Concerted regulation of ISWI by an autoinhibitory domain and the H4 N-terminal tail
Figures
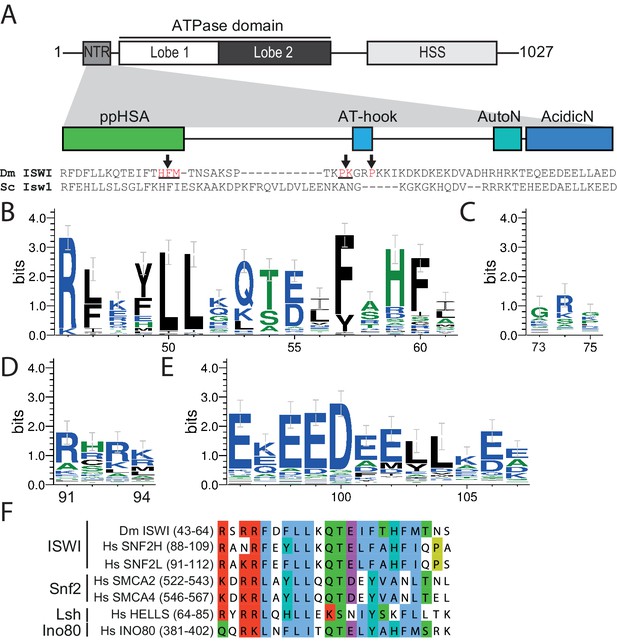
The NTR of ISWI contains several conserved sequence motifs.
(A) Schematic representation of the ISWI domain composition. The grey inset shows the sequence and motifs of the NTR. Arrows indicate amino acids within the NTR of Drosophila ISWI that crosslinked to Lobe 2 of the ATPase domain (Table 1). HSS, HAND-SANT-SLIDE domain. (B–E) Sequence logos showing the sequence conservation of (B) ppHSA, (C) AT-hook, (D) AutoN, and (E) AcidicN. X-Axis values are amino acid positions in D. melanogaster ISWI. See Figure 1—figure supplement 1 for full alignment. (F) Alignment of the ppHSA motif of Drosophila (Dm) ISWI with the human (Hs) ISWI homologs SNF2H and SNF2L and representatives of unrelated remodeler families.
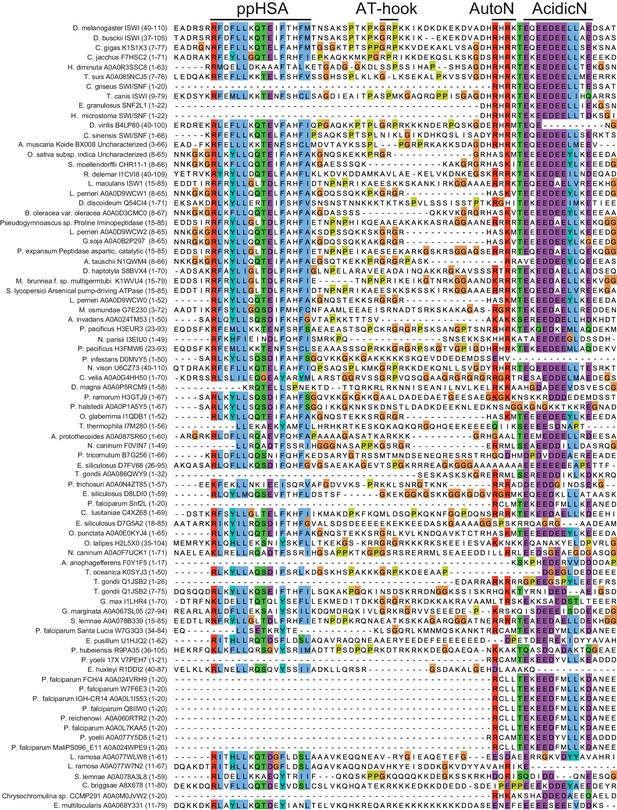
Alignment of ISWI homologs from various organisms.
Search for homologous proteins and alignment was done using HHblits (toolkit.tuebingen.mpg.de/hhblits). 26 Sequences lacking an NTR were manually deleted from the alignment.
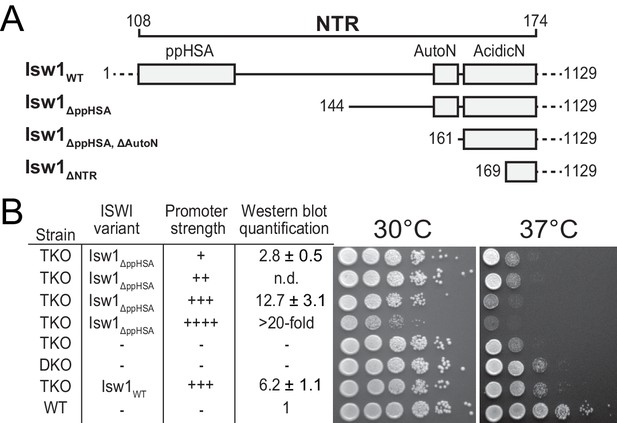
Functional importance of the NTR of yeast Isw1 in vivo.
(A) Successive N-terminal truncation mutants of Isw1. Note that Isw1ΔNTR lacked the entire N-terminus up to the first seven residues of AcidicN (Figure 1E). (B) Complementation assay with Isw1ΔppHSA. A yeast strain lacking ISW1, ISW2 and CHD1 (TKO) was transformed with Isw1 derivatives under control of promoters of varying strengths. In comparison to a strain lacking only ISW2 and CHD1 (DKO), Isw1WT fully complemented the growth phenotype at elevated temperatures (37°C). In contrast, Isw1ΔppHSA did not complement at any expression level. Results for other Isw1 variants can be found in Figure 2—figure supplement 1. Growth was assessed by spotting tenfold serial dilutions of liquid cultures.
