Bacterial flagellar capping proteins adopt diverse oligomeric states
Figures
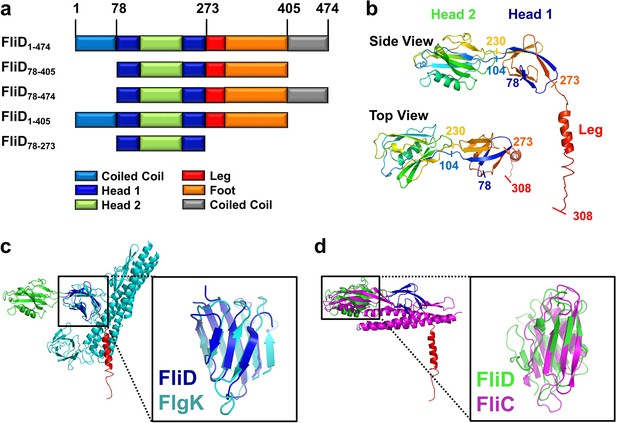
Crystal structure of Pseudomonas FliD reveals structural similarity to other flagellar proteins.
(a) Schematic representation of the FliD proteins used in these studies. Protein domain/region boundaries are labeled and are drawn approximately to scale. (b) Crystal structure of the Pseudomonas FliD78–405 monomer subunit with spectrum coloring from the N-terminus (blue) to the C-terminus (red). Head domain 1, head domain 2 and the leg region are indicated. (c) Superposition of the FliD78–405 crystal structure (domain coloring as in panel (a)) and Burkholderia FlgK/HAP1/hook filament capping protein (cyan). (d) Superposition of the FliD78–405 crystal structure (domain coloring as in panel (a)) and Pseudomonas flagellin/FliC (magenta).
-
Figure 1—source data 1
Crystallographic data collection, phasing and refinement statistics.
- https://doi.org/10.7554/eLife.18857.004

Protein sequence of FliD1–474.
The protein sequence of FliD from P. aeruginosa PAO1 is shown. The tertiary domain structure based on the presented X-ray crystal structure and the predicted secondary structure is indicated.
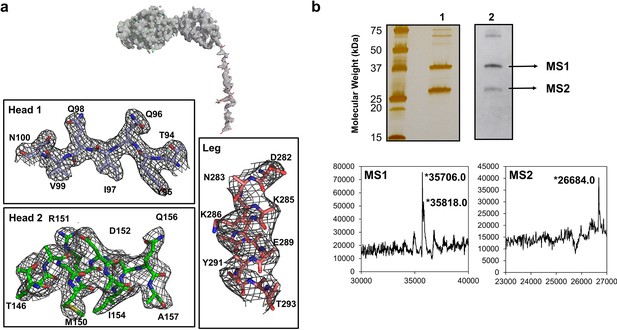
Electron density and protein degradation of FliD crystals.
(a) The overall electron density observed in the crystal structure of FliD is shown. Close-up views of representative residues in the head 1, head 2 and leg regions reveal low-quality and missing side-chain density in the leg region in comparison to the well-structured head domains. (b) SDS-PAGE (lane 1) and anti-His6-horseradish peroxidase (lane 2) Western blot analysis reveal C-terminal degradation of FliD78–405 in crystals. LC-MS analysis indicates the presence of the native FliD78–405 protein and degradation products in crystals of FliD78–405.
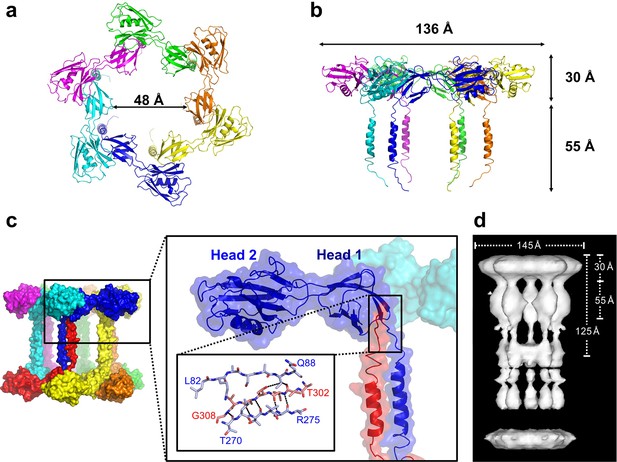
Pseudomonas FliD forms hexamers in crystals.
