Phospholipase D activity couples plasma membrane endocytosis with retromer dependent recycling
Figures
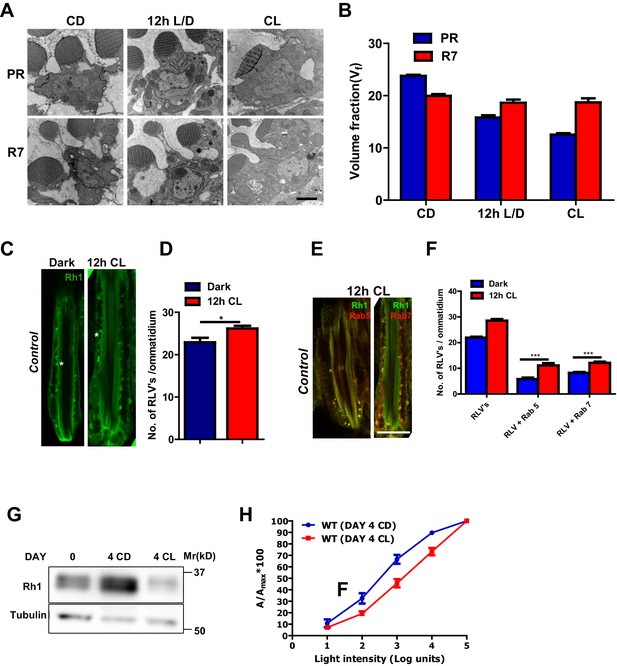
Rhabdomere size regulation during illumination in Drosophila photoreceptors.
(A) TEM images of single rhabdomere from wild-type photoreceptors (PRs) of 2-day-old flies post-eclosion reared in constant dark (CD), 12 hr light, 12 hr dark (12 h L/D) and constant light (CL). Scale bar: 1 µm. (B) Quantification of rhabdomere volume in PRs reared in various conditions. The peripheral PRs represent R1 to R6 rhabdomeres. The X-axis represents the rearing condition and the Y-axis represents the volume fraction (Vf) of rhabdomere expressed as a % with respect to total cell volume. n = 90 rhabdomeres taken from three separate flies. (C) Longitudinal section (LS) of retinae from control stained with rhodopsin 1 (Rh1) antibody. Flies were dissected after 0–6 hr (day 0) and 12 hr of bright light illumination (12 h CL) post-eclosion. Scale bar: 5 µm. (D) Quantification of RLVs from LS of retinae from control. The X-axis represents the time point and rearing condition. Y-axis shows the number of RLV’s per ommatidium. n = 10 ommatidia taken from three separate preps. (E) LS of retinae from control stained with Rh1 and Rab5; Rh1 and GFP (for Rh1>GFP::Rab7). Rearing condition is same as mentioned in (panel C). Scale bar: 5 µm. (F) Quantification of RLVs from LS of retinae from control. The X-axis represents the population of vesicles positive for mentioned protein. Y-axis shows the number of RLVs per ommatidium. n = 10 ommatidia taken from three separate preps. (G) Western blot from head extracts of control flies reared in various conditions as indicated on the top of the blot. The blot was probed with antibody to rhodopsin. Tubulin levels were used as a loading control. (H) Intensity response function of the light response from 4-day-constant-light (DAY 4 CL) and 4-day-constant-dark (DAY 4 CD) old control flies. The X-axis represents increasing light intensity in log units and Y-axis the peak response amplitude at each intensity normalized to the response at the maximum intensity. n=separate flies. Data are presented as mean ± SEM.
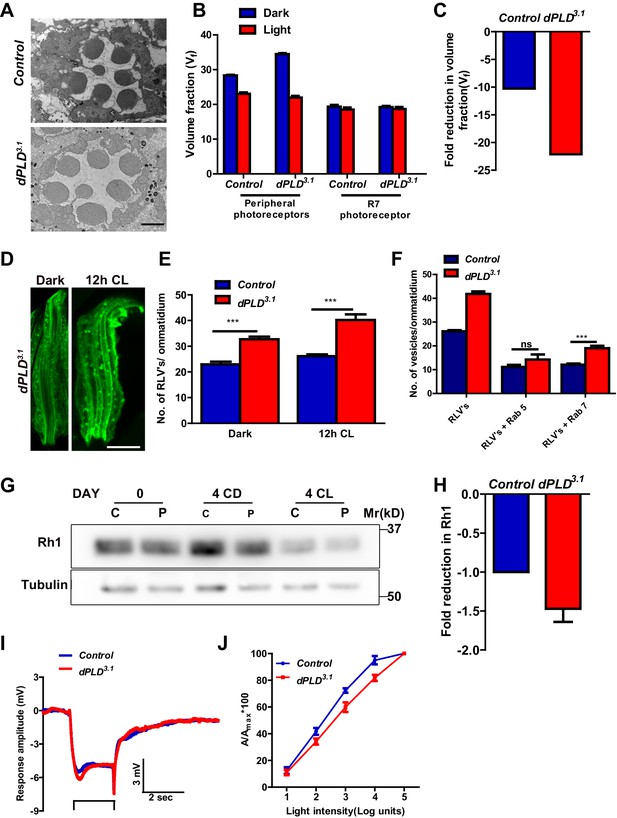
dPLD is required to support rhabdomere volume during illumination.