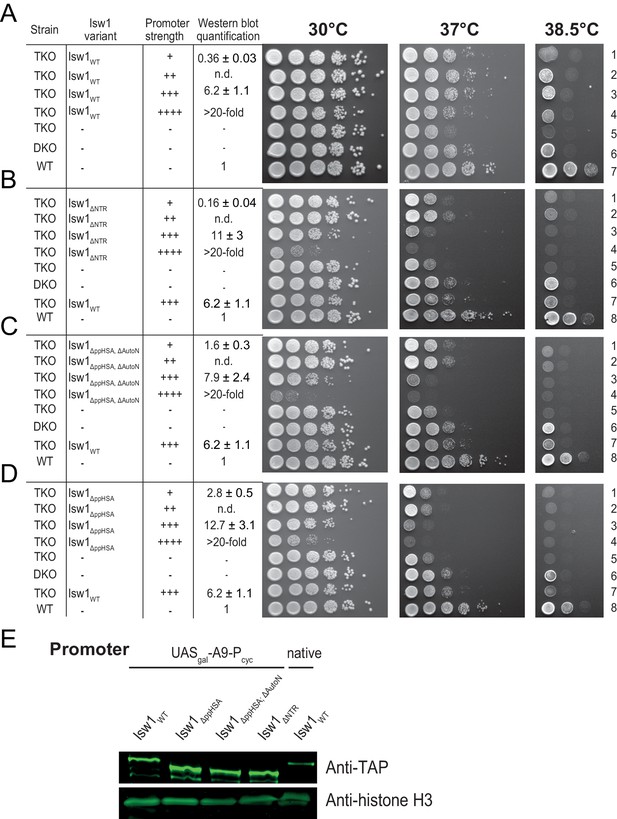

Complementation assay with N-terminal truncation variants of Isw1.
(A–D) Growth assays as in Figure 2B. Expression levels were estimated by Western analysis (see panel E). Results in D (30°C and 37°C) are replotted from Figure 2. (E) Exemplary Western blot to determine relative expression levels (tabulated in A–D) using an Anti-TAP antibody. TAP-tagged Isw1 variants under control of the indicated promoter were expressed. Their expression level was normalized against genomically TAP-tagged wild-type Isw1. Errors are minimal and maximal values of two technical replicates. H3 served as a loading control.
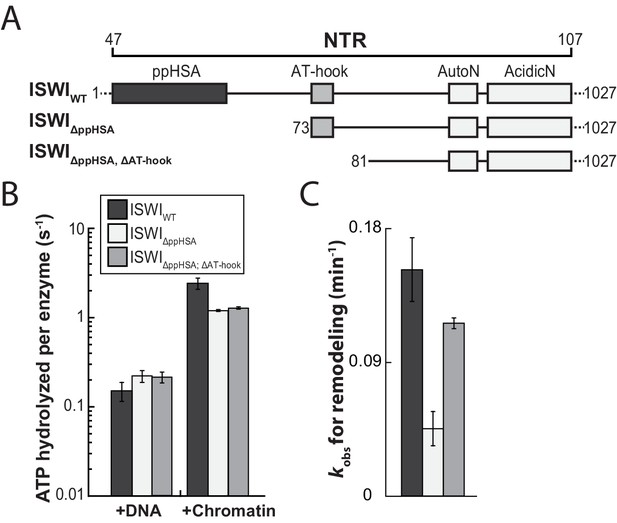
The ppHSA motif is largely dispensable for catalysis.
(A) N-terminal truncation mutants of Drosophila ISWI. (B) DNA- and nucleosome-stimulated ATP turnover. ATPase rates were measured in the presence of saturating concentrations of ATP (1 mM), DNA (0.2 g/l) or nucleosomes (0.1 g/l). Errors for nucleosome-stimulated rates of ISWI deletion mutants are minimal and maximal values of two independent measurements, and s.d. for all other measurements (n ≥ 4). ATPase rates in absence of nucleic acids were <0.022 s−1 for all ISWI variants (data not shown). (C) Remodeling activity was determined by measuring the accessibility changes of a unique KpnI restriction site in a 25-mer nucleosomal array (100 nM nucleosomes, 300 nM enzyme). Errors are s.d. (n ≥ 3) except for ISWIΔppHSA; ΔAT-hook for which minimal and maximal values of two independent measurements are shown. Raw data of the remodeling assay can be found in Figure 3—figure supplement 3. Color code as in panel B.
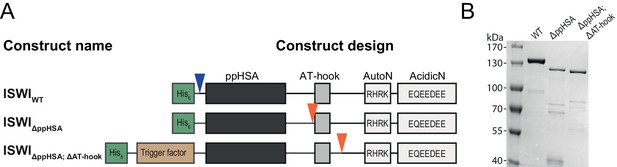
Cloning and purification of N-terminal truncation variants of Drosophila ISWI.
(A) Construct design. Only the NTR region and N-terminally fused affinity (His6) and solubility tags (Trigger factor) are shown (not to scale). Blue arrowheads indicate a TEV cleavage site, orange arrowheads indicate a 3C protease cleavage site that were used to cleave off the tag. (B) Coomassie stained SDS-PAGE of purified recombinant ISWI variants.
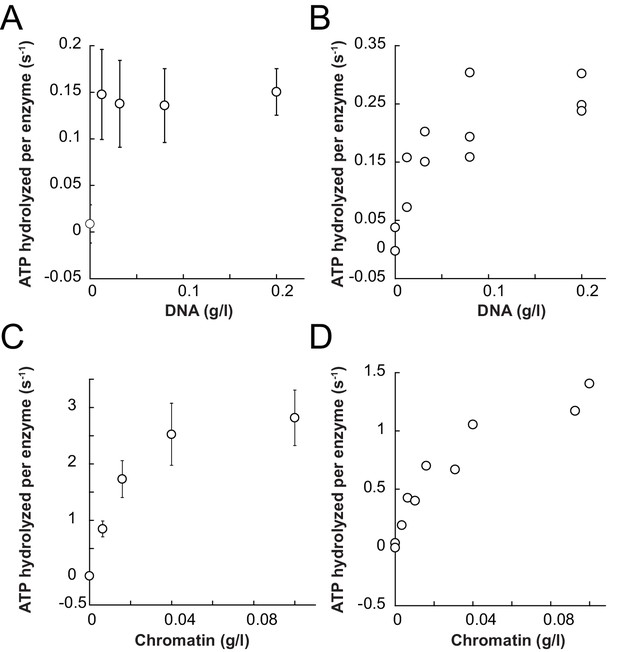
Saturation controls for ISWIWT and ISWIΔppHSA in ATPase assays.
(A,B) Linearized pT7blue DNA was titrated over a 16-fold range. 0.2 mg/ml were saturating for ISWIWT. (A) and ISWIΔppHSA (B). (C,D) Titration of chromatin assembled on linearized pT7blue DNA. 0.1 mg/ml were close to saturation for ISWIWT (C) and ISWIΔppHSA (D). Errors are s.d. (n ≥ 7).