(a) Top view, cartoon representation of the FliD78–405 hexamer. Each monomer subunit is colored distinctly and inner diameter dimension is indicated. (b) Side view, cartoon representation of the FliD78–405 hexamer. Each monomer subunit is colored distinctly. Outer dimensions are indicated. (c) FliD78–405 hexamers as arranged in the crystal are stacked head-to-head and leg-to-leg (shown) in an alternating fashion, with residues 303–308 assembling in the Head 1 domain of an opposing molecule (close-up views) leading to a dodecameric crystal packing. (d) Cryo-EM structure of Salmonella FliD from (Maki-Yonekura et al., 2003) for comparison.
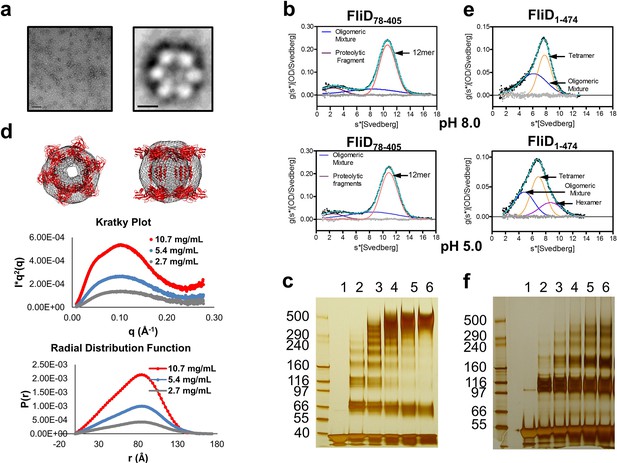
Pseudomonas FliD oligomerization.
(a) Negative stain EM image of FliD78–405: left, single particles (scale bar=1000 Å); right, class-averaged particle (scale bar=50 Å). (b) AUC analysis of FliD78–405 proteins at pH 8.0 (upper panel) and pH 5.0 (lowel panel) indicates that it forms dodecamers in solution. (c) Silver-stained SDS-PAGE after chemical crosslinking of FliD78–405. (d) SAXS analysis of FliD78–405. Kratky plot (I*q2 versus q) and radial distribution function calculated by GNOM, and SAXS envelopes calculated by DAMMIF, with superimposed crystal structures are shown for FliD78–405 at 10.7 mg/mL (red, used to calculate envelope), 5.4 mg/mL (blue) and 2.7 mg/mL (grey). (e) AUC analysis of full length FliD1–474 proteins at pH 8.0 (upper panel) and pH 5.0 (lower panel) indicates a mixture of oligomers, including tetramers and hexamers. (f) Silver-stained SDS-PAGE after chemical crosslinking of FliD1–474.
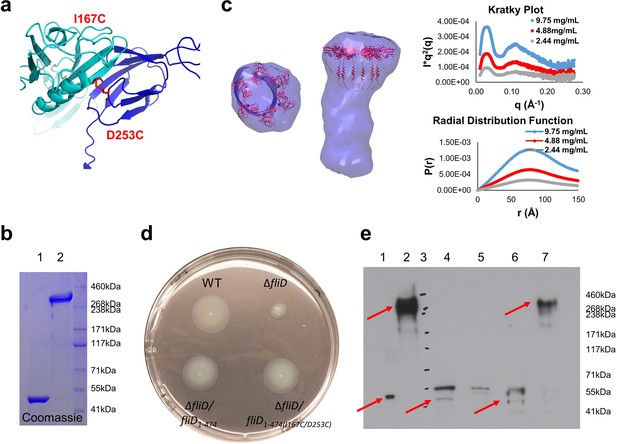
Stable hexameric DM1-FliD1–474 complements P. aeruginosa PAO1 dFliD transposon strain.