(A) TEM images showing single ommatidium from control and dPLD3.1. PRs of 0- to 12-hr-old flies post eclosion. Scale bar: 1 µm (B) Quantification of the rhabdomere volume of control and dPLD3.1. PRs reared in constant dark and constant light for 2 days post-eclosion. n = 90 rhabdomeres taken from three separate flies. (C) Quantification of fold reduction in rhabdomere volume of control and dPLD3.1 in light compared to dark. Genotypes are indicated on the X-axis and the Y-axis represents the percentage volume fraction (Vf) of the rhabdomere with respect to cell. (D) LS of retinae stained with rhodopsin one from dPLD3.1. Rearing conditions are indicated. Scale bar: 5 µm. (E) Quantification of RLVs from LS of retinae from control and dPLD3.1. n = 10 ommatidia taken from three separate preps. (F) Quantification of RLVs from LS of retinae from control and dPLD3.1 reared in 12 hr CL. The X-axis represents the population of vesicles positive for mentioned protein. Y-axis shows the number of RLVs per ommatidium. n = 10 ommatidia taken from three separate preps. (G) Western blot from head extracts of control (C) and dPLD3.1 (P) of matched eye color. Rearing conditions as indicated on the top of the blot. The blot was probed with antibody to rhodopsin. Tubulin levels were used as a loading control. (H) Quantification of fold reduction of rhodopsin seen in dPLD3.1 normalized to controls. The X-axis shows the genotype. Y-axis represents the fold reduction in rhodopsin. n = 3. (I) Representative ERG responses of 0- to 12-hr-old flies to a single 2 s flash of green light. Genotypes are indicated. X-axis represents the time in seconds (s) and the Y-axis represents the amplitude of response in mV. The duration of light pulse is indicated. (J) Intensity response function of the light response of 0- to 12-hr-old flies. Responses from control and dPLD3.1 flies with matched eye color are shown. The X-axis represents increasing light intensity in log units and Y-axis the peak response amplitude at each intensity normalized to the response at the maximum intensity. n= five separate flies. Data are presented as mean ± SEM.
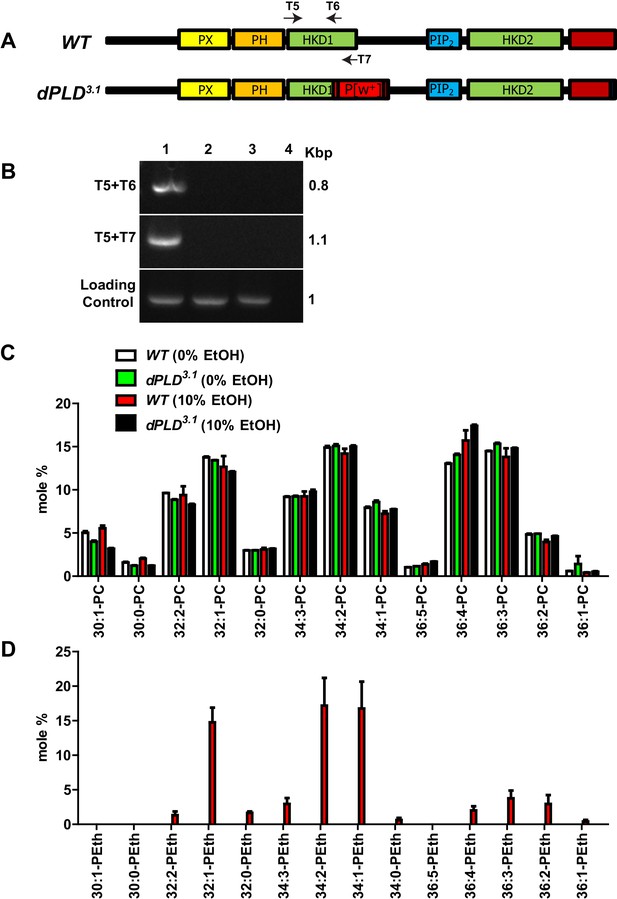
Characterization of dPLD3.1 loss-of-function allele.