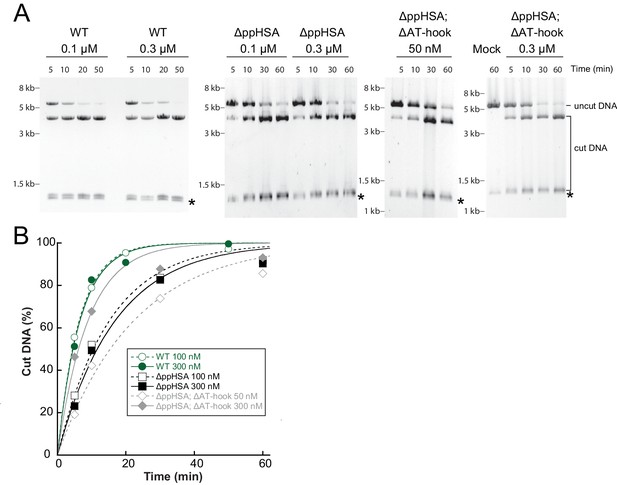
Determination of the rate constants for remodeling (kobs; Figure 3C) for ISWIWT and N-terminal truncation mutants of ISWI.
(A) Exemplary remodeling time courses for ISWIWT, ISWIΔppHSA and ISWIΔppHSA; ΔAT-hook. Asterisks mark a contaminating non-nucleosomal DNA (competitor DNA) that was not completely removed during preparation of nucleosomal arrays. Mock: Sample lacking ISWI. (B) Quantification of time courses shown in (A). Data were fit to a single exponential function to extract the rate constant kobs. The reactions progressed similarly fast when 100 nM and 300 nM enzyme were employed, suggesting saturation of chromatin. Because ISWIΔppHSA; ΔAT-hook at 50 nM was substoichiometric to nucleosomes (100 nM), it remodeled noticeably more slowly than at 300 nM.
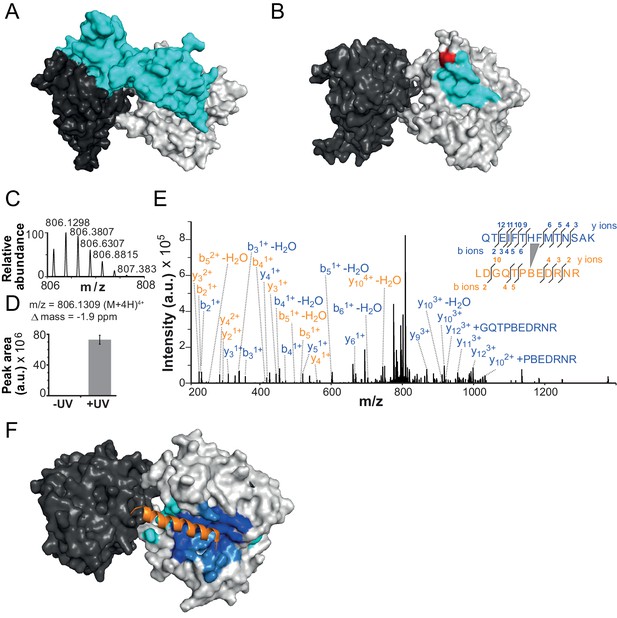
The NTR contacts Lobe 2 of the ATPase domain.
(A) Surface representation of the Chd1 crystal structure (PDB code 3MWY) (Hauk et al., 2010). ATPase Lobe 1 and 2 are colored dark and light grey, respectively, and the N-terminal chromo domains cyan. (B) Homology model of the ISWI ATPase domain (Forné et al., 2012). Cyan: hypothetical binding interface of the ISWI NTR (see main text), red: position of Bpa substitution (H483). (C–E) Mass spectrometric validation of the crosslink XL1 (Table 1) formed between Bpa at position 483 and an NTR peptide. (C) Isotopic distribution of the crosslinked peptide. (D) UV-dependent increase of the signal for the crosslinked peptide. Extracted ion chromatograms of the ions were used for the quantification. (E) High resolution, high accuracy MS2 fragmentation spectrum. Top right: summary of observed product ions mapped onto the sequence of the crosslinked peptide. B: Bpa. (F) Predicted docking interface of AcidicN (blue and dark blue), AutoN (cyan and dark blue) and overlapping regions (dark blue) in the structural model of ISWI. The predicted interface for AcidicN overlaps with the interface for the acidic helix of the N-terminal chromo domains of Chd1 (orange) (Hauk et al., 2010).
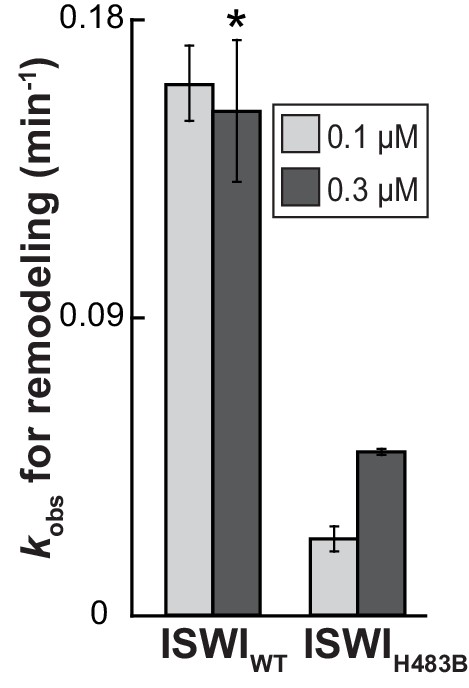
The effect of the H483B mutation on chromatin remodeling.
With 0.3 µM enzyme, the observed rate constant for remodeling (kobs) was ~threefold affected by the mutation. Note, however, that ISWIH483B, in contrast to ISWIWT, was not fully saturating at this concentration, as suggested by the saturation control (0.1 µM enzyme). Error bars are s.d. (n ≥ 3) for ISWIWT and minimal and maximal values of two independent measurements for ISWIH483B. Data marked with an asterisk (*) was replotted from Figure 3C.
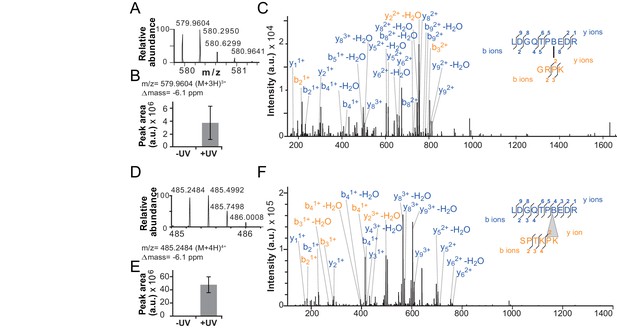
Validation of additional crosslinks detected in the ISWIH483B dataset.
(A–C) Crosslink XL7 (Table 1) to P75. (D–F) Crosslink XL6 to P71K72. For description of data, see Figure 4.
