-hook-like) motif in cyan, and Cbox (chromobox) in emerald. (B) Schematic representation of highly inclined and laminated optical sheet (HILO) microscopy. (C) Live-cell single-molecule visualization of HaloTag-Cbx7 molecules in mES cells during a 30-ms exposure. Oval white dash circle outlines the nucleus of the cell. The individual white points represent single HaloTag-Cbx7 molecules. Scale bar, 2 µm. (D) Normalized histograms of the log maximum likelihood diffusion coefficient for H2A-HaloTag (N = 19 cells, n = 2675 trajectories) and HaloTag-NLS (N = 69 cells, n = 2087 trajectories) in wild-type mES cells. The H2A-HaloTag histogram was fitted with a three-component Gaussian and the HaloTag-NLS histogram a two-component Gaussian. The color bars indicate that the fraction of proteins in the chromatin-bound (CB, red), intermediate (ID, cyan), and fast diffusion (FD, green) population. NLS, nuclear localization sequence. (E) Normalized histograms of the log maximum likelihood diffusion coefficient for HaloTag-Cbx2 (N = 44 cells, n = 2833 trajectories), HaloTag-Cbx4 (N = 34 cells, n = 11,343 trajectories), HaloTag-Cbx6 (N = 33 cells, n = 7457 trajectories), HaloTag-Cbx7 (N = 51 cells, n = 3097 trajectories), and HaloTag-Cbx8 (N = 36 cells, n = 3351 trajectories) in wild-type mES cells. The histograms were fitted with a three-component Gaussian. (F) Fraction of the CB (red), ID (cyan), and FD (green) population for H2A-HaloTag, HaloTag-NLS, HaloTag-Cbx2, HaloTag-Cbx4, HaloTag-Cbx6, HaloTag-Cbx7, and HaloTag-Cbx8. The data were obtained from Figure 1D and E fitted with a Gaussian. Results are means ± SD.
-
Figure 1—source data 1
Source data for Figure 1D–Eand Figure 1—figure supplement 1B and 2A.
- https://doi.org/10.7554/eLife.17667.003
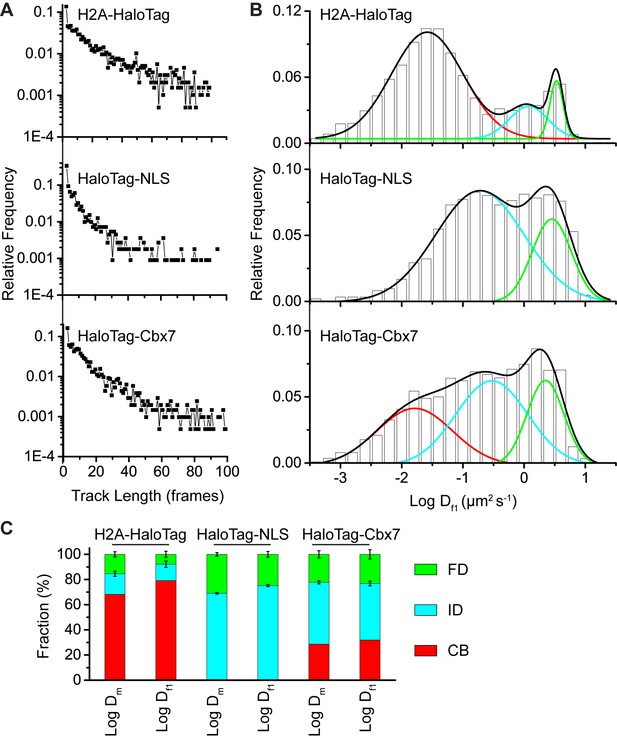
Comparison of the analysis.
(A) Distributions of the track (trajectory) length (in frames) for H2A-HaloTag, HaloTag-NLS, and HaloTag-Cbx7. (B) Normalized histograms of the log diffusion coefficient calculated from the first-step displacement of individual tracks for H2A-HaloTag, HaloTag-NLS, and HaloTag-Cbx7 in wild-type mES cells. (C) Fractions calculated from the analysis of H2A-HaloTag, HaloTag-NLS, and HaloTag-Cbx7 in wild-type mES cells.
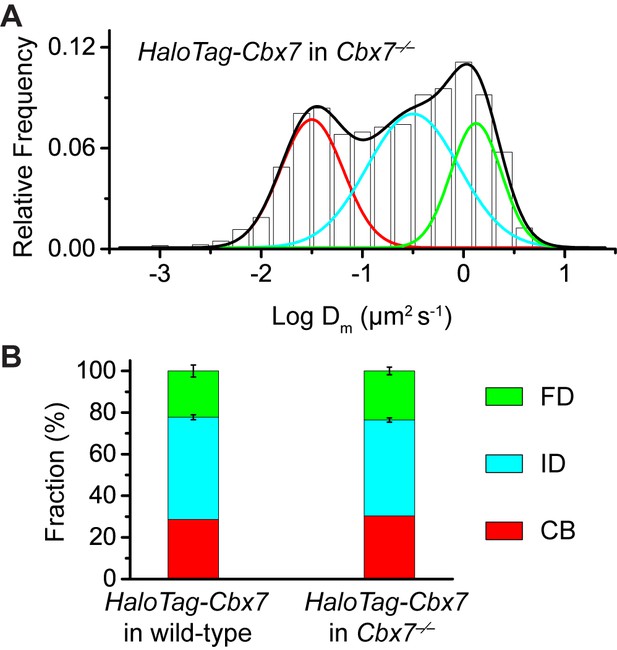
Control experiments for testing the effects of the endogenous Cbx7 protein on the kinetic fractions of the exogenous HaloTag-Cbx7 fusion.
(A) Normalized histograms of the log maximum likelihood diffusion coefficient for HaloTag-Cbx7 in Cbx7−/− mES cells (N = 25 cells, n = 5119 trajectories). (B) Fraction of the CB (red), ID (cyan), and FD (green) population for HaloTag-Cbx7 in wild-type mES cells replicated from Figure 1F and for HaloTag-Cbx7 in Cbx7−/− mES cells. Results are means ± SD.