(A) Schematic diagram representing the method used to generate the knockout of Drosophila phospholipase D (dPLD3.1) using homologous recombination. The mutant allele generated with respect to the wild-type locus is shown. The domains of dPLD (PX, PH, catalytic HKD1 and HKD2 and PIP2-binding domains) are shown. The C-terminal domain is marked in red. A Pw+ insertion (red box) that disrupts the HKD1 domain with stop codons in all three frames on both strands (white boxes) is indicated as P[w+]. The last three amino acids at the C-terminus that have been mutated are shown as a black box. T5, T6 and T7 marks the primers designed in the HKD1 motif of dPLD. (B) PCR analysis for presence of HKD1 motif from crude genomic DNA extracts of WT (1) dPLD3.1 (2) dPLD3.1/Df(2R)ED1612 (3) and water control (4). One the left side of the gel picture, primer pairs used are mentioned and on the right side, the product lengths are indicated. (C) Total amounts of various phosphatidylcholine (PC) species extracted and measured from flies used for the transphosphatidylation assay experiment. The X-axis shows acyl chains species that were detected. Y-axis represents the mole percent of phosphatidylethanol species. Species measured from wild type and dPLD3.1 with (10%) and without (0%) ethanol are shown. (D) The generation of phosphatidylethanol(P-EtOH) by dPLD (via the enzyme’s transphosphatidylation activity) was measured. The X-axis shows acyl chains species that were detected. Y-axis represents the mole percent of phosphatidylethanol species. Species measured from wild type and dPLD3.1 with (10%) and without (0%) ethanol are shown.
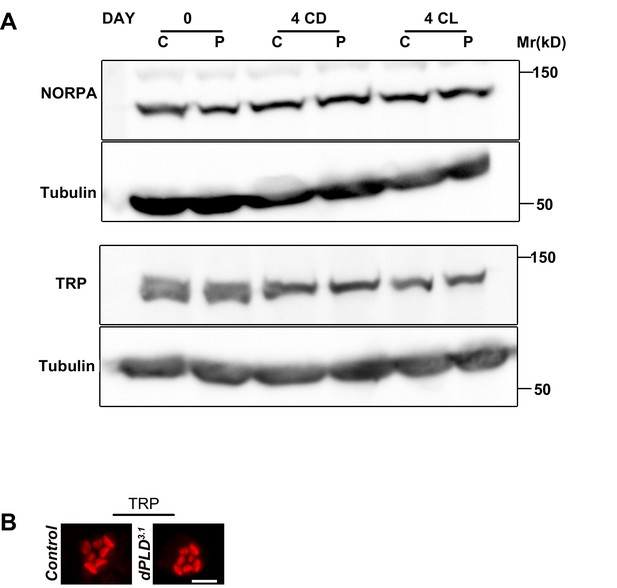
Normal levels and localization of other apical domain proteins in dPLD3.1.
(A) Western blot from head extracts of control (C) and dPLD3.1 (P) reared in various conditions as indicated on the top of the blot. The blot was probed with antibody to NORPA and TRP. Tubulin was used as loading control. (B) Confocal images of transverse section of retinae stained with an antibody to TRP in control and dPLD3.1. Cross-sections of the rhabdomere stained in red are shown. Scale bar 5 µm
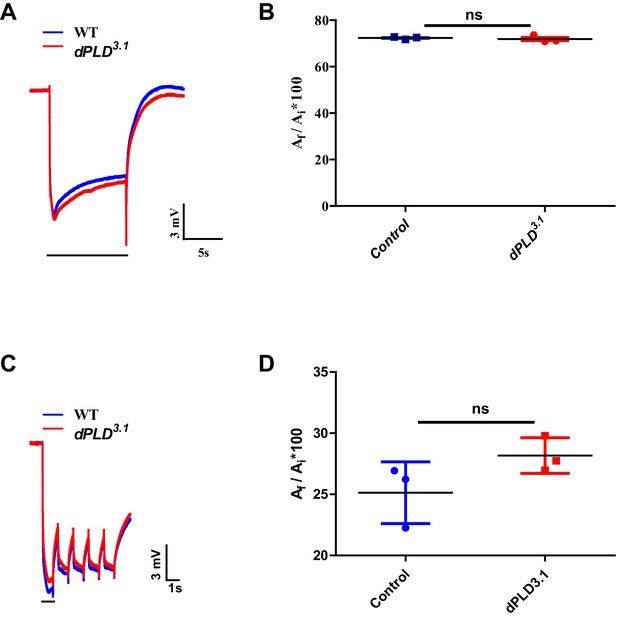
Electrophysiological characterization of dPLD3.1.
