Eukaryotic translation initiation factor 3 plays distinct roles at the mRNA entry and exit channels of the ribosomal preinitiation complex
Figures
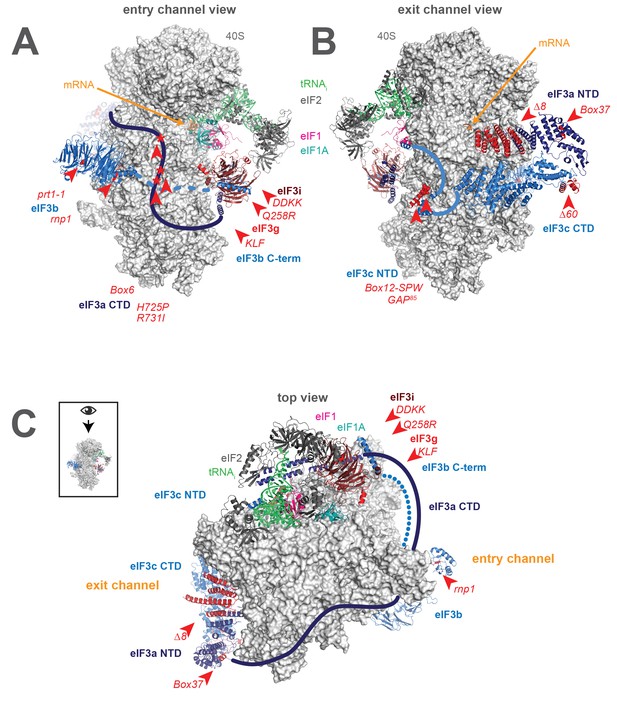
A library of S. cerevisiae eIF3 functional variants affected by mutations spanning all five essential subunits.
Three views of the cryo-EM structure of eIF3 bound to the partial yeast 48S PIC in the closed conformation (py48S-closed), as reported by Llacer, et al. The 40S subunit is shown in light grey, with resolved domains of eIF3 shown as ribbons and unresolved regions cartooned as solid or dotted lines: eIF3a is shown in dark blue, eIF3c and eIF3b in light blue, eIF3i in maroon, and eIF3g in red. The location of mutations throughout eIF3 are shown colored in red and indicated by red arrowheads; the eIF3i and eIF3g subunits are also depicted in red hues owing to their absence in the a/b/c subcomplex resulting from the DDKK mutation in eIF3i. The mRNA (orange), Met-tRNAi (green), eIF2 (charcoal), eIF1 (pink), and eIF1A (teal) are also visible in this structure. (A) View looking towards the mRNA entry channel, with the 40S intersubunit face on the right, and the solvent face on the left. (B) Opposite view looking towards the mRNA exit channel, with the 40S intersubunit face on the left, and the solvent face on the right. (C) View of the PIC looking down at the 40S head, showing the relative orientation of the entry- and exit-channel arms of eIF3 and its contacts on the 40S solvent face. The PIC is oriented such that the intersubunit face appears at the top, while the solvent face appears at the bottom.
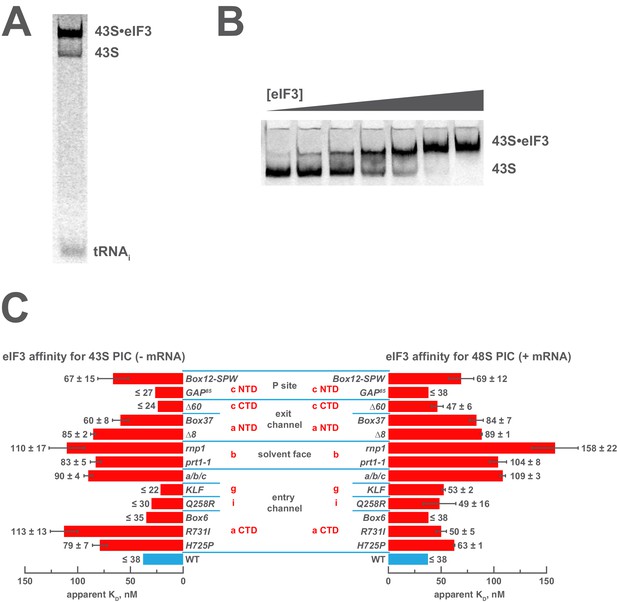
Mutations in the mRNA entry- or exit-channel arms of eIF3 or in eIF3b destabilize eIF3 binding to the PIC.
(A) Binding of ternary complex containing [35S]-Met-tRNAiMet to 40S ribosomal subunits in the presence of eIF1, eIF1A, and WT or mutant eIF3 was measured using a native gel-shift assay, which separates free [35S]-Met-tRNAiMet from that bound to 43S PICs alone or 43S PICs containing eIF3 (43S·eIF3). (B) The titration of eIF3 into reactions containing 43S PICs produces a well-resolved gel-shift that monitors the binding of eIF3 to the PIC. The amounts of [35S]-Met-tRNAiMet free, bound to 43S PICs, and bound to 43S∙eIF3 complexes were quantified and analyzed as described in Figure 2—figure supplement 1. (C) The apparent equilibrium dissociation constant (KD) of WT (blue bars) and eIF3 variants (red bars) for the PIC, both in the absence (left) and presence (right) of mRNA, obtained by fitting the data with the Hill equation (see Figure 2—figure supplement 2B). Bars and errors represent mean and SEM, respectively (as determined by individual fitting of each experiment), of ≥2 experiments. Owing to the conditions of these assays, apparent affinities ≤ 30 nM likely represent upper limits.
-
Figure 2—source data 1
Individual eIF3:43S and eIF3:48S dissociation constant measurements.
- https://doi.org/10.7554/eLife.20934.006
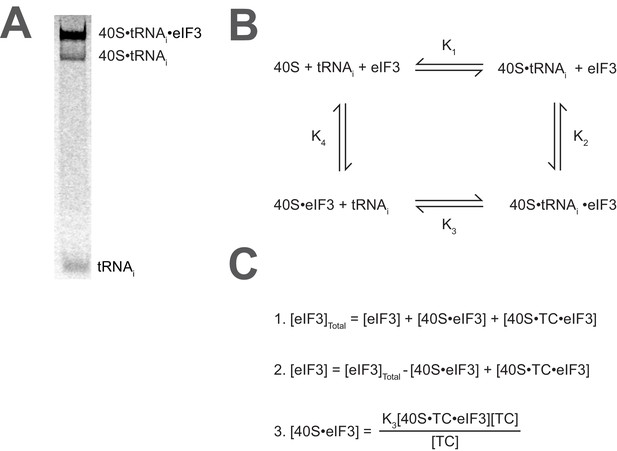
Calculating the affinity of eIF3 for PICs under conditions where free 40S subunits compete for binding.