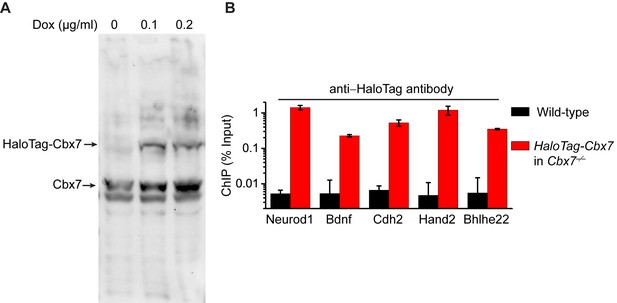
Control experiments for analyzing the protein level of HaloTag-Cbx7 and for testing whether HaloTag-Cbx7 occupies Polycomb target promoters.
(A) Western blotting of nuclear extracts from wild-type mES cells expressing HaloTag-Cbx7 in the presence or absence of Dox indicated above lanes. The endogenous and fusion proteins are marked at the left of gel. (B) ChIP analysis. Chromatin was prepared from wild-type and HaloTag-Cbx7/Cbx7−/− mES cells, respectively, and precipitated using antibody directed against HaloTag. DNA was quantified by qPCR. Results are means ± S.D.
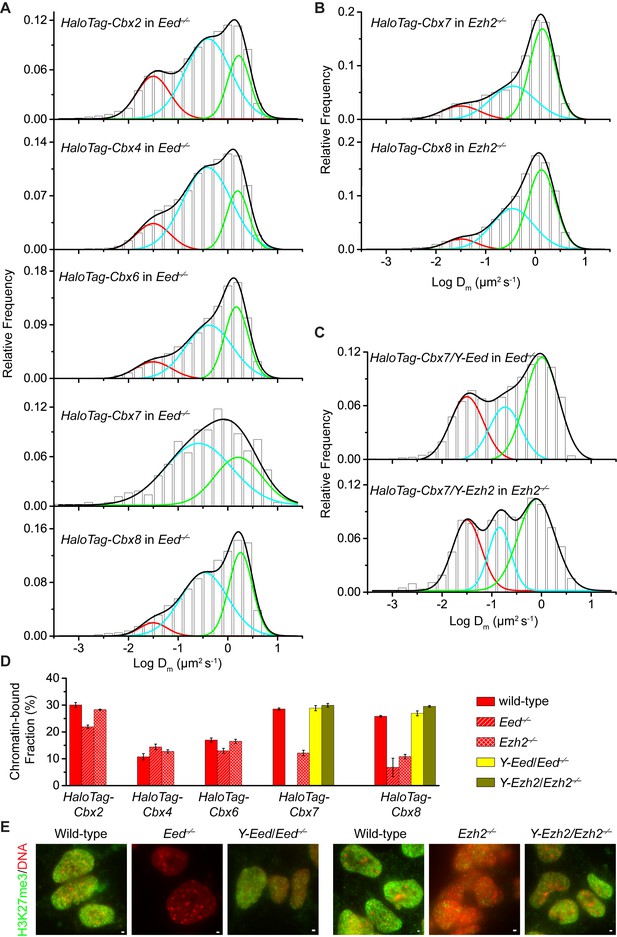
H3K27me3 is important for the targeting of Cbx7 and Cbx8 to chromatin, but plays a less important role for Cbx2, Cbx4, and Cbx6.
(A) Normalized histograms of the log maximum likelihood diffusion coefficient for HaloTag-Cbx2 (N = 27 cells, n = 2471 trajectories), HaloTag-Cbx4 (N = 21 cells, n = 3254 trajectories), HaloTag-Cbx6 (N = 11 cells, n = 4860 trajectories), HaloTag-Cbx7 (N = 25 cells, n = 453 trajectories), and HaloTag-Cbx8 (N = 47 cells, n = 5825 trajectories) in Eed−/− mES cells. The distributions for HaloTag-Cbx2, HaloTag-Cbx4, HaloTag-Cbx6, and HaloTag-Cbx8 were fitted with three populations while the distribution for HaloTag-Cbx7 with two populations. (B) Normalized histograms of the log maximum likelihood diffusion coefficient for HaloTag-Cbx7 (N = 26 cells, n = 3874 trajectories) and HaloTag-Cbx8 (N = 42 cells, n = 9220 trajectories) in Ezh2−/− mES cells. The distributions were fitted with three components. (C) Normalized histograms of the log maximum likelihood diffusion coefficient for HaloTag-Cbx7 in Y-Eed/Eed−/− (N = 16 cells, n = 1733 trajectories) and Y-Ezh2/Ezh2−/− (N = 14 cells, n = 846 trajectories) mES cells. The histograms were fitted with a three-component Gaussian. (D) Chromatin-bound fraction for HaloTag-Cbx2, HaloTag-Cbx4, HaloTag-Cbx6, HaloTag-Cbx7, and HaloTag-Cbx8 in wild-type (red solid), Eed−/− (red strip), and Ezh2−/− (red cross-strip) mES cells, and for HaloTag-Cbx7 and HaloTag-Cbx8 in Y-Eed/Eed−/− (yellow solid) andY-Ezh2/Ezh2−/− (dark yellow solid) mES cells. The data were obtained from Figure 1E, Figure 2A–C, and Figure 2—figure supplement 1B–C fitted with a Gaussian function. Results are means ± SD. (E) Immunostaining of H3K27me3 in wild-type, Eed−/−, Ezh2−/−, Y-Eed/Eed−/−, and Y-Ezh2/Ezh2−/− mES cells by using antibody directed against H3K27me3 (green). DNA was stained with hoechst (red). Overlay images are shown. Note that H3K27me3 staining is visible in Ezh2−/− mES cells because of the redundancy of Ezh1. Scale bar is 5 µm.