(A) Representative ERG responses of 0- to 12-hr-old flies to a single 10 s flash of green light. Genotypes are indicated. X-axis represents the time in seconds (s) and the Y-axis represents the amplitude of response in mV. The duration of light pulse is indicated. (B) Quantification of the light response. Y-axis represents the ratio of final (Af) and initial (Ai) amplitude of single trace during the stimulus in percentage. X-axis represents the genotypes. n = 3 separate flies (C) Representative ERG responses of 0- to 12-hr-old flies to a 1 s flash of green light train - five pulses. Genotypes are indicated. X-axis represents the time in seconds (s) and the Y-axis represents the amplitude of response in mV. The duration of light pulse is indicated. (D) Quantification of the light response. Y-axis represents the ratio of final (Af # 5 pulse) and initial (Ai # 1 pulse) amplitude during the stimulus in percentage. X-axis represents the genotypes. n = 3 separate flies Data are presented as mean ± SEM.
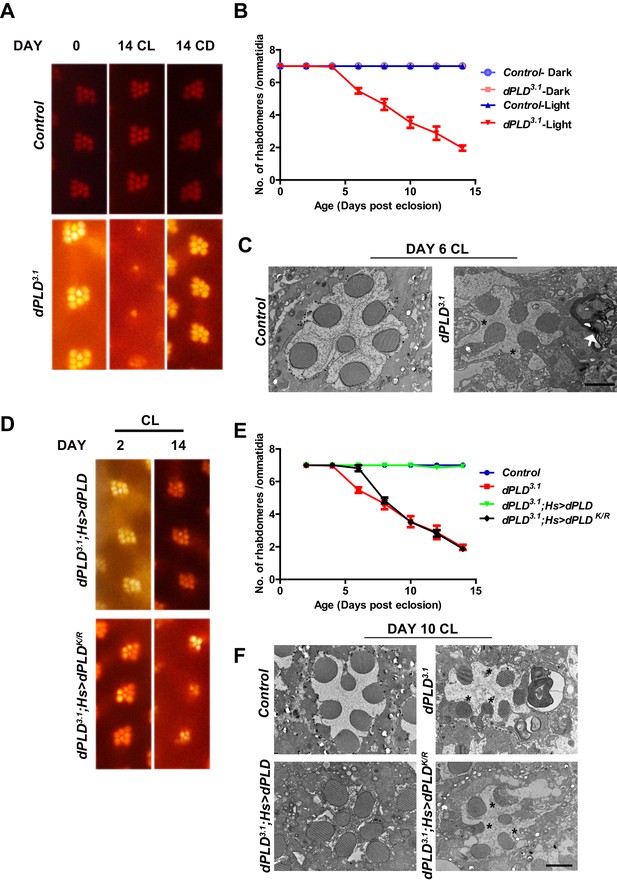
dPLD is essential to support rhabdomere structure during illumination.
(A) Representative optical neutralization (ON) images showing rhabdomere structure from control and dPLD3.1. The age and rearing conditions are mentioned on the top of the panels. (B) Quantification of rate of PR degeneration of control and dPLD3.1 reared in bright light. The X-axis represents age of the flies and the Y-axis represents the number of intact rhabdomeres visualized in each ommatidium. n = 50 ommatidia taken from at least five separate flies. (C) TEM images showing a single ommatidium from control and dPLD3.1 PRs reared in bright illumination for 6 days post eclosion. * indicates the collapsed rhabdomere and the arrow head indicate whorl like membranes accumulated in the cell body. Scale bar 1 µm. (D) Representative ON images showing ommatidia from dPLD3.1;Hs>dPLD and dPLD3.1;Hs>dPLDK/R. The age and rearing conditions are indicated on the top of the image. (E) Quantification of rate of PR degeneration of control, dPLD3.1, dPLD3.1;Hs>dPLD and dPLD3.1;Hs>dPLDK/R reared in bright light. n = 50 ommatidia taken from at least five separate flies. (F) TEM images showing a single ommatidium from control, dPLD3.1, dPLD3.1;Hs>dPLD and dPLD3.1;Hs>dPLDK/R PRs reared in light for 10 days post-eclosion. Scale bar 1 µm. Data are presented as mean ± SEM.