(A) Native gel electrophoresis of PICs formed in the presence of [35S]-Met-tRNAiMet resolves free tRNAi and all other TC-bound species. (B) Thermodynamic cycle for the binding of both eIF3 and tRNAi (as TC) to 40S (as 40S•eIF1•eIF1A). At 40S concentrations near or below the dissociation constant for tRNAi binding to 40S (K1), free 40S subunits compete with 40S•tRNAi for eIF3 such that the concentration of total eIF3 is not equivalent to the concentration of free eIF3 necessary to determine its affinity for 40S•tRNAi (K2) (C) The concentration of free eIF3 ([eIF3]) can be determined from the total concentration of eIF3 ([eIF3]Total) and the concentration of all other eIF3-containing species. The concentration of eIF3 bound to PICs ([40S•TC•eIF3]) can be determined empirically, as this species contains radiometrically-labeled tRNAi and can thus be measured. The concentration of eIF3 bound to 40S subunits alone ([40S•eIF3]) cannot be measured directly, but can be calculated from the measured concentrations of eIF3 bound to PICs ([40S•TC•eIF3]), the concentration of free tRNAi ([TC]) (both of which also contain radiometrically-labeled tRNAi and can thus also be measured empirically), and the measured dissociation constant for the binding of tRNAi to the 40S•eIF3 complex (K3). This value can then be used, together with the measured concentrations of other eIF3-bound species, to determine the concentration of free eIF3 and model the data with a Langmuir isotherm to determine K2.
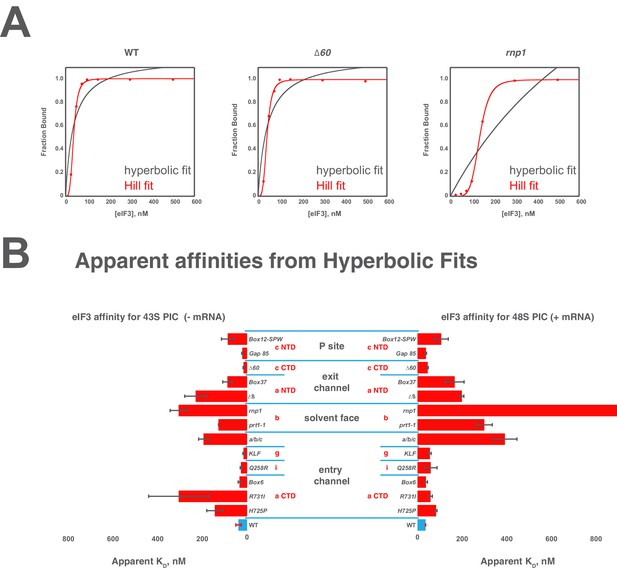
Comparison of hyperbolic and Hill equations for modeling eIF3 binding to the PIC.
(B) Comparison of representative hyperbolic (grey) and Hill (red) fits obtained upon modeling the binding of either WT, ∆60, or rnp1 eIF3 to 43S PICs. In all cases, the Hill fit more accurately models the binding data, and in the case of more severe binding defects, such as that observed for rnp1 eIF3, the hyperbolic (Langmuir) fit is unable to model the data. (A) Apparent affinities of WT (blue bars) and eIF3 variants (red bars) for 43S (left) and 48S (right) PICs, as determined by modeling binding data with a standard hyperbolic (Langmuir) binding isotherm. Bars and errors represent mean values and SEM, respectively (as determined by individual fitting of each experiment), of ≥2 experiments, with the exception of rnp1 eIF3 binding to 48S PICs (right), which could not be accurately modeled with a hypergolic (Langmuir) binding isotherm.
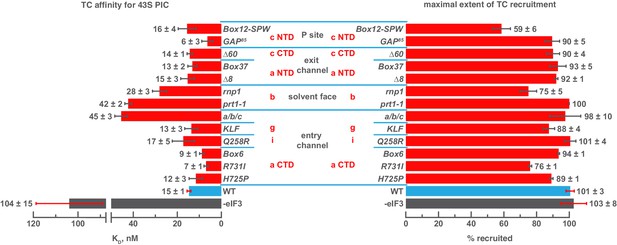
Mutations in the mRNA entry-channel arm of eIF3 or in eIF3b destabilize binding of TC to the PIC.
The KD of TC (left) or maximal extent of TC recruitment (right) observed for PICs assembled either in the absence of eIF3 (grey bar), or the presence of either WT (blue bar) or variant eIF3 (red bars). Bars and errors represent mean and SEM, respectively (as determined by individual fitting of each experiment), of ≥2 experiments.
-
Figure 3—source data 1
Individual measurements of ternary complex binding (dissociation constants and reaction extents) to 43S PICs.
- https://doi.org/10.7554/eLife.20934.010
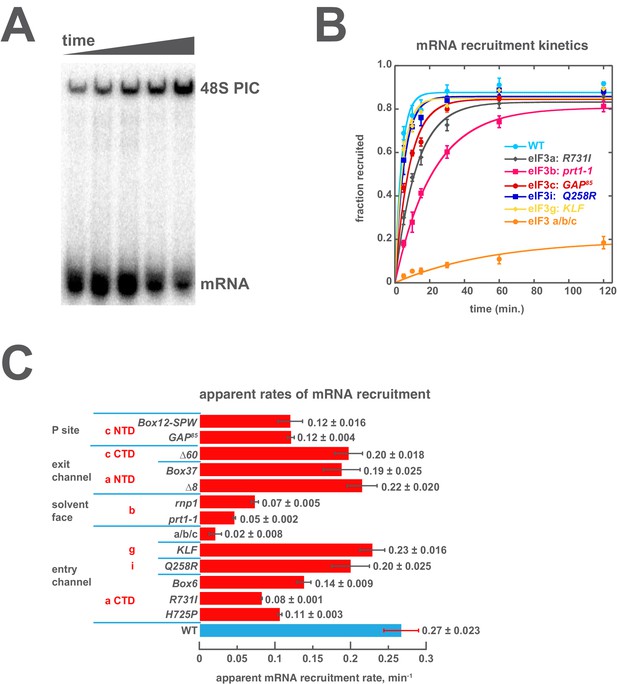
Mutations throughout the eIF3 complex compromise its ability to accelerate recruitment of capped native RPL41A mRNA to the PIC.