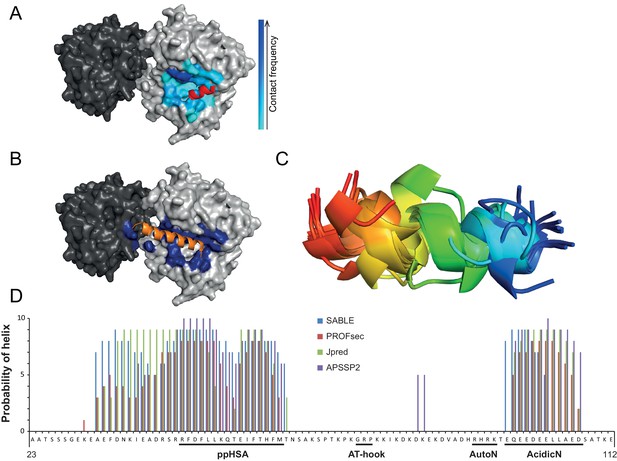
Structural predictions of NTR elements.
(A) Predicted docking interface of AcidicN. The cyan to blue color scale denotes low to high contact probabilities. The location of AcidicN in the MtISWI crystal structure (PDB 5JXR), which became available during the revision of this study, is shown in red. (B) Predicted docking interface (blue) of the AutoN-AcidicN peptide in the structural model of ISWI. During docking, both Lobe 1 and Lobe 2 were present (cf. the docking interface shown in Figure 4F). The acidic helix of the N-terminal chromo domains of Chd1 is shown in orange for reference. (C) Structure prediction of a peptide comprising AutoN and AcidicN (DHRHRKTEQEEDEELL) by PEP-FOLD and I-TASSER. (D) Helical Propensity of amino acids 23–112 of ISWI predicted by four different algorithms (see legend).
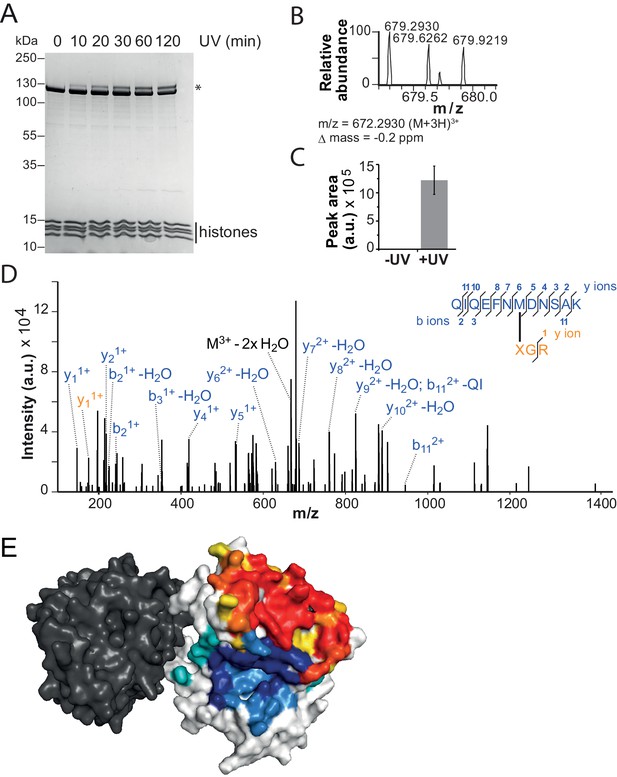
The binding sites of the NTR and the H4-tail on Lobe 2 are proximal.
(A–D) Crosslinking of nucleosomes containing benzophenone-labeled H4 to ISWI. (A) Crosslinking time course analyzed by SDS-PAGE and Coomassie staining. The asterisk marks a UV-irradiation dependent band of lower mobility containing the crosslink mapped in B–D. (B–D) Mapping and validation of a crosslink (XL11; Table 2) formed in the upshifted band in A. Isotopic distribution of the crosslinked peptide, MS2 spectrum and quantification as in Figure 4. (E) Crosslink-guided in silico docking of an H4 peptide (amino acids 1–20) to ISWI. The predicted docking interface of the H4 tail on Lobe 2 is illustrated in a yellow and red color scale, which indicates low to high contact probabilities between the docked H4 tail and Lobe 2. The contact probabilities were calculated from a family of 383 docked structures (see Materials and methods). For comparison, the predicted docking interface of the NTR is shown in shades of blue (see Figure 4F).
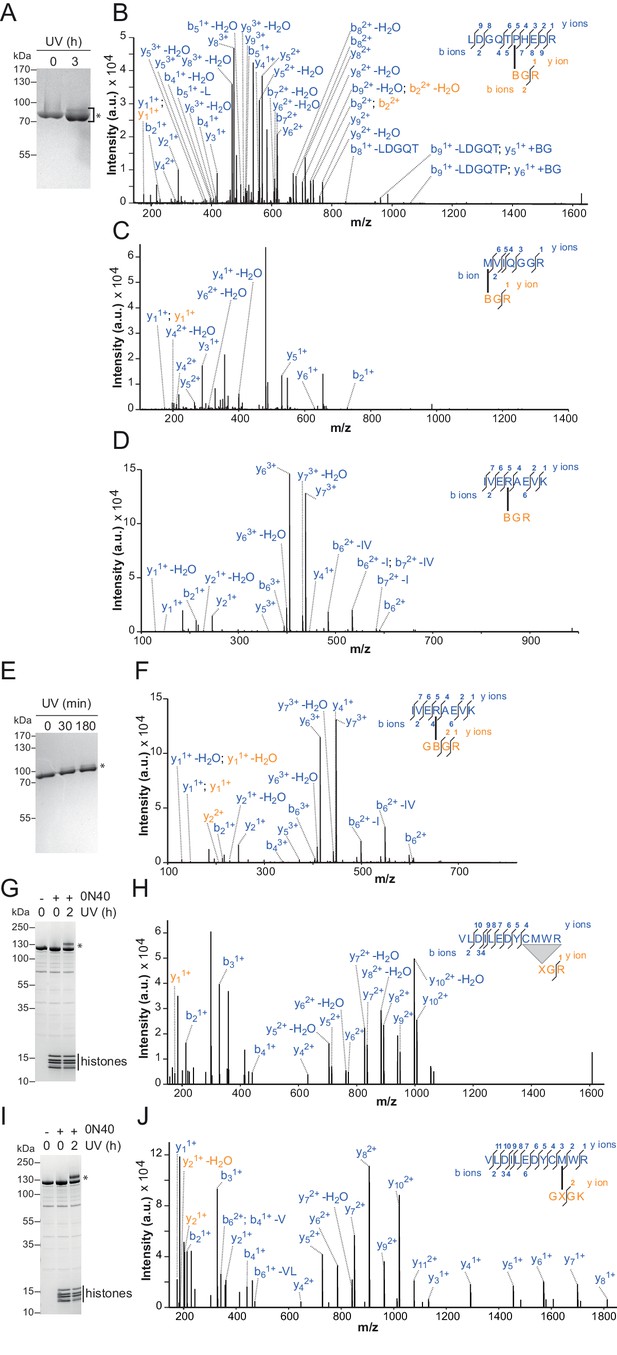
Additional crosslinks between the H4 tail and ISWI or SNF2H.