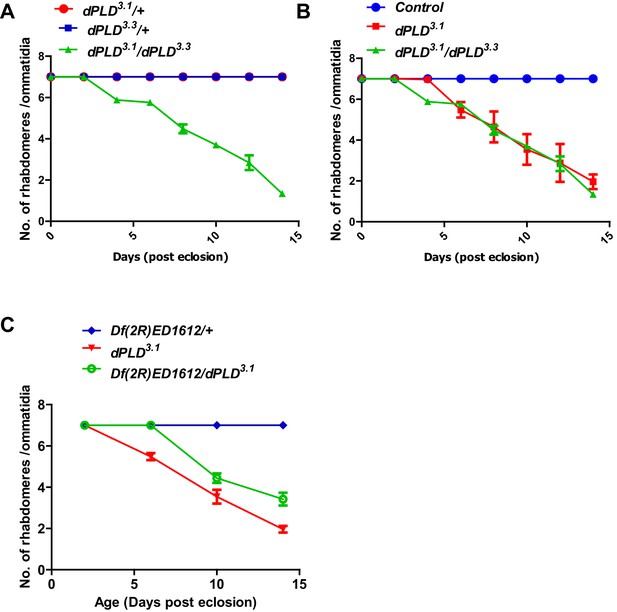
Genetic validation of the retinal degeneration phenotype of dPLD3.1.
(A) Quantification of retinal degeneration in dPLD3.1/+, dPLD1.1/+, dPLD3.1/ dPLD1.1. The X-axis represents age of the flies and the Y-axis represents the number of rhabdomere visualized in each ommatidium. Error bars represents mean ± SEM from 50 ommatidia taken from at least five separate flies. (B) Quantification of retinal degeneration in control, dPLD3.1, dPLD3.1/ dPLD1.1. The X-axis represents age of the flies and the Y-axis represents the number of rhabdomere visualized in each ommatidium. Error bars represents mean ± SEM from 50 ommatidia taken from at least five separate flies. (C) Quantification of retinal degeneration in Df(2R)ED1612/+, dPLD3.1, dPLD3.1/Df(2R)ED1612. The X-axis represents age of the flies and the Y-axis represents the number of rhabdomere visualized in each ommatidium. Error bars represents mean ± SEM from 50 ommatidia taken from at least five separate flies.
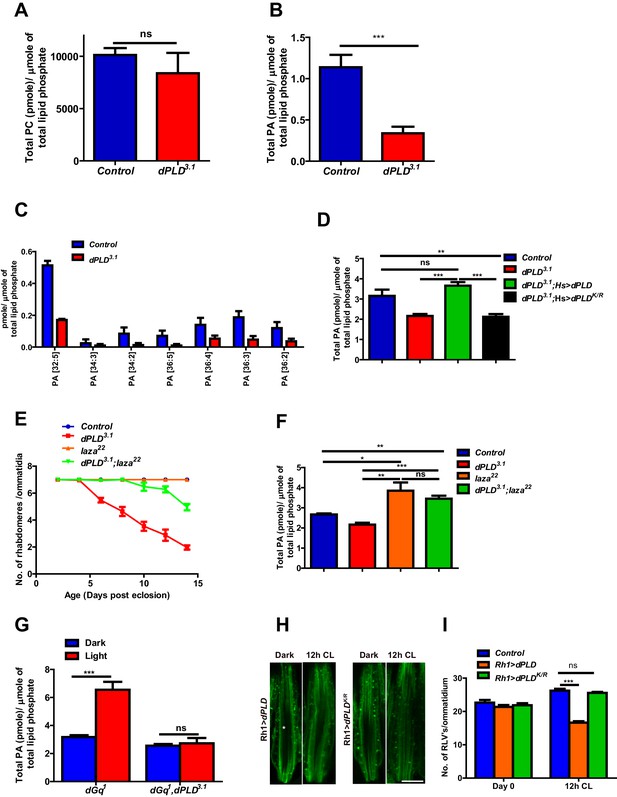
Phosphatidic acid levels and retinal degeneration in dPLD3.1.
(A) Total PC level in retinae of control and dPLD3.1. The X-axis represents the genotypes and the Y-axis shows the level of PC as pmole/µmole of total lipid phosphate present in the sample. n = 3. (B) Total PA level in retinae of control and dPLD3.1. The X-axis represents the genotypes and the Y-axis shows the level of PA as pmole/µmole of total lipid phosphate present in the sample. n = 3. (C) Molecular species of PA in retinae of control and dPLD3.1. X-axis shows the acyl chain composition of each species predicted from its monoisotopic peaks and corresponding elemental composition constraints. Y-axis shows the abundance of each species as pmole/µmole of total lipid phosphate present in the sample. n = 3. (D) PA levels in heads extracts of control, dPLD3.1, dPLD3.1;Hs>dPLD and dPLD3.1;Hs>dPLDK/R . n = 3. (E) Quantification of retinal degeneration seen in control, laza22, dPLD3.1 and dPLD3.1;laza22. n= 50 ommatidia taken from at least five separate flies. (F) PA levels in heads extracts of control, laza22, dPLD3.1 and dPLD3.1;laza22 n = 3. (G) PA levels from retinal extracts of Gq1 and Gq1,dPLD3.1. Flies were reared in complete darkness and post ecclosion one set of flies were shifted to bright illumination for 12 hr while the others kept in darkness for 12 hr. n = 3. (H) LS of retinae stained with Rh1 from Rh1>dPLD and Rh1>dPLDK/R. Rearing conditions are indicated at the top of panels. Scale bar:5 µm. (I) Quantification of RLVs from LS of retinae from control, Rh1>dPLD and Rh1>dPLDK/R. n = 10 ommatidia taken from three separate preps. Data are presented as mean ± SEM.