(A) mRNA recruitment reactions were quenched at appropriate time points on a running native gel to separate free [32P]-capped RPL41A mRNA from mRNA recruited to form 48S PICs. The bands were quantified to determine the fraction of total mRNA bound at each time point. (B) Individual time courses were fit with single-exponential rate equations to determine the observed rate constant for each experiment. (C) The observed rates of mRNA recruitment of capped RPL41A mRNA measured in the presence of WT (blue bar) and variants of eIF3 (red bars). Bars and errors represent mean values and SEM, respectively (as determined by individual fitting of each experiment), of ≥2 experiments.
-
Figure 4—source data 1
Individual observed apparent rates for RPL41A mRNA recruitment.
- https://doi.org/10.7554/eLife.20934.012
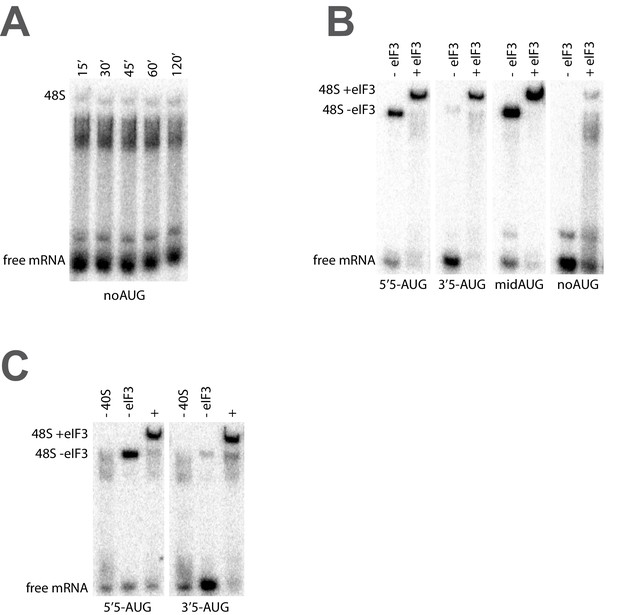
The recruitment of unstructured model mRNAs to the PIC depends on the presence of an AUG codon and 40S subunits.
(A) Representative mRNA recruitment gel showing recruitment time course for the noAUG unstructured model mRNA, which lacks a start codon. (B) Comparison of mRNA recruitment gels at 120’, both in the presence and absence of eIF3, for the 5’5-AUG, 3’5-AUG, and midAUG mRNAs, all of which contain an AUG codon, and the noAUG mRNA, which lacks a start codon. (C) mRNA recruitment gels for the 5’5-AUG (left panel) and 3’5-AUG mRNAs (right panel), showing lanes containing reactions lacking either 40S subunits (- 40S, left lane) or eIF3 (- eIF3, middle lane), or in the presence of both (+, right lane).
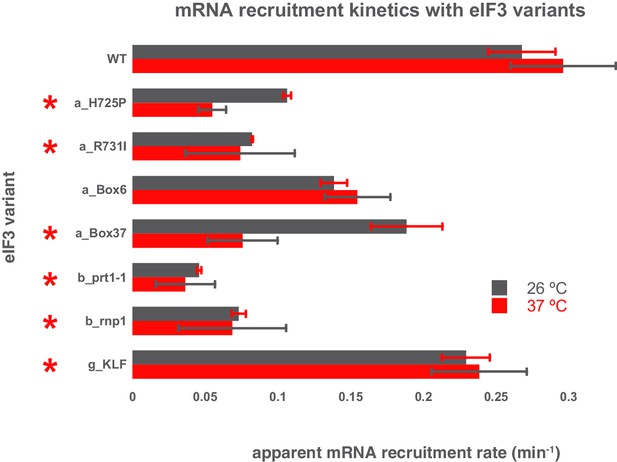
Comparison of the kinetics of mRNA recruitment for WT and variant eIF3 at 26°C and 37°C.
The apparent rates of mRNA recruitment of capped RPL41A mRNA observed in the presence of WT and eIF3 variants that confer Ts- phenotypes in vivo (denoted with asterisks) as well as the box6 variant, which does not. Bars represent apparent recruitment rates at either 26°C (grey bars) or 37°C (red bars). Bars and errors represent mean values and SEM, respectively (as determined by individual fitting of each experiment), of ≥2 experiments.
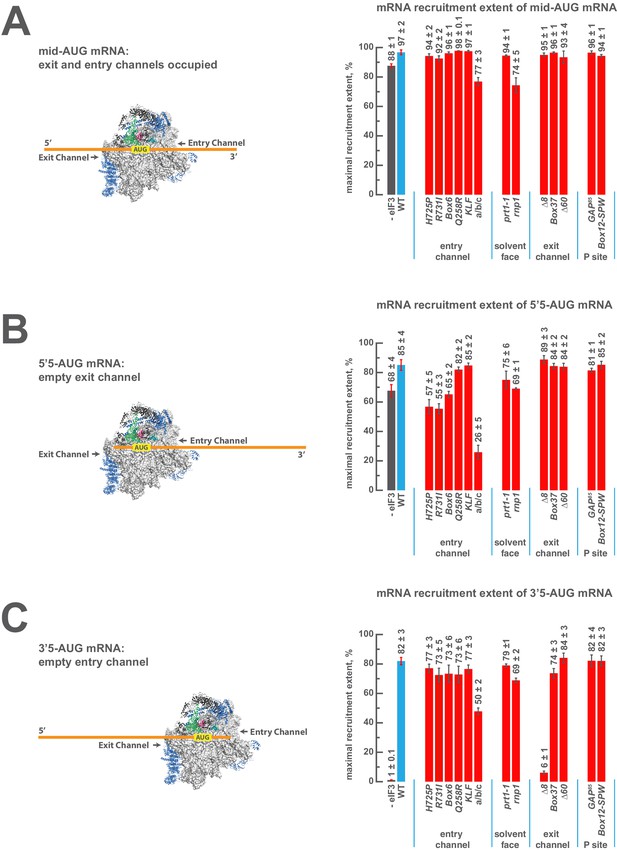
eIF3 strongly stabilizes binding of mRNA at the exit channel of the PIC in a manner dependent on the eIF3a NTD.
(A) The maximal extent of mid-AUG mRNA recruited in the absence of eIF3 (grey bar), or in the presence of either WT (blue bar) or variant eIF3 (red bars). The locations of sequences 5’ or 3’ of the AUG in the PIC, with AUG in the P site, are shown schematically on the left, indicating that both the mRNA entry and exit channels of the PIC are fully occupied. Bars and errors represent mean and SEM, respectively, of ≥2 experiments. (B) The maximal extent of 5’5-AUG mRNA recruited in the absence of eIF3, or in the presence of either WT or variant eIF3. As shown on the left, 5’5 AUG mRNA programs a recruited complex in which only the mRNA entry channel is fully occupied, and thus is sensitized to changes in that channel. (C) The maximal extent of 3’5-AUG mRNA recruited in the absence of eIF3, or in the presence of either WT or variant eIF3. The 3’5-AUG programs a recruited complex in which only the mRNA exit channel is occupied (left), and thus is sensitized to changes in that channel.