-
Figure 2—source data 1
Source data for Figure 2A–C and Figure 2—figure supplementary 1B–C.
- https://doi.org/10.7554/eLife.17667.016
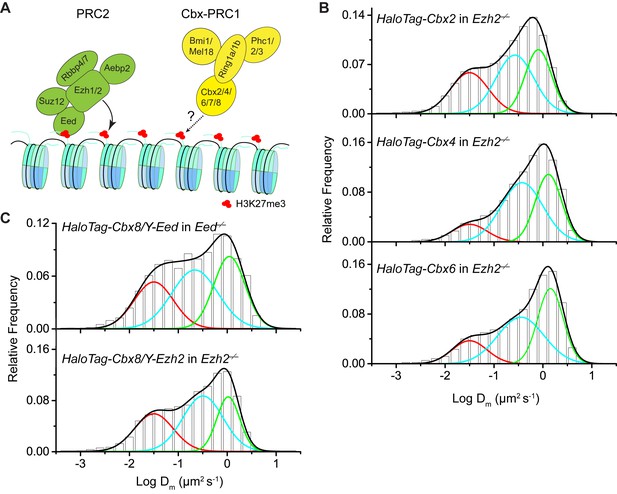
Additional experiments for HaloTag-Cbx in Eed−/−and Ezh2−/−mES cells.
(A) Schematic representation of the hypothetical model for the targeting of Cbx-PRC1 to chromatin via the Cbx CD interaction with H3K27me3 mediated by PRC2. The current research tests whether H3K27me3 is required for the targeting of Cbx-PRC1 to chromatin (mark by '?' and 'dash lines'). (B) Normalized histograms of the log maximum likelihood diffusion coefficient for HaloTag-Cbx2 (N = 45 cells, n = 8151 trajectories), HaloTag-Cbx4 (N = 24 cells, n = 4534 trajectories), HaloTag-Cbx6 (N = 18 cells, n = 5720 trajectories) in Ezh2−/− mES cells. The distributions were fitted with three populations. (C) Normalized histograms of the log maximum likelihood diffusion coefficient for HaloTag-Cbx8 in Y-Eed/Eed−/− (N = 24 cells, n = 1979 trajectories) and Y-Ezh2/Ezh2−/− (N = 28 cells, n = 7523 trajectories) mES cells. The histograms were fitted with three populations.
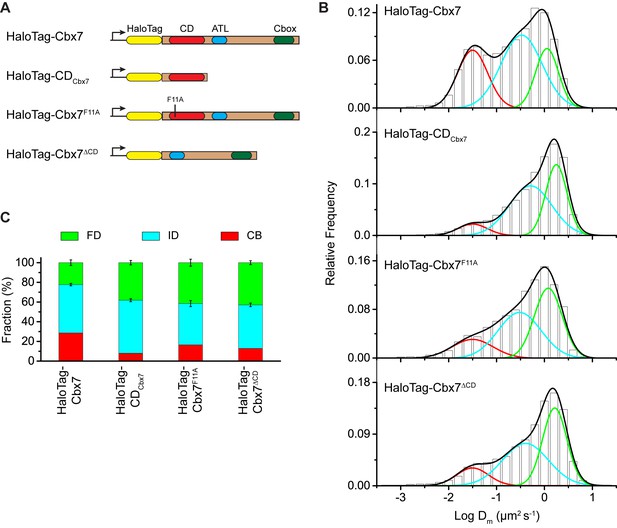
The Cbx7 CD is not efficient for the targeting of Cbx7 to chromatin.
(A) Schematic representation of Cbx7 variants. (B) Normalized histograms of the log maximum likelihood diffusion coefficient for HaloTag-Cbx7 replicated from Figure 1E and for HaloTag-CDCbx7 (N = 24 cells, n = 6600 trajectories), HaloTag-Cbx7F11A (N = 22 cells, n = 1882 trajectories), and HaloTag-Cbx7△CD (N = 15 cells, n = 5215 trajectories) in wild-type mES cells. The histograms of HaloTag-CDCbx7, HaloTag-Cbx7F11A, and HaloTag-Cbx7△CD were fitted with a three-component Gaussian. (C) Fraction of the CB, ID, and FD population for HaloTag-Cbx7 replicated from Figure 1F, HaloTag-CDCbx7, HaloTag-Cbx7F11A, and HaloTag-Cbx7△CD. The data were obtained from Figure 3B fitted with a Gaussian. Results are means ± SD.
-
Figure 3—source data 1
Source data for Figure 3B
- https://doi.org/10.7554/eLife.17667.034
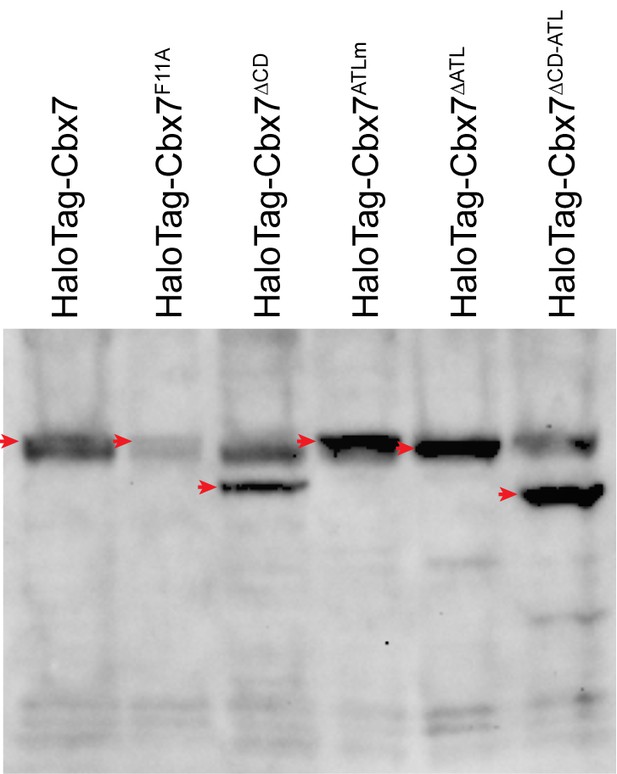
Control experiments for analyzing the protein levels of the HaloTag-Cbx7 variants.