(A–D) SDS-PAGE analysis and MS mapping of crosslinks formed between a synthetic H4 tail peptide containing Bpa at amino acid one to the following amino acids in ISWI26-648: H482 (A; XL13a in Table 2); M578 (B; XL15); R568 (C; XL16). (E–F) MS mapping and SDS-PAGE analysis of crosslinks formed between a synthetic H4 tail peptide containing Bpa at amino acid 10 to R568 of ISWI26-648 (XL17). Asterisks next to SDS gels mark upshifted bands indicative of successful crosslinking. (G,H) SDS-PAGE analysis and MS mapping of crosslinks formed by benzophenone-labeled T1C nucleosomes to C519M520W521R522 of SNF2H (XL14). (I,J) SDS-PAGE analysis and MS mapping of crosslinks formed by benzophenone-labeled L10C nucleosomes to position 520 of SNF2H (XL12). B: Bpa, X: Benzophenone-labeled cysteine, 0N40: Mononucleosomes with 40 bp of DNA flanking one side of the nucleosome.
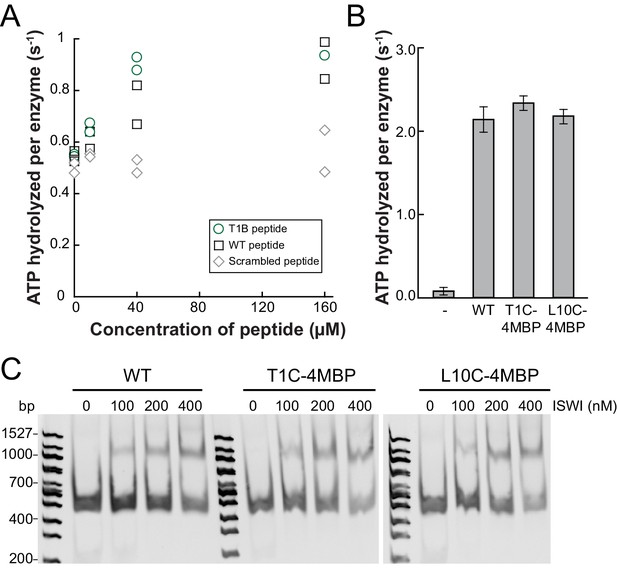
Controls for possible adversary effects of covalent modifications of the H4 tail.
(A) H4 tail peptides (amino acid 1–24) carrying a T1B substitution stimulated the ATPase of ISWIWT (0.5 µM) activity similarly well as WT tail peptides in the presence of 1.2 g/l salmon sperm DNA. In contrast, a scrambled sequence with a Bpa moiety at position one did not noticeably stimulate the ATPase. The ATP concentration was 1 mM. Scrambled and WT peptide data were replotted from ref. 15. (B) T1C and L10C mononucleosomes labeled with 4-(N-Maleimido)benzophenone (4MBP) stimulated the ATPase activity of ISWIWT similarly well as WT nucleosomes. A reaction without nucleosomes (−) served as a control. (C) WT and 4MBP-labeled T1C and L10C mononucleosomes (200 nM) bound ISWIWT (0 to 400 nM) similarly well in an electrophoretic mobility shift assay. Samples were resolved on a 5% native polyacrylamide gel.
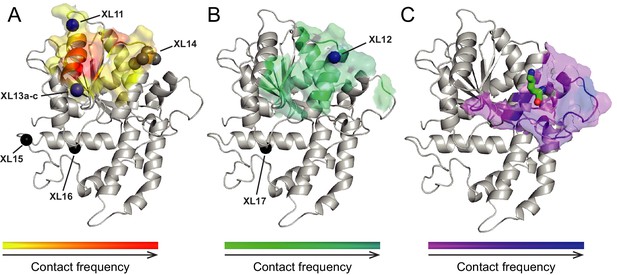
Surfaces on Lobe 2 that were sampled by selected amino acids in the H4 tail during crosslink-guided structural docking.
(A) Interaction surface of histone H4 T1 on Lobe 2 of ISWI. Crosslinked amino acids (Table 2) are shown as spheres. Blue, high confidence crosslink positions used for modeling; black and grey, lower confidence crosslink positions. Precise attachment sites are not available for XL14 (grey). (B) Interaction surface of H4 L10. Coloring of spheres as in (A). (C) Predicted interaction surface of H4 K16. H4 K16 from the crystal structure of MtISWI Lobe 2 in complex with an H4 peptide (PDB 5JXT) is shown as stick representation for reference. The color scales indicate contact probabilities between individual amino acids in the H4 tail and Lobe 2 across a family of 383 structural models.
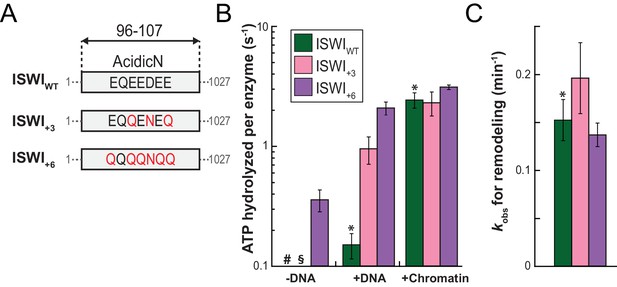
AcidicN is a strong negative regulator of the ATPase.
(A) Design of AcidicN derivatives of ISWI (see also Figure 6—figure supplement 1A). (B) Effects of AcidicN mutation on ATP hydrolysis in absence or presence of saturating concentrations of DNA and chromatin. In absence of DNA, ATPase activities of ISWIWT (#) and ISWI+3 (§) were ≤0.06 s−1. Errors are s.d. (n ≥ 4). (C) Effects of AcidicN mutation on the remodeling activities. Nucleosomal arrays containing wild-type H4 were used. Errors are s.d. (n ≥ 3) except for ISWI+3 for which minimal and maximal values of two independent measurements are shown. Color code as in panel (B). Raw data of the remodeling assay can be found in Figure 8—figure supplement 1. Results for ISWIWT (*) are replotted for comparison from Figure 3B,C.