-
Figure 5—source data 1
Individual measurements of recruitment extent.
- https://doi.org/10.7554/eLife.20934.016
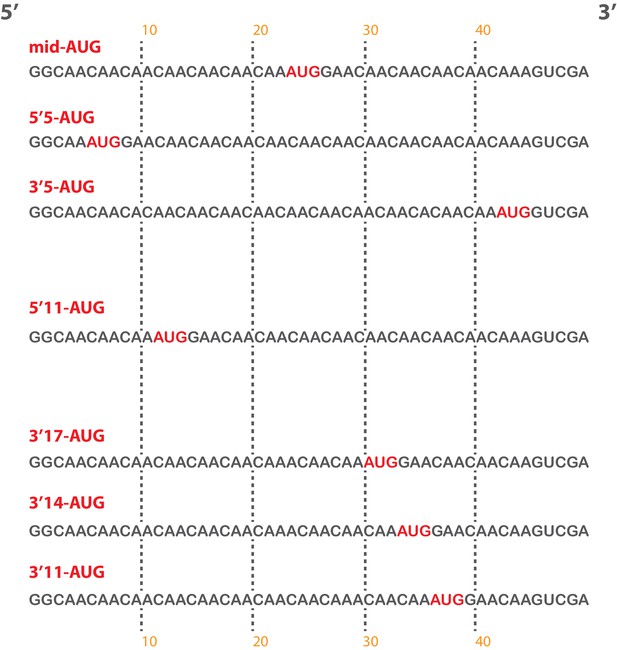
Short, unstructured model mRNAs with distinct start codon positions.
Specific sequences of the unstructured model mRNAs used in this study, with the AUG start codon highlighted in red. All model mRNAs are 50 nucleotides in length, and primarily contain unstructured CAA repeats. Each mRNA was in-vitro transcribed and capped, as described in the materials and methods.
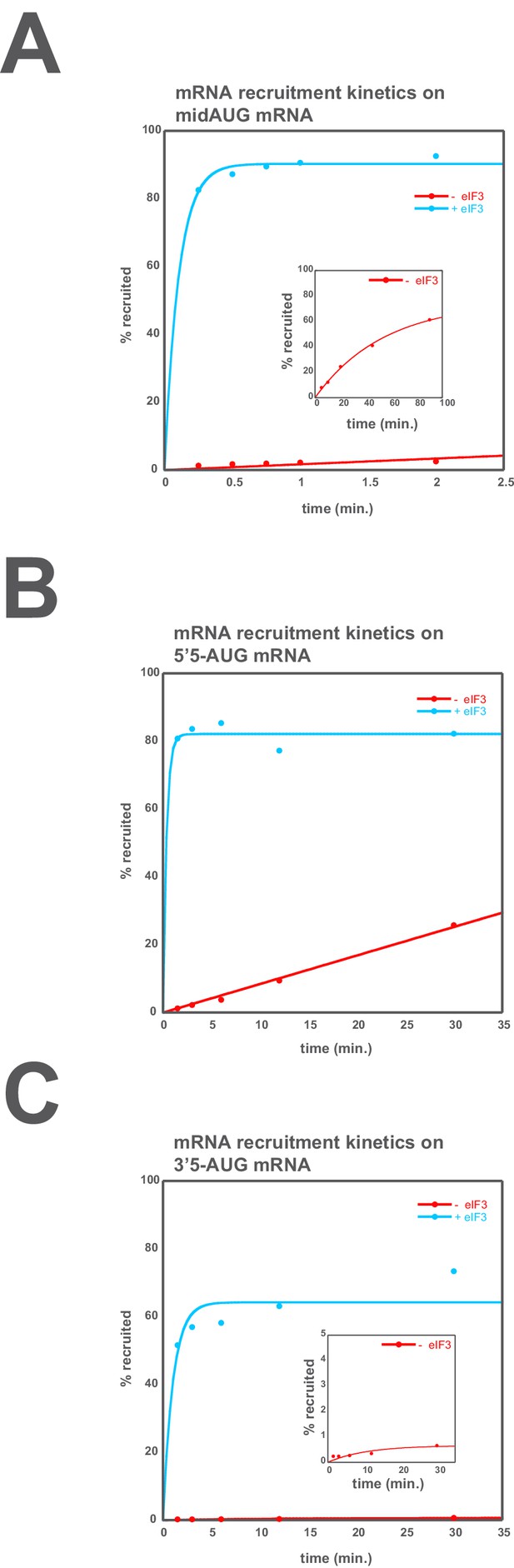
eIF3 accelerates the recruitment of unstructured model mRNAs.
Apparent mRNA recruitment kinetics observed in both the presence (blue) and absence (red) of eIF3, for the mid-AUG (A), 5’5-AUG (B), and 3’5-AUG (C) mRNAs. Extended time courses are shown (as insets) for both the midAUG and 3’5-AUG mRNAs.
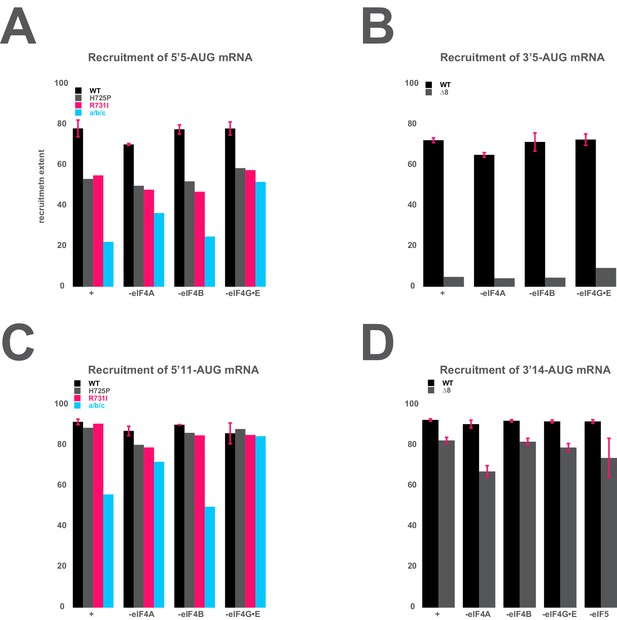
The mRNA recruitment extent defects observe for entry- and exit-channel variants of eIF3 are not exacerbated in the absence of the eIF4 factors.