(a) Location of residues I167 and D253, which were predicted by the web server Disulfide by Design 2.0 (Craig and Dombkowski, 2013) to form stable disulfide bridges after mutation to cysteines. (b) FliD1–474(I167C/D253C) analyzed under reducing (lane 1) and non-reducing (lane 2) conditions by SDS-PAGE. (c) SAXS analysis of FliD1–474(I167C/D253C). Kratky plot (I*q2 versus q) and radial distribution function calculated by GNOM for 9.75 mg/mL (blue, used to calculate envelope), 4.88 mg/mL (red) and 2.44 mg/mL (grey). SAXS envelope calculated by DAMMIF with superimposed FliD78–405 crystal structure. (d) Swimming motility assay of wildtype PAO1 (WT), FliD transposon strain PW2975 (ΔfliD), ΔfliD complemented with FliD1–474 (ΔfliD/fliD1–474) or FliD1–474(I167C/D253C) (ΔfliD/fliD1–474(I167C/D253C)), respectively. (e) Western blot using anti-FliD scFv-Fc SH1579-B7 showing purified protein FliD1–474(I167C/D253C) under reducing (lane 1) and under non-reducing (lane 2) conditions. The presence of FliD in flagella preparations from wildtype PAO1 (lane 4), ΔfliD (lane 5), ΔfliD/fliD1–474(lane 6) and ΔfliD/fliD1–474(I167C/D253C) (lane 7) was analyzed under non-reducing conditions. The molecular weight standard is shown in lane 3 and the corresponding molecular weights are indicated on the right side of the blot. The 50 kDa and the 300 kDa bands representing FliD1–474 or hexameric FliD1–474(I167C/D253C), respectively, are indicated by red arrows.
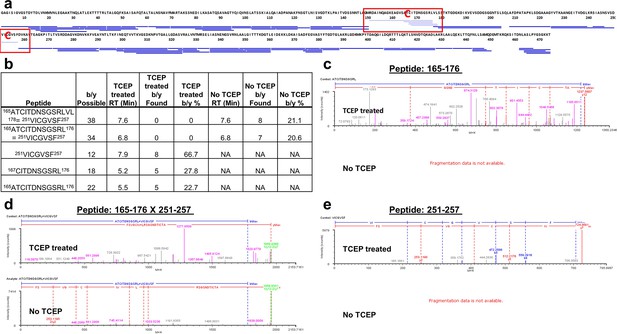
Analysis of FliD1–474(I167C/D253C) peptides following pepsin digestion under reducing and non-reducing conditions.
(a) Comparison of peptide coverage maps determined by LC/MS after pepsin digestion in the presence or absence of TCEP. Shown in light blue are the peptides that are identified only after treatment with TCEP, in dark blue are peptides identified both in the presence and in the absence of TCEP. The introduced cysteine residues are highlighted in red. (b) Peptides containing the introduced cysteines that were identified by Biopharmalynx in the presence and/or absence of TCEP are tabulated along with retention time (RT), and the number and percentage of b/y ions identified. (c–e) b/y fragment ions for three peptides surrounding the introduced cysteines as identified by Biopharmalynx in the presence and absence of TCEP. (c) and (e) show the b/y ions for peptides 165–176 and 251–257. (d) shows the b/y ions identified for the disulfide-bridged peptide 165–176 x 251–257.
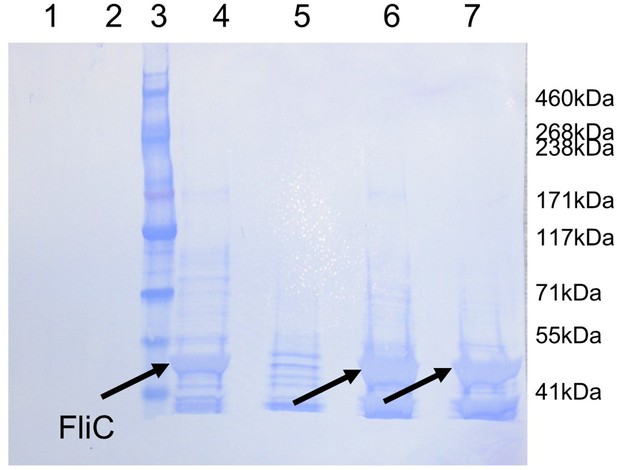
Western blot analysis of PAO1 strain flagella preparations.
Coomassie-stained membrane of the Western blot displayed in Figure 4e showing purified protein FliD1–474(I167C/D253C) under reducing (lane 1) and under non-reducing (lane 2) conditions. The presence of FliC in flagella preparations from wildtype PAO1 (lane 4), ΔfliD (lane5), ΔfliD/fliD1–474 (lane 6) and ΔfliD/fliD1–474(I167C/D253C)(lane 7) was analyzed under non-reducing conditions. The molecular weight standard is shown in lane 3 and the molecular weights are listed on the right side of the membrane. The 50 kDa FliC bands are indicated by black arrows.