To determine whether the Cbx7 variants are correctly expressed in wild-type mES cells, we performed immunoblotting of nuclear extracts from wild-type mES cells stably expressing HaloTag-Cbx7 and its variant fusions. The membrane was probed with antibody directed against with HaloTag. Red arrow indicates bands corresponding to the Cbx7 fusions indicated above the gel.
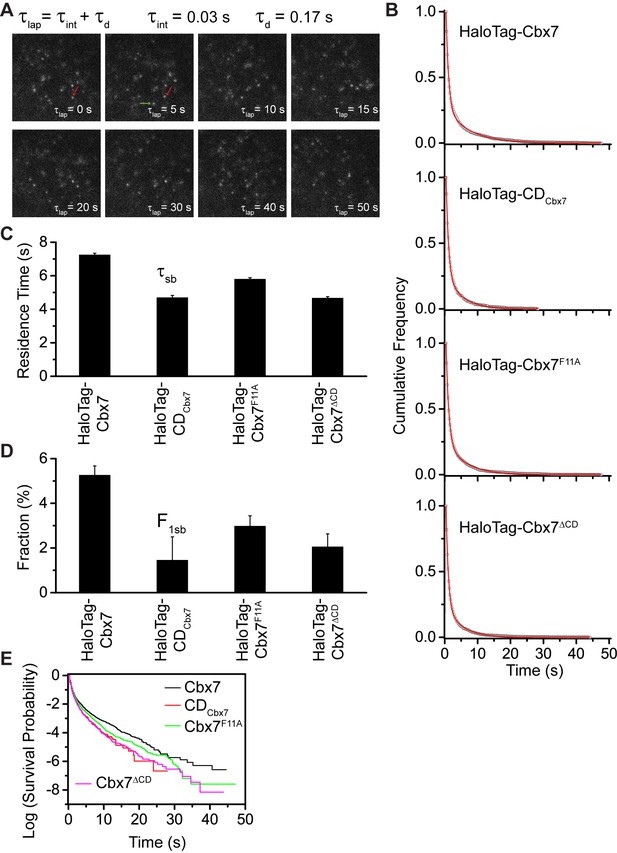
Effects of the Cbx7 CD on the residence time of Cbx7 at chromatin.
(A) Time-lapse imaging of HaloTag-Cbx7 at constant integration (τint = 30 ms) and dark (τd = 170 ms) time in wild-type mES cells. The red arrow indicates a molecule that binds to chromatin. The green arrow indicates a diffusing molecule. Molecules with < 0.10 µm2/s were selected to calculate residence time and survival probability. (B) Cumulative frequency distribution of the dwell times for HaloTag-Cbx7 (N = 17 cells, n = 2169 trajectories), HaloTag-CDCbx7 (N = 18 cells, n = 790 trajectories), HaloTag-Cbx7F11A (N = 25 cells, n = 3956 trajectories), and HaloTag-Cbx7△CD (N = 21 cells, n = 3471 trajectories) in wild-type mES cells. The histograms were fitted with a two-component exponential decay model. (C) Residence time (τsb) of the stable chromatin-bound population for HaloTag-Cbx7, HaloTag-CDCbx7, HaloTag-Cbx7F11A, and HaloTag-Cbx7△CD in wild-type mES cells. (D) Fraction (F1sb) of the stable chromatin-bound population for HaloTag-Cbx7, HaloTag-CDCbx7, HaloTag-Cbx7F11A, and HaloTag-Cbx7△CD in wild-type mES cells. (E) Survival probability for HaloTag-Cbx7, HaloTag-CDCbx7, HaloTag-Cbx7F11A, and HaloTag-Cbx7△CD in wild-type mES cells.
-
Figure 4—source data 1
Source data for Figure 4B and Figure 4—figure supplementary 1
- https://doi.org/10.7554/eLife.17667.039
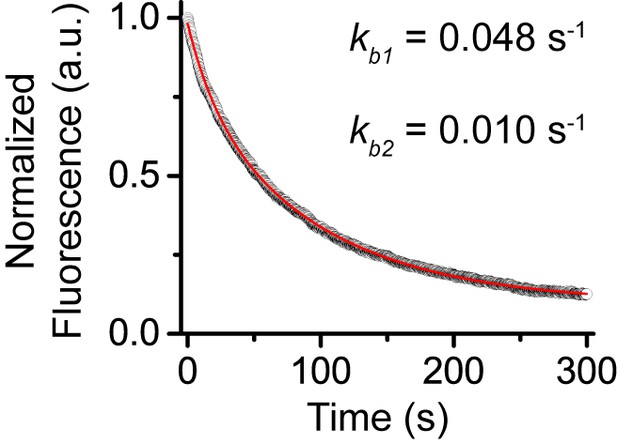
Control experiments for determine photobleaching constant of JF549.
We performed photobleaching of H2A-HaloTag labelled with JF549 dye. Wild-type mES cells stably expressing H2A-HaloTag were incubated with 500 nM of JF549 dye. Live-cell image stacks were taken using the same power and integration and dark time as that for single-molecule experiments. We obtained 9 photobleaching curves. After background correction, the curves were normalized to 1 and averaged. The averaged curve was better described by a two-component exponential decay function based the significance of the F-test.