(A) The extent of 5’5-AUG mRNA recruited to the PIC in the presence of WT eIF3 (black bars) and eIF3 entry-channel variants (grey, pink, and blue bars), in either the presence all factors (+), or the absence of either eIF4A, eIF4B, or eIF4G•E. (B) The extent of 3’5-AUG mRNA recruited to the PIC in the presence of WT (black bars) and ∆8 (grey bars) eIF3, in either the presence of all factors (+), or the absence of either eIF4A, eIF4B, or eIF4G•E. (C) The extent of 5’11-AUG mRNA recruited to the PIC in the presence of WT eIF3 (black bars) and eIF3 entry-channel variants (grey, pink, and blue bars), in either the presence of all factors (+), or the absence of either eIF4A, eIF4B, or eIF4G•E. (D) The extent of 3’14-AUG mRNA recruited to the PIC in the presence of WT (black bars) and ∆8 (grey bars) eIF3, in either the presence of all factors (+), or the absence of either eIF4A, eIF4B, or eIF4G•E.
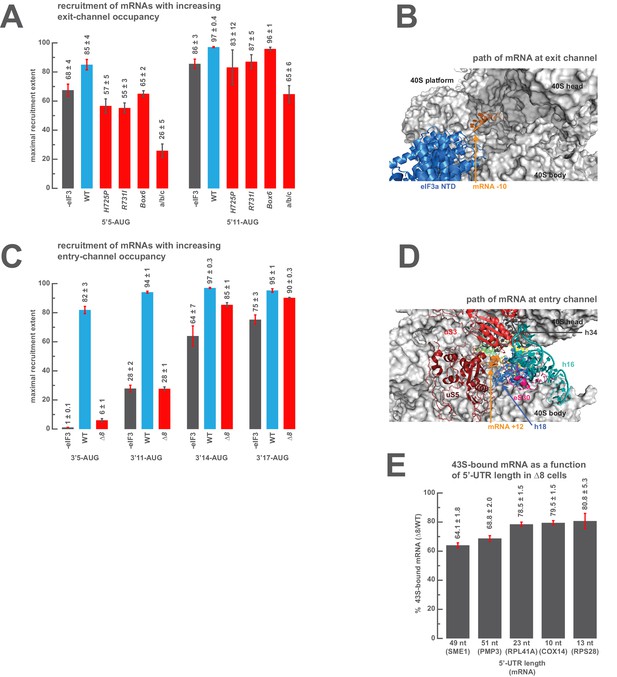
The presence of sufficient mRNA in the opposite channel rescues the destabilization of mRNA binding caused by mutations in the entry- or exit-channel arms of eIF3.
(A) Bars represent the maximal extent of recruitment of mRNAs that program complexes with either 5 (5’5-AUG) or 11 (5’11-AUG) nucleotides in the mRNA exit channel, in the absence of eIF3 (grey bars) or in the presence of either WT eIF3 (blue bars) or variants with mutations near the mRNA entry channel (red bars). Bars and errors represent mean and SEM, respectively, of ≥2 experiments. (B) The path of mRNA (orange) at the exit channel of the 40S subunit (grey) in the py48S-closed structure (Llácer et al., 2015). mRNA is visualized up to the −10 nucleotide, where it emerges from the exit channel pore and is located approximately 15 Å from the eIF3a NTD (blue). (C) Bars represent the maximal extent of recruitment of mRNAs with 5 (3’5-AUG), 11 (3’11-AUG), 14 (3’14-AUG), or 17 (3’17-AUG) nucleotides in the entry channel in the absence of eIF3 (grey bars) or in the presence of either WT eIF3 (blue bars) or the ∆8 variant (red bars). Bars and errors represent mean and SEM, respectively, of ≥2 experiments. (D) The path of mRNA (orange) at the entry channel of the 40S subunit (grey) in the py48S PIC lacking eIF3 (Hussain et al., 2014). mRNA is visualized up to the +12 nucleotide, where it emerges from the 40S latch composed of h34 (charcoal), h18 (blue), and uS3 (red), and projects towards a constriction formed by uS5 (burgundy), uS3 (red), and eS30 (pink), near h16 (teal). (E) The amount of mRNA cross-linked to PICs in ∆8 cells, as a percentage of mRNA cross-linked to PICs in WT cells, determined for five mRNAs with decreasing 5’-UTR lengths. Bars and errors represent means and SD from two independent biological replicates.
-
Figure 6—source data 1
Individual measurements of recruitment extent and of the amount of mRNA associated with 43S PICs in vivo.
- https://doi.org/10.7554/eLife.20934.021
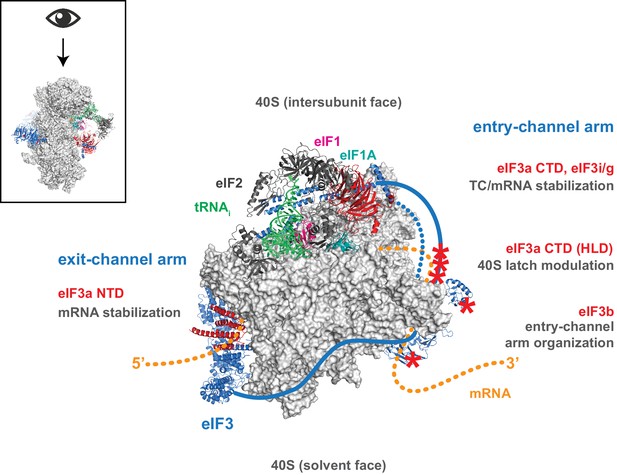
Distinct roles for the mRNA entry- and exit-channel arms of eIF3.
A proposed model for the roles of the eIF3 entry- and exit-channel arms during translation initiation. The py48S-closed complex is shown from above (boxed schematic), looking down at the head of the 40S (light grey), with the solvent face at the bottom, and the intersubunit face at the top with Met-tRNAi (green), eIF2 (dark grey), eIF1 (pink), and eIF1A (teal) bound to the decoding center. eIF3 is shown in blue, with areas of interest highlighted in red or denoted with red asterisks; resolved regions of eIF3 are depicted as ribbons, with unresolved regions cartooned as solid or dashed lines. The mRNA is cartooned in orange, entering the PIC though the entry channel at right, and exiting at the pore near the platform at left. Our results suggest that the eIF3 exit-channel arm, and specifically the eIF3a NTD, is critical for stabilizing mRNA binding to the PIC at the exit channel, whereas the eIF3a CTD (unresolved regions appear as solid blue line) and eIF3i/g enhance the stability of mRNA interactions at the entry-channel. The entry-channel arm and its attachment to the solvent face via eIF3b (unresolved regions appear as dashed blue line) is also important for stabilizing TC binding to the PIC and promoting steps that control the kinetics of mRNA recruitment, perhaps as a function of modulating the 40S latch.