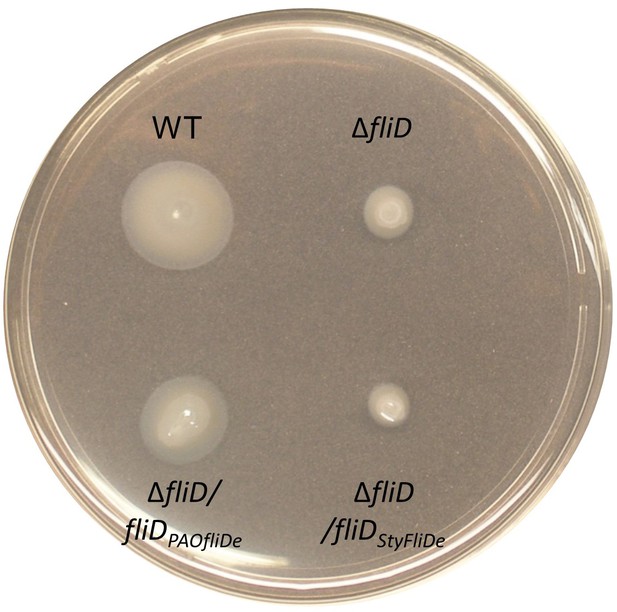
Swimming motility assay.
Swimming motility of wildtype PAO1 (WT), FliD transposon strain ΔfliD, ΔfliD complemented with FliD1–474 with an optimized codon usage for Echerichia coli expression (ΔfliD/fliDPAOfliDe) or Salmonella typhimurium FliD1–467 with an optimized codon usage for E. coli expression (ΔfliD/fliDStyFliDe).
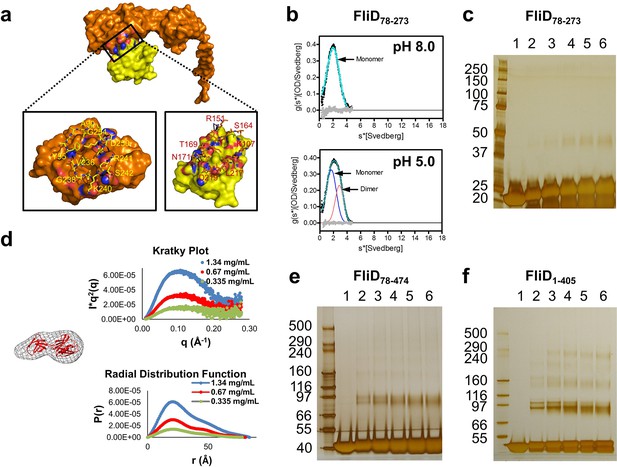
Molecular determinants of Pseudomonas FliD oligomerization reside outside of the stable head region.
(a) Intermolecular interface formed between head region monomer subunits, with an 'open book' rendering of the interface expanded below. Head domain 1 is yellow; domain 2 is orange; interface oxygen and nitrogen atoms are red and blue, respectively. (b) AUC analysis of the head region alone, FliD78–273, at pH 8.0 (upper panel) and pH 5.0 (lower panel) reveals a monomeric species at pH 8.0 and the additional minor presence of a dimeric species at pH 5.0. (c) Silver-stained SDS-PAGE after chemical crosslinking of FliD78–273. (d) SAXS analysis of FliD78–273. Kratky plot (I*q2 versus q) and radial distribution function calculated by GNOM and SAXS envelopes calculated by DAMMIF with superimposed crystal structures are shown for FliD7–273 at 1.34 mg/mL (blue), 0.67 mg/mL (red, used to calculate the envelope) and 0.335 mg/mL (green). (e) Silver-stained SDS-PAGE after chemical crosslinking of FliD78–474. (f) Silver-stained SDS-PAGE after chemical crosslinking of FliD1–405.
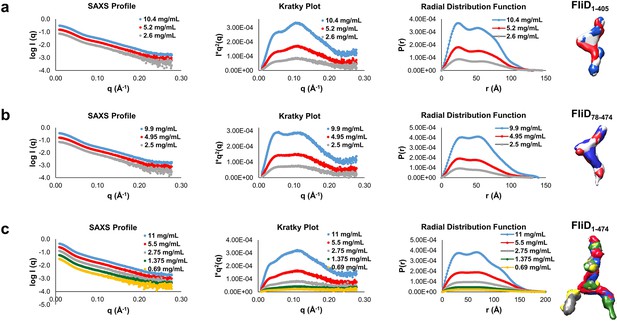
Small angle X-ray scattering (SAXS) data of FliD1–405, FliD78–474 and FliD1–474.