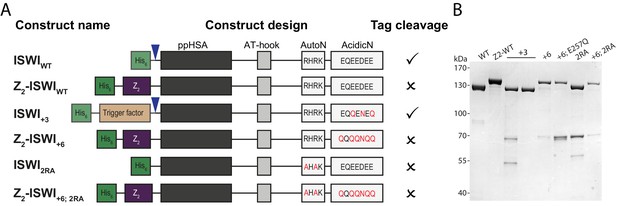
AcidicN and AutoN mutants.
(A) Construct design. Only the NTR region of ISWI including affinity and solubility tags are shown (not to scale). Blue arrowheads indicate a TEV cleavage site. Tags were removed by protease cleavage as indicated. (B) Coomassie-stained SDS-PAGE of purified recombinant proteins.
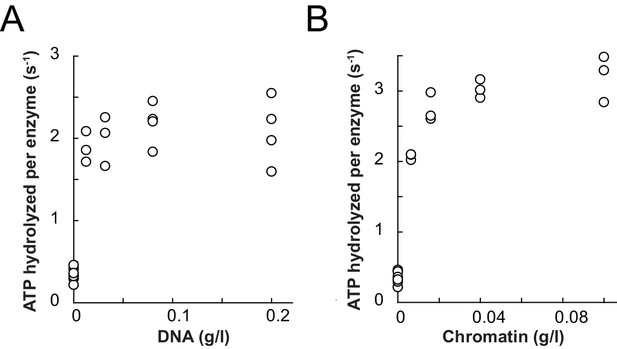
Comparison of ATPase and remodeling activities of ISWI control variants used in this study.
(A) The Z2 solubility tag did not interfere with DNA- and chromatin-stimulated ATPase activities. Saturating amounts of nucleic acid ligands (0.2 mg/ml of linearized pT7blue and 0.1 mg/ml of chromatin assembled on the same DNA, respectively) and ATP (1 mM) were used. The unstimulated basal activity was ≤0.05 s−1. Errors are s.d. (n ≥ 3). (B) The Z2 solubility tag did not interfere with remodeling rates on wild-type H4 containing chromatin and tail-less H4 chromatin. Z2-tagged ISWI+6; E257Q, which contained a point mutation in the ATPase domain rendering it catalytically inactive, was included as a control. Its activity on tail-less H4 arrays was undetectable (§). Errors are s.d. (n ≥ 3) except for the ATPase-dead construct (ISWI+6; E257Q), which was tested once. (C) DNA- and chromatin-stimulated ATP hydrolysis rates of the ATPase dead double mutant ISWI+6; E257Q were negligible (≤0.04 s−1). Errors are s.d. (n ≥ 3) for ISWIWT and minimal and maximal values of two independent measurements for ISWI+6; E257Q. The asterisks (*) mark data that were replotted for comparison from Figure 3B,C.
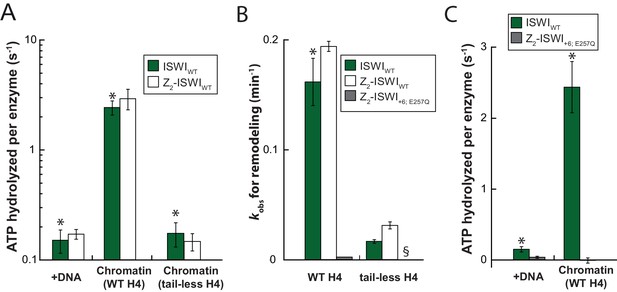
Saturation controls for ISWI+6 in ATPase assays.
(A) Linearized pT7blue DNA was titrated over a 16-fold range. 0.2 mg/ml were saturating. (B) Titration of chromatin assembled on linearized pT7blue DNA. 0.1 mg/ml were close to saturation.
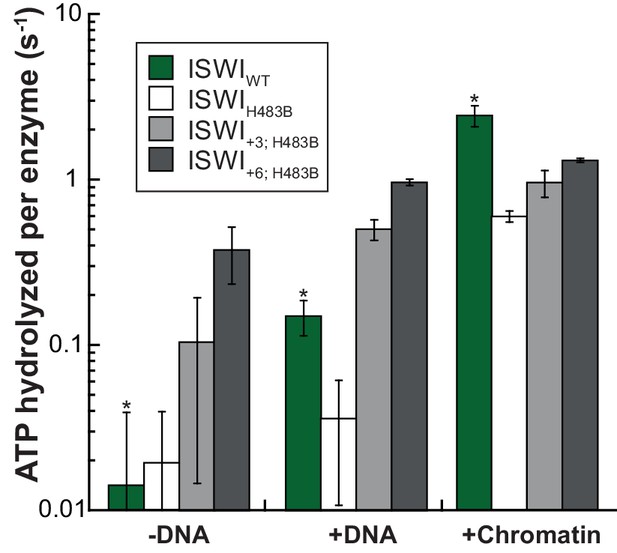
AcidicN mutations upregulate the ATPase activity of ISWIH483B.
Relative to ISWIWT, ISWIH483B had a ~fourfold diminished DNA- and chromatin-stimulated ATPase activity. Additional mutation of AcidicN (+3; +6) strongly activated both DNA- and chromatin-stimulated ATP turnover. Errors are s.d. for ISWIWT and minimal and maximal values of two independent measurements for all other constructs. Data for ISWIWT (*) were replotted for comparison from Figure 3B.
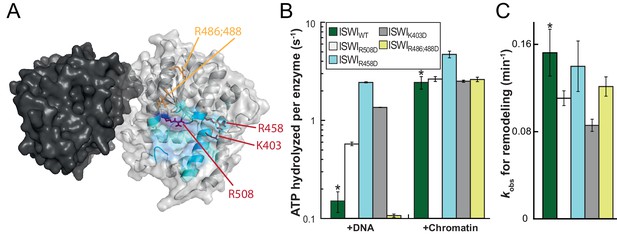
Validation of the predicted binding interface of AcidicN on Lobe 2.
(A) Homology model of the ISWI ATPase domain. Dark and light grey, ATPase lobes 1 and 2, respectively; blue, hypothetical binding interface of AcidicN as in Figure 4—figure supplement 3A. Positively charged residues selected for mutagenesis are shown in red (AcidicN interface mutant) and orange (control mutant). (B) Mutation of the AcidicN interface (K403D, R458D and R508D) strongly upregulated DNA-stimulated ATP hydrolysis relative to ISWIWT, whereas the nucleosome-stimulated ATP turnover was similar. In contrast, a control mutation (R486; 488D) had little effect on ATP hydrolysis. Saturating concentrations of DNA and chromatin were used. Errors are s.d. for ISWIWT and minimal and maximal values of two independent measurements for all other constructs. (C) AcidicN interface variants of ISWI robustly remodeled nucleosomes within twofold of ISWIWT. Nucleosomal arrays containing wild-type H4 were used. Errors are s.d. (n ≥ 3) for ISWIWT and minimal and maximal values of two independent measurements for all other constructs. Raw data of the remodeling assay can be found in Figure 7—figure supplement 2. Color code as in (B). Results for ISWIWT (*) were replotted for comparison from Figure 3B,C.