Tables
Putative mechanistic defects attributed to eIF3 variants. Table describing the putative mechanistic defects attributed to each eIF3 variant within our library. Variants are organized according to their general location within the PIC (left column) and by the subunit or subunit domain (second column from left) in which the associated mutation occurs (allelic designation in third column from left). The specific identity of the mutation affecting each variant (second column from right) and the putative mechanistic defects previously attributed to each variant (final column, see text for references) are described.
| Location | Subunit | Variant | Mutation | Putative defects |
|---|---|---|---|---|
| | eIF3a/Tif32 CTD | H725P | H725P | mRNA recruitment scanning codon recognition |
| R731I | R731I | |||
| Box6 | Alanine substitution of 692LDLDTIKQVI701 | |||
eIF3i/Tif34 | Q258R | Q258R | scanning | |
DDKK | D207K/D224K | 40S binding 43S PIC formation scanning start codon recognition | ||
| eIF3g/Tif35 | KLF | K194A/L235A/F237A | scanning | |
eIF3b/Prt1 | prt1-1 | S555F | 40S binding 43S PIC formation scanning start codon recognition | |
| rnp1 | Alanine substitution of 124KGFLVE130 | |||
| | eIF3c/Nip1 CTD | ∆60 | C-terminal truncation of aa 753–813 | 40S binding 43S PIC formation |
| eIF3a/Tif32 NTD | ∆8 | N-terminal truncation of aa 1–200 | 40S binding 43S PIC formation re-initiation | |
Box37 | Alanine substitution of 361PTRKEMLQSI370 | mRNA recruitment | ||
| eIF3c/Nip1 NTD | GAP85 | Deletion of aa 60–144 | eIF1 binding start codon recognition | |
| Box12 -SPW | K111S/K114P/K116W |
Yeast strains used in this study.
Strain | Genotype | Source or reference |
|---|---|---|
BY4741 | MATa his3∆0 leu2∆0 met15∆0 ura3∆0 | |
BJ5465 | MATa ura3-52 trp1 leu2- Δ1 his3-Δ200 pep4::HIS3 prb1-Δ1.6 can1 GAL+ | Elizabeth Jones |
WLCY12 | MATa ura3-52 trp1 leu2-Δ1 his3-Δ200 pep4::HIS3 prb1-Δ1.6 can1 GAL+ tif32Δ::kanMX4 p3908 sc[TIF32, NIP1, URA3] | This study |
PBH42 | MATa ura3-52 trp1 leu2-Δ1 his3-Δ200 pep4::HIS3 prb1-Δ1.6R can1 TIF35-del GAL+ (YEp-TIF35) | This study |
PBH44 | MATa ura3-52 trp1 leu2-Δ1 his3-Δ200 pep4::HIS3 prb1-Δ1.6R can1 prt1-Δ GAL+ (YEpPRT1-U) | This study |
PBH51 | MATa ura3-52 trp1 leu2-Δ1 his3-Δ200 pep4::HIS3 prb1-Δ1.6R can1 GAL+ tif32Δ::KanMX4 (YEpPRT1-His-TIF34-TIF35-W, YEptif32-Δ8-NIP1-L) | This study |
PBH143 | MATa ura3-52 trp1 leu2-Δ1 his3-Δ200 pep4::HIS3 prb1-Δ1.6R can1 GAL+ tif32Δ::KanMX4 (YEpPRT1-His-TIF34-TIF35-W, Yep3tif32_BOX37-NIP-L) | This study |
PBH54 | MATa ura3-52 trp1 leu2-Δ1 his3-Δ200 pep4::HIS3 prb1-Δ1.6R can1 prt1-Δ GAL+ (YEpprt1-rnp1-His-TIF34-TIF35-W, YEpTIF32-NIP1-L) | This study |
PBH55 | MATa ura3-52 trp1 leu2-Δ1 his3-Δ200 pep4::HIS3 prb1-Δ1.6R can1 prt1-Δ GAL+ (YEpprt1-1-His-TIF34-TIF35-W, YEpTIF32-NIP1-L) | This study |
PBH56 | MATa ura3-52 trp1 leu2-Δ1 his3-Δ200 pep4::HIS3 prb1-Δ1.6R can1 nip1-Δ GAL+ (YEpNIP1-His-U) | This study |
PBH57 | MATa ura3-52 trp1 leu2-Δ1 his3-Δ200 pep4::HIS3 prb1-Δ1.6R can1 tif34-ΔGAL+ (YEp-TIF34) | This study |
PBH65 | MATa ura3-52 trp1 leu2-Δ1 his3-Δ200 pep4::HIS3 prb1-Δ1.6R can1 nip1-Δ GAL+ (YEpPRT1-His-TIF34-TIF35-W, YEpTIF32-nip1-Δ60-L) | This study |
PBH66 | MATa ura3-52 trp1 leu2-Δ1 his3-Δ200 pep4::HIS3 prb1-Δ1.6R can1 nip1-Δ GAL+ (YEpPRT1-His-TIF34-TIF35-W, Yep3TIF32-nip_GAP85-L) | This study |
PBH67 | MATa ura3-52 trp1 leu2-Δ1 his3-Δ200 pep4::HIS3 prb1-Δ1.6R can1 nip1-Δ GAL+ (YEpPRT1-His-TIF34-TIF35-W, Yep3TIF32-nip_Box12-SPW-L) | This study |
PBH71 | MATa ura3-52 trp1 leu2-Δ1 his3-Δ200 pep4::HIS3 prb1-Δ1.6R can1 tif34-Δ GAL+ (YEpPRT1-His-tif34_DDKK-TIF35-W, YEpTIF32-NIP1-L) | This study |
PBH73 | MATa ura3-52 trp1 leu2-Δ1 his3-Δ200 pep4::HIS3 prb1-Δ1.6R can1 tif34-Δ GAL+ (YEpPRT1-His-tif34_Q258R-TIF35-W, YEpTIF32-NIP1-L) | This study |
PBH76 | MATa ura3-52 trp1 leu2-Δ1 his3-Δ200 pep4::HIS3 prb1-Δ1.6R can1 TIF35-Δ GAL+ (YEpPRT1-His-TIF34-tif35_KLF-W, YEpTIF32-NIP1-L) | This study |
WLCY11 | MATα ura3-52 trp1 leu2-Δ1 his3-Δ200 pep4::HIS3 prb1-Δ1.6R can1 GAL+ tif32Δ::kanMX4 p3908 sc[TIF32, NIP1, URA3] | This study |