Log-scale intensity SAXS profiles, Kratky Plot (I × q2 versus q), radial distribution function calculated by GNOM and SAXS envelopes calculated by DAMMIF are shown for: (a) FliD1–405 at 10.4 mg/mL (blue), 5.2 mg/mL (red) and 2.6 mg/mL (grey); (b) FliD78–474 at 9.9 mg/mL (blue), 4.95 mg/mL (red) and 2.5 mg/mL (grey); and (c) FliD1–474 at 11 mg/mL (blue), 5.5 mg/mL (red), 2.7 mg/mL (grey), 1.38 mg/mL (green) and 0.69 mg/mL (yellow).
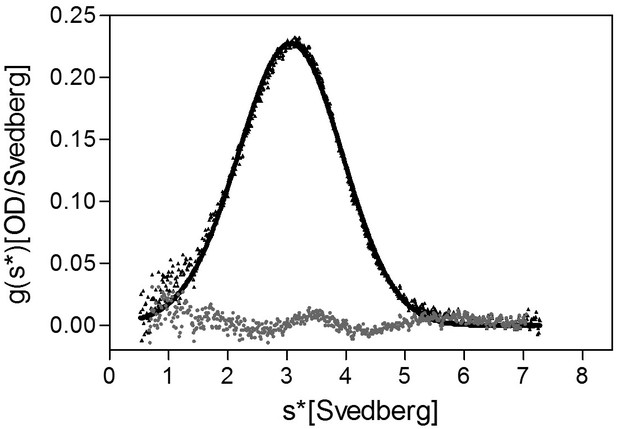
Analytical ultracentrifugation (AUC) analysis of FliD1–474 at pH 11.0.
AUC analysis of FliD1–474 at pH 11.0 indicates that the protein is monomeric at pH 11.0.
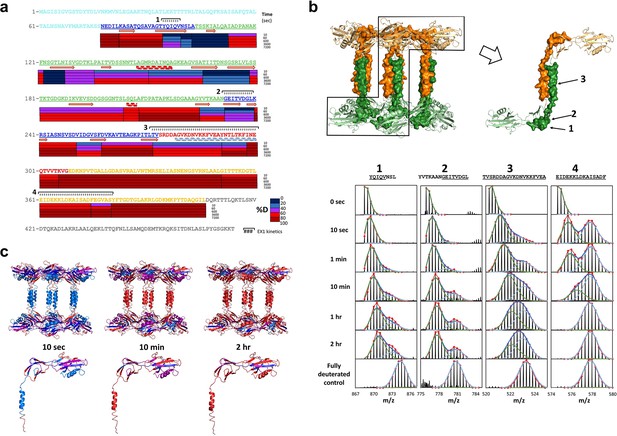
Regions of Pseudomonas FliD outside of the head domains and initial leg helix are highly dynamic.
(a) Hydrogen/deuterium exchange analysis of FliD78–405. Percent deuteration (%D) heat map is shown. Peptides exhibiting EX1 kinetics are indicated. (b) Mass spectra of four FliD peptides exhibit double isotopic envelopes characteristic of EX1 kinetics (below). Three of these peptides are mapped to the crystal structure (above; FliD hexamers are in green and gold). (c) Conformational stability as determined by hydrogen/deuterium exchange mapped to the crystal structure of FliD78–405 using the same color coding for %D as shown in (a).
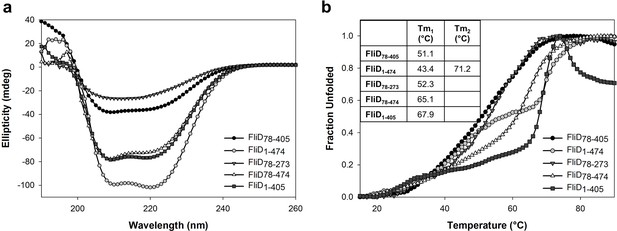
Circular dichroism analysis of FliD variants.
(a) CD spectra recorded from 190 to 260 nm and (b) melting curves recorded from 15 to 90°C are shown for FliD78–405, FliD1–474, FliD78–273, FliD78–474 and FliD1–405. Melting curves were recorded at 222 nm for all proteins except for FliD78–273, which was analyzed at 205 nm.