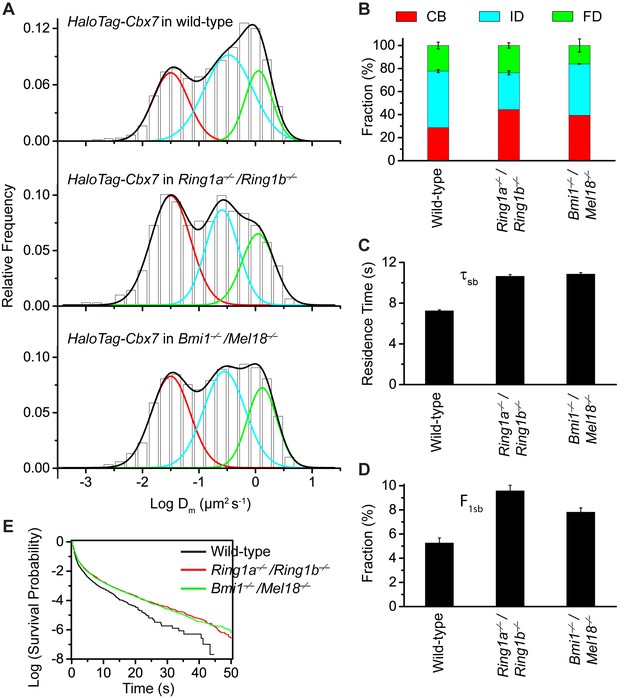
Disruption of the complex formation of Cbx7-PRC1 facilitates the targeting of Cbx7 to chromatin.
(A) Normalized histograms of the log maximum likelihood diffusion coefficient for HaloTag-Cbx7 in wild-type mES cells replicated from Figure 1E and for HaloTag-Cbx7 in Ring1a−/−/Ring1b−/− (N = 29 cells, n = 2600 trajectories) and Bmi1−/−/Mel18−/− (N = 27 cells, n = 6859 trajectories) mES cells. The histograms were fitted with a three-component Gaussian. (B) Fraction of the CB (red), ID (cyan), and FD (green) population for HaloTag-Cbx7 in wild-type mES cells replicated from Figure 1F and for HaloTag-Cbx7 in Ring1a−/−/Ring1b−/− and Bmi1−/−/Mel18−/− mES cells. Results are means ± SD. (C–D) Residence time (C) and fraction (D) of the stable chromatin-bound population for HaloTag-Cbx7 in wild-type mES cells replicated from Figure 4C and D and for HaloTag-Cbx7 in Ring1a−/−/Ring1b−/− (N = 18 cells, n = 4849 trajectories) and Bmi1−/−/Mel18−/− (N = 27 cells, n = 3484 trajectories) mES cells. (E) Survival probability for HaloTag-Cbx7 in wild-type mES cells replicated from Figure 4E, and for HaloTag-Cbx7 in Ring1a−/−/Ring1b−/− and Bmi1−/−/Mel18−/− mES cells.
-
Figure 5—source data 1
Source data for Figure 5A and Figure 5—figure supplementary 1
- https://doi.org/10.7554/eLife.17667.046

Cumulative frequency distribution of the dwell times for determining the residence times of HaloTag-Cbx7 in Ring1a−/−/Ring1b−/− and Bmi1−/−/Mel18−/− mES cells.
Cumulative frequency distribution of the dwell times for HaloTag-Cbx7 replicated from Figure 4B and for HaloTag-Cbx7 in Ring1a−/−/Ring1b−/− (N = 38 cells, n = 7825 trajectories) and Bmi1−/−/Mel18−/− (N = 47 cells, n = 8142 trajectories) mES cells. The cumulative frequency distributions were fitted with a two-component exponential decay model.
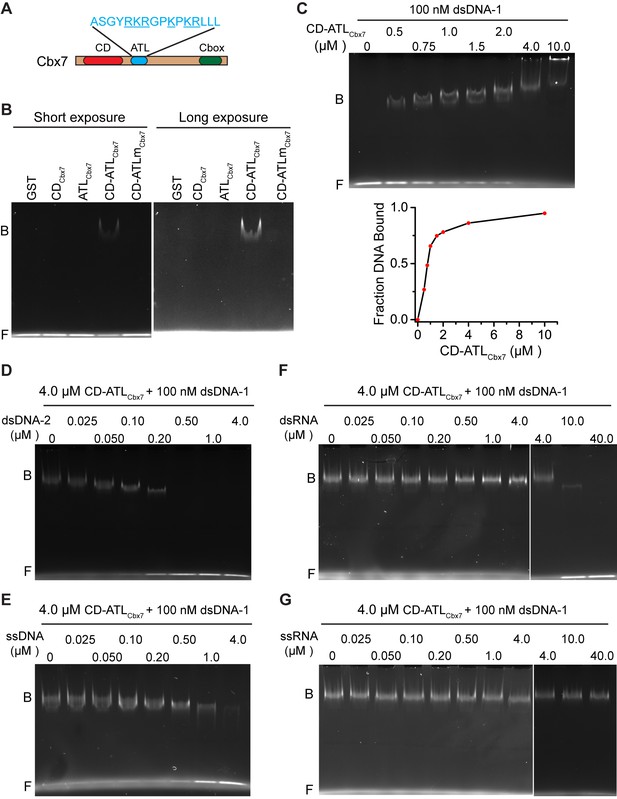
CDCbx7 and ATLCbx7 together constitute a DNA-binding entity.
(A) Schematic representation of Cbx7. The sequence of amino acids of ATL motif is shown. The basic amino acids are underlined and mutated to alanine to generate ATLm. (B) EMSA for the determination of Cbx7 variants binding to dsDNA-1. dsDNA-1 was labelled with Alexa Fluor 488 dye. Left: short-time exposure. Right: long-time exposure. B: bound DNA-protein complex. F: free DNA. (C) EMSA for the determination of the dissociation constant (Kd) of the CD-ATLCbx7 cassette binding to dsDNA-1. Bottom: binding curve for the CD-ATLCbx7 cassette. (D–G) EMSA for the determination of the relative affinities for the CD-ATLCbx7 cassette binding to dsDNA-2, ssDNA, dsRNA, and ssRNA. dsDNA-1 within dsDNA-1/CD-ATLCbx7 complexes was competed with competitors, dsDNA-2 (D), ssDNA (E), dsRNA (F), and ssRNA (G), respectively. ds: double-strand. ss: single-strand.