WLCY13 | MATa ura3-52 trp1 leu2-Δ1 his3-Δ200 pep4::HIS3 prb1-Δ1.6 can1 GAL+ tif32Δ::kanMX4 p3908 sc[TIF32, NIP1, URA3] p3127 hc[PRT1-His, TIF34-HA, TIF35-FLAG, LEU2] | This study |
WLCY14 | MATα ura3-52 trp1 leu2-Δ1 his3-Δ200 pep4::HIS3 prb1-Δ1.6R can1 GAL+ tif32Δ::kanMX4 p5237 hc[TIF32, NIP1, TRP1] | This study |
WLCY15 | MATα ura3-52 trp1 leu2-Δ1 his3-Δ200 pep4::HIS3 prb1-Δ1.6R can1 GAL+ tif32Δ::kanMX4 p5238 hc[tif32-H725P, NIP1, TRP1] | This study |
WLCY16 | MATα ura3-52 trp1 leu2-Δ1 his3-Δ200 pep4::HIS3 prb1-Δ1.6R can1 GAL+ tif32Δ::kanMX4 p5239 hc[tif32-R731I, NIP1, TRP1] | This study |
WLCY17 | MATα ura3-52 trp1 leu2-Δ1 his3-Δ200 pep4::HIS3 prb1-Δ1.6R can1 GAL+ tif32Δ::kanMX4 p5240 hc[tif32-box6, NIP1, TRP1] | This study |
WLCY18 | MATa/MATα ura3-52/ura3-52 trp1/trp1 leu2-Δ1/leu2-Δ1 his3-Δ200/his3-Δ200 pep4::HIS3/pep4::HIS3 prb1-Δ1.6R/prb1-Δ1.6R can1/can1 GAL+/GAL+ tif32Δ::kanMX4/tif32Δ::kanMX p5237 hc[TIF32, NIP1, TRP1] p3127 hc[PRT1-His, TIF34-HA, TIF35-FLAG, LEU2] | This study |
WLCY19 | MATa/MATα ura3-52/ura3-52 trp1/trp1 leu2-Δ1/leu2-Δ1 his3-Δ200/his3-Δ200 pep4::HIS3/pep4::HIS3 prb1-Δ1.6R/prb1-Δ1.6R can1/can1 GAL+/GAL+ tif32Δ::kanMX4/tif32Δ::kanMX p5238 hc[tif32-H725P, NIP1, TRP1] p3127 hc[PRT1-His, TIF34-HA, TIF35-FLAG, LEU2] | This study |
WLCY20 | MATa/MATα ura3-52/ura3-52 trp1/trp1 leu2-Δ1/leu2-Δ1 his3-Δ200/his3-Δ200 pep4::HIS3/pep4::HIS3 prb1-Δ1.6R/prb1-Δ1.6R can1/can1 GAL+/GAL+ tif32Δ::kanMX4/tif32Δ::kanMX p5239 hc[tif32-R731I, NIP1, TRP1] p3127 hc[PRT1-His, TIF34-HA, TIF35-FLAG, LEU2] | This study |
WLCY21 | MATa/MATα ura3-52/ura3-52 trp1/trp1 leu2-Δ1/leu2-Δ1 his3-Δ200/his3-Δ200 pep4::HIS3/pep4::HIS3 prb1-Δ1.6R/prb1-Δ1.6R can1/can1 GAL+/GAL+ tif32Δ::kanMX4/tif32Δ::kanMX p5240 hc[tif32-box6, NIP1, TRP1] p3127 hc[PRT1-His, TIF34-HA, TIF35-FLAG, LEU2] | This study |
Plasmids used in this study. High-copy, low-copy, and single-copy plasmids are designated hc, lc. and sc, respectively.
Plasmid | Description | Source of reference |
|---|---|---|
pLV10 | hc vector containing nip1Δ::hisG::URA3::hisG disruption cassette | |
p4473 (pRS315-prt1-rnp1-His) | lc LEU2 vector containing prt1-rnp1-His | |
pTZ-tif34Δ | hc vector containing tif34Δ::hisG::URA3::hisG disruption cassette | |
B3289 | hc vector containing prt1Δ::hisG::URA3::hisG disruption cassette | |
B444 | hc vector containing tif35Δ::hisG::URA3::hisG disruption cassette | |
YCpMJ-MET-NIP1-W | sc TRP1 vector containing NIP1 under MET3 promotor | |
YCpAH-MET-PRT1-W | sc TRP1 vector containing PRT1 under MET3 promotor | This study |
pGAD-PRT1 | PRT1 ORF cloned into pGAD424 | |
YCplac22MET-W | sc cloning vector with conditional MET3 promoter, TRP1 plasmid from YCplac22 | K. Nasmyth |
pLPY202 | lc URA3 vector containing prt1-1-His | |
YEp112 | hc TRP1 vector | |
YEp111-MET-TIF34-L | hc LEU2 vector containing TIF34 under MET3 promotor | This study |
pGAD-TIF34 | TIF34 ORF cloned into pGAD424 | |
YCpLV06 | sc LEU2 vector containing TIF32 under MET3 promotor | |
YCplac111 | sc LEU2 vector | |
pGAD-TIF35 | TIF35 ORF cloned into pGAD424 | |
YEp111-MET-TIF35-L | hc LEU2 vector containing TIF35 under MET3 promotor | This study |
YEpNIP1-His-U | hc URA3 vector containing NIP1-His | |
YEpprt1-1-His-TIF34-TIF35-W | hc TRP1 vector containing prt1-1-His, TIF34 and TIF35 | This study |
YEpPRT1-His-TIF34-TIF35-W | hc TRP1 vector containing PRT1-His, TIF34 and TIF35 | This study |
YEpprt1-rnp1-His-TIF34-TIF35-W | hc TRP1 vector containing prt1-rnp1-His, TIF34 and TIF35 | This study |
YEpPRT1-His-TIF34-TIF35-U | hc URA3 vector containing PRT1-His, TIF34 and TIF35 | This study |
YEpPRT1-U | hc URA3 vector containing PRT1 | This study |
YEplac195 | hc URA3 vector | |
YEpPRT1-TIF35-U | hc URA3 vector containing PRT1 and TIF35 | This study |
YEpPRT1-TIF34-TIF35-U | hc URA3 vector containing PRT1, TIF34 and TIF35 | This study |
YEpTIF32-NIP1-L | hc LEU2 vector containing TIF32 and NIP1 | This study |
YEpTIF32-NIP1-His-L | hc LEU2 vector containing TIF32 and NIP1-His | This study |
YEpNIP1-His-L | hc LEU2 vector containing NIP1-His | |