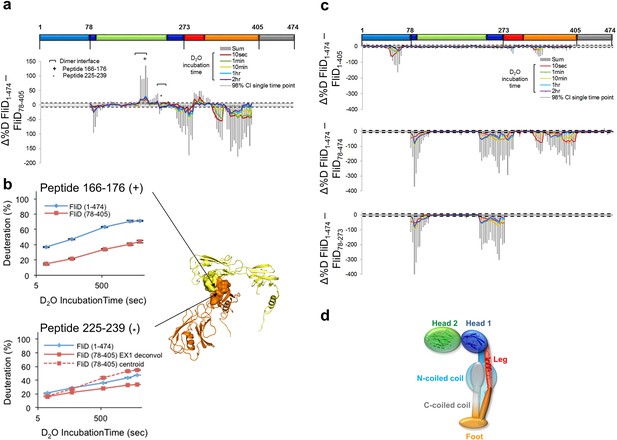
Interaction of Pseudomonas FliD regions.
(a) Difference plot of hydrogen/deuterium exchange data from full length FliD1–474 and the crystallized fragment, FliD78–405. (b) Hydrogen/deuterium exchange for peptides corresponding to residues 166–176 (top, marked by + in (a) and (b)) and residues 225–239 (bottom, marked by * in (a) and (b)). Positions of peptides 166–176 and 225–239 in the FliD78–405 crystal structure (right). (c) Difference plots of hydrogen/deuterium exchange data from full length FliD1–474 and the fragments missing only the C-terminal coiled coil, FliD1–405, only the N-terminal coiled coil, FliD78–474, or both the N- and C-terminal coiled coils and the leg domain, FliD78–273 (top, middle and bottom, respectively). (d) Schematic model of the FliD monomeric subunit showing the N-terminal coiled coil stabilizing the head 1 and foot domains and also interacting with the C-terminal coiled coil.

Hydrogen-deuterium exchange-mass spectrometry analysis of FliD variants.
Heat maps of hydrogen/deuterium exchange for FliD1–474, FliD78–405, FliD78–474, FliD1–405 and FliD78–273, from top to bottom, respectively.
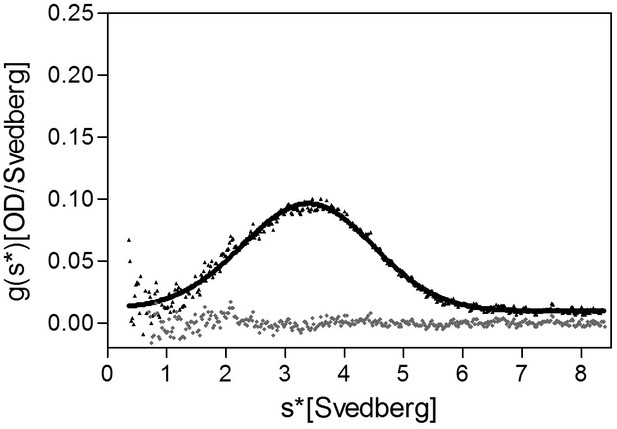
Analytical ultracentrifugation (AUC) analysis of FliD1–474 at 4 μM.
AUC analysis of FliD1–474 at a concentration of 4 μM at pH 8.0 indicates that the protein is predominantly monomeric in an equilibrium with dimeric species at 4 μM and pH 8.0.
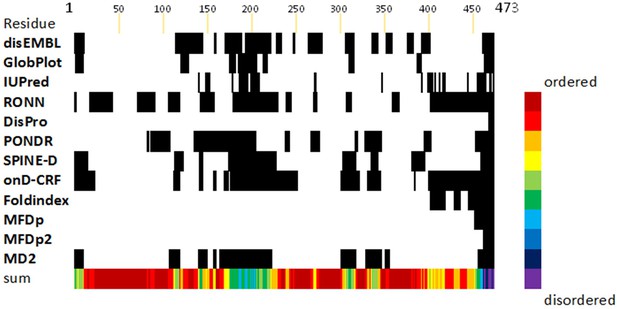
FliD intrinsic disorder analysis.
Intrinsic disorder prediction of FliD1–474 using publicly available algorithms (as indicated, left column). Averaging of the results (color, bottom row) was done giving all servers equal weight.