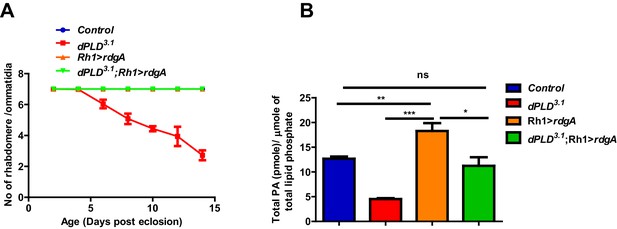
Rescue of dPLD3.1 phenotypes by overexpression of DGK.
(A) Quantification of retinal degeneration in control, Rh1>rdgA, dPLD3.1, dPLD3.1;Rh1>rdgA. The X-axis represents age of the flies and the Y-axis represents the number of rhabdomere visualized in each ommatidium. Error bars represents mean ± SEM from 50 ommatidia taken from at least five separate flies. (B) PA levels in heads extracts. Genotypes indicated on X-axis. Y-axis shows the total PA as pmole/µmole of total lipid phosphate present in the sample. Error bars indicate the mean ± SEM from three separate analyses.
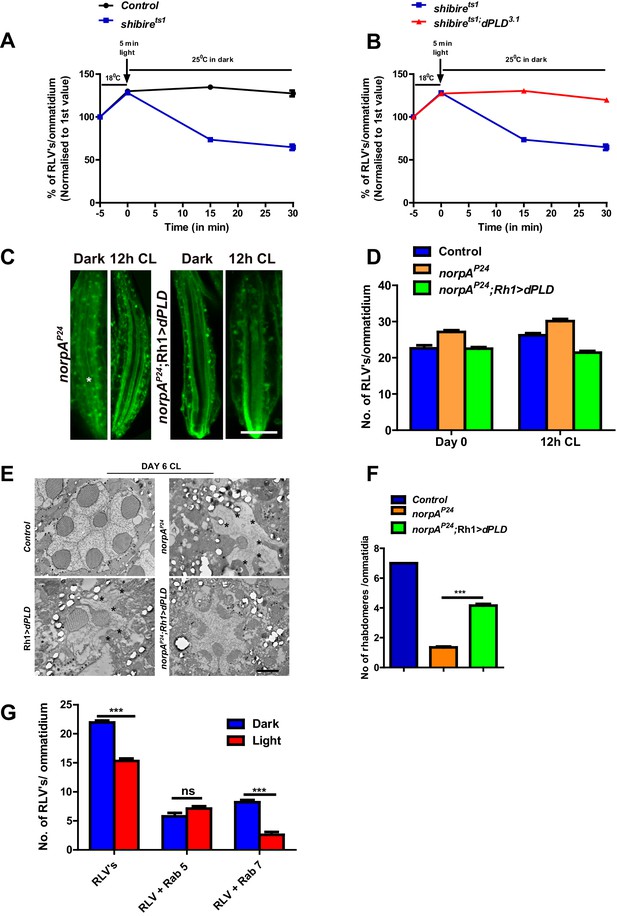
dPLD activity supports the removal of RLVs from the cell body during illumination.
(A) Quantification of RLVs from LS of retinae from control and shits1. n = 10 ommatidia taken from three separate preps. (B) Quantification of RLVs from LS of retinae from shits1 and shits1;dPLD3.1. n = 10 ommatidia taken from three separate preps. (C) LS of retinae stained with Rh1 from norpAP24 and norpAP24;Rh1>dPLD. Rearing condition is indicated at the top of each panel. Scale bar: 5 µm. (D) Quantification of RLVs from LS of retinae from control, norpAP24 and norpAP24;Rh1>dPLD. n = 10 ommatidia taken from three separate preps. (E) TEM images showing single ommatidium from control, norpAP24, Rh1>dPLD and norpAP24;Rh1>dPLD PRs of flies. * indicates the degenerated rhabdomere. Rearing condition is indicated on the top of the image. Scale bar: 1 µm. (F) Quantification of retinal degeneration in control, norpAP24 and norpAP24;Rh1>dPLD done using TEM images. The Y-axis represents the number of rhabdomeres visualized in each ommatidium. n = 50 ommatidia taken from at least two separate flies. (G) Quantification of RLVs from LS of retinae from Rh1>dPLD in dark vs light(12 h CL). The X-axis represents the population of vesicles positive for mentioned protein. Y-axis shows the number of RLV’s per ommatidium. n = 10 ommatidia taken from three separate preps. Data are presented as mean ± SEM.