YEpTIF32-nip1-Δ60-L | hc LEU2 vector containing TIF32 and NIP1 truncated by 60 amino acid residues in its CTD | This study |
YEptif32-Δ8-NIP1-His-L | hc LEU2 vector containing TIF32 with the deletion of first 200aa and NIP1-His | This study |
pRSTIF32-∆8-His | lc TIF32-∆8-His[200–964], LEU2 plasmid from pRS315 | |
YEptif32-Δ8-NIP1-L | hc LEU2 vector containing TIF32 truncated by 200 amino acid residues in its NTD and NIP1 | This study |
YEp-TIF34 | hc URA3 vector containing TIF34 | |
YEp-TIF35 | hc URA3 vector containing TIF35 | |
Yep3tif32_BOX37-NIP-L | hc LEU2 vector containing TIF32 with ten alanine substitution in its NTD and NIP1 | This study |
YCp-a/tif32-Box37-H | sc tif32-Box37-His in LEU2 plasmid, from YCplac111 | |
YEp3TIF32-nip_Box12-SPW-L | hc LEU2 vector containing TIF32 and NIP1 containing K113S/K116P/K118W substitutions in Box12 | This study |
YCpNIP1-Box12-SPW | sc NIP1-His containing K113S/K116P/K118W substitutions in Box12; LEU2 plasmid from YCplac111 | |
Yep3TIF32-nip_GAP85-L | hc LEU2 vector containing TIF32 and NIP1 containing deletion of 85 residues (Val60- Asn144) | This study |
YCpNIP1-GAP85 | sc NIP1-His containing deletion of 85 residues (Val60- Asn144); LEU2 plasmid from YCplac111 | {Karaskova:2012ce} |
YEpPRT1-His-tif34_Q258R-TIF35-W | hc TRP1 vector containing PRT1-His, tif34-Q258R and TIF35 | This study |
YCpL-i/tif34-HA-3 | sc tif34-HA-Q258R in LEU2 plasmid from YCplac111 | |
YEpPRT1-His-tif34_DDKK-TIF35-W | hc TRP1 vector containing PRT1-His, TIF34 containing D207K and D224K mutations and TIF35 | This study |
YCp-i/TIF34-D207K-D224K-HA | sc TIF34-HA containing D207K and D224K mutations in LEU2 plasmid from YCplac111 | |
YEpPRT1-His-TIF34-tif35_KLF-W | hc TRP1 vector containing PRT1-His, TIF34 and tif35-KLF | This study |
YCp22-g/TIF35-KLF | sc TIF35-KLF-His inTRP1 plasmid from YCplac22 | |
pWCB27 | hc NIP1, tif32-H725P, URA3 plasmid, from YEplac195 | This study |
pWCB28 | hc NIP1, tif32-R731I, URA3 plasmid, from YEplac195 | This study |
pWCB29 | hc NIP1, tif32-box6, URA3 plasmid, from YEplac195 | This study |
pWLCB01 | tif32-H725P-His in lc LEU2 plasmid, from pRS315 | |
p4577 (pRS-a/tif32-R731I-His-L) | tif32-R731I-His in lc LEU2 plasmid, from pRS315 | |
pRS-a/tif32-box6-His-L | tif32-box6-His in lc LEU2 plasmid, from pRS315 | |
p3131 (pLPY-NIP1-TIF32) | hc NIP1, TIF32, URA3 plasmid, from YEplac195 | |
p3908 (YCp-a/TIF32-His-U) | sc TIF32-His URA3 plasmid | |
pWCB23 | hc NIP1, TIF32, TRP1 plasmid, from YEplac195 | This study |
pWCB24 | hc NIP1, tif32-H725P, TRP1 plasmid, from YEplac195 | This study |
pWCB25 | hc NIP1, tif32-R731I, TRP1 plasmid, from YEplac195 | This study |
pWCB26 | hc NIP1, tif32-box6, TRP1 plasmid, from YEplac195 | This study |
pUT11 | URA3 to TRP1 marker swap plasmid | |
p3127 (pLPY-PRT1His-TIF34HA-TIF35FL-Leu) | hc PRT1-His, TIF34-HA, TIF35-FLAG, LEU2 plasmid from YEplac 181 |
Primers used in this study.
Primer name | Primer sequence (5´to 3´) |
|---|---|
MJNIP1Sal-Xba | AGCAAAGAGTCAAGAAAGTTTCTA |
NIP1-noHis-BamHI-R | TTATTGGATCCTCAACGACGATTTGATGGTGGGTTAAGACG |
PBNIPD60-BamHI-R | CCCCGGATCCTCACACCTTATTTTCTGGAAGATC |
PRT1-8xHis-F | ATTTCATGAACTTACGGGCTTGTATGTAAA |
PRT1-8xHis-PstI-R | TTTACATACAAGCCCGTAAGTTCATGAAATCTGCAGTTAGTGGTGGTGGTGGTGGTGGTGGTGTTCGACCTTTTCCTTTGTTTCTTCCAAAAC |
PRT1-SpeI | GACGTGAAGACTAGTGTGTTC |
PRT1-SphI-R | TGCGTCTACTTGTGCATGCAT |
PBTIF34SacI | AGTGAATTCGAGCTCCTTATTCAGCGG |
TIF34-SmaI-R | TTATTCCCGGGTTAATTAGCTTCTTGCATGTGCTC |
TIF35-SphI | AATAAGCATGCACAAGTAGACGCACCTAAAAG |
TIF35-SmaI-R | TTATTCCCGGGCTATTCCTTAACCTTAGGTTTGGA |
AH-PRT1-SalI | ATATATGTCGACATGACTACCGAGACTTTCGAA |
AH-PRT1-HindIII-R | GTCCAAAGCTTCTGAATAAGCCCT |
OJ-TIF34 | attcggatccatatgaaggctatcaaattaacaggt |
OJ-TIF34-R | atctctgcagttaattagcttcttgcatgtgctcttt |
LVPRT1-5’ | aataaAAGCTTAGGGCGATCTGCTACAGGAAGCTA |
LVPRT1-3’R | aataaCTGCAGGCATGCATACAAAGATAATAGAGCCTATT |
LVTIF35-5’ | aataaGCATGCACAAGTAGACGCACCTAAAAGTCC |
LVTIF35-3’R | aataaGTCGACACATATTCACGACAGCCTCTGAGC |
LVTIF34-5’ | aataaGAGCTCAAActcgagGAGCTGATAAAACCCTACACTACGGTGTAA |
LVTIF34-3’R | CGCCTATGCCCGGGAACTGCATAC |