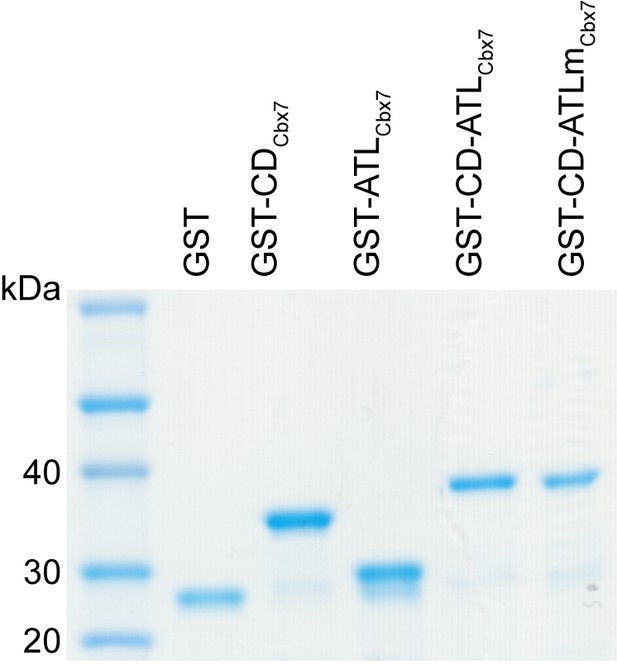
Control experiments for analyzing the Cbx7 variants purified from BL21 cells by SDS-PAGE.
https://doi.org/10.7554/eLife.17667.053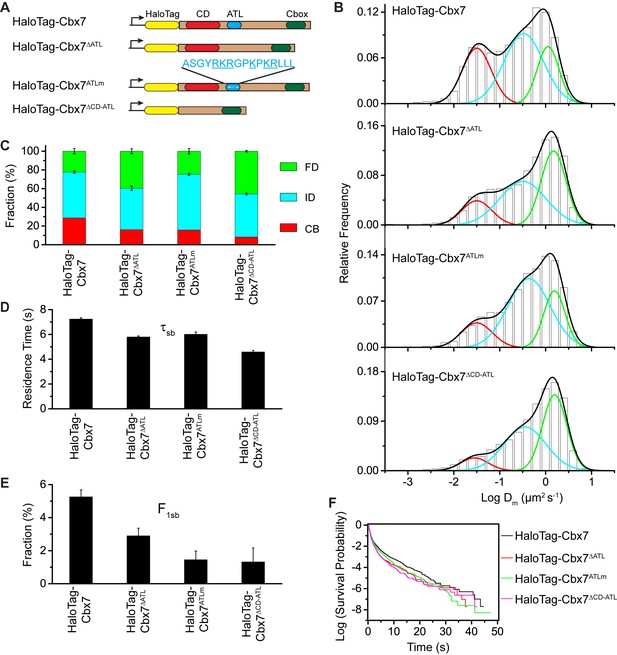
CDCbx7 and ATLCbx7 together control the targeting of Cbx7 to chromatin.
(A) Schematic representation of Cbx7 variants. The underlined ATL amino acids were mutated into alanine or glycine. (B) Normalized histograms of the log maximum likelihood diffusion coefficient for HaloTag-Cbx7 replicated from Figure 1E and for HaloTag-Cbx7△ATL (N = 12 cells, n = 3065 trajectories), HaloTag-Cbx7ATLm (N = 13 cells, n = 2257 trajectories), and HaloTag-Cbx7△CD-ATL (N = 35 cells, n = 8329 trajectories) in wild-type mES cells. The histograms were fitted with a three-component Gaussian. (C) Fraction of the CB, ID, and FD population for HaloTag-Cbx7 replicated from Figure 1F, HaloTag-Cbx7△ATL, HaloTag-Cbx7ATLm, and HaloTag-Cbx7△CD-ATL. The data were obtained from Figure 7B fitted with a Gaussian. Results are means ± SD. (D–E) Residence time (D) and fraction (E) of the stable chromatin-bound population for HaloTag-Cbx7 replicated from Figure 4C and D, HaloTag-Cbx7△ATL (N = 17 cells, n = 2384 trajectories), HaloTag-Cbx7ATLm (N = 24 cells, n = 2957 trajectories), and HaloTag-Cbx7△CD-ATL (N = 22 cells, n = 4908 trajectories). Results are means ± SD. (F) Survival probability for HaloTag-Cbx7 replicated from Figure 4D, HaloTag-Cbx7△ATL, HaloTag-Cbx7ATLm, and HaloTag-Cbx7△CD-ATL.
-
Figure 7—source data 1
Source data for Figure 7B and Figure 7—figure supplementary 1.