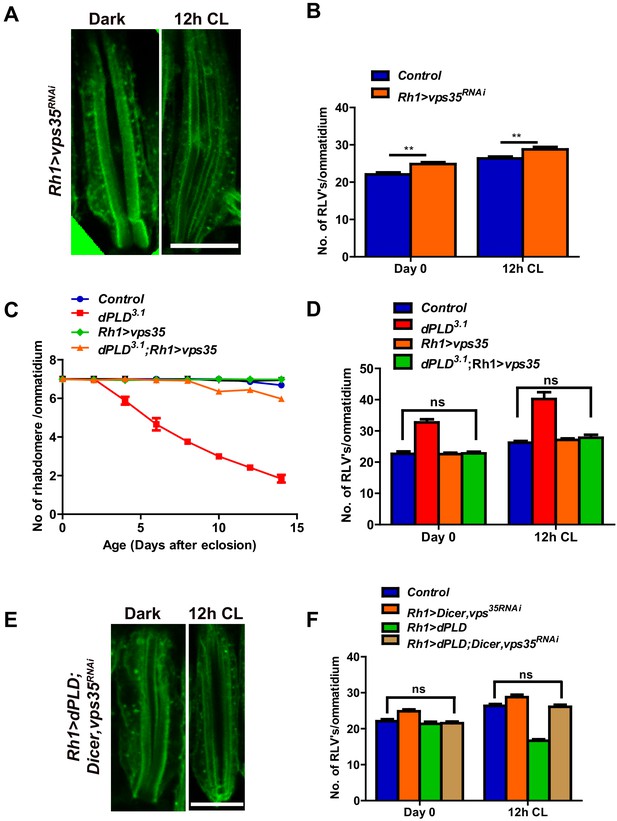
dPLD regulates clearance of RLVs via retromer function.
(A) LS of retinae stained with Rh1 from Rh1>Dicer,vps35RNAi. Rearing conditions are indicated at the top of panels. Scale bar:5 µm. (B) Quantification of RLVs from LS of retinae control and Rh1>Dicer,vps35RNAi. n = 10 ommatidia taken from three separate preps. (C) Quantification of retinal degeneration in control, dPLD3.1, Rh1>vps35 and dPLD3.1;Rh1>vps35. n = 50 ommatidia taken from at least five separate flies. (D) Quantification of RLVs from LS of retinae from control, dPLD3.1, Rh1>vps35 and dPLD3.1;Rh1>vps35. n = 10 ommatidia taken from three separate preps. (E) Longitudinal section of retinae stained with Rh1 Rh1>dPLD; Dicer,vps35RNAi. Rearing condition is indicated at the top of each panel. Scale bar: 5 µm. (F) Quantification of RLVs from longitudinal section of retinae from control, Rh1>Dicer,vps35RNAi, Rh1>dPLD and Rh1>dPLD; Dicer,vps35RNAi. n = 10 ommatidia taken from three separate preps. Data are presented as mean ± SEM.
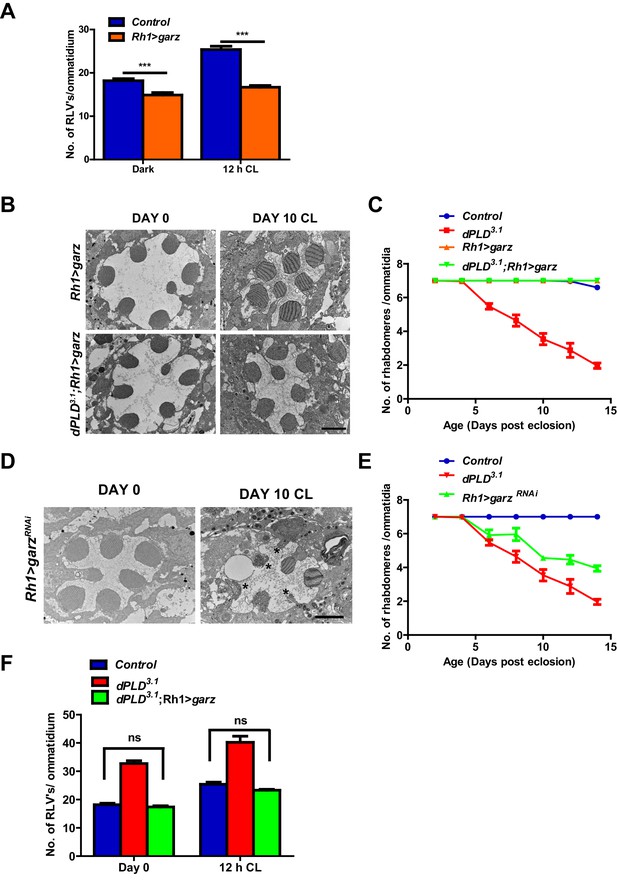
Arf1 activity and retinal degeneration in dPLD3. .