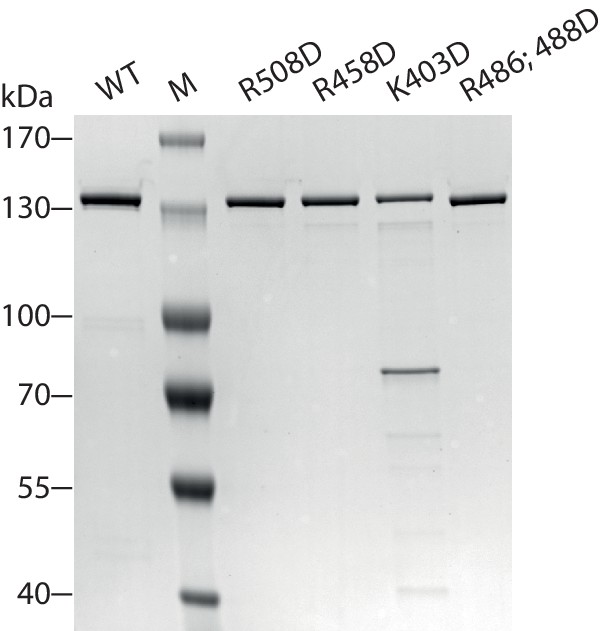
Coomassie-stained SDS-PAGE of purified recombinant ISWI constructs analyzed in Figure 7.
https://doi.org/10.7554/eLife.21477.027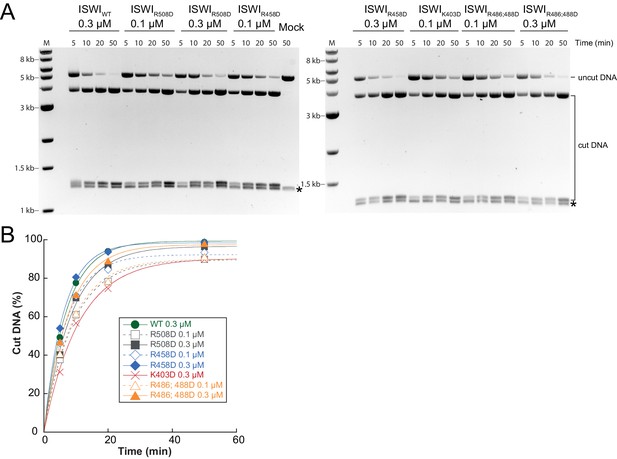
Determination of rate constants for remodeling of AcidicN interface mutants.
(A) Exemplary remodeling time courses on WT H4-arrays for interface mutants. Asterisks mark a contaminating non-nucleosomal DNA (competitor DNA) that was not completely removed during preparation of nucleosomal arrays. Mock: Sample lacking ISWI. (B) Quantification of time courses shown in (A). Data were fit to a single exponential function to extract kobs (see Figure 7C).
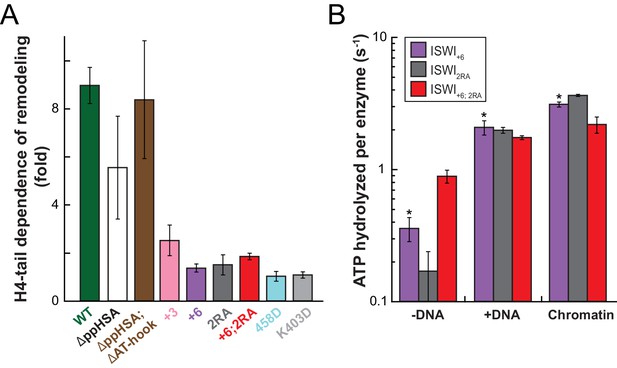
Mutation of AcidicN, the AcidicN binding interface or AutoN suppresses dependence on the H4-tail.
(A) H4-tail dependence of the remodeling activities of ISWI variants. Values were calculated from the observed remodeling rate constants obtained for WT and tail-less H4 chromatin (Figure 8—figure supplement 1E). (B) ATP hydrolysis measurements of ISWT+6, ISWI2RA and ISWI+6; 2RA in absence or presence of saturating concentrations of DNA and chromatin.
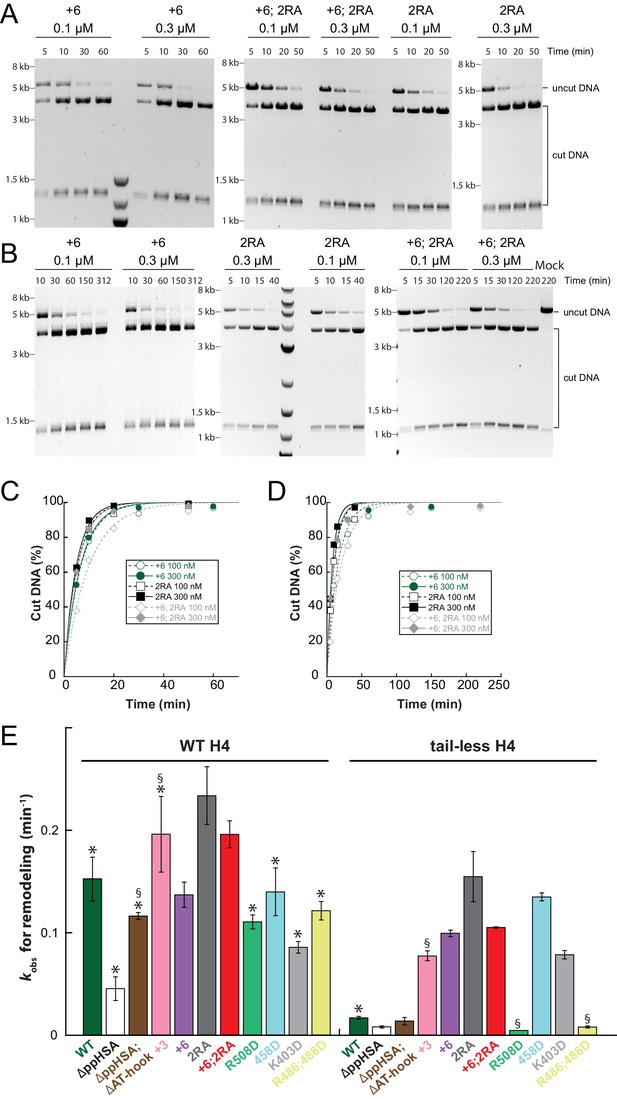
Raw data of the remodeling assays.