- https://doi.org/10.7554/eLife.17667.055
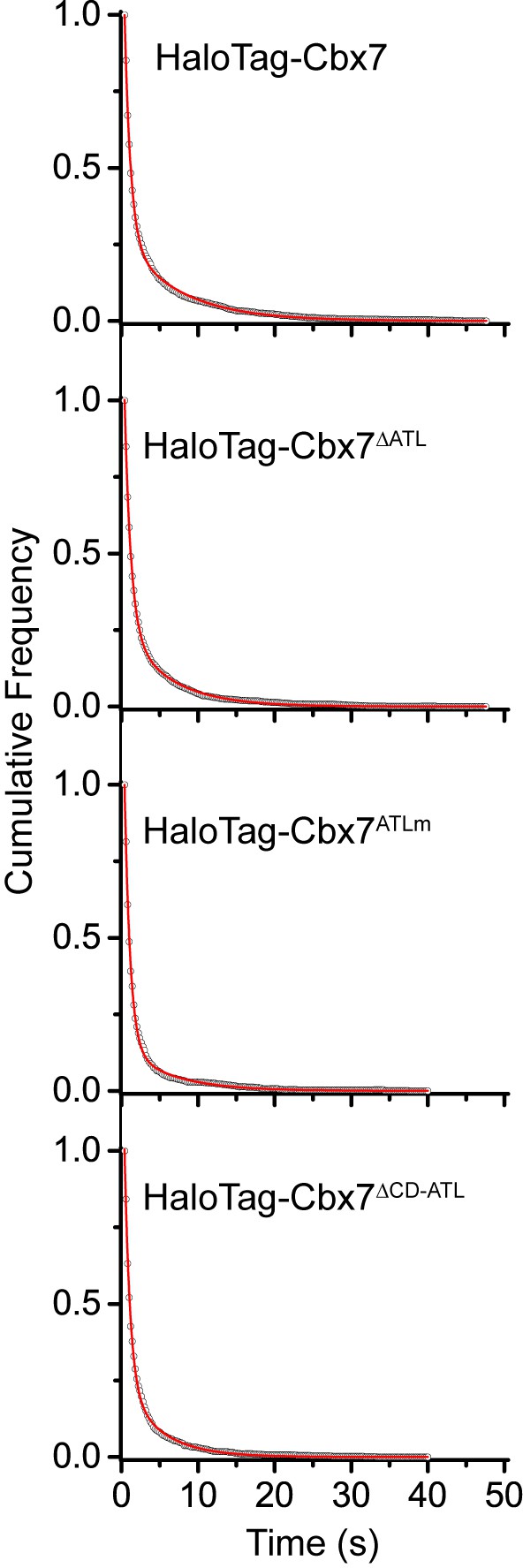
Cumulative frequency distribution of the dwell times for determining the residence times of HaloTag-Cbx7△ATL, HaloTag-Cbx7ATLm, and HaloTag-Cbx7△CD-ATL in wild-type mES cells.
Cumulative frequency distribution of the dwell times for HaloTag-Cbx7 replicated from Figure 4B and for HaloTag-Cbx7△ATL (N = 17 cells, n = 3956 trajectories), HaloTag-Cbx7ATLm (N = 24 cells, n = 2115 trajectories), and HaloTag-Cbx7△CD-ATL (N = 22 cells, n = 2252 trajectories) in wild-type mES cells. The cumulative frequency distributions were fitted with a two-component exponential decay model.
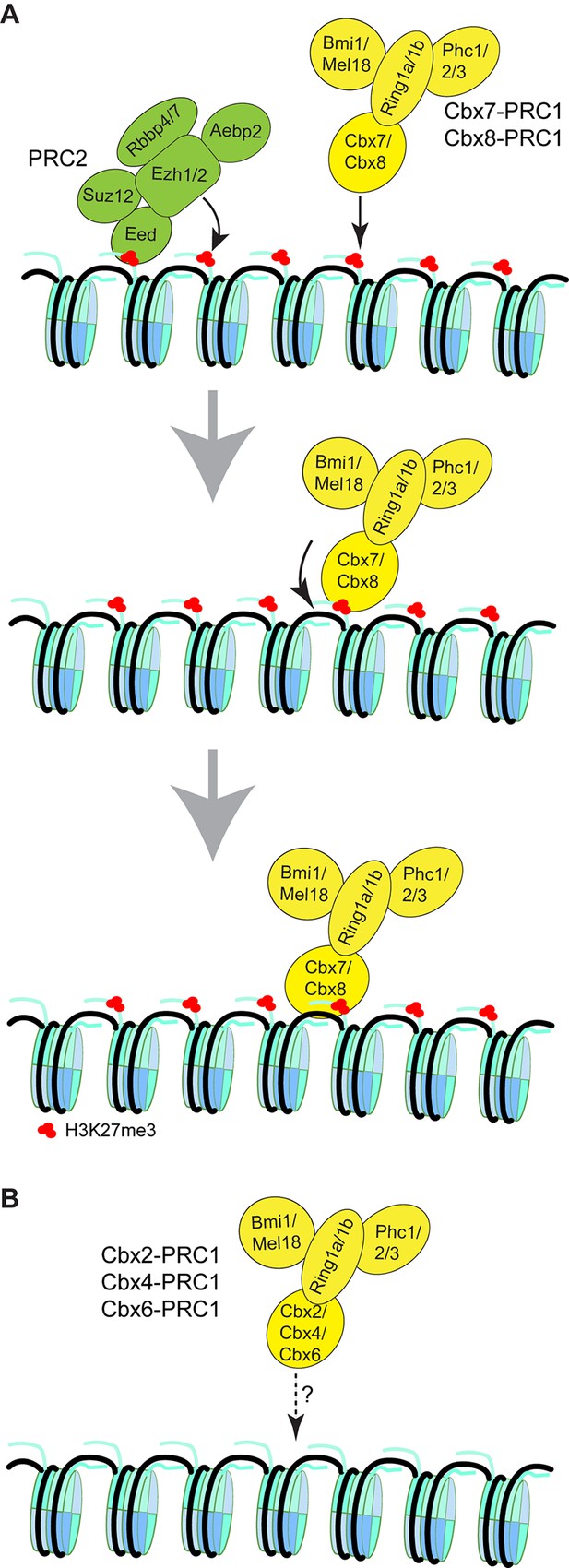
Proposed models for the targeting of Cbx-PRC1 to chromatin.
(A) The Cbx7-PRC1 and Cbx8-PRC1 complexes are targeted to chromatin by co-recognition of H3K27me3 and DNA. The Cbx7- and Cbx8-PRC1 complexes are guided to genomic loci by the CD interaction with H3K27me3. The interaction triggers conformational changes of the Cbx7- and Cbx8-PRC1 complexes and induces the CD-ATL cassette interaction with DNA. Multivalent engagement of DNA and H3K27me3 by the CD-ATL cassette stabilizes the Cbx7- and Cbx8-PRC1 complexes at chromatin. (B) Molecular mechanisms for the targeting of Cbx2-PRC1, Cbx4-PRC1, and Cbx6-PRC1 complexes to chromatin remain unknown.