(A) Quantification of RLVs from longitudinal section of retinae from control and Rh1>garz. n = 10 ommatidia taken from three separate preps. (B) TEM images showing single ommatidium from Rh1>garz and dPLD3.1;Rh1>garz PRs of flies. Rearing condition is indicated on the top of the image. Scale bar: 1 µm. (C) Quantification of retinal degeneration in control, dPLD3.1, Rh1>garz and dPLD3.1;Rh1>garz. n = 50 ommatidia taken from at least five separate flies. (D) TEM images showing single ommatidium from Rh1>garzRNAi PRs of flies. Rearing condition is indicated on the image. Scale bar:1 µm. (E) Quantification showing the retinal degeneration in control, dPLD3.1 and Rh1>garzRNAi. n = 50 ommatidia taken from at least five separate flies. G) Quantification of RLVs from longitudinal section of retinae from control, dPLD3.1, dPLD3.1;Rh1>garz. n = 10 ommatidia taken from three separate preps. Data are presented as mean ± SEM.
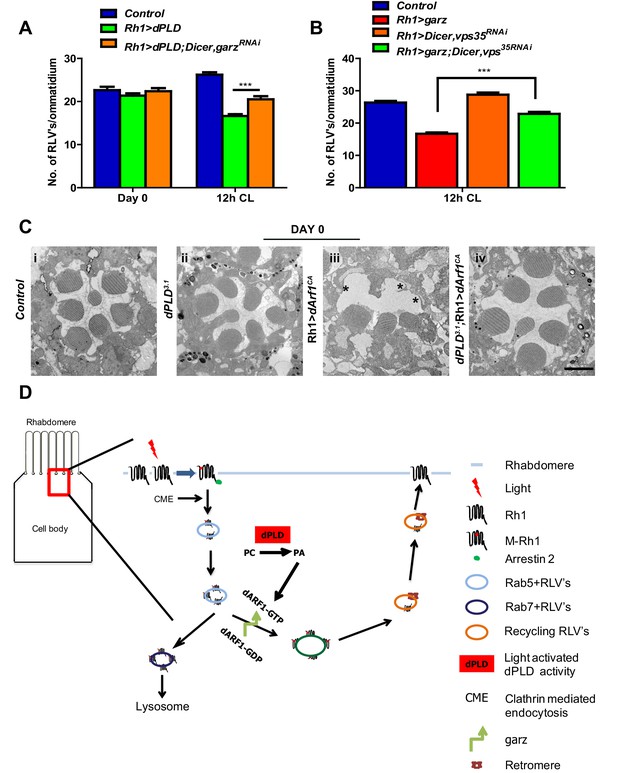
dPLD and garz are required for RLV clearance during illumination.
(A) Quantification of RLVs from longitudinal section of retinae from control, Rh1>dPLD, Rh1>dPLD; Dicer,Rh1>garzRNAi. n = 10 ommatidia taken from three separate preps. (B) Quantification of RLVs from longitudinal section of retinae from control, Rh1>garz, Rh1>Dicer,vps35RNAi and Rh1>garz;Dicer,vps35RNAi. n = 10 ommatidia taken from three separate preps. (C) TEM images showing single ommatidium from control and dPLD3.1,Rh1>Arf1CA and dPLD3.1;Rh1>Arf1CA PRs of day 0-old flies post-eclosion. Scale bar: 1 µm (D) A model of the light activated turnover of rhabdomere membranes in Drosophila photoreceptors. The cross-section of a PR is shown. The area indicated by the red box is enlarged to the left. PC-phosphatidylcholine, PA-phosphatidic acid, dARF1-GTP- GTP bound active ARF1, dARF1-GDP-GDP bound inactive Arf1, brown star indicates retromer, blue RLVs indicate endocytic compartment while orange RLVs indicate recycling compartment.