(A–D) Determination of rate constants (kobs) from remodeling assays for ISWI+6, ISWI2RA and ISWI+6; 2RA. Shown are exemplary time courses on nucleosomal arrays containing wild-type (A,C) and tail-less H4 (B,D). Data were fit to a single exponential function to extract the rate constant kobs. (E) Rate constants for remodeling of ISWI variants used in this study (all 300 nM). Errors are s.d. (n ≥ 3) for ISWIWT, ISWIΔppHSA, and ISWI+6 and minimal and maximal values of two independent measurements for all other variants. Samples, in which the enzyme concentration was not saturating, are indicated (§). Data marked with asterisks (*) were replotted from previous figures for better overview.
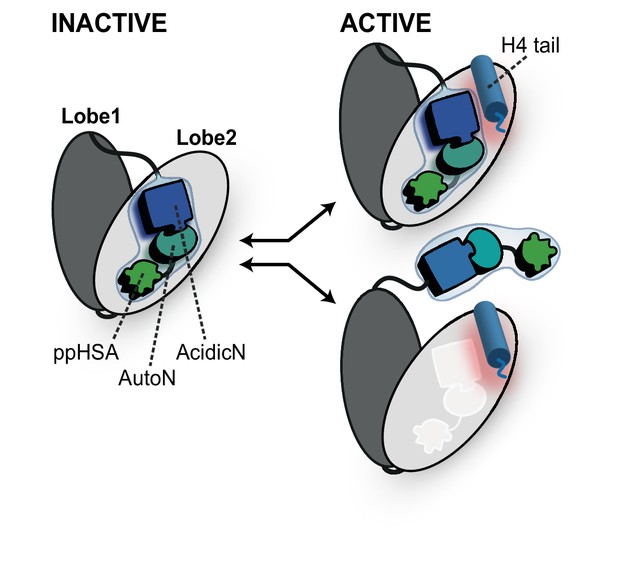
Proposed models for autoregulation imposed by the NTR and the recognition process of the H4 tail.
The ppHSA motif, AcidicN and AutoN dock against Lobe 2 of the ATPase domain, promoting an overall structural architecture of the ATPase module that is reminiscent of Chd1 (Figure 4A). AcidicN and AutoN functionally collaborate in the H4 tail recogniton process. The docking site of AutoN-AcidicN is adjacent to the H4 tail potentially allowing simultaneous binding (top). Alternatively, the H4 tail may displace the NTR as suggested previously (bottom) (Clapier and Cairns, 2012).
Tables
Overview of crosslinks formed by ISWIH483B.
| ID | Mass (D a) | Error (ppm) | Bpa peptide | Target peptide | ||
|---|---|---|---|---|---|---|
| Sequence*,† | Site | Sequence† | Site | |||
| XL1 | 3220.4946 | −1.9 | LDGQTPBEDRNR | 483 | QTEIFTHFMoxTNSAK‡ | 59–60 |
| XL2 | 3204.5056 | −3.7 | LDGQTPBEDRNR | 483 | QTEIFTHFMTNSAK‡ | 60–61 |
| XL3 | 2950.3474 | −1.0 | LDGQTPBEDR | 483 | QTEIFTHFMoxTNSAK‡ | 55–59 |
| XL4 | 2934.3571 | −2.6 | LDGQTPBEDR | 483 | QTEIFTHFMTNSAK‡ | 59–61 |
| XL5 | 2207.0968 | +0.2 | LDGQTPBEDRNR | 483 | SPTKPK‡ | 69–72 |
| XL6 | 1936.9645 | −6.1 | LDGQTPBEDR | 483 | SPTKPK‡ | 71–72 |
| XL7 | 1736.8594 | −6.1 | LDGQTPBEDR | 483 | GRPK | 75 |
-
*B symbolizes Bpa.
-
†Crosslinked amino acids are underlined; oxindicates oxidized methionine (+15.9949 Da).
-
‡Precise attachment sites not distinguishable from data.
Overview of H4-tail mediated crosslinks.
| ID | Reliability | H4 | Remodeler construct | Mass (Da) | Error (ppm) | H4 peptide | Remodeler peptide | ||
|---|---|---|---|---|---|---|---|---|---|
| Sequence* | Site | Sequence† | Site | ||||||
| XL11 | high | nucleosomal | ISWIWT | 2034.8571 | −0.2 | XGR | 1 | QIQEFNMDNSAK | 495 |
| XL12 | high | nucleosomal | SNF2H | 2251.9753 | −0.4 | GXGK | 10 | VLDILEDYCMWR | 520 |
| XL13a | high | peptide | ISWI26-648 | 1648.7601 | −0.4 | BGR | 1 | LDGQTPHEDR | 482 |
| XL13b | high | peptide | ISWI26-648 | 1918.9052 | −0.9 | BGR | 1 | LDGQTPHEDRNR | 482 |
| XL13c | high | peptide | ISWI26-648 | 3340.5374 | −2.8 | BGR | 1 | LDGQTPHEDRNRQIQEFNMDNSAK | 482 |
| XL14 | medium | nucleosomal | SNF2H | 2222.9624 | −1.5 | XGR | 1 | VLDILEDYCMWR‡ | 519–22 |
| XL15 | medium | peptide | ISWI26-648 | 1257.6261 | +2.3 | BGR | 1 | MVIQGGR | 578 |
| XL16 | medium | peptide | ISWI26-648 | 1424.7832 | −3.9 | BGR | 1 | IVERAEVK | 568 |
| XL17 | medium | peptide | ISWI26-648 | 1453.7998 | −4.6 | GBGK | 10 | IVERAEVK | 568 |
-
*B symbolizes Bpa; X symbolizes Benzophenone-labeled cysteine.
-
†Crosslinked amino acids are underlined.
-
‡Precise attachment sites not distinguishable from data.
Yeast strains used in this study.
Strain | Genotype | Reference |
|---|---|---|
W1588-4C | MATa ade2-1 his3-11,15 leu2-3,112 trp1-1 ura3-1 can1-100 but RAD5 | |
YTT227 | MATa ade2-1 his3-11,15 leu2-3,112 trp1-1 ura3-1 can1-100 but RAD5 isw1::ADE2 isw2::LEU2 chd1::TRP1 | |
YTT225 | MATa ade2-1 his3-11,15 leu2-3,112 trp1-1 ura3-1 can1-100 but RAD5 isw2::LEU2 chd1::TRP1 | |
YFMP047 | MATa his3Δ1 leu2Δ0 met15Δ0 ura3Δ0 ISW1-TAP::HIS3MX6 | Open Biosystems |