Videos
H2A-HaloTag in wild-type mES cells (Fractional studies).
https://doi.org/10.7554/eLife.17667.007HaloTag-NLS in wild-type mES cells (Fractional studies).
https://doi.org/10.7554/eLife.17667.008HaloTag-Cbx2 in wild-type mES cells (Fractional studies).
https://doi.org/10.7554/eLife.17667.009HaloTag-Cbx4 in wild-type mES cells (Fractional studies).
https://doi.org/10.7554/eLife.17667.010HaloTag-Cbx6 in wild-type mES cells (Fractional studies).
https://doi.org/10.7554/eLife.17667.011HaloTag-Cbx7 in wild-type mES cells (Fractional studies).
https://doi.org/10.7554/eLife.17667.012HaloTag-Cbx8 in wild-type mES cells (Fractional studies).
https://doi.org/10.7554/eLife.17667.013HaloTag-Cbx7 in Cbx7 KO mES cells (Fractional studies).
https://doi.org/10.7554/eLife.17667.014HaloTag-Cbx2 in Eed KO mES cells (Fractional studies).
https://doi.org/10.7554/eLife.17667.018HaloTag-Cbx4 in Eed KO mES cells (Fractional studies).
https://doi.org/10.7554/eLife.17667.019HaloTag-Cbx6 in Eed KO mES cells (Fractional studies).
https://doi.org/10.7554/eLife.17667.020HaloTag-Cbx7 in Eed KO mES cells (Fractional studies)
https://doi.org/10.7554/eLife.17667.021HaloTag-Cbx8 in Eed KO mES cells (Fractional studies).
https://doi.org/10.7554/eLife.17667.022HaloTag-Cbx2 in Ezh2 KO mES cells (Fractional studies).
https://doi.org/10.7554/eLife.17667.023HaloTag-Cbx4 in Ezh2 KO mES cells (Fractional studies).
https://doi.org/10.7554/eLife.17667.024HaloTag-Cbx6 in Ezh2 KO mES cells (Fractional studies).
https://doi.org/10.7554/eLife.17667.025HaloTag-Cbx7 in Ezh2 KO mES cells (Fractional studies).
https://doi.org/10.7554/eLife.17667.026HaloTag-Cbx8 in Ezh2 KO mES cells (Fractional studies).
https://doi.org/10.7554/eLife.17667.027HaloTag-Cbx7/Y-Eed in Eed KO mES cells (Fractional studies).
https://doi.org/10.7554/eLife.17667.028HaloTag-Cbx8/Y-Eed in Eed KO mES cells (Fractional studies).
https://doi.org/10.7554/eLife.17667.029HaloTag-Cbx7/Y-Ezh2 in Ezh2 KO mES cells (Fractional studies).
https://doi.org/10.7554/eLife.17667.030HaloTag-Cbx8/Y-Ezh2 in Ezh2 KO mES cells (Fractional studies).
https://doi.org/10.7554/eLife.17667.031HaloTag-CDCbx7 in wild-type mES cells (Fractional studies).
https://doi.org/10.7554/eLife.17667.032HaloTag-Cbx7F11A in wild-type mES cells (Fractional studies).
https://doi.org/10.7554/eLife.17667.036HaloTag-Cbx7△CD in wild-type mES cells (Fractional studies).
https://doi.org/10.7554/eLife.17667.037HaloTag-Cbx7 in wild-type mES cells (Residence time studies).
https://doi.org/10.7554/eLife.17667.041HaloTag-CDCbx7 in wild-type mES cells (Residence time studies).
https://doi.org/10.7554/eLife.17667.042HaloTag-Cbx7F11A in wild-type mES cells (Residence time studies).
https://doi.org/10.7554/eLife.17667.043HaloTag-Cbx7△CD in wild-type mES cells (Residence time studies).
https://doi.org/10.7554/eLife.17667.044HaloTag-Cbx7 in Ring1a/Ring1b dKO mES cells (Fractional studies).
https://doi.org/10.7554/eLife.17667.048HaloTag-Cbx7 in Bmi1/Mel18 dKO mES cells (Fractional studies).
https://doi.org/10.7554/eLife.17667.049HaloTag-Cbx7 in Ring1a/Ring1b dKO mES cells (Residence time studies).
https://doi.org/10.7554/eLife.17667.050HaloTag-Cbx7 in Bmi1/Mel18 dKO mES cells (Residence time studies).
https://doi.org/10.7554/eLife.17667.051HaloTag-Cbx7△ATL in wild-type mES cells (Fractional studies).
https://doi.org/10.7554/eLife.17667.057HaloTag-Cbx7ATLm in wild-type mES cells (Fractional studies).
https://doi.org/10.7554/eLife.17667.058HaloTag-Cbx7△CD-ATL in wild-type mES cells (Fractional studies).
https://doi.org/10.7554/eLife.17667.059HaloTag-Cbx7△ATL in wild-type mES cells (Residence time studies).
https://doi.org/10.7554/eLife.17667.060HaloTag-Cbx7ATLm in wild-type mES cells (Residence time studies).
https://doi.org/10.7554/eLife.17667.061HaloTag-Cbx7△CD-ATL in wild-type mES cells (Residence time studies).
https://doi.org/10.7554/eLife.17667.062Additional files
-
Supplementary file 1
Fractional sizes and diffusion constants of the CB, ID, and FD populations obtained from live-cell SMT analysis of the Cbx family proteins and their variants.
- https://doi.org/10.7554/eLife.17667.064
-
Supplementary file 2
Residence times, transient (F1tb) and stable (F1sb) chromatin-binding fractions of Cbx7 and its variants.
- https://doi.org/10.7554/eLife.17667.065
-
Supplementary file 3
U-track parameters used in this research.
- https://doi.org/10.7554/eLife.17667.066





